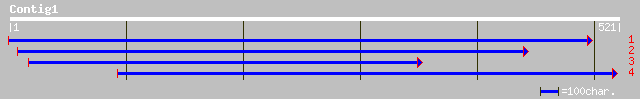
Nr search
BLASTX 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= KMC002138A_C01 KMC002138A_c01
(517 letters)
Database: nr
1,393,205 sequences; 448,689,247 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAA36585.1| adenylyl cyclase associated protein [Gossypium h... 100 2e-23
ref|NP_195175.1| putative cyclase associated protein CAP; protei... 105 2e-22
sp|P54654|CAP_DICDI ADENYLYL CYCLASE-ASSOCIATED PROTEIN (CAP) gi... 80 2e-14
ref|NP_524806.1| capulet CG5061-PA gi|18466908|ref|XP_079396.1| ... 67 9e-11
gb|AAD27865.2|AF132566_1 LD24380p [Drosophila melanogaster] 67 9e-11
>dbj|BAA36585.1| adenylyl cyclase associated protein [Gossypium hirsutum]
Length = 471
Score = 99.8 bits (247), Expect(2) = 2e-23
Identities = 52/67 (77%), Positives = 55/67 (81%)
Frame = -2
Query: 516 QGSAPTISVDNTSGCQLYLSKDSLETSISTAKSSEINVLVPGAEPDGDLVEHSLPQPYIH 337
QGSAPTISVDNTSGCQLYLSKDSL TSI++AKSSEINVLVP PDGD EHSLPQ YIH
Sbjct: 396 QGSAPTISVDNTSGCQLYLSKDSLGTSITSAKSSEINVLVP-TGPDGDWGEHSLPQQYIH 454
Query: 336 ALQGWSF 316
+ F
Sbjct: 455 VFKDGQF 461
Score = 30.8 bits (68), Expect(2) = 2e-23
Identities = 12/14 (85%), Positives = 13/14 (92%)
Frame = -3
Query: 329 KDGRFETTPASHSG 288
KDG+FETTP SHSG
Sbjct: 457 KDGQFETTPVSHSG 470
>ref|NP_195175.1| putative cyclase associated protein CAP; protein id: At4g34490.1,
supported by cDNA: gi_3169135 [Arabidopsis thaliana]
gi|7447122|pir||T05269 adenylyl cyclase-associated
protein [imported] - Arabidopsis thaliana
gi|3096918|emb|CAA18828.1| putative cyclase associated
protein CAP [Arabidopsis thaliana]
gi|3169136|dbj|BAA28621.1| Atcap1 [Arabidopsis thaliana]
gi|7270399|emb|CAB80166.1| putative cyclase associated
protein CAP [Arabidopsis thaliana]
Length = 476
Score = 105 bits (263), Expect = 2e-22
Identities = 50/67 (74%), Positives = 57/67 (84%)
Frame = -2
Query: 516 QGSAPTISVDNTSGCQLYLSKDSLETSISTAKSSEINVLVPGAEPDGDLVEHSLPQPYIH 337
QGSAPT+SVDNT+GCQLYL+KDSLET+I+TAKSSEINV+VPGA PDGD VEH+LPQ Y H
Sbjct: 400 QGSAPTVSVDNTTGCQLYLNKDSLETAITTAKSSEINVMVPGATPDGDWVEHALPQQYNH 459
Query: 336 ALQGWSF 316
F
Sbjct: 460 VFTEGKF 466
>sp|P54654|CAP_DICDI ADENYLYL CYCLASE-ASSOCIATED PROTEIN (CAP) gi|1150975|gb|AAB09713.1|
cyclase associated protein
Length = 464
Score = 79.7 bits (195), Expect = 2e-14
Identities = 36/63 (57%), Positives = 51/63 (80%)
Frame = -2
Query: 513 GSAPTISVDNTSGCQLYLSKDSLETSISTAKSSEINVLVPGAEPDGDLVEHSLPQPYIHA 334
G P+I++D TSGCQ+YLSKDSLET I ++KSSE+NVL+PGA + DLVE ++P+ Y +
Sbjct: 391 GRVPSIAIDKTSGCQIYLSKDSLETEIVSSKSSEMNVLIPGATENDDLVELAIPEQYKTS 450
Query: 333 LQG 325
++G
Sbjct: 451 VKG 453
>ref|NP_524806.1| capulet CG5061-PA gi|18466908|ref|XP_079396.1| similar to capulet;
Adenylyl cyclase-associated protein; Act up (D.
melanogaster) [Drosophila melanogaster]
gi|22945519|gb|AAF51408.2| CG5061-PA [Drosophila
melanogaster]
Length = 424
Score = 67.4 bits (163), Expect = 9e-11
Identities = 32/63 (50%), Positives = 43/63 (67%)
Frame = -2
Query: 513 GSAPTISVDNTSGCQLYLSKDSLETSISTAKSSEINVLVPGAEPDGDLVEHSLPQPYIHA 334
GS PT+S+D T GCQ+YLSKDSL I +KSSE+N+L+P + GD E +LP+ Y
Sbjct: 352 GSVPTVSIDKTDGCQMYLSKDSLGVEIVNSKSSEMNILLP--DDSGDYTELALPEQYKTT 409
Query: 333 LQG 325
+ G
Sbjct: 410 IAG 412
>gb|AAD27865.2|AF132566_1 LD24380p [Drosophila melanogaster]
Length = 528
Score = 67.4 bits (163), Expect = 9e-11
Identities = 32/63 (50%), Positives = 43/63 (67%)
Frame = -2
Query: 513 GSAPTISVDNTSGCQLYLSKDSLETSISTAKSSEINVLVPGAEPDGDLVEHSLPQPYIHA 334
GS PT+S+D T GCQ+YLSKDSL I +KSSE+N+L+P + GD E +LP+ Y
Sbjct: 456 GSVPTVSIDKTDGCQMYLSKDSLGVEIVNSKSSEMNILLP--DDSGDYTELALPEQYKTT 513
Query: 333 LQG 325
+ G
Sbjct: 514 IAG 516
Database: nr
Posted date: Apr 1, 2003 2:05 AM
Number of letters in database: 448,689,247
Number of sequences in database: 1,393,205
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 465,902,031
Number of Sequences: 1393205
Number of extensions: 9856572
Number of successful extensions: 27547
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 26749
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 27535
length of database: 448,689,247
effective HSP length: 115
effective length of database: 288,470,672
effective search space used: 16154357632
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)