

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149576.6 - phase: 0
(597 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
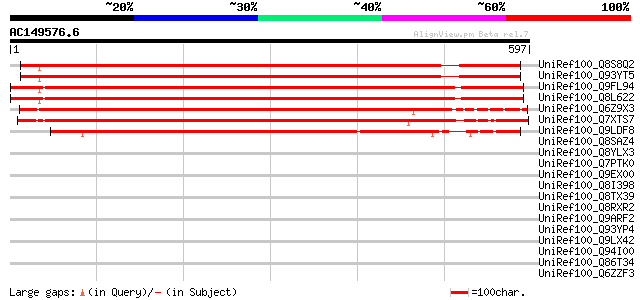
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8S8Q2 Expressed protein [Arabidopsis thaliana] 812 0.0
UniRef100_Q93YT5 Hypothetical protein At2g35155 [Arabidopsis tha... 810 0.0
UniRef100_Q9FL94 Gb|AAC61821.1 [Arabidopsis thaliana] 798 0.0
UniRef100_Q8L622 Hypothetical protein At5g45030 [Arabidopsis tha... 795 0.0
UniRef100_Q6Z9X3 Hypothetical protein P0453D01.16-1 [Oryza sativa] 791 0.0
UniRef100_Q7XTS7 OSJNBa0008M17.6 protein [Oryza sativa] 747 0.0
UniRef100_Q9LDF8 Gb|AAF35944.1 [Arabidopsis thaliana] 652 0.0
UniRef100_Q8SAZ4 Putative polyprotein [Oryza sativa] 40 0.15
UniRef100_Q8YLX3 Two-component hybrid sensor and regulator [Anab... 39 0.33
UniRef100_Q7PTK0 ENSANGP00000013418 [Anopheles gambiae str. PEST] 37 1.6
UniRef100_Q9EX00 Hypothetical protein SCO1126 [Streptomyces coel... 36 3.7
UniRef100_Q8I398 Hypothetical protein PFI0250c [Plasmodium falci... 35 4.8
UniRef100_Q8TX39 Uncharacterized protein conserved in archaea [M... 35 4.8
UniRef100_Q8RXR2 Chloride channel protein CLC-f [Arabidopsis tha... 35 4.8
UniRef100_Q9ARF2 Hypothetical protein [Capsella rubella] 35 6.3
UniRef100_Q93YP4 Epsin-like protein [Arabidopsis thaliana] 35 8.2
UniRef100_Q9LX42 Epsin-like protein [Arabidopsis thaliana] 35 8.2
UniRef100_Q94I00 Hypothetical protein OSJNBa0034E23.25 [Oryza sa... 35 8.2
UniRef100_Q86T34 Hypothetical protein DKFZp451A047 [Homo sapiens] 35 8.2
UniRef100_Q6ZZF3 Putative Ssy5 homologue [Pichia jadinii] 35 8.2
>UniRef100_Q8S8Q2 Expressed protein [Arabidopsis thaliana]
Length = 579
Score = 812 bits (2098), Expect = 0.0
Identities = 413/584 (70%), Positives = 467/584 (79%), Gaps = 29/584 (4%)
Query: 13 SGSTQSEESALDLERNYYGH-----PSSSPLHMQTFAVGVQHSEGNAAYFSWPTLNRWND 67
+ S++SE+SALDLERN++ + SSSP +Q F + +QH+E NA YFSWPTL+R ND
Sbjct: 14 AASSESEDSALDLERNHHCNHLSLPSSSSPSPLQPFTLNIQHAESNAPYFSWPTLSRLND 73
Query: 68 AAEDRANYFGNLQKGVLPETLGRLPSGQQATTLLELMTIRAFHSKILRRFSLGTAIGFRI 127
EDRANYFGNLQKGVLPET+GRLPSGQQATTLLELMTIRAFHSKILRRFSLGTA+GFRI
Sbjct: 74 TVEDRANYFGNLQKGVLPETVGRLPSGQQATTLLELMTIRAFHSKILRRFSLGTAVGFRI 133
Query: 128 RGGVLTDIPAILVFVAHKVHRQWLNHVQCLPAALEGPGGVWCDVDVVEFSYYGAPAPTPK 187
GVLT++PAILVFVA KVHRQWLN +QCLP+ALEGPGGVWCDVDVVEF YYGAPA TPK
Sbjct: 134 SRGVLTNVPAILVFVARKVHRQWLNPMQCLPSALEGPGGVWCDVDVVEFQYYGAPAATPK 193
Query: 188 EQLYTELADGLRGSDSCVGSGSQVASQETYGTLGAIVRSRTGNREVGFLTNRHVAVDLDY 247
EQ+Y EL DGLRGSD C+GSGSQVASQETYGTLGAIV+SRTGN +VGFLTNRHVAVDLDY
Sbjct: 194 EQVYNELVDGLRGSDPCIGSGSQVASQETYGTLGAIVKSRTGNHQVGFLTNRHVAVDLDY 253
Query: 248 PNQKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTNPETFVRADGAFIPFAEDF 307
P+QKMFHPLPPSLGPGVYLGAVERATSFITDD WYGIFAGTNPETFVRADGAFIPFAEDF
Sbjct: 254 PSQKMFHPLPPSLGPGVYLGAVERATSFITDDQWYGIFAGTNPETFVRADGAFIPFAEDF 313
Query: 308 NMNNVITSIRGVGDIGEVHRIDLQSPINSLIGRQVIKVGRSSGLTTGTIMAYALEYNDEK 367
N +NV T I+G+G+IG+VH IDLQSPI+SLIG+QV+KVGRSSG TTGTIMAYALEYNDEK
Sbjct: 314 NTSNVTTLIKGIGEIGDVHVIDLQSPIDSLIGKQVVKVGRSSGYTTGTIMAYALEYNDEK 373
Query: 368 GICFLTDFLVVGENQQTFDLEGDSGSLILLTGQNREKPRPVGIIWGGTANRGRLKLRVGQ 427
GICFLTDFLV+GENQQTFDLEGDSGSLILLTG N +KPRPVGIIWGGTANRGRLKL GQ
Sbjct: 374 GICFLTDFLVIGENQQTFDLEGDSGSLILLTGPNGQKPRPVGIIWGGTANRGRLKLIAGQ 433
Query: 428 PPENWTSGVDLGRLLDLLELDLVTTNETLQ--DSGQEQMNGSTAGIGSTVGESSP--TVP 483
PENWTSGVDLGRLLDLLELDL+T+N L+ + +E+ N S + STV +SSP VP
Sbjct: 434 EPENWTSGVDLGRLLDLLELDLITSNHELEAAAAAREERNTSVTALDSTVSQSSPPDPVP 493
Query: 484 IKEKLEESFEPFCLNMEHVPVEEPSTIVKPSLRPCEFHIRNEIETVPNVEHQFIRTSFAG 543
+K +ESFEPF P EFHI I+ VE +
Sbjct: 494 SGDKQDESFEPFI--------------------PPEFHIEEAIKPTLEVEEHIFIAPISV 533
Query: 544 KSPVHQSFLKEDMQFKSLSELRNEPDEDNFVSLHLGEPEAKRRK 587
+E + +L L+N +E+ +SLHLGEP+ K+ K
Sbjct: 534 NESTSAIKGQEIPKLDNLMALKNSSEEEVNISLHLGEPKLKKPK 577
>UniRef100_Q93YT5 Hypothetical protein At2g35155 [Arabidopsis thaliana]
Length = 579
Score = 810 bits (2091), Expect = 0.0
Identities = 412/584 (70%), Positives = 466/584 (79%), Gaps = 29/584 (4%)
Query: 13 SGSTQSEESALDLERNYYGH-----PSSSPLHMQTFAVGVQHSEGNAAYFSWPTLNRWND 67
+ S++SE+SALDLERN++ + SSSP +Q F + +QH+E NA YFSWPTL+R ND
Sbjct: 14 AASSESEDSALDLERNHHCNHLSLPSSSSPSPLQPFTLNIQHAESNAPYFSWPTLSRLND 73
Query: 68 AAEDRANYFGNLQKGVLPETLGRLPSGQQATTLLELMTIRAFHSKILRRFSLGTAIGFRI 127
EDRANYFGNLQKGVLPET+GRLPSGQQATTLLELMTIRAFHSKILRRFSLGTA+GFRI
Sbjct: 74 TVEDRANYFGNLQKGVLPETVGRLPSGQQATTLLELMTIRAFHSKILRRFSLGTAVGFRI 133
Query: 128 RGGVLTDIPAILVFVAHKVHRQWLNHVQCLPAALEGPGGVWCDVDVVEFSYYGAPAPTPK 187
GVLT++PAILVFVA KVHRQWLN +QCLP+ALEGPGGVWCDVDVVEF YYGAPA TPK
Sbjct: 134 SRGVLTNVPAILVFVARKVHRQWLNPMQCLPSALEGPGGVWCDVDVVEFQYYGAPAATPK 193
Query: 188 EQLYTELADGLRGSDSCVGSGSQVASQETYGTLGAIVRSRTGNREVGFLTNRHVAVDLDY 247
EQ+Y EL DGLRGSD C+GSGSQVASQETYGTLGAIV+SRTGN +VGFLTNRHVAVDLDY
Sbjct: 194 EQVYNELVDGLRGSDPCIGSGSQVASQETYGTLGAIVKSRTGNHQVGFLTNRHVAVDLDY 253
Query: 248 PNQKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTNPETFVRADGAFIPFAEDF 307
P+QKMFHPLPPSLGPGVYLGAVERATSFITDD WYGIFAGTNPETFVRADGAFIPFAED
Sbjct: 254 PSQKMFHPLPPSLGPGVYLGAVERATSFITDDQWYGIFAGTNPETFVRADGAFIPFAEDV 313
Query: 308 NMNNVITSIRGVGDIGEVHRIDLQSPINSLIGRQVIKVGRSSGLTTGTIMAYALEYNDEK 367
N +NV T I+G+G+IG+VH IDLQSPI+SLIG+QV+KVGRSSG TTGTIMAYALEYNDEK
Sbjct: 314 NTSNVTTLIKGIGEIGDVHVIDLQSPIDSLIGKQVVKVGRSSGYTTGTIMAYALEYNDEK 373
Query: 368 GICFLTDFLVVGENQQTFDLEGDSGSLILLTGQNREKPRPVGIIWGGTANRGRLKLRVGQ 427
GICFLTDFLV+GENQQTFDLEGDSGSLILLTG N +KPRPVGIIWGGTANRGRLKL GQ
Sbjct: 374 GICFLTDFLVIGENQQTFDLEGDSGSLILLTGPNGQKPRPVGIIWGGTANRGRLKLIAGQ 433
Query: 428 PPENWTSGVDLGRLLDLLELDLVTTNETLQ--DSGQEQMNGSTAGIGSTVGESSP--TVP 483
PENWTSGVDLGRLLDLLELDL+T+N L+ + +E+ N S + STV +SSP VP
Sbjct: 434 EPENWTSGVDLGRLLDLLELDLITSNHELEAAAAAREERNTSVTALDSTVSQSSPPDPVP 493
Query: 484 IKEKLEESFEPFCLNMEHVPVEEPSTIVKPSLRPCEFHIRNEIETVPNVEHQFIRTSFAG 543
+K +ESFEPF P EFHI I+ VE +
Sbjct: 494 SGDKQDESFEPFI--------------------PPEFHIEEAIKPTLEVEEHIFIAPISV 533
Query: 544 KSPVHQSFLKEDMQFKSLSELRNEPDEDNFVSLHLGEPEAKRRK 587
+E + +L L+N +E+ +SLHLGEP+ K+ K
Sbjct: 534 NESTSAIKGQEIPKLDNLMALKNSSEEEVNISLHLGEPKLKKPK 577
>UniRef100_Q9FL94 Gb|AAC61821.1 [Arabidopsis thaliana]
Length = 607
Score = 798 bits (2060), Expect = 0.0
Identities = 424/605 (70%), Positives = 486/605 (80%), Gaps = 21/605 (3%)
Query: 1 MNRNRLGLSAHHSGSTQSEESA-LDLERNYYGH---PSSSPLHMQTFAVGVQHSEGNAA- 55
M RL L HHS S+QS ESA LDL++N Y H SSSPL Q F G QH E +AA
Sbjct: 1 MEGKRLDLRFHHSTSSQSVESAALDLDKNVYNHIKLASSSPL--QPFPSGAQHPETSAAA 58
Query: 56 -YFSWPTLNRWNDAAEDRANYFGNLQKGVLPETLGRLPSGQQATTLLELMTIRAFHSKIL 114
YFSWPT +R ND+AEDRANYF NLQKGVLPE+ LP+G++ATTLLELM IRAFHSK L
Sbjct: 59 AYFSWPTSSRLNDSAEDRANYFANLQKGVLPESFDGLPTGKKATTLLELMMIRAFHSKNL 118
Query: 115 RRFSLGTAIGFRIRGGVLTDIPAILVFVAHKVHRQWLNHVQCLPAALEGPGGVWCDVDVV 174
RRFSLGTAIGFRIR GVLT+I AILVFVA KVH+QWLN +QCLP ALEGPGGVWCDVDVV
Sbjct: 119 RRFSLGTAIGFRIRRGVLTNIAAILVFVARKVHKQWLNPLQCLPTALEGPGGVWCDVDVV 178
Query: 175 EFSYYGAPAPTPKEQLYTELADGLRGSDSCVGSGSQVASQETYGTLGAIVRSRTGNREVG 234
EF YYGAPA TPKEQ+YTEL D LRGS S +GSGSQVASQETYGTLGAIV+S+TG R+VG
Sbjct: 179 EFQYYGAPAQTPKEQVYTELVDDLRGSGSSIGSGSQVASQETYGTLGAIVKSKTGIRQVG 238
Query: 235 FLTNRHVAVDLDYPNQKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTNPETFV 294
FLTNRHVAVDLDYP+QKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTNPETFV
Sbjct: 239 FLTNRHVAVDLDYPSQKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTNPETFV 298
Query: 295 RADGAFIPFAEDFNMNNVITSIRGVGDIGEVHRIDLQSPINSLIGRQVIKVGRSSGLTTG 354
RADGAFIPFAEDFN NNV T+++G+G+IG++H DLQSP+NSLIGR+V+KVGRSSGLTTG
Sbjct: 299 RADGAFIPFAEDFNTNNVTTTVKGIGEIGDIHATDLQSPVNSLIGRKVVKVGRSSGLTTG 358
Query: 355 TIMAYALEYNDEKGICFLTDFLVVGENQQTFDLEGDSGSLILLTG--QNREKPRPVGIIW 412
TIMAYALEYNDEKGICFLTDFLVVGENQQTFDLEGDSGSLILL + EKPRPVGIIW
Sbjct: 359 TIMAYALEYNDEKGICFLTDFLVVGENQQTFDLEGDSGSLILLAAGDEKNEKPRPVGIIW 418
Query: 413 GGTANRGRLKLRVGQPPENWTSGVDLGRLLDLLELDLVTTNETLQDSGQEQMNG-STAGI 471
GGTANRGRLKL+VG+ PENWTSGVDLGR+L+LLELDL+T+NE LQ + EQ NG A +
Sbjct: 419 GGTANRGRLKLKVGEQPENWTSGVDLGRVLNLLELDLITSNEGLQAAVLEQRNGIMCAAV 478
Query: 472 GSTVGESSPTV--PIKEKLEESFEPFCLNMEHVPVEEPSTIVKPSLRPCEFHIRNEIETV 529
STV ESSP V + K E+FEP LN++ V +E+ ++ + P EF I + +E+V
Sbjct: 479 DSTVVESSPGVCNISRCKTGENFEPINLNVQQVLIEDDNSNIHP-----EFQIEDVLESV 533
Query: 530 PNV-EHQFIRTSFAGKSPVHQS-FLKEDMQFKSLSELRNEPDEDNF-VSLHLGEPEAKRR 586
+ EHQFI +S S +HQ E+++ K+LS L+ D SL LGE + K+R
Sbjct: 534 AVIEEHQFIPSSSNNGSALHQKPNGPENLESKNLSSLKTSSSGDEIGFSLQLGESDTKKR 593
Query: 587 KHSNS 591
K ++S
Sbjct: 594 KRTDS 598
>UniRef100_Q8L622 Hypothetical protein At5g45030 [Arabidopsis thaliana]
Length = 607
Score = 795 bits (2054), Expect = 0.0
Identities = 423/605 (69%), Positives = 485/605 (79%), Gaps = 21/605 (3%)
Query: 1 MNRNRLGLSAHHSGSTQSEESA-LDLERNYYGH---PSSSPLHMQTFAVGVQHSEGNAA- 55
M RL L HHS S+QS ESA LDL++N Y H SSSPL Q F G QH E +AA
Sbjct: 1 MEGKRLDLRFHHSTSSQSVESAALDLDKNVYNHIKLASSSPL--QPFPSGAQHPETSAAA 58
Query: 56 -YFSWPTLNRWNDAAEDRANYFGNLQKGVLPETLGRLPSGQQATTLLELMTIRAFHSKIL 114
YFSWPT +R ND+AEDRANYF NLQKGVLPE+ LP+G++ATTLLELM IRAFHSK L
Sbjct: 59 AYFSWPTSSRLNDSAEDRANYFANLQKGVLPESFDGLPTGKKATTLLELMMIRAFHSKNL 118
Query: 115 RRFSLGTAIGFRIRGGVLTDIPAILVFVAHKVHRQWLNHVQCLPAALEGPGGVWCDVDVV 174
RRFSLGTAIGFRIR GVLT+I AILVFVA KVH+QWLN +QCLP ALEGPGGVWCDVDVV
Sbjct: 119 RRFSLGTAIGFRIRRGVLTNIAAILVFVARKVHKQWLNPLQCLPTALEGPGGVWCDVDVV 178
Query: 175 EFSYYGAPAPTPKEQLYTELADGLRGSDSCVGSGSQVASQETYGTLGAIVRSRTGNREVG 234
EF YYGAPA TPKEQ+YTEL D LRGS S +GSGSQVASQE YGTLGAIV+S+TG R+VG
Sbjct: 179 EFQYYGAPAQTPKEQVYTELVDDLRGSGSSIGSGSQVASQERYGTLGAIVKSKTGIRQVG 238
Query: 235 FLTNRHVAVDLDYPNQKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTNPETFV 294
FLTNRHVAVDLDYP+QKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTNPETFV
Sbjct: 239 FLTNRHVAVDLDYPSQKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTNPETFV 298
Query: 295 RADGAFIPFAEDFNMNNVITSIRGVGDIGEVHRIDLQSPINSLIGRQVIKVGRSSGLTTG 354
RADGAFIPFAEDFN NNV T+++G+G+IG++H DLQSP+NSLIGR+V+KVGRSSGLTTG
Sbjct: 299 RADGAFIPFAEDFNTNNVTTTVKGIGEIGDIHATDLQSPVNSLIGRKVVKVGRSSGLTTG 358
Query: 355 TIMAYALEYNDEKGICFLTDFLVVGENQQTFDLEGDSGSLILLTG--QNREKPRPVGIIW 412
TIMAYALEYNDEKGICFLTDFLVVGENQQTFDLEGDSGSLILL + EKPRPVGIIW
Sbjct: 359 TIMAYALEYNDEKGICFLTDFLVVGENQQTFDLEGDSGSLILLAAGDEKNEKPRPVGIIW 418
Query: 413 GGTANRGRLKLRVGQPPENWTSGVDLGRLLDLLELDLVTTNETLQDSGQEQMNG-STAGI 471
GGTANRGRLKL+VG+ PENWTSGVDLGR+L+LLELDL+T+NE LQ + EQ NG A +
Sbjct: 419 GGTANRGRLKLKVGEQPENWTSGVDLGRVLNLLELDLITSNEGLQAAVLEQRNGIMCAAV 478
Query: 472 GSTVGESSPTV--PIKEKLEESFEPFCLNMEHVPVEEPSTIVKPSLRPCEFHIRNEIETV 529
STV ESSP V + K E+FEP LN++ V +E+ ++ + P EF I + +E+V
Sbjct: 479 DSTVVESSPGVCNISRCKTGENFEPINLNVQQVLIEDDNSNIHP-----EFQIEDVLESV 533
Query: 530 PNV-EHQFIRTSFAGKSPVHQS-FLKEDMQFKSLSELRNEPDEDNF-VSLHLGEPEAKRR 586
+ EHQFI +S S +HQ E+++ K+LS L+ D SL LGE + K+R
Sbjct: 534 AVIEEHQFIPSSSNNGSALHQKPNGPENLESKNLSSLKTSSSGDEIGFSLQLGESDTKKR 593
Query: 587 KHSNS 591
K ++S
Sbjct: 594 KRTDS 598
>UniRef100_Q6Z9X3 Hypothetical protein P0453D01.16-1 [Oryza sativa]
Length = 590
Score = 791 bits (2043), Expect = 0.0
Identities = 411/589 (69%), Positives = 478/589 (80%), Gaps = 17/589 (2%)
Query: 12 HSGSTQSEESALDLERNYYGHPSSSPLHMQTFAVGVQHSEGNAAYFSWPTLNRWNDAAED 71
H+GS+QSE SALD+ERN H + P +Q A G QHSE +AAYFSWPT + +AE
Sbjct: 10 HAGSSQSEGSALDMERNGCNH-NCCPSPLQPIASGGQHSESSAAYFSWPTSTLMHGSAEG 68
Query: 72 RANYFGNLQKGVLPETLGRLPSGQQATTLLELMTIRAFHSKILRRFSLGTAIGFRIRGGV 131
RANYFGNLQKGVLP LGRLP+GQ+ATTLL+LM IRAFHSKILRRFSLGTAIGFRI+ G
Sbjct: 69 RANYFGNLQKGVLPGHLGRLPTGQRATTLLDLMIIRAFHSKILRRFSLGTAIGFRIKKGT 128
Query: 132 LTDIPAILVFVAHKVHRQWLNHVQCLPAALEGPGGVWCDVDVVEFSYYGAPAPTPKEQLY 191
LTD PAILVFVA KVHR+WL+ QCLPA LEGPGGVWCDVDVVEFSYYGAPAPTPKEQLY
Sbjct: 129 LTDTPAILVFVARKVHRKWLSPTQCLPAHLEGPGGVWCDVDVVEFSYYGAPAPTPKEQLY 188
Query: 192 TELADGLRGSDSCVGSGSQVASQETYGTLGAIVRSRTGNREVGFLTNRHVAVDLDYPNQK 251
EL DGLRGSD +GSGSQVAS ETYGTLGAIV+SRTGN++VGFLTNRHVAVDLDYPNQK
Sbjct: 189 DELVDGLRGSDPSIGSGSQVASLETYGTLGAIVKSRTGNKQVGFLTNRHVAVDLDYPNQK 248
Query: 252 MFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTNPETFVRADGAFIPFAEDFNMNN 311
MFHPLPP+LGPGVYLGAVERATSFITDD+WYGI+AGTNPETFVRADGAFIPFA+D+++ +
Sbjct: 249 MFHPLPPNLGPGVYLGAVERATSFITDDVWYGIYAGTNPETFVRADGAFIPFADDYDITS 308
Query: 312 VITSIRGVGDIGEVHRIDLQSPINSLIGRQVIKVGRSSGLTTGTIMAYALEYNDEKGICF 371
V TS++GVG IG+V IDLQSPI+SLIGRQV+KVGRSSGLTTGT++AYALEYNDEKGICF
Sbjct: 309 VNTSVKGVGVIGDVKAIDLQSPISSLIGRQVVKVGRSSGLTTGTVVAYALEYNDEKGICF 368
Query: 372 LTDFLVVGENQQTFDLEGDSGSLILLTGQNREKPRPVGIIWGGTANRGRLKLRVGQPPEN 431
TDFLVVGENQQTFDLEGDSGSLI+LTG++ EKP+P+GIIWGGTANRGRLKL+ GQ PEN
Sbjct: 369 FTDFLVVGENQQTFDLEGDSGSLIILTGKDGEKPQPIGIIWGGTANRGRLKLKSGQGPEN 428
Query: 432 WTSGVDLGRLLDLLELDLVTTNETLQDSGQEQ---MNGSTAGIGSTVGESSPTV--PIKE 486
WTSGVDLGRLLDLLELDL+TT+E LQ++ +EQ + + A ST GESSP E
Sbjct: 429 WTSGVDLGRLLDLLELDLITTSEGLQEALEEQRIILAAAAAAANSTAGESSPVAGPQENE 488
Query: 487 KLEESFEPFCLNMEHVPVEEPSTIVKPSLRPCEFHIRNEIETVPNVEHQFIRTSFAGKSP 546
K+++ +EP +N++ +P + +T S P EFH+ + +E V NVE R G SP
Sbjct: 489 KVDKIYEPLGINIQQLPRDNSAT----STGPDEFHV-DTVEGVTNVEE---RQFLIGMSP 540
Query: 547 VHQSFLKEDMQFKSLSELRNEPDEDNFVSLHLGEPEAKRRKHSNSSLSL 595
+ + + +L+EL N P ED SLHLGE E KR + S+SSL +
Sbjct: 541 AREG-QEANGDLNNLAELENSP-EDICFSLHLGEREPKRLR-SDSSLDI 586
>UniRef100_Q7XTS7 OSJNBa0008M17.6 protein [Oryza sativa]
Length = 588
Score = 747 bits (1929), Expect = 0.0
Identities = 396/597 (66%), Positives = 458/597 (76%), Gaps = 27/597 (4%)
Query: 10 AHHSGSTQSEESALDLERNYYGHPSSSPLHMQTFAVGVQHSEGNAAYFSWPTLNRWNDAA 69
A SG QSEES+LD++ + P S + Q A G H+E +AAYF WPT N + AA
Sbjct: 8 AQLSGLAQSEESSLDVDHQSF--PCSPSI--QPVASGCTHTENSAAYFLWPTSNLQHCAA 63
Query: 70 EDRANYFGNLQKGVLPETLGRLPSGQQATTLLELMTIRAFHSKILRRFSLGTAIGFRIRG 129
E RANYFGNLQKG+LP GRLP GQQA +LL+LMTIRAFHSKILRRFSLGTA+GFRIR
Sbjct: 64 EGRANYFGNLQKGLLPRHPGRLPKGQQANSLLDLMTIRAFHSKILRRFSLGTAVGFRIRK 123
Query: 130 GVLTDIPAILVFVAHKVHRQWLNHVQCLPAALEGPGGVWCDVDVVEFSYYGAPAPTPKEQ 189
G LTDIPAILVFVA KVH++WLN QCLPA LEGPGGVWCDVDVVEFSYYGAPA TPKEQ
Sbjct: 124 GDLTDIPAILVFVARKVHKKWLNPAQCLPAILEGPGGVWCDVDVVEFSYYGAPAQTPKEQ 183
Query: 190 LYTELADGLRGSDSCVGSGSQVASQETYGTLGAIVRSRTGNREVGFLTNRHVAVDLDYPN 249
+++EL D L GSD C+GSGSQVAS ET+GTLGAIV+ RTGN++VGFLTN HVAVDLDYPN
Sbjct: 184 MFSELVDKLCGSDECIGSGSQVASHETFGTLGAIVKRRTGNKQVGFLTNHHVAVDLDYPN 243
Query: 250 QKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTNPETFVRADGAFIPFAEDFNM 309
QKMFHPLPP+LGPGVYLGAVERATSFITDD+WYGI+AGTNPETFVRADGAFIPFA+DF++
Sbjct: 244 QKMFHPLPPNLGPGVYLGAVERATSFITDDVWYGIYAGTNPETFVRADGAFIPFADDFDI 303
Query: 310 NNVITSIRGVGDIGEVHRIDLQSPINSLIGRQVIKVGRSSGLTTGTIMAYALEYNDEKGI 369
+ V T +RGVGDIG+V IDLQ P+NSLIGRQV KVGRSSG TTGT+MAYALEYNDEKGI
Sbjct: 304 STVTTVVRGVGDIGDVKVIDLQCPLNSLIGRQVCKVGRSSGHTTGTVMAYALEYNDEKGI 363
Query: 370 CFLTDFLVVGENQQTFDLEGDSGSLILLTGQNREKPRPVGIIWGGTANRGRLKLRVGQPP 429
CF TD LVVGEN+QTFDLEGDSGSLI+LT Q+ EKPRP+GIIWGGTANRGRLKL P
Sbjct: 364 CFFTDILVVGENRQTFDLEGDSGSLIILTSQDGEKPRPIGIIWGGTANRGRLKLTSDHGP 423
Query: 430 ENWTSGVDLGRLLDLLELDLVTTNETLQ------DSGQEQMNGSTAGIGSTVGESS--PT 481
ENWTSGVDLGRLLD LELD++ TNE+LQ D+ Q+Q A + S VGESS P
Sbjct: 424 ENWTSGVDLGRLLDRLELDIIITNESLQEFAYYKDAVQQQRFALVAAVTSAVGESSGVPV 483
Query: 482 VPIKEKLEESFEPFCLNMEHVPVEEPSTIVKPSLRPCEFHIRNEIETVPNV-EHQFIRTS 540
+EK+EE FEP + ++ +P + + TV NV EHQFI ++
Sbjct: 484 AIPEEKIEEIFEPLGIQIQQLPRHDVAASGTEG--------EEASNTVVNVEEHQFI-SN 534
Query: 541 FAGKSPVHQSFLKEDMQFKSLSELRNEPDEDNFVSLHLGEPEAKR-RKHSNSSLSLK 596
F G SPV +D +S++ L N +E+ +SLHLG+ E KR R S SSL L+
Sbjct: 535 FVGMSPVRDD---QDAP-RSITNLNNPSEEELAMSLHLGDREPKRLRSDSGSSLDLE 587
>UniRef100_Q9LDF8 Gb|AAF35944.1 [Arabidopsis thaliana]
Length = 558
Score = 652 bits (1683), Expect = 0.0
Identities = 353/564 (62%), Positives = 429/564 (75%), Gaps = 50/564 (8%)
Query: 48 QHSEGNAA-YFSWPTLNRWNDAAEDRANYFGNLQKG------VLPETLGRLPSGQQATTL 100
QH E AA YFSWPT +R ++AAE+RANYF NLQK V PE + P GQ+ATTL
Sbjct: 9 QHCEFTAASYFSWPTSSRLSNAAEERANYFSNLQKEEDDDDEVSPEPVSTEPKGQRATTL 68
Query: 101 LELMTIRAFHSKILRRFSLGTAIGFRIRGGVLTDIPAILVFVAHKVHRQWLNHVQCLPAA 160
LELMTIRAFHSK+LR +SLGTAIGFRIR GVLTDIPAI+VFV+ KVH+QWL+ +QCLP A
Sbjct: 69 LELMTIRAFHSKMLRCYSLGTAIGFRIRRGVLTDIPAIIVFVSRKVHKQWLSPLQCLPTA 128
Query: 161 LEGPGGVWCDVDVVEFSYYGAP--APTPKEQLYTELADGLRGSDSCVGSGSQVASQETYG 218
LEG GG+WCDVDVVEFSY+G P PTPK+ T++ D L+GSD +GSGSQVASQET G
Sbjct: 129 LEGAGGIWCDVDVVEFSYFGEPDHQPTPKQTFTTDIVDHLQGSDPFIGSGSQVASQETCG 188
Query: 219 TLGAIVRSRTGNREVGFLTNRHVAVDLDYPNQKMFHPLPPSLGPGVYLGAVERATSFITD 278
TLGAIVRS+TG R+VGF+TNRHVAV+LDYP+QKMFHPLPP+LGPGVYLGAVERATSFITD
Sbjct: 189 TLGAIVRSQTGGRQVGFVTNRHVAVNLDYPSQKMFHPLPPALGPGVYLGAVERATSFITD 248
Query: 279 DLWYGIFAGTNPETFVRADGAFIPFAEDFNMNNVITSIR-GVGDIGEVHRIDLQSPINSL 337
DLW+GIFAGTNPETFVRADGAFIPFA+D++++ V TS++ GVG+IGEV I+LQSP+ SL
Sbjct: 249 DLWFGIFAGTNPETFVRADGAFIPFADDYDLSRVTTSVKGGVGEIGEVKAIELQSPVGSL 308
Query: 338 IGRQVIKVGRSSGLTTGTIMAYALEYNDEKGICFLTDFLVVGENQQT-FDLEGDSGSLIL 396
+G+QV+KVGRSSGLTTGT++AYALEYNDE+G+CFLTDFLVVGEN ++ FDLEGDSGSLI+
Sbjct: 309 VGKQVVKVGRSSGLTTGTVLAYALEYNDERGVCFLTDFLVVGENHRSPFDLEGDSGSLIV 368
Query: 397 LTGQNREKPRPVGIIWGGTANRGRLKLRVGQPPENWTSGVDLGRLLDLLELDLVTTNETL 456
+ G+ EK RP+GIIWGGT +RGRLKL+VG+ PE+WT+GVDLGRLL L+LDL+TT+E L
Sbjct: 369 MKGE--EKARPIGIIWGGTGSRGRLKLKVGECPESWTTGVDLGRLLTHLQLDLITTDEGL 426
Query: 457 QDSGQEQMNGSTAGIGSTVGESSPT-VPIK-------EKLEESFEPFCLNMEHVPVEEPS 508
+ + QEQ ST G+ S V +SSP V +K EKLE S P L ++H+ +EE
Sbjct: 427 KAAVQEQRAASTTGMSSMVADSSPPYVNLKKEKRSPEEKLEASLGP--LQVQHIDLEE-- 482
Query: 509 TIVKPSLRPCEFHIRNEIET---VPNVEHQFIRTSFAGK--SPVHQSFLKEDMQFKSLSE 563
IET P+VEHQF+ T F+G+ + +ED+
Sbjct: 483 ----------------RIETKGGAPSVEHQFMPT-FSGQCSASAWPETAREDL---VAGF 522
Query: 564 LRNEPDEDNFVSLHLGEPEAKRRK 587
D D V L LG+ AKRR+
Sbjct: 523 TNGSCDGDLCVGLRLGDDGAKRRR 546
>UniRef100_Q8SAZ4 Putative polyprotein [Oryza sativa]
Length = 1819
Score = 40.4 bits (93), Expect = 0.15
Identities = 35/104 (33%), Positives = 42/104 (39%), Gaps = 26/104 (25%)
Query: 181 APAPTPKEQLYTELADGLRGSDSCVG---------------SGSQVASQETY-------- 217
APAP P L A +RG S +G SGSQ S ETY
Sbjct: 231 APAPAPAFVLSFAPASAVRGPASGIGGWLPATPSGSRDRSSSGSQTGSSETYLPGSRSRS 290
Query: 218 ---GTLGAIVRSRTGNREVGFLTNRHVAVDLDYPNQKMFHPLPP 258
G L +V +RTGNR G +N VD +PN + PP
Sbjct: 291 EGPGCLFGMVNTRTGNRTTGEGSNGEERVDGMHPNGDPGNGPPP 334
>UniRef100_Q8YLX3 Two-component hybrid sensor and regulator [Anabaena sp.]
Length = 965
Score = 39.3 bits (90), Expect = 0.33
Identities = 26/106 (24%), Positives = 48/106 (44%), Gaps = 6/106 (5%)
Query: 447 LDLVTTNETLQDSGQEQMNGSTAGIGSTVGESSPTVPIKEKLEESFEPFCLNMEHVPVEE 506
L L LQ G + ST+G GST + P P ++ EE F L++ H ++
Sbjct: 658 LGLAICRSILQHHGGQIWAESTSGEGSTFSFTLPIFPEEQNKEEEF----LHLSHAAPQQ 713
Query: 507 PSTIVKPSLRPC--EFHIRNEIETVPNVEHQFIRTSFAGKSPVHQS 550
P+ + P + C + +R ++ + + + T +GK V ++
Sbjct: 714 PTESISPLILHCDDDLSVRAVVQVILEKQGYRVMTVASGKEAVEKA 759
>UniRef100_Q7PTK0 ENSANGP00000013418 [Anopheles gambiae str. PEST]
Length = 1700
Score = 37.0 bits (84), Expect = 1.6
Identities = 26/84 (30%), Positives = 39/84 (45%), Gaps = 1/84 (1%)
Query: 450 VTTNETLQDSGQEQMNGSTAGIGSTVGESSPTVPIKEKLEESFEPFCLNMEHVPVEEPST 509
V+ +T++ SG E +A ES P + E+ E+SF+ C+ +V VE PS
Sbjct: 979 VSETDTVRSSGFEYRKLVSAANAEDEDESLPDDELGERSEKSFQLPCIPTINVVVEPPSP 1038
Query: 510 IVKPSLRPCEF-HIRNEIETVPNV 532
+ LR IR E PN+
Sbjct: 1039 AISDELRSLRVPDIRRHSEHTPNL 1062
>UniRef100_Q9EX00 Hypothetical protein SCO1126 [Streptomyces coelicolor]
Length = 384
Score = 35.8 bits (81), Expect = 3.7
Identities = 32/125 (25%), Positives = 49/125 (38%), Gaps = 16/125 (12%)
Query: 89 GRLPSGQQATTL----LELMTIRAFHSKILRRFSLGTAIGFRIRGGVLTDIPAILVFVAH 144
GR PSG L L + R ++ F LG + +R L +P++++ +
Sbjct: 91 GRRPSGTPLARLAGDRLRELAARTGADVVVPVFHLGAQLTGHLRDRGLLPVPSVVLVIDF 150
Query: 145 KVHRQWL----NHVQCL--PAALEGPGGVWCDVDVV------EFSYYGAPAPTPKEQLYT 192
++HRQWL +H CL AA E G + EFS P + +
Sbjct: 151 ELHRQWLHPGNDHCLCLTEEAAREARGNTGTPAETCGPVVAPEFSAGRVPGAAQWRETFD 210
Query: 193 ELADG 197
L G
Sbjct: 211 RLGPG 215
>UniRef100_Q8I398 Hypothetical protein PFI0250c [Plasmodium falciparum]
Length = 2112
Score = 35.4 bits (80), Expect = 4.8
Identities = 32/158 (20%), Positives = 66/158 (41%), Gaps = 9/158 (5%)
Query: 330 LQSPINSLIGRQVIKVGRSSGLTTGTIMAYALEYNDEKGICFLTDFLVVGENQQTFDLEG 389
+Q P N+L+ + + +G SSG +TG ++ Y++ L + G + + +G
Sbjct: 76 IQQPNNALVNKSIFNLGGSSGTSTGLSGGKSI-YDNMNSQSNLNTKNIFGSTNVSNNTQG 134
Query: 390 DSGSLILLTGQNREKPRPVGIIWGG-------TANRGRLKLRVGQPPENWTS-GVDLGRL 441
+ G L N + + I+G T+N G + G P N T+ ++
Sbjct: 135 NMGGNSLFMNANNQNNLNMKNIFGSSSGLNNQTSNLGNKSIFGGLQPSNQTTPSNNIFGN 194
Query: 442 LDLLELDLVTTNETLQDSGQEQMNGSTAGIGSTVGESS 479
+ + + L + Q + N G+G++ +S+
Sbjct: 195 MSSNQTNSSNIFGNLSSTSQNKSNSIFGGLGTSTNQST 232
>UniRef100_Q8TX39 Uncharacterized protein conserved in archaea [Methanopyrus
kandleri]
Length = 288
Score = 35.4 bits (80), Expect = 4.8
Identities = 34/104 (32%), Positives = 43/104 (40%), Gaps = 9/104 (8%)
Query: 147 HRQWLNHVQCLPAALEGPGGVWCDVDVVEFSYYGAPAPTPKEQLYTELADGL---RGSDS 203
H+Q + L ALE CDV V YY AP P +++ EL + L RG D
Sbjct: 114 HKQMILEEFGLLHALEPVSEAGCDVGNVP-GYYAAPVPESADRVARELREELRRRRGVDV 172
Query: 204 CVGSGSQVASQETYGTLGAIVRSR-----TGNREVGFLTNRHVA 242
V G + E G L V TG +GFL R +A
Sbjct: 173 SVIVGDTDKTYEILGILFTTVARAHPDIVTGTGVIGFLVGRLLA 216
>UniRef100_Q8RXR2 Chloride channel protein CLC-f [Arabidopsis thaliana]
Length = 781
Score = 35.4 bits (80), Expect = 4.8
Identities = 28/101 (27%), Positives = 42/101 (40%), Gaps = 20/101 (19%)
Query: 412 WGGTANRGRLKLRVGQPPENW--------TSGVDLGRLLDLLE-LDLVTTNETLQDSGQE 462
W GT N G LR+ + + W T GV +G + LLE LD + + + Q G +
Sbjct: 161 WAGTPNEGAAWLRLQRLADTWHRILLIPVTGGVIVGMMHGLLEILDQIRQSNSSQRQGLD 220
Query: 463 QMNG-----------STAGIGSTVGESSPTVPIKEKLEESF 492
+ G T G G ++G P+V I + F
Sbjct: 221 FLAGIYPVIKAIQAAVTLGTGCSLGPEGPSVDIGKSCANGF 261
>UniRef100_Q9ARF2 Hypothetical protein [Capsella rubella]
Length = 780
Score = 35.0 bits (79), Expect = 6.3
Identities = 28/101 (27%), Positives = 42/101 (40%), Gaps = 20/101 (19%)
Query: 412 WGGTANRGRLKLRVGQPPENW--------TSGVDLGRLLDLLE-LDLVTTNETLQDSGQE 462
W GT N G LR+ + + W T GV +G + LLE LD + + + Q G +
Sbjct: 160 WAGTPNEGAAWLRLQRLADTWHRILLIPVTGGVIVGMMHGLLEILDQIRQSTSSQRQGLD 219
Query: 463 QMNG-----------STAGIGSTVGESSPTVPIKEKLEESF 492
+ G T G G ++G P+V I + F
Sbjct: 220 FLAGIYPVIKAIQAAVTLGTGCSLGPEGPSVDIGKSCANGF 260
>UniRef100_Q93YP4 Epsin-like protein [Arabidopsis thaliana]
Length = 1024
Score = 34.7 bits (78), Expect = 8.2
Identities = 33/112 (29%), Positives = 44/112 (38%), Gaps = 23/112 (20%)
Query: 393 SLILLTG------QNREKPRPVGIIWGGTANRGRLKLRVGQPPENWTS--GVD---LGRL 441
SL LTG QN++K P IW T +RG + + P N + GVD + R
Sbjct: 805 SLTPLTGAIEIVPQNQKKFEPKSTIWADTLSRGLVNFNISGPKTNPLADIGVDFEAINRK 864
Query: 442 LDLLELDLVTTNETLQDSGQEQMNGSTAGIG------------STVGESSPT 481
LE +T + + GS G+G S VG S PT
Sbjct: 865 EKRLEKPTITQQQVTSTINMGKAMGSGTGLGRAGAGAMRPPTNSMVGSSMPT 916
>UniRef100_Q9LX42 Epsin-like protein [Arabidopsis thaliana]
Length = 1023
Score = 34.7 bits (78), Expect = 8.2
Identities = 33/112 (29%), Positives = 44/112 (38%), Gaps = 23/112 (20%)
Query: 393 SLILLTG------QNREKPRPVGIIWGGTANRGRLKLRVGQPPENWTS--GVD---LGRL 441
SL LTG QN++K P IW T +RG + + P N + GVD + R
Sbjct: 804 SLTPLTGAIEIVPQNQKKFEPKSTIWADTLSRGLVNFNISGPKTNPLADIGVDFEAINRK 863
Query: 442 LDLLELDLVTTNETLQDSGQEQMNGSTAGIG------------STVGESSPT 481
LE +T + + GS G+G S VG S PT
Sbjct: 864 EKRLEKPTITQQQVTSTINMGKAMGSGTGLGRAGAGAMRPPTNSMVGSSMPT 915
>UniRef100_Q94I00 Hypothetical protein OSJNBa0034E23.25 [Oryza sativa]
Length = 1446
Score = 34.7 bits (78), Expect = 8.2
Identities = 41/163 (25%), Positives = 63/163 (38%), Gaps = 30/163 (18%)
Query: 129 GGVLTDIPAILVFVAHKVHRQ----WLNHVQCLPAALEGPGGVWCDVDVVEFS-YYGAPA 183
G + T P ++ + H H Q W+N V+C A GP +D E YG
Sbjct: 231 GNLKTGSPVLVNHLQHGFHVQHFNGWMNQVECQSAPHVGPASSAASLDPTEEKILYGDDN 290
Query: 184 PTPKEQLYTELADGLRGSDSCVGSG-SQVASQETYGTLGAIVR-------SRTG------ 229
+ +DG+ G D+ G+G S + G+ A+++ S+ G
Sbjct: 291 NFTGLLGEDDSSDGVPGHDNSSGNGNSSIPVSAQGGSWSALMQEALQSTSSKNGLQEEWS 350
Query: 230 -----NREVGFLTNRHVAVDLDYP-----NQKMFHPLPPSLGP 262
NR+ F TN+ + DL+ N H PPS P
Sbjct: 351 SVNFQNRDQAF-TNKMTSPDLEQRQHATLNSMNLHSAPPSAQP 392
>UniRef100_Q86T34 Hypothetical protein DKFZp451A047 [Homo sapiens]
Length = 1163
Score = 34.7 bits (78), Expect = 8.2
Identities = 32/132 (24%), Positives = 59/132 (44%), Gaps = 12/132 (9%)
Query: 452 TNETLQDSGQEQMNGSTAGIGSTVGESSPTVPIKEKL-EESFEPFCLNMEHVPVEEPSTI 510
T+ L+ + + +G+ G+G+ E + VP++ EE F +++ +EEP +
Sbjct: 439 THHALERADEAGSHGN--GVGNASPEVNLNVPVQVSFPEEEFASGATHVQETSLEEPKIL 496
Query: 511 VKPSLRPCEFHIRNEIETVPNVEHQFIRTSFAGKSPVHQSFLKEDMQFKSLSELRNEPDE 570
V P P E +RN V+ ++ T V Q L E++ +S E +
Sbjct: 497 VPP--EPSEERLRNS-----PVQDEYEFTESLHNEVVPQDILSEELSSESTPEDVLSQGK 549
Query: 571 DNFVSLHLGEPE 582
++F H+ E E
Sbjct: 550 ESFE--HISENE 559
>UniRef100_Q6ZZF3 Putative Ssy5 homologue [Pichia jadinii]
Length = 303
Score = 34.7 bits (78), Expect = 8.2
Identities = 21/56 (37%), Positives = 32/56 (56%), Gaps = 1/56 (1%)
Query: 341 QVIKVGRSSGLTTGTIMAYALEYNDEKGICFLTDFLVVGENQQTFDLEGDSGSLIL 396
+V K+G SS TTG + + + Y + G ++F+V + F + GDSGSLIL
Sbjct: 200 RVFKIGASSNYTTGYLNSVKMVYWLD-GSIQTSEFVVSNSAKSMFAMGGDSGSLIL 254
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,053,034,335
Number of Sequences: 2790947
Number of extensions: 48071504
Number of successful extensions: 98681
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 98636
Number of HSP's gapped (non-prelim): 22
length of query: 597
length of database: 848,049,833
effective HSP length: 133
effective length of query: 464
effective length of database: 476,853,882
effective search space: 221260201248
effective search space used: 221260201248
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 78 (34.7 bits)
Medicago: description of AC149576.6


