

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
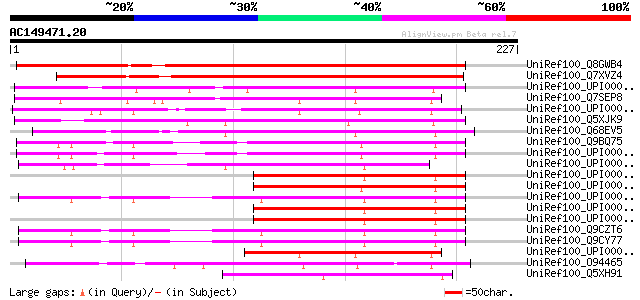
Query= AC149471.20 + phase: 0
(227 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8GWB4 Hypothetical protein At2g43110 [Arabidopsis tha... 217 2e-55
UniRef100_Q7XVZ4 OSJNBa0020I02.12 protein [Oryza sativa] 171 1e-41
UniRef100_UPI000042FF92 UPI000042FF92 UniRef100 entry 81 2e-14
UniRef100_Q7SEP8 Predicted protein [Neurospora crassa] 72 8e-12
UniRef100_UPI000023468A UPI000023468A UniRef100 entry 72 1e-11
UniRef100_Q5XJK9 Zgc:101814 [Brachydanio rerio] 71 2e-11
UniRef100_Q68EV5 MGC84286 protein [Xenopus laevis] 71 2e-11
UniRef100_Q9BQ75 Hypothetical protein MGC4308 [Homo sapiens] 71 2e-11
UniRef100_UPI000036B65A UPI000036B65A UniRef100 entry 69 1e-10
UniRef100_UPI000034D7EC UPI000034D7EC UniRef100 entry 68 2e-10
UniRef100_UPI000036621A UPI000036621A UniRef100 entry 68 2e-10
UniRef100_UPI00001D0704 UPI00001D0704 UniRef100 entry 67 3e-10
UniRef100_UPI0000192CDB UPI0000192CDB UniRef100 entry 67 3e-10
UniRef100_UPI00003A9B9B UPI00003A9B9B UniRef100 entry 67 3e-10
UniRef100_UPI00003A9B9A UPI00003A9B9A UniRef100 entry 67 3e-10
UniRef100_Q9CZT6 Mus musculus 10 days embryo whole body cDNA, RI... 67 3e-10
UniRef100_Q9CY77 Mus musculus 8 days embryo whole body cDNA, RIK... 67 3e-10
UniRef100_UPI000023DFD0 UPI000023DFD0 UniRef100 entry 64 4e-09
UniRef100_O94465 SPBC23G7.07c protein [Schizosaccharomyces pombe] 60 3e-08
UniRef100_Q5XH91 Hypothetical protein [Xenopus tropicalis] 52 9e-06
>UniRef100_Q8GWB4 Hypothetical protein At2g43110 [Arabidopsis thaliana]
Length = 288
Score = 217 bits (552), Expect = 2e-55
Identities = 111/201 (55%), Positives = 147/201 (72%), Gaps = 6/201 (2%)
Query: 4 RKGRENNDDRRKKKKKKTETTGSEQLRFFVNEFQSANDVQLSSIELESLKADSSILELSN 63
R + D+ ++ + SEQL +F+N SA +++SS+ELE +K D+ I+ELS
Sbjct: 55 RSNDQKTDNDEDEQLYSEPVSASEQLNYFLNHLDSAIGIKVSSLELEPIK-DTCIVELSQ 113
Query: 64 SQNSRDTDLDVKLLGGDIKGAFGNCWRQVLCESEVVEGKIPPGSPSVLIVSPSALRSIHL 123
D DV LG IK + G+ WR+ LCE E +E K+ PG+PSVL++S SALRS+ L
Sbjct: 114 G-----LDQDVSNLGEHIKLSCGSSWRETLCEGESLERKVEPGNPSVLVISSSALRSLEL 168
Query: 124 LKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNRVNIASGTPSRIKKLIDVEALGLSRLQV 183
L+G +TKQC AVKLFSKH+K++EQ+SLLK RVNI SGTP+RIKKL+D+EALGLSRL +
Sbjct: 169 LRGLHSLTKQCPAVKLFSKHLKVEEQVSLLKKRVNIGSGTPNRIKKLVDIEALGLSRLDM 228
Query: 184 LVLDMHPDVKGYSLFTLPQVR 204
+V+DMHPDVKG+SLFTLPQVR
Sbjct: 229 IVIDMHPDVKGFSLFTLPQVR 249
>UniRef100_Q7XVZ4 OSJNBa0020I02.12 protein [Oryza sativa]
Length = 283
Score = 171 bits (434), Expect = 1e-41
Identities = 85/182 (46%), Positives = 129/182 (70%), Gaps = 6/182 (3%)
Query: 22 ETTGSEQLRFFVNEFQSANDVQLSSIELESLKADSSILELSNSQNSRDTDLDVKLLGGDI 81
E + QL F + F+ A ++LS +EL++ ++ ++ L+ + DV+ G +
Sbjct: 66 EMPPARQLEFLLRSFERAAKMRLSPLELDAY-SEGCMVPLAEGASQ-----DVEGFGDHV 119
Query: 82 KGAFGNCWRQVLCESEVVEGKIPPGSPSVLIVSPSALRSIHLLKGFRFMTKQCSAVKLFS 141
KGAFG+ W++ LCE E+ G + GSP++L++ +ALRS+ LL+G + TK+C +KLF+
Sbjct: 120 KGAFGSSWKEELCEGELDGGAVDAGSPALLVICSAALRSLELLRGLKMFTKECRPIKLFA 179
Query: 142 KHIKLQEQISLLKNRVNIASGTPSRIKKLIDVEALGLSRLQVLVLDMHPDVKGYSLFTLP 201
KH+K++EQ++LLK RVNIA GTPSRIKKLID+EAL LSR++++VLDM D K ++LFTLP
Sbjct: 180 KHMKVEEQVALLKTRVNIACGTPSRIKKLIDMEALSLSRVKLVVLDMQRDAKSFTLFTLP 239
Query: 202 QV 203
QV
Sbjct: 240 QV 241
>UniRef100_UPI000042FF92 UPI000042FF92 UniRef100 entry
Length = 330
Score = 81.3 bits (199), Expect = 2e-14
Identities = 67/213 (31%), Positives = 110/213 (51%), Gaps = 20/213 (9%)
Query: 3 KRKGRENNDDRRKKKKKKTETTGSEQLRFFVNEFQSANDVQLSSIELESLKAD---SSIL 59
KR+ +E +RR K+++ E S + LSS+ L S++ SS +
Sbjct: 101 KRRKKEKEKERRAKRRQGAELVPSTTAPTHLAT------TDLSSVLLRSIRESFPSSSAI 154
Query: 60 ELSNSQNSRDTDLDVKLLGG---DIKGAFGNCWRQVLCESEVVEGKIPP---GSPSVLIV 113
E+ L+ D AF +++ SE+ +GK P G+P+V+++
Sbjct: 155 EIEEVLIPPANLLEPPTYPAPKADPSNAFKPLQQRI---SELTKGKKEPKGNGAPAVIVL 211
Query: 114 SPSALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLL-KNRVNIASGTPSRIKKLID 172
S LR +++G R + + KLF+KH KL +QI L + +V+IA GTP+R+ KL+
Sbjct: 212 GISGLRCADIVRGVRGVKGKGEVAKLFAKHFKLADQIKYLERTKVSIAVGTPARVGKLLA 271
Query: 173 VEALGLSRLQVLVLDM-HPDVKGYSLFTLPQVR 204
A+ +S+ VL+LD+ H D K +L TLP+VR
Sbjct: 272 EGAIKISKDTVLLLDIGHQDSKTRTLLTLPEVR 304
>UniRef100_Q7SEP8 Predicted protein [Neurospora crassa]
Length = 248
Score = 72.4 bits (176), Expect = 8e-12
Identities = 68/203 (33%), Positives = 102/203 (49%), Gaps = 14/203 (6%)
Query: 3 KRKGRENNDDRRKKKKKKT-ETTGSEQLRFFVNEFQSANDVQLSSIELES----LKADSS 57
KRK + ++KK+ KK E G + +N D QL + + D S
Sbjct: 9 KRKADDEGGAKKKKRSKKAREDEGDLDVEAGLNRAFERMDGQLLADHIAQKVTRFGTDLS 68
Query: 58 ILELSN---SQNS-RDTDLDVKLLGGDIKGAFGNCWRQVLCESEVVEGKIPPGSPSVLIV 113
+ELS+ S N+ +DT K D F + + +++ E GSP L+V
Sbjct: 69 SVELSDLYISANAIKDTTSFQKPRNKDNLPDFLEEFSEN--PAKLSEAPKSNGSPHTLVV 126
Query: 114 SPSALRSIHLLKGFR-FMTKQCSAVKLFSKHIKLQEQISLL-KNRVNIASGTPSRIKKLI 171
+ + LR+ L++ R F TK S KLF+KH K++EQ+S L K+R IA GTP R+ LI
Sbjct: 127 AGAGLRAADLVRSLRKFQTKGNSVAKLFAKHFKVEEQVSFLKKSRTGIAVGTPQRLIDLI 186
Query: 172 DVEALGLSRLQVLVLD-MHPDVK 193
+ E+L L L+ +V+D H D K
Sbjct: 187 ENESLSLDSLKRIVVDASHIDQK 209
>UniRef100_UPI000023468A UPI000023468A UniRef100 entry
Length = 298
Score = 72.0 bits (175), Expect = 1e-11
Identities = 66/221 (29%), Positives = 110/221 (48%), Gaps = 31/221 (14%)
Query: 2 GKRKGRENNDDRRKKKKKKTETTGSEQLRFFVNE--------------FQSA--NDVQLS 45
G++ ++ ND K K + E E+ VNE Q A ++ +L+
Sbjct: 59 GRKDKKDKNDKNDKPKAVRGEPLEKERKGISVNEAIGKMDGRLLADHFVQKAKRHNKELT 118
Query: 46 SIELESLKA-DSSILELSNSQNSRDTDLDVKLLGGDIKGAFGNCWRQVLCESEVVEGKIP 104
++EL L +S+ L+ S+ +SR+ + KL D AF + SE
Sbjct: 119 AVELSDLSVPESAFLDTSSFTSSRNLE---KL--PDFLKAFSPKGVNLSDSSE------Q 167
Query: 105 PGSPSVLIVSPSALRSIHLLKGFR-FMTKQCSAVKLFSKHIKLQEQISLLKN-RVNIASG 162
G+P ++VSP+ LR+ +++ R F TK+ KLF+KHIKL+E L+ R+ I +G
Sbjct: 168 KGTPHTIVVSPAGLRAADVVRALRMFQTKESPIGKLFAKHIKLEEAKQFLERARMGIGAG 227
Query: 163 TPSRIKKLIDVEALGLSRLQVLVLD-MHPDVKGYSLFTLPQ 202
TP+RI LI+ +L L L+ +V+D + D K +F + +
Sbjct: 228 TPARISDLINSGSLKLDELKRIVIDGSYVDQKQRGIFDMKE 268
>UniRef100_Q5XJK9 Zgc:101814 [Brachydanio rerio]
Length = 291
Score = 71.2 bits (173), Expect = 2e-11
Identities = 64/206 (31%), Positives = 104/206 (50%), Gaps = 14/206 (6%)
Query: 3 KRKGRENNDDRRKKKKKKTETTGSEQLRFFVNEFQSANDVQLSSIELESLKADSSILELS 62
++ E+N +++++KKKKT T + S+ V S ++L SL S
Sbjct: 71 EKPDNESNKNKKRRKKKKTIT----------DVLTSSKPVPGSPVDLVSLLKTYHSQTRS 120
Query: 63 NSQNSRDTDLDVKLLG-GDIKGAFGNCWRQVLCE-SEVVEGKIPPGSPSVLIVSPSALRS 120
+ T D L D+ + + ++V + +++ + S +LIV SALR+
Sbjct: 121 VIEQEELTLQDSCFLSCNDLTHSLSSYLKEVCPKWAKMQKQHTQTSSVVLLIVCGSALRT 180
Query: 121 IHLLKGFRFMTKQCSAVKLFSKHIKLQEQI-SLLKNRVNIASGTPSRIKKLIDVEALGLS 179
I L+K Q +KLF+KHIK++EQI SL K +IA GTP RI L++ E L +
Sbjct: 181 IDLIKQLVTFKGQAKVLKLFAKHIKVEEQIKSLSKGVTHIAVGTPGRICALLEKEGLTVQ 240
Query: 180 RLQVLVLDM-HPDVKGYSLFTLPQVR 204
L+ LVLD + D K + +P+V+
Sbjct: 241 GLRYLVLDWNYRDQKQRRMVDVPEVK 266
>UniRef100_Q68EV5 MGC84286 protein [Xenopus laevis]
Length = 277
Score = 70.9 bits (172), Expect = 2e-11
Identities = 60/201 (29%), Positives = 102/201 (49%), Gaps = 9/201 (4%)
Query: 11 DDRRKKKKKKTETTGSEQLRFFVNEFQSANDVQLSSIELESLKADSSILELSNSQNSRDT 70
+D KKK++ + T S+ L + D+Q + LE + S++EL Q DT
Sbjct: 62 EDSAPKKKRRKKKTISDVLAQHAPTAGTPEDMQ--KLVLEHFENKRSVIELEELQIP-DT 118
Query: 71 DLDVKLLGGDIKGAFGNCWRQVLCE-SEVVEGKIPPGSPSVLIVSPSALRSIHLLKGFRF 129
+ D+ + +++ + S++ + S +L+V SA R++ L+K
Sbjct: 119 CF---VKENDLTHTLSSYLKEICPKWSKLSKNHKEKKSVLLLVVCSSAHRTLELIKLINA 175
Query: 130 MTKQCSAVKLFSKHIKLQEQISLL-KNRVNIASGTPSRIKKLIDVEALGLSRLQVLVLDM 188
+KLF+KHIK+++QI+LL KN +I GTP RIK LID + L L ++ LV D
Sbjct: 176 FKADTKVMKLFAKHIKIKDQINLLEKNVTHIGIGTPGRIKALIDQDGLSLESMKYLVFDW 235
Query: 189 H-PDVKGYSLFTLPQVRCASL 208
+ D K + +P+V+ +L
Sbjct: 236 NWRDQKLRRMMDIPEVKKETL 256
>UniRef100_Q9BQ75 Hypothetical protein MGC4308 [Homo sapiens]
Length = 279
Score = 70.9 bits (172), Expect = 2e-11
Identities = 64/213 (30%), Positives = 104/213 (48%), Gaps = 31/213 (14%)
Query: 4 RKGRENNDDRRKKKKKK-TETTGS--------EQLRFFVNEFQSANDVQLSSIELESLKA 54
++ +EN RK++KKK T+ E L+ + ++ S+ + IELE L
Sbjct: 61 KERKENTTKTRKRRKKKITDVLAKSEPKPGLPEDLQKLMKDYYSSRRLV---IELEELNL 117
Query: 55 -DSSILELSNSQNSRDTDLDVKLLGGDIKGAFGNCWRQVLCESEVVEGKIPPGSPSVLIV 113
DS L+ ++ +S + L G C + V E K S +LI+
Sbjct: 118 PDSCFLKANDLTHSLSSYLK------------GICPKWVKLRKNHSEKK----SVLMLII 161
Query: 114 SPSALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNR-VNIASGTPSRIKKLID 172
SA+R++ L++ +KLF+KHIK+Q Q+ LL+ R V++ GTP RIK+L+
Sbjct: 162 CSSAVRALELIRSMTAFRGDGKVIKLFAKHIKVQAQVKLLEKRVVHLGVGTPGRIKELVK 221
Query: 173 VEALGLSRLQVLVLDMH-PDVKGYSLFTLPQVR 204
L LS L+ LV D + D K + +P++R
Sbjct: 222 QGGLNLSPLKFLVFDWNWRDQKLRRMMDIPEIR 254
>UniRef100_UPI000036B65A UPI000036B65A UniRef100 entry
Length = 279
Score = 68.6 bits (166), Expect = 1e-10
Identities = 63/213 (29%), Positives = 104/213 (48%), Gaps = 31/213 (14%)
Query: 4 RKGRENNDDRRKKKKKK-TETTGS--------EQLRFFVNEFQSANDVQLSSIELESLKA 54
++ +EN RK++KKK T+ E L+ + ++ S+ + S IELE L
Sbjct: 61 KERKENTTKTRKRRKKKITDVLAKSEPKPGLPEDLQKLMKDYYSS---RRSVIELEELNL 117
Query: 55 -DSSILELSNSQNSRDTDLDVKLLGGDIKGAFGNCWRQVLCESEVVEGKIPPGSPSVLIV 113
DS L+ ++ +S + L C + V E K S +LI+
Sbjct: 118 PDSCFLKANDLTHSLSSYLKEI------------CPKWVKLRKNHSEKK----SVLMLII 161
Query: 114 SPSALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNR-VNIASGTPSRIKKLID 172
SA+R++ L++ +KLF+KHIK+Q Q+ LL+ R V++ GTP RIK+L+
Sbjct: 162 CSSAIRALELIRSMTAFRGDGKVIKLFAKHIKVQAQVKLLEKRVVHLGVGTPGRIKELVK 221
Query: 173 VEALGLSRLQVLVLDMH-PDVKGYSLFTLPQVR 204
L L+ L+ LV D + D K + +P++R
Sbjct: 222 QGGLNLNPLKFLVFDWNWRDQKLRRMMDIPEIR 254
>UniRef100_UPI000034D7EC UPI000034D7EC UniRef100 entry
Length = 296
Score = 68.2 bits (165), Expect = 2e-10
Identities = 61/196 (31%), Positives = 94/196 (47%), Gaps = 31/196 (15%)
Query: 5 KGRENNDDRRKKKKKKTET---------TGSE-QLRFFVNEFQSANDVQLSSIELESLKA 54
+G +N +RK+KKKKT T GS L+ V ++ S + S IE E L+
Sbjct: 78 EGGTSNKPKRKRKKKKTITDVVAASGPRAGSPADLQKLVTQYFSD---KRSVIEQEELRL 134
Query: 55 DSSILELSNSQNSRDTDLDVKLLGGDIKGAFGNCWRQVLCE-SEVVEGKIPPGSPSVLIV 113
S SN D+ + +QV + +++ + S +LIV
Sbjct: 135 PDSCFLSSN----------------DLTHTLASYLKQVCPKWAKIQKTHTEKSSVVLLIV 178
Query: 114 SPSALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNRV-NIASGTPSRIKKLID 172
SALR+I L+K +KLF+KHIK++EQ+ LL+ V +I GTP RI L++
Sbjct: 179 CSSALRTIELIKQLTAFRGAAKVLKLFAKHIKIEEQVKLLQKGVTHIGVGTPGRISALVE 238
Query: 173 VEALGLSRLQVLVLDM 188
+AL + L+ LV +
Sbjct: 239 KDALNVRSLRYLVFGL 254
>UniRef100_UPI000036621A UPI000036621A UniRef100 entry
Length = 206
Score = 67.8 bits (164), Expect = 2e-10
Identities = 40/97 (41%), Positives = 59/97 (60%), Gaps = 2/97 (2%)
Query: 110 VLIVSPSALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNRV-NIASGTPSRIK 168
+LIV SALR+I LLK +KLF+KHIK++EQ+ LL+ V +I GTP R+
Sbjct: 85 LLIVCSSALRAIELLKQLTTFKGAAKTIKLFAKHIKIEEQVKLLQKGVTHIGVGTPGRVS 144
Query: 169 KLIDVEALGLSRLQVLVLDMH-PDVKGYSLFTLPQVR 204
LI+ + L L L+ LVLD + D K + +P+++
Sbjct: 145 ALIENDGLNLKSLRYLVLDWNWRDQKLRRMVDIPEIK 181
>UniRef100_UPI00001D0704 UPI00001D0704 UniRef100 entry
Length = 457
Score = 67.4 bits (163), Expect = 3e-10
Identities = 36/97 (37%), Positives = 61/97 (62%), Gaps = 2/97 (2%)
Query: 110 VLIVSPSALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNR-VNIASGTPSRIK 168
+LI+ SA+R++ L++ +KLF+KHIK+QEQ+ LL+ R +++ GTP RIK
Sbjct: 336 MLILCSSAVRALELIRSLTAFKGDAKVMKLFAKHIKVQEQVKLLEKRAIHLGVGTPGRIK 395
Query: 169 KLIDVEALGLSRLQVLVLDMH-PDVKGYSLFTLPQVR 204
+L+ + L L+ L+ LV D + D K + +P++R
Sbjct: 396 ELVKQDGLNLNPLKFLVFDWNWRDQKLRRMMDIPEIR 432
>UniRef100_UPI0000192CDB UPI0000192CDB UniRef100 entry
Length = 256
Score = 67.4 bits (163), Expect = 3e-10
Identities = 62/213 (29%), Positives = 102/213 (47%), Gaps = 34/213 (15%)
Query: 5 KGRENNDDRRKKKKKKTETTGS--------EQLRFFVNEFQSANDVQLSSIELESLKA-D 55
K ++ R+ KKKK ++ E L+ + + S++ S IELE L D
Sbjct: 40 KAEDSRKTRKWKKKKISDILAKSEPKPGTPEDLQKLIRDHYSSSR---SVIELEELHLPD 96
Query: 56 SSILELSNSQNSRDTDLDVKLLGGDIKGAFGNCWRQVLCESEVVEGKIPPGSPSVL--IV 113
S L+ ++ +S + L + +C V K SVL I+
Sbjct: 97 SCFLKANDLTHSLSSYL------------------KEICPKWVKLRKTHNEKKSVLMLIL 138
Query: 114 SPSALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNRV-NIASGTPSRIKKLID 172
SA+R++ L++ +KLF+KHIK+QEQ+ LL+ RV ++ GTP RIK+L+
Sbjct: 139 CSSAVRALELIRSLTAFKGDAKVMKLFAKHIKVQEQVKLLEKRVIHLGVGTPGRIKELVK 198
Query: 173 VEALGLSRLQVLVLDMH-PDVKGYSLFTLPQVR 204
+ L L+ L+ LV D + D K + +P++R
Sbjct: 199 QDGLHLNPLKFLVFDWNWRDQKLRRMMDIPEIR 231
>UniRef100_UPI00003A9B9B UPI00003A9B9B UniRef100 entry
Length = 264
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/97 (36%), Positives = 62/97 (63%), Gaps = 2/97 (2%)
Query: 110 VLIVSPSALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNRV-NIASGTPSRIK 168
+L++ SALRS+ L+K C +KLF+KHIK++EQ+++L+ V +I GTP RIK
Sbjct: 143 MLVICSSALRSLELIKSMTAFKGDCRVLKLFAKHIKIKEQMNMLEKGVFHIGVGTPGRIK 202
Query: 169 KLIDVEALGLSRLQVLVLDMH-PDVKGYSLFTLPQVR 204
L++ + L L+ + ++LD + D K + +P+++
Sbjct: 203 ALVEQDGLCLNSTKYIILDWNWRDQKLRRMMDIPEIK 239
>UniRef100_UPI00003A9B9A UPI00003A9B9A UniRef100 entry
Length = 281
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/97 (36%), Positives = 62/97 (63%), Gaps = 2/97 (2%)
Query: 110 VLIVSPSALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNRV-NIASGTPSRIK 168
+L++ SALRS+ L+K C +KLF+KHIK++EQ+++L+ V +I GTP RIK
Sbjct: 160 MLVICSSALRSLELIKSMTAFKGDCRVLKLFAKHIKIKEQMNMLEKGVFHIGVGTPGRIK 219
Query: 169 KLIDVEALGLSRLQVLVLDMH-PDVKGYSLFTLPQVR 204
L++ + L L+ + ++LD + D K + +P+++
Sbjct: 220 ALVEQDGLCLNSTKYIILDWNWRDQKLRRMMDIPEIK 256
>UniRef100_Q9CZT6 Mus musculus 10 days embryo whole body cDNA, RIKEN full-length
enriched library, clone:2610528E23 product:hypothetical
P-loop containing nucleotide triphosphate hydrolases
structure containing protein, full insert sequence [Mus
musculus]
Length = 276
Score = 67.4 bits (163), Expect = 3e-10
Identities = 62/213 (29%), Positives = 102/213 (47%), Gaps = 34/213 (15%)
Query: 5 KGRENNDDRRKKKKKKTETTGS--------EQLRFFVNEFQSANDVQLSSIELESLKA-D 55
K ++ R+ KKKK ++ E L+ + + S++ S IELE L D
Sbjct: 60 KAEDSRKTRKWKKKKISDILAKSEPKPGTPEDLQKLIRDHYSSSR---SVIELEELHLPD 116
Query: 56 SSILELSNSQNSRDTDLDVKLLGGDIKGAFGNCWRQVLCESEVVEGKIPPGSPSVL--IV 113
S L+ ++ +S + L + +C V K SVL I+
Sbjct: 117 SCFLKANDLTHSLSSYL------------------KEICPKWVKLRKTHNEKKSVLMLIL 158
Query: 114 SPSALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNRV-NIASGTPSRIKKLID 172
SA+R++ L++ +KLF+KHIK+QEQ+ LL+ RV ++ GTP RIK+L+
Sbjct: 159 CSSAVRALELIRSLTAFKGDAKVMKLFAKHIKVQEQVKLLEKRVIHLGVGTPGRIKELVK 218
Query: 173 VEALGLSRLQVLVLDMH-PDVKGYSLFTLPQVR 204
+ L L+ L+ LV D + D K + +P++R
Sbjct: 219 QDGLHLNPLKFLVFDWNWRDQKLRRMMDIPEIR 251
>UniRef100_Q9CY77 Mus musculus 8 days embryo whole body cDNA, RIKEN full-length
enriched library, clone:5730589J07 product:hypothetical
P-loop containing nucleotide triphosphate hydrolases
structure containing protein, full insert sequence [Mus
musculus]
Length = 276
Score = 67.4 bits (163), Expect = 3e-10
Identities = 62/213 (29%), Positives = 102/213 (47%), Gaps = 34/213 (15%)
Query: 5 KGRENNDDRRKKKKKKTETTGS--------EQLRFFVNEFQSANDVQLSSIELESLKA-D 55
K ++ R+ KKKK ++ E L+ + + S++ S IELE L D
Sbjct: 60 KAEDSRKTRKWKKKKISDILAKSEPKPGTPEDLQKLIRDHYSSSR---SVIELEELHLPD 116
Query: 56 SSILELSNSQNSRDTDLDVKLLGGDIKGAFGNCWRQVLCESEVVEGKIPPGSPSVL--IV 113
S L+ ++ +S + L + +C V K SVL I+
Sbjct: 117 SCFLKANDLTHSLSSYL------------------KEICPKWVKLRKTHNEKKSVLMLIL 158
Query: 114 SPSALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNRV-NIASGTPSRIKKLID 172
SA+R++ L++ +KLF+KHIK+QEQ+ LL+ RV ++ GTP RIK+L+
Sbjct: 159 CSSAVRALELIRSLTAFKGDAKVMKLFAKHIKVQEQVKLLEKRVIHLGVGTPGRIKELVK 218
Query: 173 VEALGLSRLQVLVLDMH-PDVKGYSLFTLPQVR 204
+ L L+ L+ LV D + D K + +P++R
Sbjct: 219 QDGLHLNPLKFLVFDWNWRDQKLRRMMDIPEIR 251
>UniRef100_UPI000023DFD0 UPI000023DFD0 UniRef100 entry
Length = 265
Score = 63.5 bits (153), Expect = 4e-09
Identities = 38/91 (41%), Positives = 57/91 (61%), Gaps = 3/91 (3%)
Query: 106 GSPSVLIVSPSALRSIHLLKGFR-FMTKQCSAVKLFSKHIKLQEQISLL-KNRVNIASGT 163
GSP LIV+ + LR+ +++ R F K + KLF+KH K++EQ+ L ++R I+ GT
Sbjct: 136 GSPHTLIVTGAGLRAADIVRSMRKFQNKDNAIAKLFAKHFKIEEQVKFLGEHRTGISVGT 195
Query: 164 PSRIKKLIDVEALGLSRLQVLVLD-MHPDVK 193
P+R+ L+D AL L L+ LV+D H D K
Sbjct: 196 PARLMDLMDNGALSLDGLKRLVVDASHIDQK 226
>UniRef100_O94465 SPBC23G7.07c protein [Schizosaccharomyces pombe]
Length = 277
Score = 60.5 bits (145), Expect = 3e-08
Identities = 61/216 (28%), Positives = 100/216 (46%), Gaps = 25/216 (11%)
Query: 8 ENNDDRRKKKKKKTETTGSEQLRFFVNEFQSANDVQLSSIELESLKADSSILELSNSQNS 67
+NND +++K+KK E+ R + E Q N S I+ +D L+N S
Sbjct: 46 QNNDSAKREKRKKQRQKQKERKRAKLLELQDTN---ASIIQSPDTLSDL----LNNYIKS 98
Query: 68 RDTDL-DVKLLGGDIKGAF-------------GNCWRQVLCESEVVEGKIPPGSPSVLIV 113
+DL DV+L IK ++ N + + + +P +L++
Sbjct: 99 IYSDLTDVELSDKVIKASYIEDTISFSKPKTVDNYPEYIQHLPGFTKKVVQNSNPEILVL 158
Query: 114 SPSALRSIHLLKGFRFM-TKQCSAVKLFSKHIKLQEQISLLK-NRVNIASGTPSRIKKLI 171
SALR+I +LK + + K KLF KHI+L+E I+ K N++ + GT RI +L
Sbjct: 159 CISALRAIDVLKPTKSLQNKNFKVAKLFGKHIRLEEHINYCKANKIGVGIGTTPRIAQLT 218
Query: 172 DVEALGLSRLQVLVLD-MHPDVKGYSLFTLPQVRCA 206
+ E L+ ++LD D+K S+ T + R A
Sbjct: 219 N-ECFTCENLKYIILDYSFRDIKNNSILTSKESRKA 253
>UniRef100_Q5XH91 Hypothetical protein [Xenopus tropicalis]
Length = 740
Score = 52.4 bits (124), Expect = 9e-06
Identities = 37/109 (33%), Positives = 56/109 (50%), Gaps = 6/109 (5%)
Query: 96 SEVVEGKIPPGSPSVLIVSPSALRSIHLLKGFRFMTKQCSAVKL----FSKHIKLQEQIS 151
++V + K P +P LI+ PS + L + K + KL + +EQ++
Sbjct: 275 AQVSQAKNLPNAPKALIIEPSRELAEQTLNNVKQFKKYVDSPKLRELLIIGGVAAKEQLT 334
Query: 152 LLKNRVNIASGTPSRIKKLIDVEALGLSRLQVLVLDMHPDV--KGYSLF 198
LL+N V+I GTP RI LI L LS+++ LVLD + +GYS F
Sbjct: 335 LLENGVDIVVGTPGRIDDLISTGKLNLSQVRFLVLDEADGLLSQGYSDF 383
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.134 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 354,672,118
Number of Sequences: 2790947
Number of extensions: 13756064
Number of successful extensions: 55301
Number of sequences better than 10.0: 368
Number of HSP's better than 10.0 without gapping: 162
Number of HSP's successfully gapped in prelim test: 206
Number of HSP's that attempted gapping in prelim test: 55018
Number of HSP's gapped (non-prelim): 416
length of query: 227
length of database: 848,049,833
effective HSP length: 123
effective length of query: 104
effective length of database: 504,763,352
effective search space: 52495388608
effective search space used: 52495388608
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 72 (32.3 bits)
Medicago: description of AC149471.20


