|
Order Kazusa clone(s) from : |
|
Order Kazusa clone(s) from : |
| Product ID | ORK04592 |
|---|---|
| Accession No | AB020709 |
| Description | connector enhancer of kinase suppressor of Ras 2 |
| Clone name | hk09777 |
| Vector information | |
| cDNA sequence | DNA sequence (4349 bp) Predicted protein sequence (910 aa) |
| Source | Human adult brain |
| Rouge ID |
mKIAA0902
by Kazusa Mouse cDNA Project
|
 Length: 4349 bp
Length: 4349 bp Physical map
Physical map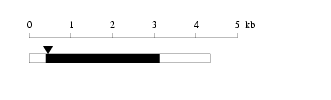
 Restriction map
Restriction map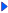 Prediction of protein coding region (GeneMark analysis).
Prediction of protein coding region (GeneMark analysis).
| N-terminal truncation | Coding interruption | |
|---|---|---|
| cloned DNA seq | No warning | Warning |
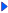 Integrity of 3' end
Integrity of 3' end
| Length of 3'UTR | 1211 bp |
|---|---|
| Genome contig ID | gi89161218f_21202481 |
| PolyA signal sequence (AATAAA,-27) |
+----*----+----*----+----*----+---- |
| Flanking genome sequence (368294 - 368343) |
----+----*----+----*----+----*----+----*----+----* |
 Length: 910 aa
Length: 910 aa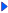 Result of homology search against nr database
(FASTA output,
Multiple alignment)
Result of homology search against nr database
(FASTA output,
Multiple alignment)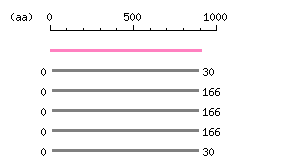 |
|
The numbers on the left and right sides of a black line in the graphical overview indicate the lengths (in amino acid residues) of the non-homologous N-terminal and C-terminal portions flanking the homologous region (indicated by the black line), respectively.
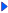 Result of homology search against HUGE database
(FASTA output,
Multiple alignment)
Result of homology search against HUGE database
(FASTA output,
Multiple alignment)The numbers on the left and right sides of a black line in the graphical overview indicate the lengths (in amino acid residues) of the non-homologous N-terminal and C-terminal portions flanking the homologous region (indicated by the black line), respectively.
 Result of motif / domain search (InterProScan and SOSUI)
Result of motif / domain search (InterProScan and SOSUI) Result of InterProScan
Result of InterProScan
| Search method | interpro_ID | From | To | Entry | Definition |
|---|---|---|---|---|---|
| HMMPfam | IPR001660 | 26 | 90 | PF00536 | Sterile alpha motif SAM |
| IPR001478 | 234 | 311 | PF00595 | PDZ/DHR/GLGF | |
| IPR010599 | 349 | 532 | PF06663 | Connector enhancer of kinase suppressor of ras 2 | |
| IPR001849 | 588 | 686 | PF00169 | Pleckstrin-like | |
| HMMSmart | IPR001660 | 25 | 93 | SM00454 | Sterile alpha motif SAM |
| IPR001478 | 242 | 314 | SM00228 | PDZ/DHR/GLGF | |
| IPR001849 | 588 | 688 | SM00233 | Pleckstrin-like | |
| ProfileScan | IPR001660 | 28 | 93 | PS50105 | Sterile alpha motif SAM |
| IPR001478 | 232 | 314 | PS50106 | PDZ/DHR/GLGF | |
| IPR001849 | 587 | 686 | PS50003 | Pleckstrin-like |
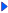 RT-PCR-ELISA
RT-PCR-ELISA
|
 Experimental conditions
Experimental conditions| Primer_f | AATCTATAGGCTGTGGGTTTC |
|---|---|
| Primer_r | ATTGATTGAGCCATAGGGGAC |
| PCR conditions | 95 °C 30 sec 30 sec 55 °C 55 °C 30 sec 30 sec 72 °C 72 °C 60 sec 60 sec 30 cycles 30 cycles |
 Chromosome No. 1
Chromosome No. 1 Experimental conditions
Experimental conditions| Panel name | GeneBridge 4 |
|---|---|
| Primer_f | AATCTATAGGCTGTGGGTTTC |
| Primer_r | ATTGATTGAGCCATAGGGGAC |
| PCR product length | 113 bp |
| PCR conditions | 95 °C 15 sec 15 sec 62 °C 62 °C 60 sec 60 sec 30 cycles 30 cycles |