

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149581.3 + phase: 0
(194 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
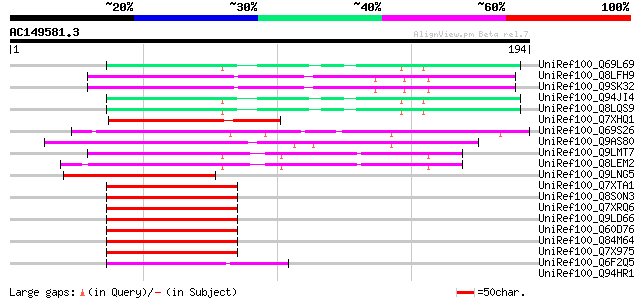
Sequences producing significant alignments: (bits) Value
UniRef100_Q69L69 Calcineurin-like phosphoesterase-like protein [... 56 5e-07
UniRef100_Q8LFH9 Hypothetical protein [Arabidopsis thaliana] 55 1e-06
UniRef100_Q9SK32 Expressed protein [Arabidopsis thaliana] 54 2e-06
UniRef100_Q94JI4 P0638D12.5 protein [Oryza sativa] 54 2e-06
UniRef100_Q8LQS9 P0702H08.15 protein [Oryza sativa] 54 2e-06
UniRef100_Q7XHQ1 Calcineurin-like phosphoesterase-like [Oryza sa... 53 4e-06
UniRef100_Q69S26 Serine/threonine phosphatase PP7-related-like [... 52 9e-06
UniRef100_Q9AS80 P0028E10.12 protein [Oryza sativa] 51 1e-05
UniRef100_Q9LMT7 F2H15.15 protein [Arabidopsis thaliana] 50 3e-05
UniRef100_Q8LEM2 Hypothetical protein [Arabidopsis thaliana] 49 6e-05
UniRef100_Q9LNG5 F21D18.16 [Arabidopsis thaliana] 47 2e-04
UniRef100_Q7XTA1 OSJNBa0008A08.9 protein [Oryza sativa] 47 3e-04
UniRef100_Q8S0N3 Putative mutator-like transposase [Oryza sativa] 47 4e-04
UniRef100_Q7XRQ6 OSJNBb0096E05.18 protein [Oryza sativa] 47 4e-04
UniRef100_Q9LD66 Similar to maize transposon MuDR mudrA protein ... 47 4e-04
UniRef100_Q60D76 Putative polyprotein [Oryza sativa] 47 4e-04
UniRef100_Q84M64 Putative mutator-like transposase [Oryza sativa] 47 4e-04
UniRef100_Q7X975 Putative Mutator protein [Oryza sativa] 47 4e-04
UniRef100_Q6F2Q5 Hypothetical protein OSJNBa0088M06.4 [Oryza sat... 46 6e-04
UniRef100_Q94HR1 Hypothetical protein OSJNBa0065C16.8 [Oryza sat... 45 0.001
>UniRef100_Q69L69 Calcineurin-like phosphoesterase-like protein [Oryza sativa]
Length = 766
Score = 56.2 bits (134), Expect = 5e-07
Identities = 55/185 (29%), Positives = 75/185 (39%), Gaps = 45/185 (24%)
Query: 37 AMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGR----ILDHDEAMNQDCAI 92
A+I A+ ERW ET SFHL GEMT+ L D+ LL +PI GR DHD A
Sbjct: 132 ALITALVERWRPETHSFHLASGEMTVTLQDVSMLLALPIDGRPVCSTTDHDYA------- 184
Query: 93 DLMIRLLGMSEVDVRVEVRSESAGHISYPTLKRVYEHHLTEARRLEEPQTREK------- 145
++I LG + +S + Y LK+ H E + QT E+
Sbjct: 185 QMVIDCLGHDPRGPSMPGKS----FLHYKWLKK----HFYELPEGTDDQTVERHVRAYIL 236
Query: 146 ------LQERGRKR-------------AIGKWSWGGMALAYLYDYLDDSVILNNRTMAGS 186
L G R +G +SWG LA+LY L +N G
Sbjct: 237 SLLCGVLFPDGTGRMSLIYLPLIADLSLVGTYSWGSAVLAFLYRSLCSVASSHNIKNIGG 296
Query: 187 TILFM 191
++L +
Sbjct: 297 SLLLL 301
>UniRef100_Q8LFH9 Hypothetical protein [Arabidopsis thaliana]
Length = 509
Score = 54.7 bits (130), Expect = 1e-06
Identities = 49/183 (26%), Positives = 85/183 (45%), Gaps = 27/183 (14%)
Query: 30 GYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILDHDEAMNQD 89
G ++ +++I A+ ERW ET++FHL +GEMTI LD++ +L + I G + + ++ +
Sbjct: 67 GPMSLNNSLISALVERWRRETNTFHLPLGEMTITLDEVALVLGLEIDGDPIVGSK-VDDE 125
Query: 90 CAIDLMIRLLGMSEVDVRVEVRSESAGHISYPTLKRVYEHHLTEAR-RLEEPQTREKL-- 146
A+D+ RLLG EV + + LKR + +A + + TR L
Sbjct: 126 VAMDMCGRLLGKLPSAANKEV---NCSRVKLNWLKRTFSECPEDASFDVVKCHTRAYLLY 182
Query: 147 --------QERGRKRAI------------GKWSWGGMALAYLYDYLDDSVILNNRTMAGS 186
G K ++ G+++WG ALA LY L ++ + + + G
Sbjct: 183 LIGSTIFATTDGDKVSVKYLPLFEDFDQAGRYAWGAAALACLYRALGNASLKSQSNICGC 242
Query: 187 TIL 189
L
Sbjct: 243 LTL 245
>UniRef100_Q9SK32 Expressed protein [Arabidopsis thaliana]
Length = 509
Score = 54.3 bits (129), Expect = 2e-06
Identities = 49/183 (26%), Positives = 84/183 (45%), Gaps = 27/183 (14%)
Query: 30 GYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILDHDEAMNQD 89
G ++ +++I A+ ERW ET++FHL +GEMTI LD++ +L + I G + + + +
Sbjct: 67 GPMSLNNSLISALVERWRRETNTFHLPLGEMTITLDEVALVLGLEIDGDPIVGSK-VGDE 125
Query: 90 CAIDLMIRLLGMSEVDVRVEVRSESAGHISYPTLKRVYEHHLTEAR-RLEEPQTREKL-- 146
A+D+ RLLG EV + + LKR + +A + + TR L
Sbjct: 126 VAMDMCGRLLGKLPSAANKEV---NCSRVKLNWLKRTFSECPEDASFDVVKCHTRAYLLY 182
Query: 147 --------QERGRKRAI------------GKWSWGGMALAYLYDYLDDSVILNNRTMAGS 186
G K ++ G+++WG ALA LY L ++ + + + G
Sbjct: 183 LIGSTIFATTDGDKVSVKYLPLFEDFDQAGRYAWGAAALACLYRALGNASLKSQSNICGC 242
Query: 187 TIL 189
L
Sbjct: 243 LTL 245
>UniRef100_Q94JI4 P0638D12.5 protein [Oryza sativa]
Length = 761
Score = 53.9 bits (128), Expect = 2e-06
Identities = 54/185 (29%), Positives = 74/185 (39%), Gaps = 45/185 (24%)
Query: 37 AMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGR----ILDHDEAMNQDCAI 92
A+I A+ ERW ET SFHL GEM + L D+ LL +PI GR DHD A
Sbjct: 149 ALITALVERWRPETHSFHLASGEMAVTLQDVAMLLALPIDGRPVCSTTDHDYA------- 201
Query: 93 DLMIRLLGMSEVDVRVEVRSESAGHISYPTLKRVYEHHLTEARRLEEPQTREK------- 145
++I LG + +S + Y LK+ H E + QT E+
Sbjct: 202 QMVIDCLGHDPRGPSMPGKS----FLHYKWLKK----HFYELPEGADDQTVERHVRAYIL 253
Query: 146 ------LQERGRKR-------------AIGKWSWGGMALAYLYDYLDDSVILNNRTMAGS 186
L G R +G +SWG LA+LY L +N G
Sbjct: 254 SLLCGVLFPDGTGRMSLIYLPLIADLSRVGTYSWGSAVLAFLYRSLCSVASSHNIKNIGG 313
Query: 187 TILFM 191
++L +
Sbjct: 314 SLLLL 318
>UniRef100_Q8LQS9 P0702H08.15 protein [Oryza sativa]
Length = 718
Score = 53.9 bits (128), Expect = 2e-06
Identities = 54/185 (29%), Positives = 74/185 (39%), Gaps = 45/185 (24%)
Query: 37 AMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGR----ILDHDEAMNQDCAI 92
A+I A+ ERW ET SFHL GEM + L D+ LL +PI GR DHD A
Sbjct: 163 ALITALVERWRPETHSFHLASGEMAVTLQDVAMLLALPIDGRPVCSTTDHDYA------- 215
Query: 93 DLMIRLLGMSEVDVRVEVRSESAGHISYPTLKRVYEHHLTEARRLEEPQTREK------- 145
++I LG + +S + Y LK+ H E + QT E+
Sbjct: 216 QMVIDCLGHDPRGPSMPGKS----FLHYKWLKK----HFYELPEGADDQTVERHVRAYIL 267
Query: 146 ------LQERGRKR-------------AIGKWSWGGMALAYLYDYLDDSVILNNRTMAGS 186
L G R +G +SWG LA+LY L +N G
Sbjct: 268 SLLCGVLFPDGTGRMSLIYLPLIADLSRVGTYSWGSAVLAFLYRSLCSVASSHNIKNIGG 327
Query: 187 TILFM 191
++L +
Sbjct: 328 SLLLL 332
>UniRef100_Q7XHQ1 Calcineurin-like phosphoesterase-like [Oryza sativa]
Length = 327
Score = 53.1 bits (126), Expect = 4e-06
Identities = 28/64 (43%), Positives = 40/64 (61%), Gaps = 3/64 (4%)
Query: 38 MICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILDHDEAMNQDCAIDLMIR 97
+I A+ ERW +T++FHL VGEMTI L D+ L +PIHGR + +D++ R
Sbjct: 70 LISALVERWRPKTNTFHLPVGEMTITLQDVSCLWGLPIHGRPI---TGQADGSWVDMIER 126
Query: 98 LLGM 101
LLG+
Sbjct: 127 LLGI 130
>UniRef100_Q69S26 Serine/threonine phosphatase PP7-related-like [Oryza sativa]
Length = 619
Score = 52.0 bits (123), Expect = 9e-06
Identities = 53/192 (27%), Positives = 86/192 (44%), Gaps = 25/192 (13%)
Query: 24 HDLIYTGYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILD-- 81
H + TG+ T+ +++ A CERW+ ET++ H + EM L D+ +L IP+ G ++
Sbjct: 62 HLALTTGF-TIDRSLLTAFCERWNNETNTAHFMGFEMAPSLRDVSYILGIPVTGHVVTAE 120
Query: 82 --HDEAMNQDCAIDL------------MIRLLGMSEVDVRVEVRSESAGHISYPTLKRVY 127
DEA+ + C L +IRL + ++ + + I+Y T R Y
Sbjct: 121 PIGDEAVRRMCLHFLGESPGNGEQLCGLIRLTWLYRKFHQLP-ENPTINEIAYST--RAY 177
Query: 128 EHHLTEARRLEEPQ----TREKLQERGRKRAIGKWSWGGMALAYLYDYLDDSVILNNRT- 182
+L + + + L R I +++WG ALA+LY L +V N T
Sbjct: 178 LLYLVGSTLFPDTMRGFVSPRYLPLLADFRKIREYAWGSAALAHLYRGLSVAVTPNATTQ 237
Query: 183 MAGSTILFMVII 194
GS L M I
Sbjct: 238 FLGSATLLMAWI 249
>UniRef100_Q9AS80 P0028E10.12 protein [Oryza sativa]
Length = 792
Score = 51.2 bits (121), Expect = 1e-05
Identities = 46/179 (25%), Positives = 73/179 (40%), Gaps = 20/179 (11%)
Query: 14 FWEPVEGFDLHDLIYTGYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHI 73
+ P L ++ G ++ A++ A+ +RW ET +FH+ GE+TI L D+ +L +
Sbjct: 237 YLRPAGLMGLANICRAGLPSIDRALVSALVDRWRPETHTFHMPCGEITITLQDVAMILGL 296
Query: 74 PIHGRILDHDEAMNQDCAIDLMIRLLGMSEVD-----------VRVEVRS--ESAGHISY 120
PI G + + Q+ +L+ R LG + VR E + E A +
Sbjct: 297 PIAGHAVTVNPTEPQN---ELVERYLGKAPPPDRPRPGLRVSWVRAEFNNCPEDADEETI 353
Query: 121 PTLKRVYEHHLTEARRLEEPQ----TREKLQERGRKRAIGKWSWGGMALAYLYDYLDDS 175
R Y L + T IG +SWG LAYLY + D+
Sbjct: 354 KQHARAYILSLISGLLFPDASGDLYTFYPFPLIADLENIGSYSWGSATLAYLYRAMCDA 412
>UniRef100_Q9LMT7 F2H15.15 protein [Arabidopsis thaliana]
Length = 478
Score = 50.1 bits (118), Expect = 3e-05
Identities = 44/162 (27%), Positives = 74/162 (45%), Gaps = 28/162 (17%)
Query: 30 GYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGR----ILDHDEA 85
G ++ +++I A+ ERW ET++FH GEMTI LD++ +L + + G+ + + DE
Sbjct: 58 GSISLNNSLISALVERWRRETNTFHFPCGEMTITLDEVSLILGLAVDGKPVVGVKEKDED 117
Query: 86 MNQDCAIDLMIRLLG------MSEVDVRVEVRSESAGHISYPTLKRVYEHHLTEARRLEE 139
+Q C +RLLG +S V + ES + E+H T A +
Sbjct: 118 PSQVC-----LRLLGKLPKGELSGNRVTAKWLKESFAECPKGATMKEIEYH-TRAYLIYI 171
Query: 140 PQTREKLQERGRKRAI------------GKWSWGGMALAYLY 169
+ K ++ G+++WG ALA+LY
Sbjct: 172 VGSTIFATTDPSKISVDYLILFEDFEKAGEYAWGAAALAFLY 213
>UniRef100_Q8LEM2 Hypothetical protein [Arabidopsis thaliana]
Length = 478
Score = 49.3 bits (116), Expect = 6e-05
Identities = 47/172 (27%), Positives = 78/172 (45%), Gaps = 30/172 (17%)
Query: 20 GFDLHDLIYTGYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGR- 78
GF LI G ++ +++I A+ ERW ET++FH GEMTI LD++ +L + + G+
Sbjct: 50 GFGWFRLI--GSISLNNSLISALVERWRRETNTFHFPCGEMTITLDEVSLILGLAVDGKP 107
Query: 79 ---ILDHDEAMNQDCAIDLMIRLLG------MSEVDVRVEVRSESAGHISYPTLKRVYEH 129
+ + DE +Q C ++LLG +S V + ES + E+
Sbjct: 108 VVGVKEKDEDPSQVC-----LKLLGKLPKGELSGNRVTAKWLKESFAECPKGATMKEIEY 162
Query: 130 HLTEARRLEEPQTREKLQERGRKRAI------------GKWSWGGMALAYLY 169
H T A + + K ++ G+++WG ALA+LY
Sbjct: 163 H-TRAYLIYIVGSTIFATTDPSKISVDYLILFEDFEKAGEYAWGAAALAFLY 213
>UniRef100_Q9LNG5 F21D18.16 [Arabidopsis thaliana]
Length = 1340
Score = 47.4 bits (111), Expect = 2e-04
Identities = 21/57 (36%), Positives = 35/57 (60%)
Query: 21 FDLHDLIYTGYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHG 77
F L+ + + + +A+I A+ ERW ET +FHL GE+T+ L D++ LL + + G
Sbjct: 67 FGLYGVYKVAFIQLDYALITALVERWRPETHTFHLPAGEITVTLQDVNILLGLRVDG 123
>UniRef100_Q7XTA1 OSJNBa0008A08.9 protein [Oryza sativa]
Length = 1560
Score = 47.0 bits (110), Expect = 3e-04
Identities = 20/49 (40%), Positives = 32/49 (64%)
Query: 37 AMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILDHDEA 85
A + A+ +RW ET +FHL GE+T+ L+D+ +L +PI G+ + D A
Sbjct: 875 AALTALVDRWRPETHTFHLPCGELTVTLEDVAMILGLPIRGQAVTSDTA 923
>UniRef100_Q8S0N3 Putative mutator-like transposase [Oryza sativa]
Length = 1556
Score = 46.6 bits (109), Expect = 4e-04
Identities = 20/49 (40%), Positives = 32/49 (64%)
Query: 37 AMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILDHDEA 85
A + A+ +RW ET +FHL GE+T+ L+D+ +L +PI G+ + D A
Sbjct: 964 AALTALVDRWRPETHTFHLPCGELTVTLEDVAMILGLPIRGQAVTGDTA 1012
>UniRef100_Q7XRQ6 OSJNBb0096E05.18 protein [Oryza sativa]
Length = 567
Score = 46.6 bits (109), Expect = 4e-04
Identities = 20/49 (40%), Positives = 32/49 (64%)
Query: 37 AMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILDHDEA 85
A + A+ +RW ET +FHL GE+T+ L+D+ +L +PI G+ + D A
Sbjct: 12 AALTALVDRWRPETHTFHLPCGELTVTLEDVAMILGLPIRGQAVTGDTA 60
>UniRef100_Q9LD66 Similar to maize transposon MuDR mudrA protein [Oryza sativa]
Length = 1230
Score = 46.6 bits (109), Expect = 4e-04
Identities = 20/49 (40%), Positives = 32/49 (64%)
Query: 37 AMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILDHDEA 85
A + A+ +RW ET +FHL GE+T+ L+D+ +L +PI G+ + D A
Sbjct: 574 AALTALVDRWRPETHTFHLPCGELTVTLEDVAMILGLPIRGQAVTGDTA 622
>UniRef100_Q60D76 Putative polyprotein [Oryza sativa]
Length = 1754
Score = 46.6 bits (109), Expect = 4e-04
Identities = 20/49 (40%), Positives = 32/49 (64%)
Query: 37 AMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILDHDEA 85
A + A+ +RW ET +FHL GE+T+ L+D+ +L +PI G+ + D A
Sbjct: 956 AALTALVDRWRPETHTFHLPCGELTVTLEDVAMILGLPIRGQAVTGDTA 1004
>UniRef100_Q84M64 Putative mutator-like transposase [Oryza sativa]
Length = 1204
Score = 46.6 bits (109), Expect = 4e-04
Identities = 20/49 (40%), Positives = 32/49 (64%)
Query: 37 AMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILDHDEA 85
A + A+ +RW ET +FHL GE+T+ L+D+ +L +PI G+ + D A
Sbjct: 833 AALTALVDRWRPETHTFHLPCGELTVTLEDVAMILGLPIRGQAVTGDTA 881
>UniRef100_Q7X975 Putative Mutator protein [Oryza sativa]
Length = 1456
Score = 46.6 bits (109), Expect = 4e-04
Identities = 20/49 (40%), Positives = 32/49 (64%)
Query: 37 AMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILDHDEA 85
A + A+ +RW ET +FHL GE+T+ L+D+ +L +PI G+ + D A
Sbjct: 895 AALTALVDRWRPETHTFHLPCGELTVTLEDVAMILGLPIRGQAVTGDTA 943
>UniRef100_Q6F2Q5 Hypothetical protein OSJNBa0088M06.4 [Oryza sativa]
Length = 1635
Score = 45.8 bits (107), Expect = 6e-04
Identities = 26/68 (38%), Positives = 38/68 (55%), Gaps = 1/68 (1%)
Query: 37 AMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILDHDEAMNQDCAIDLMI 96
A++ A+ +RW ET +FHL VGEM L D+ LL +PI G ++ +N A DL+
Sbjct: 928 ALLSALVDRWRPETHTFHLTVGEMVPTLQDVSYLLGLPIAGPVVG-PTMVNAGWADDLLA 986
Query: 97 RLLGMSEV 104
G+ V
Sbjct: 987 SFGGVLPV 994
>UniRef100_Q94HR1 Hypothetical protein OSJNBa0065C16.8 [Oryza sativa]
Length = 681
Score = 45.1 bits (105), Expect = 0.001
Identities = 19/42 (45%), Positives = 28/42 (66%)
Query: 39 ICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRIL 80
+ A+ +RW ET SFHL GEMTI L D+ +L +P+ G ++
Sbjct: 172 LTALVDRWRPETHSFHLPSGEMTITLQDVAMILALPLQGHVV 213
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.323 0.139 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 327,704,347
Number of Sequences: 2790947
Number of extensions: 12829692
Number of successful extensions: 29348
Number of sequences better than 10.0: 123
Number of HSP's better than 10.0 without gapping: 98
Number of HSP's successfully gapped in prelim test: 25
Number of HSP's that attempted gapping in prelim test: 29188
Number of HSP's gapped (non-prelim): 184
length of query: 194
length of database: 848,049,833
effective HSP length: 120
effective length of query: 74
effective length of database: 513,136,193
effective search space: 37972078282
effective search space used: 37972078282
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 71 (32.0 bits)
Medicago: description of AC149581.3


