

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
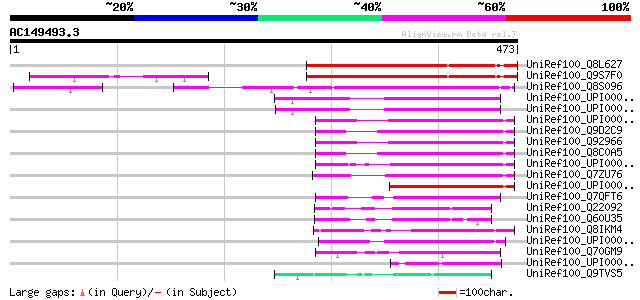
Query= AC149493.3 - phase: 0 /pseudo
(473 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8L627 Hypothetical protein At1g28560 [Arabidopsis tha... 256 1e-66
UniRef100_Q9S7F0 F1K23.20 [Arabidopsis thaliana] 256 1e-66
UniRef100_Q8S096 P0470A12.23 protein [Oryza sativa] 238 3e-61
UniRef100_UPI00002F9F48 UPI00002F9F48 UniRef100 entry 130 1e-28
UniRef100_UPI0000364690 UPI0000364690 UniRef100 entry 126 1e-27
UniRef100_UPI000018321C UPI000018321C UniRef100 entry 124 4e-27
UniRef100_Q9D2C9 Mus musculus 11 days pregnant adult female ovar... 124 8e-27
UniRef100_Q92966 snRNA activating protein complex 50 kDa subunit... 122 3e-26
UniRef100_Q8C0A5 Mus musculus adult male medulla oblongata cDNA,... 120 8e-26
UniRef100_UPI00003A973B UPI00003A973B UniRef100 entry 119 2e-25
UniRef100_Q7ZU76 Similar to small nuclear RNA activating complex... 115 4e-24
UniRef100_UPI00003A973C UPI00003A973C UniRef100 entry 105 4e-21
UniRef100_Q7QFT6 ENSANGP00000017886 [Anopheles gambiae str. PEST] 79 3e-13
UniRef100_Q22092 Hypothetical protein T02C12.2 [Caenorhabditis e... 72 3e-11
UniRef100_Q60U35 Hypothetical protein CBG20195 [Caenorhabditis b... 67 1e-09
UniRef100_Q8IKM4 Hypothetical protein [Plasmodium falciparum] 67 1e-09
UniRef100_UPI00004687FF UPI00004687FF UniRef100 entry 66 2e-09
UniRef100_Q70GM9 Small nuclear RNA gene activation protein 50 [T... 62 5e-08
UniRef100_UPI000049882B UPI000049882B UniRef100 entry 59 4e-07
UniRef100_Q9TVS5 Hypothetical protein F18A11.6 [Caenorhabditis e... 58 5e-07
>UniRef100_Q8L627 Hypothetical protein At1g28560 [Arabidopsis thaliana]
Length = 375
Score = 256 bits (654), Expect = 1e-66
Identities = 122/196 (62%), Positives = 149/196 (75%), Gaps = 4/196 (2%)
Query: 278 QKKGQHDPSGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQK 337
QK G++DPSGYFLIEDVF+ DLR+PSA D + PILDWL NSK+EA KKWE ++ GELQ+K
Sbjct: 183 QKAGKYDPSGYFLIEDVFHNDLRNPSAKDYSYPILDWLWNSKDEALKKWECVLTGELQKK 242
Query: 338 QKAIVGEASVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHAD 397
QK ++GEA LPR+ + +M HFCD+ FR+GA Y+YCHQGDC HT+VIRDMR+ H +
Sbjct: 243 QKLVLGEAKSVDLPRYRTADMQSTHFCDIRFRVGASYVYCHQGDCKHTIVIRDMRMSHPE 302
Query: 398 DVQNWAVYPIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAED 457
DVQN A YPI+ F K R QKCGVCKI RA+KV VDDKW +N YFCD CF LLH +E+
Sbjct: 303 DVQNRAAYPIM-FWPKRRIQKCGVCKIKRASKVAVDDKWASENSSYFCDVCFELLH-SEE 360
Query: 458 GGSPMYTDFIEYDYNH 473
G P+ DF +DY H
Sbjct: 361 G--PLNCDFPVFDYVH 374
Score = 37.4 bits (85), Expect = 0.94
Identities = 16/24 (66%), Positives = 20/24 (82%)
Query: 162 KVEEVVRIKQKQEEDKAQVRLHSF 185
KVE++ ++KQKQEEDKA V LH F
Sbjct: 70 KVEQLAKLKQKQEEDKAAVTLHCF 93
>UniRef100_Q9S7F0 F1K23.20 [Arabidopsis thaliana]
Length = 482
Score = 256 bits (654), Expect = 1e-66
Identities = 122/196 (62%), Positives = 149/196 (75%), Gaps = 4/196 (2%)
Query: 278 QKKGQHDPSGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQK 337
QK G++DPSGYFLIEDVF+ DLR+PSA D + PILDWL NSK+EA KKWE ++ GELQ+K
Sbjct: 290 QKAGKYDPSGYFLIEDVFHNDLRNPSAKDYSYPILDWLWNSKDEALKKWECVLTGELQKK 349
Query: 338 QKAIVGEASVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHAD 397
QK ++GEA LPR+ + +M HFCD+ FR+GA Y+YCHQGDC HT+VIRDMR+ H +
Sbjct: 350 QKLVLGEAKSVDLPRYRTADMQSTHFCDIRFRVGASYVYCHQGDCKHTIVIRDMRMSHPE 409
Query: 398 DVQNWAVYPIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAED 457
DVQN A YPI+ F K R QKCGVCKI RA+KV VDDKW +N YFCD CF LLH +E+
Sbjct: 410 DVQNRAAYPIM-FWPKRRIQKCGVCKIKRASKVAVDDKWASENSSYFCDVCFELLH-SEE 467
Query: 458 GGSPMYTDFIEYDYNH 473
G P+ DF +DY H
Sbjct: 468 G--PLNCDFPVFDYVH 481
Score = 77.8 bits (190), Expect = 6e-13
Identities = 59/184 (32%), Positives = 88/184 (47%), Gaps = 42/184 (22%)
Query: 19 IPRGGPIYVSNMSGPIVRVPLFQDSILTQLHSLQSELPPDS------NHDISVDDLKVFT 72
+PRGGPIY+ NM G I P F+ S L L L++ L DS D+S+D LK++T
Sbjct: 22 VPRGGPIYLPNMVGNISTTPEFKSSFLNVLQDLETHLSLDSTSSSSQQFDVSIDSLKIYT 81
Query: 73 EDDLMDMALKQVFQGRDNNQDPPNAELVTSLPLFFFIL*YH*LIMMDVVVLNNID*ILNL 132
+++L +MA+K+ F +Q+ EL SL N+ N
Sbjct: 82 DEELTEMAMKEAFPEDYLSQE----ELEPSL---------------------NVSHHENP 116
Query: 133 VAG-------VRRSNSGESLV*QINPFLMLQSNCKE----KVEEVVRIKQKQEEDKAQVR 181
+AG V+ + + + M + +E KVE++ ++KQKQEEDKA V
Sbjct: 117 LAGRAKRKRTVKNTEVKKRTLKNTEVMKMTEKKTEEAYLVKVEQLAKLKQKQEEDKAAVT 176
Query: 182 LHSF 185
LH F
Sbjct: 177 LHCF 180
>UniRef100_Q8S096 P0470A12.23 protein [Oryza sativa]
Length = 545
Score = 238 bits (607), Expect = 3e-61
Identities = 138/348 (39%), Positives = 188/348 (53%), Gaps = 65/348 (18%)
Query: 154 MLQSNCKEKVEEVVRIKQKQEEDKAQVRLHSFQLVFLIFIHFIMCKITSI*SNSAIFL*S 213
+LQ + K ++ IK KQEEDK LHSF
Sbjct: 230 ILQGSYLTKAVKMAEIKAKQEEDKHAASLHSFS--------------------------- 262
Query: 214 *LQDQSICQQINQNSKDDVS*VHKFRQKG----QHTGSSGAHTCAGFRSCSLCRDFP*FS 269
S+ ++++ S + V R T G H + LC + +
Sbjct: 263 ---GDSVLAKVSKPSAEKVDVAKSLRYISTTWKNKTFKPGEHRPVVYPEVLLCVEV--YE 317
Query: 270 KRCQAIQCQK--------------------------KGQHDPSGYFLIEDVFYTDLRDPS 303
KR +++ Q+ QH SGYFLIED FY D R S
Sbjct: 318 KRYGSVKSQEFLVLGSQLLTDLRDNIYCFKDKLMNVAKQHVHSGYFLIEDTFYNDTRR-S 376
Query: 304 AIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEASVSHLPRFASFEMHKIHF 363
+D ++PILDW++NS+ EA++KW+ I +G L+++QK ++ +VS++P F S +M K F
Sbjct: 377 TVDYSKPILDWIKNSRNEAEEKWDAITSGVLKKRQKDLLMGLNVSNVPDFKSAKMEKTRF 436
Query: 364 CDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVYPIVTFQLKIRFQKCGVCK 423
DL FRLGAGYLYCHQG+C H +VIRDMRLIH +D QN A YP++TFQ++ R QKC VC+
Sbjct: 437 SDLNFRLGAGYLYCHQGNCKHMIVIRDMRLIHPEDTQNQAEYPLMTFQMQRRLQKCSVCQ 496
Query: 424 IFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTDFIEYDY 471
IF ATK+TVDDKWT +NPCYFCD+C+ LLH ED S +Y + YDY
Sbjct: 497 IFHATKMTVDDKWTLNNPCYFCDKCYYLLHYKED-NSLLYHHTV-YDY 542
Score = 55.8 bits (133), Expect = 3e-06
Identities = 30/86 (34%), Positives = 52/86 (59%), Gaps = 3/86 (3%)
Query: 4 DELELEFSIDDPSISIPRGGPIYVSNMSGPIVRVPLFQDSILTQLHSLQSEL--PPDS-N 60
+E E + + + RGGP++V M GP+ VP F S L +L SL+ EL P D +
Sbjct: 8 EEAESSAAGEQRRMPFARGGPVFVPFMVGPVSTVPEFMSSALHELQSLKDELGDPGDEFD 67
Query: 61 HDISVDDLKVFTEDDLMDMALKQVFQ 86
++ VD+L+V +E++L++ AL++ +
Sbjct: 68 EELCVDELRVLSEEELVERALREAME 93
>UniRef100_UPI00002F9F48 UPI00002F9F48 UniRef100 entry
Length = 274
Score = 130 bits (326), Expect = 1e-28
Identities = 67/215 (31%), Positives = 101/215 (46%), Gaps = 34/215 (15%)
Query: 248 SGAHTCAGFRSCSLC----RDFP*FSKRCQAIQCQKKGQHDPSGYFLIEDVFYTDLRDPS 303
+G+HT A R C + FS A H S +F E VFY D+R P
Sbjct: 72 TGSHTLAELRDAICCVSDLQVCGEFSNNPDAAPAFISKDHFKSAFFYFEGVFYNDMRFPE 131
Query: 304 AIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEASVSHLPRFASFEMHKIHF 363
+D++R ++W ++ + P F+ +M F
Sbjct: 132 CVDISRTTIEWAESH------------------------------NFPPFSQAKMEDTRF 161
Query: 364 CDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVYPIVTFQLKIRFQKCGVCK 423
DL ++G YLYCHQGDC H ++I D+RL+H D + +YP++T + + QKC C
Sbjct: 162 VDLKVKVGFPYLYCHQGDCEHLVIITDIRLVHKSDCLDRKLYPLLTHKHRFSTQKCSACH 221
Query: 424 IFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDG 458
+F VT D++ P +PC FCD+CF +LH +G
Sbjct: 222 LFIGRWVTTGDRFAPSDPCLFCDKCFRMLHYDAEG 256
>UniRef100_UPI0000364690 UPI0000364690 UniRef100 entry
Length = 375
Score = 126 bits (317), Expect = 1e-27
Identities = 67/214 (31%), Positives = 97/214 (45%), Gaps = 34/214 (15%)
Query: 249 GAHTCAGFRSCSLC----RDFP*FSKRCQAIQCQKKGQHDPSGYFLIEDVFYTDLRDPSA 304
G+HT A R C + FS A H S +F E VFY D+R P
Sbjct: 174 GSHTLAELRDAICCVSDLQVCGEFSNNPDAAPSFLSKDHFKSAFFYFEGVFYNDMRFPEC 233
Query: 305 IDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEASVSHLPRFASFEMHKIHFC 364
+D++R ++W ++ + P F+ +M F
Sbjct: 234 VDISRTTIEWAESH------------------------------NFPPFSQAKMEDTRFV 263
Query: 365 DLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVYPIVTFQLKIRFQKCGVCKI 424
DL ++G YLYCHQGDC H ++I D+RL H D + +YP++T + + QKC VC +
Sbjct: 264 DLKVKVGFPYLYCHQGDCEHLIIITDIRLAHRSDCLDKKLYPLLTHKHRFTTQKCSVCHV 323
Query: 425 FRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDG 458
F T D+ P +PC FCD+CF +LH G
Sbjct: 324 FIGRWFTTGDRLAPSDPCLFCDKCFRMLHYDAQG 357
>UniRef100_UPI000018321C UPI000018321C UniRef100 entry
Length = 407
Score = 124 bits (312), Expect = 4e-27
Identities = 65/186 (34%), Positives = 94/186 (49%), Gaps = 30/186 (16%)
Query: 286 SGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEA 345
S +F E FY D R P DL+R I++W ++ K
Sbjct: 245 SAFFYFEGTFYNDKRYPECRDLSRTIIEWSESHDRGYGK--------------------- 283
Query: 346 SVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVY 405
F + M F DL +LG YLYCHQGDC H +VI D+RL+H DD + +Y
Sbjct: 284 -------FQTARMEDFTFNDLNIKLGFPYLYCHQGDCEHVVVITDIRLVHHDDCLDRTLY 336
Query: 406 PIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTD 465
P++T + + +KC VCK++ A VT +D + P++PC+FCD CF +LH +G +
Sbjct: 337 PLLTKKHWLWTRKCFVCKMYTARWVTNNDTFAPEDPCFFCDVCFRMLHYDSEGNK--LGE 394
Query: 466 FIEYDY 471
F+ Y Y
Sbjct: 395 FLAYPY 400
>UniRef100_Q9D2C9 Mus musculus 11 days pregnant adult female ovary and uterus cDNA,
RIKEN full-length enriched library, clone:5031401C21
product:SNRNA ACTIVATING PROTEIN COMPLEX 50 kDa SUBUNIT
(SNAPC 50 kDa SUBUNIT) (PROXIMAL SEQUENCE
ELEMENT-BINDING TRANSCRIPTION FACTOR BETA SUBUNIT)
(PSE-BINDING FACTOR BETA SUBUNIT) (PTF BETA SUBUNIT)
homolog (Small nuclear RNA activating complex,
polypeptide 3) (Mus musculus adult male hippocampus
cDNA, RIKEN full-length enriched library,
clone:C630012M08 product:SNRNA ACTIVATING PROTEIN
COMPLEX 50 kDa SUBUNIT (SNAPC 50 kDa SUBUNIT) (PROXIMAL
SEQUENCE ELEMENT-BINDING TRANSCRIPTION FACTOR BETA
SUBUNIT) (PSE-BINDING FACTOR BETA SUBUNIT) (PTF BETA
SUBUNIT) homolog) [Mus musculus]
Length = 407
Score = 124 bits (310), Expect = 8e-27
Identities = 65/186 (34%), Positives = 94/186 (49%), Gaps = 30/186 (16%)
Query: 286 SGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEA 345
S +F E FY D R P DL+R I++W E+
Sbjct: 245 SAFFYFEGTFYNDRRYPECRDLSRTIIEW----------------------------SES 276
Query: 346 SVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVY 405
+F + M F DL +LG YLYCHQGDC H +VI D+RL+H DD + +Y
Sbjct: 277 HDRGYGKFQTARMEDFTFNDLHIKLGFPYLYCHQGDCEHVVVITDIRLVHHDDCLDRTLY 336
Query: 406 PIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTD 465
P++T + + +KC VCK++ A VT +D + P++PC+FCD CF +LH +G +
Sbjct: 337 PLLTKKHWLWTRKCFVCKMYTARWVTNNDTFAPEDPCFFCDVCFRMLHYDSEGNK--LGE 394
Query: 466 FIEYDY 471
F+ Y Y
Sbjct: 395 FLAYPY 400
>UniRef100_Q92966 snRNA activating protein complex 50 kDa subunit [Homo sapiens]
Length = 411
Score = 122 bits (305), Expect = 3e-26
Identities = 64/186 (34%), Positives = 93/186 (49%), Gaps = 30/186 (16%)
Query: 286 SGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEA 345
S +F E FY D R P DL+R I++W ++ K
Sbjct: 249 SAFFYFEGTFYNDKRYPECRDLSRTIIEWSESHDRGYGK--------------------- 287
Query: 346 SVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVY 405
F + M F DL +LG YLYCHQGDC H +VI D+RL+H DD + +Y
Sbjct: 288 -------FQTARMEDFTFNDLCIKLGFPYLYCHQGDCEHVIVITDIRLVHHDDCLDRTLY 340
Query: 406 PIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTD 465
P++ + + +KC VCK++ A VT +D + P++PC+FCD CF +LH +G +
Sbjct: 341 PLLIKKHWLWTRKCFVCKMYTARWVTNNDSFAPEDPCFFCDVCFRMLHYDSEGNK--LGE 398
Query: 466 FIEYDY 471
F+ Y Y
Sbjct: 399 FLAYPY 404
>UniRef100_Q8C0A5 Mus musculus adult male medulla oblongata cDNA, RIKEN full-length
enriched library, clone:6330437E01 product:SNRNA
ACTIVATING PROTEIN COMPLEX 50 kDa SUBUNIT (SNAPC 50 kDa
SUBUNIT) (PROXIMAL SEQUENCE ELEMENT-BINDING
TRANSCRIPTION FACTOR BETA SUBUNIT) (PSE-BINDING FACTOR
BETA SUBUNIT) (PTF BETA SUBUNIT) homolog [Mus musculus]
Length = 407
Score = 120 bits (301), Expect = 8e-26
Identities = 64/186 (34%), Positives = 93/186 (49%), Gaps = 30/186 (16%)
Query: 286 SGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEA 345
S +F E F D R P DL+R I++W E+
Sbjct: 245 SAFFYFEGTFCNDRRYPECRDLSRTIIEW----------------------------SES 276
Query: 346 SVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVY 405
+F + M F DL +LG YLYCHQGDC H +VI D+RL+H DD + +Y
Sbjct: 277 HDRGYGKFQTARMEDFTFNDLHIKLGFPYLYCHQGDCEHVVVITDIRLVHHDDCLDRTLY 336
Query: 406 PIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTD 465
P++T + + +KC VCK++ A VT +D + P++PC+FCD CF +LH +G +
Sbjct: 337 PLLTKKHWLWTRKCFVCKMYTARWVTNNDTFAPEDPCFFCDVCFRMLHYDSEGNK--LGE 394
Query: 466 FIEYDY 471
F+ Y Y
Sbjct: 395 FLAYPY 400
>UniRef100_UPI00003A973B UPI00003A973B UniRef100 entry
Length = 372
Score = 119 bits (298), Expect = 2e-25
Identities = 65/186 (34%), Positives = 96/186 (50%), Gaps = 30/186 (16%)
Query: 286 SGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEA 345
S +F E +FY D R P DL+R I++W + S++ G LQ
Sbjct: 210 SAFFYFEGIFYNDSRYPECRDLSRTIIEWSE-SRDRGY--------GNLQ---------- 250
Query: 346 SVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVY 405
S +M F DL R+G YL+CHQGDC H +++ D+RLIH +D + +Y
Sbjct: 251 ---------SVKMEDYTFNDLSLRIGYPYLFCHQGDCEHIVIVTDIRLIHHEDCLDRNLY 301
Query: 406 PIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTD 465
P++ + + +KC VCK++ A VT D P++PC+FCD CF +LH +G D
Sbjct: 302 PLLIKKHWLCTRKCFVCKMYTARWVTNKDSLAPEDPCFFCDVCFRMLHYDAEGNK--LGD 359
Query: 466 FIEYDY 471
F+ Y Y
Sbjct: 360 FLAYPY 365
>UniRef100_Q7ZU76 Similar to small nuclear RNA activating complex, polypeptide 3,
50kDa [Brachydanio rerio]
Length = 378
Score = 115 bits (287), Expect = 4e-24
Identities = 57/189 (30%), Positives = 89/189 (46%), Gaps = 32/189 (16%)
Query: 283 HDPSGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIV 342
H S +F FY D R P D+++ I +W ++
Sbjct: 215 HYKSAFFFFNGTFYNDTRFPECQDISKVIKEWTRSRD----------------------- 251
Query: 343 GEASVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNW 402
P F + M F DL ++G YLY HQGDC H +V+ D+RL+H DD +
Sbjct: 252 -------FPDFKTARMEDTSFNDLQMKVGFPYLYTHQGDCEHVVVLTDVRLVHQDDCLDI 304
Query: 403 AVYPIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPM 462
+YP++T + ++ +KC VC ++ + +T +D P +PC FCD+CF + H + G
Sbjct: 305 KLYPLITHKHRVMTRKCSVCHLYISRWITTNDALAPMDPCLFCDQCFRMFHYDDKGNK-- 362
Query: 463 YTDFIEYDY 471
DF+ Y Y
Sbjct: 363 VGDFLAYAY 371
>UniRef100_UPI00003A973C UPI00003A973C UniRef100 entry
Length = 155
Score = 105 bits (261), Expect = 4e-21
Identities = 48/117 (41%), Positives = 71/117 (60%), Gaps = 2/117 (1%)
Query: 355 SFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVYPIVTFQLKI 414
S +M F DL R+G YL+CHQGDC H +++ D+RLIH +D + +YP++ + +
Sbjct: 34 SVKMEDYTFNDLSLRIGYPYLFCHQGDCEHIVIVTDIRLIHHEDCLDRNLYPLLIKKHWL 93
Query: 415 RFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTDFIEYDY 471
+KC VCK++ A VT D P++PC+FCD CF +LH +G DF+ Y Y
Sbjct: 94 CTRKCFVCKMYTARWVTNKDSLAPEDPCFFCDVCFRMLHYDAEGNK--LGDFLAYPY 148
>UniRef100_Q7QFT6 ENSANGP00000017886 [Anopheles gambiae str. PEST]
Length = 244
Score = 79.0 bits (193), Expect = 3e-13
Identities = 49/173 (28%), Positives = 71/173 (40%), Gaps = 29/173 (16%)
Query: 286 SGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEA 345
SG+F + D FY D RD + D + I W +++++GE
Sbjct: 71 SGFFFVHDTFYNDFRDDANHDYSGVIRKWAD---------------------RQSLIGEL 109
Query: 346 SVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVY 405
+ M F DL FRLG +Y HQG+C H VI D RL+ A D+ + Y
Sbjct: 110 KTAR--------MEDTRFGDLKFRLGYPQMYQHQGNCEHLFVISDCRLLAATDILTRSRY 161
Query: 406 PIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDG 458
P + R C +C +A + + +P Y C+ C H EDG
Sbjct: 162 PWLNSYGFSRDVPCNICGHCQAQYIVQNSTRHIFDPAYICENCLETYHYTEDG 214
>UniRef100_Q22092 Hypothetical protein T02C12.2 [Caenorhabditis elegans]
Length = 418
Score = 72.0 bits (175), Expect = 3e-11
Identities = 45/165 (27%), Positives = 71/165 (42%), Gaps = 31/165 (18%)
Query: 285 PSGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGE 344
PS + + D FY D+ P+AID++ PI + ++++ E+ +A
Sbjct: 245 PSSFIFVHDTFYVDM-PPNAIDISHPIRN--------------FMLHREIYDPVEAC--- 286
Query: 345 ASVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAV 404
M + DL RLG Y++ H G+C H LV D+RL+H D
Sbjct: 287 ------------SMEGVRIIDLKLRLGQPYIFQHSGNCEHLLVFHDLRLLHESDPWGIDK 334
Query: 405 YPIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECF 449
YP ++ K +KC +CK V + P+ +FC CF
Sbjct: 335 YPFTLYE-KGNEKKCDICKKGHVEFVVERHELLPNTYTHFCRTCF 378
>UniRef100_Q60U35 Hypothetical protein CBG20195 [Caenorhabditis briggsae]
Length = 416
Score = 67.0 bits (162), Expect = 1e-09
Identities = 46/167 (27%), Positives = 71/167 (41%), Gaps = 35/167 (20%)
Query: 285 PSGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGE 344
PS + + + FY D +A D++ PI ++Q+ E+ A+
Sbjct: 243 PSSFIFVHNTFYVDSAPENAQDISFPIRKFMQDR--------------EIFDPVDAV--- 285
Query: 345 ASVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAV 404
M + DL RLG Y++ H G+C H L+ D+RL+H D +
Sbjct: 286 ------------PMEGVRIIDLSLRLGQPYVFQHSGNCEHVLIFHDLRLLHESDPRGIDQ 333
Query: 405 YPIVTFQLKIRFQKCGVCKIFRATKVTVDDK--WTPDNPCYFCDECF 449
YP V ++ K +KC +CK V V D+ P+ YFC CF
Sbjct: 334 YPYVLYE-KGNERKCEICK---KGHVDVVDRHELLPNTYTYFCRSCF 376
>UniRef100_Q8IKM4 Hypothetical protein [Plasmodium falciparum]
Length = 635
Score = 66.6 bits (161), Expect = 1e-09
Identities = 52/189 (27%), Positives = 84/189 (43%), Gaps = 29/189 (15%)
Query: 284 DPSGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQK-KWEYIINGELQQKQKAIV 342
D S YF I+ + Y DLR +A+D + IL++ + K + K+ Y IN + KAI+
Sbjct: 471 DGSLYF-IDGILYPDLRSNNAVDYSTSILNFYKMKKMKTNFIKYPYKIN-----QDKAIL 524
Query: 343 GEASVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNW 402
+ + P F + HQG C H +V ++R + ++
Sbjct: 525 SQIEI---PLFKKC------------------CFLHQGTCEHRIVFNNIRQYNKLRDKHL 563
Query: 403 AVYPIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPM 462
+ YP+ TF+ I + C C A K+ +D +NP Y C+ CF L L + P+
Sbjct: 564 SKYPLRTFKPNISNKYCISCHKNIAQKIVLDSYLLKENPSYMCNNCFDLF-LMDKNNQPI 622
Query: 463 YTDFIEYDY 471
T +DY
Sbjct: 623 DTSMKYFDY 631
>UniRef100_UPI00004687FF UPI00004687FF UniRef100 entry
Length = 610
Score = 66.2 bits (160), Expect = 2e-09
Identities = 45/174 (25%), Positives = 73/174 (41%), Gaps = 13/174 (7%)
Query: 289 FLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEASVS 348
+ I+ + Y D+RD A+D + I+++ +N K + N L
Sbjct: 437 YYIDGILYPDVRDKDALDYSTSIINFYKNRKNISSSTASSSSNSNLDD------------ 484
Query: 349 HLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVYPIV 408
L K D+ L + HQG+C H +V ++R + ++ + YPI
Sbjct: 485 FLKHPYKIYQDKAVLKDIKIPLYKKCCFLHQGNCEHRIVFSNVRQYNKFRDKDISQYPIR 544
Query: 409 TFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPM 462
TF+ I + C C A K+ +D +NP Y CD CF L L + G P+
Sbjct: 545 TFKPNIATKYCICCHKNIAQKIIIDSYLFKENPSYVCDGCFELF-LLDSNGKPV 597
>UniRef100_Q70GM9 Small nuclear RNA gene activation protein 50 [Trypanosoma brucei
brucei]
Length = 448
Score = 61.6 bits (148), Expect = 5e-08
Identities = 49/180 (27%), Positives = 76/180 (42%), Gaps = 23/180 (12%)
Query: 286 SGYFLIEDVFYTDLRDPSA-----IDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKA 340
+ +F I FY D R DLT PI + + + + GE +QK A
Sbjct: 265 NAFFFIGGTFYVDNRHAGEGGEDYEDLTAPIRHFDPCGEGASTE-------GETRQKNIA 317
Query: 341 IVGEASVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQ 400
G V ++ + F DL RLG + H G C H + + + D
Sbjct: 318 F-GNCPVKYVSQTT--------FGDLNLRLGEYGVMRHLGWCNHYFYLSSVTSLRGFDRD 368
Query: 401 NW--AVYPIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDG 458
+ A YP + R +C +C+ AT V +D+ +P++PC +C CF LLH ++G
Sbjct: 369 DHTRAAYPQRVMKTPTRVVRCRLCRSHPATVVCYNDEISPESPCPYCVPCFELLHATDEG 428
>UniRef100_UPI000049882B UPI000049882B UniRef100 entry
Length = 342
Score = 58.5 bits (140), Expect = 4e-07
Identities = 34/104 (32%), Positives = 49/104 (46%), Gaps = 7/104 (6%)
Query: 356 FEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVYPIVTFQLKIR 415
FE+ KI + YLY H DC H ++ D+R+ +D YP + F+ +
Sbjct: 240 FELGKIDI-----EIDEPYLYGHLLDCEHIFIVSDIRVPLQEDKNG--KYPRIIFRKRKE 292
Query: 416 FQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGG 459
Q+C +C +A D +P Y+C ECFSLLH D G
Sbjct: 293 QQRCNICDSRKADIEVTGDSAGISDPSYYCKECFSLLHNDMDIG 336
>UniRef100_Q9TVS5 Hypothetical protein F18A11.6 [Caenorhabditis elegans]
Length = 352
Score = 58.2 bits (139), Expect = 5e-07
Identities = 58/208 (27%), Positives = 83/208 (39%), Gaps = 38/208 (18%)
Query: 248 SGAHTCAGFRSCSLCR-DFP*---FSKRCQAIQCQKKGQHDPSGYFLIEDVFYTDLRDPS 303
+G HT ++ C DF FS++ + + K + PS F I D FY D +
Sbjct: 139 TGDHTLRDLKNAFSCPIDFSFSDDFSEKKPSFKDMAKNKW-PSSMFFIHDTFYIDSNTGN 197
Query: 304 A-IDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEASVSHLPRFASFEMHKIH 362
+D + I W KK++YI G + KQ M +
Sbjct: 198 QFVDPSITIRTWA--------KKFDYI--GPMHVKQ-------------------MSETR 228
Query: 363 FCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVYPIVTFQLKIRFQKCGVC 422
DL RLG Y+Y HQG C H +V D+ L D+ +P + R C C
Sbjct: 229 IGDLICRLGQPYVYIHQGVCEHLIVFNDLCL--RDESHTNVEFPRRLVERNFRRIACDTC 286
Query: 423 KIFRATKVTVD-DKWTPDNPCYFCDECF 449
K A + VD D P++P Y C C+
Sbjct: 287 KEASAHWMIVDHDNLLPNSPGYLCSSCY 314
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.334 0.146 0.467
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 778,278,102
Number of Sequences: 2790947
Number of extensions: 31530906
Number of successful extensions: 108081
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 27
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 108014
Number of HSP's gapped (non-prelim): 47
length of query: 473
length of database: 848,049,833
effective HSP length: 131
effective length of query: 342
effective length of database: 482,435,776
effective search space: 164993035392
effective search space used: 164993035392
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 77 (34.3 bits)
Medicago: description of AC149493.3


