

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149305.1 + phase: 0 /pseudo
(146 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
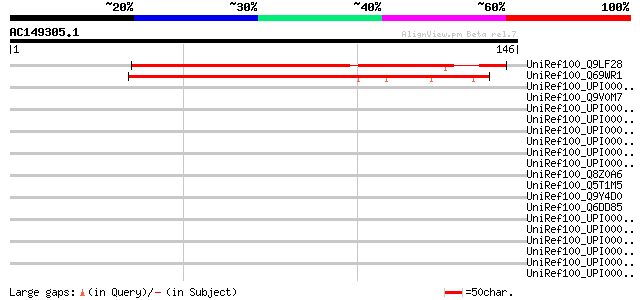
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LF28 Hypothetical protein T20K14_150 [Arabidopsis th... 107 4e-23
UniRef100_Q69WR1 Putative IDN3 protein isoform A [Oryza sativa] 96 1e-19
UniRef100_UPI000023E4A2 UPI000023E4A2 UniRef100 entry 35 0.35
UniRef100_Q9V0M7 Hypothetical protein [Pyrococcus abyssi] 35 0.45
UniRef100_UPI0000436A82 UPI0000436A82 UniRef100 entry 34 0.77
UniRef100_UPI000042EC8B UPI000042EC8B UniRef100 entry 34 0.77
UniRef100_UPI0000363B49 UPI0000363B49 UniRef100 entry 33 1.0
UniRef100_UPI0000363B48 UPI0000363B48 UniRef100 entry 33 1.0
UniRef100_UPI0000368F2B UPI0000368F2B UniRef100 entry 33 1.0
UniRef100_UPI000021AAB0 UPI000021AAB0 UniRef100 entry 33 1.0
UniRef100_Q8Z0A6 Serine/threonine kinase [Anabaena sp.] 33 1.0
UniRef100_Q5T1M5 KIAA0674 [Homo sapiens] 33 1.0
UniRef100_Q9Y4D0 KIAA0674 protein [Homo sapiens] 33 1.0
UniRef100_Q6DD85 KIAA0674 protein [Homo sapiens] 33 1.0
UniRef100_UPI000031C155 UPI000031C155 UniRef100 entry 33 1.3
UniRef100_UPI00002FD103 UPI00002FD103 UniRef100 entry 33 1.3
UniRef100_UPI00002E354D UPI00002E354D UniRef100 entry 33 1.3
UniRef100_UPI00002DECA4 UPI00002DECA4 UniRef100 entry 33 1.3
UniRef100_UPI00002D43C1 UPI00002D43C1 UniRef100 entry 33 1.3
UniRef100_UPI00002BDDFB UPI00002BDDFB UniRef100 entry 33 1.3
>UniRef100_Q9LF28 Hypothetical protein T20K14_150 [Arabidopsis thaliana]
Length = 1755
Score = 107 bits (268), Expect = 4e-23
Identities = 54/112 (48%), Positives = 78/112 (69%), Gaps = 13/112 (11%)
Query: 36 SEPPKPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIKR 95
+EP KPGD SRQS+ F++ E++ LP++ Q+L+QRYQEFKNA++EDTVD+++Y+ N+KR
Sbjct: 1642 TEPLKPGDPLSRQSVAFDLSETRTDLPSTYQDLVQRYQEFKNAMREDTVDFTIYSTNVKR 1701
Query: 96 KRPTQTPRKVRKSGPPMVGGDNDEDDDED----WAGGSRNISFSGGRRSSLR 143
KRP TPRK +S V + D+DDD++ W GG GGR ++ R
Sbjct: 1702 KRP--TPRKTSRSAKKTVAYNEDDDDDDNDDRGWHGG-------GGRGAARR 1744
>UniRef100_Q69WR1 Putative IDN3 protein isoform A [Oryza sativa]
Length = 764
Score = 96.3 bits (238), Expect = 1e-19
Identities = 51/114 (44%), Positives = 71/114 (61%), Gaps = 10/114 (8%)
Query: 35 VSEPPKPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIK 94
+ +PPK G+ S+Q+IP NI + SLP+ PQ+ + YQ+FK L+EDTVDY +YT + +
Sbjct: 643 LKDPPKSGETISKQNIPLNISNTNTSLPSCPQDAARVYQDFKTVLREDTVDYGMYTVSAQ 702
Query: 95 RKRPT-QTPRKVRK-----SGPPMVGGDNDED-DDEDWAGGSRNI---SFSGGR 138
+KRPT ++ +VR+ G GG DED DDEDW GG + S GGR
Sbjct: 703 KKRPTPRSSSRVRRPAAVTRGRGGGGGGGDEDTDDEDWTGGGARVLDFSAQGGR 756
>UniRef100_UPI000023E4A2 UPI000023E4A2 UniRef100 entry
Length = 460
Score = 35.0 bits (79), Expect = 0.35
Identities = 22/73 (30%), Positives = 33/73 (45%), Gaps = 11/73 (15%)
Query: 82 DTVDYSLYTANIKRKRPTQTPRKVRKSGPPMVGGDNDED-----DDEDWAGGSR------ 130
D+ + T N KRK+ K P GD+D+D DD+D A G
Sbjct: 75 DSENEDFNTRNEKRKKEEADAAKSGSKAPGTTAGDDDDDDMFAVDDDDAANGDNNVPEGD 134
Query: 131 NISFSGGRRSSLR 143
N+ SGG+++ +R
Sbjct: 135 NVEESGGKKNKVR 147
>UniRef100_Q9V0M7 Hypothetical protein [Pyrococcus abyssi]
Length = 602
Score = 34.7 bits (78), Expect = 0.45
Identities = 21/76 (27%), Positives = 36/76 (46%), Gaps = 9/76 (11%)
Query: 53 NIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIKRKRPTQTPRKVRKSGPPM 112
N+ E F T EL++ +E N +E V+ + K P + PR + PP
Sbjct: 497 NLLEEYFEQGTFDVELLEELEEKDNLFQEVPVEAYI------SKEPPEVPRYIE---PPE 547
Query: 113 VGGDNDEDDDEDWAGG 128
V +N+E++ +W+ G
Sbjct: 548 VPDNNEEEEKGEWSSG 563
>UniRef100_UPI0000436A82 UPI0000436A82 UniRef100 entry
Length = 448
Score = 33.9 bits (76), Expect = 0.77
Identities = 21/56 (37%), Positives = 31/56 (54%), Gaps = 6/56 (10%)
Query: 90 TANIKRKRPTQTPRKVRKSGPPMVGGDND-----EDDDEDWAGGSRNISFS-GGRR 139
+A K + P++ R RK+GPP+ G+ D ED+DE+ GS N+ GRR
Sbjct: 357 SAAAKEETPSKGRRGGRKAGPPVEEGNEDEQQEEEDEDEEEDAGSENMEVKPKGRR 412
>UniRef100_UPI000042EC8B UPI000042EC8B UniRef100 entry
Length = 796
Score = 33.9 bits (76), Expect = 0.77
Identities = 28/85 (32%), Positives = 36/85 (41%), Gaps = 17/85 (20%)
Query: 74 EFKNALKEDTVDYSLYTANIKRKRPTQTP----------RKVRKSGPPMVGGDNDEDDDE 123
+F A E V IK+KR Q+P V S P GD+D+DDDE
Sbjct: 575 DFDEADDEGVVRQEEEGRAIKKKRQRQSPPVWVTRTGATTGVGSSSSPFSIGDDDDDDDE 634
Query: 124 DWAG-------GSRNISFSGGRRSS 141
D G G R++SF+ SS
Sbjct: 635 DGEGVKNRETQGDRSMSFNHPDNSS 659
>UniRef100_UPI0000363B49 UPI0000363B49 UniRef100 entry
Length = 714
Score = 33.5 bits (75), Expect = 1.0
Identities = 28/105 (26%), Positives = 46/105 (43%), Gaps = 12/105 (11%)
Query: 43 DVFSRQSIPFNIGESQFSLPTSPQELIQRYQEF---KNALKEDTVDYSLYTANIKRKRPT 99
++ + S+P G Q TSP ++QR F N K + A + R R +
Sbjct: 326 NIMGKPSVPKGGGPPQV---TSPNPIVQRLPAFLDNHNYAKSPMQEEEDIAAGVGRTRMS 382
Query: 100 QTPRKVRKSGPPMVGGDNDEDDDEDWAGGSRNISFSGGRRSSLRN 144
P+ PP ++D +D+E+ GS S R++SLR+
Sbjct: 383 APPQ------PPYSDDEDDYEDEEEEVTGSAVTSNRFRRKASLRS 421
>UniRef100_UPI0000363B48 UPI0000363B48 UniRef100 entry
Length = 682
Score = 33.5 bits (75), Expect = 1.0
Identities = 28/105 (26%), Positives = 46/105 (43%), Gaps = 12/105 (11%)
Query: 43 DVFSRQSIPFNIGESQFSLPTSPQELIQRYQEF---KNALKEDTVDYSLYTANIKRKRPT 99
++ + S+P G Q TSP ++QR F N K + A + R R +
Sbjct: 326 NIMGKPSVPKGGGPPQV---TSPNPIVQRLPAFLDNHNYAKSPMQEEEDIAAGVGRTRMS 382
Query: 100 QTPRKVRKSGPPMVGGDNDEDDDEDWAGGSRNISFSGGRRSSLRN 144
P+ PP ++D +D+E+ GS S R++SLR+
Sbjct: 383 APPQ------PPYSDDEDDYEDEEEEVTGSAVTSNRFRRKASLRS 421
>UniRef100_UPI0000368F2B UPI0000368F2B UniRef100 entry
Length = 523
Score = 33.5 bits (75), Expect = 1.0
Identities = 26/101 (25%), Positives = 40/101 (38%), Gaps = 10/101 (9%)
Query: 37 EPP--KPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIK 94
+PP K DV S + ++ + L + +++L D D A +K
Sbjct: 423 QPPQLKKDDVTSSTGPHKELSSTEAGSTVAGAALRPSHHSQRSSLSGDEEDELFKGATLK 482
Query: 95 RKRPTQTPRKVRKSGPPMVG--------GDNDEDDDEDWAG 127
RP P + + M G GD+D+DDD DW G
Sbjct: 483 ALRPKAQPEEEDEDEVSMKGRPPPTPLFGDDDDDDDIDWLG 523
>UniRef100_UPI000021AAB0 UPI000021AAB0 UniRef100 entry
Length = 423
Score = 33.5 bits (75), Expect = 1.0
Identities = 15/40 (37%), Positives = 23/40 (57%), Gaps = 2/40 (5%)
Query: 89 YTANIKRKRPTQTPRKVRKSGPPMVGGDNDEDDDEDWAGG 128
Y N+ +K +Q P ++ P+ G D D DDD++ AGG
Sbjct: 8 YGLNLSKKSGSQKPNPAKRR--PVFGDDGDSDDDDNRAGG 45
>UniRef100_Q8Z0A6 Serine/threonine kinase [Anabaena sp.]
Length = 553
Score = 33.5 bits (75), Expect = 1.0
Identities = 14/43 (32%), Positives = 23/43 (52%)
Query: 62 PTSPQELIQRYQEFKNALKEDTVDYSLYTANIKRKRPTQTPRK 104
PT+ Q ++QR ++ K + D + YS +K P Q+P K
Sbjct: 275 PTNTQGILQRLRDIKQQVNRDEIPYSEQQTQLKSHLPLQSPTK 317
>UniRef100_Q5T1M5 KIAA0674 [Homo sapiens]
Length = 1244
Score = 33.5 bits (75), Expect = 1.0
Identities = 26/101 (25%), Positives = 40/101 (38%), Gaps = 10/101 (9%)
Query: 37 EPP--KPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIK 94
+PP K DV S + ++ + L + +++L D D A +K
Sbjct: 1144 QPPQLKKDDVTSSTGPHKELSSTEAGSTVAGAALRPSHHSQRSSLSGDEEDELFKGATLK 1203
Query: 95 RKRPTQTPRKVRKSGPPMVG--------GDNDEDDDEDWAG 127
RP P + + M G GD+D+DDD DW G
Sbjct: 1204 ALRPKAQPEEEDEDEVSMKGRPPPTPLFGDDDDDDDIDWLG 1244
>UniRef100_Q9Y4D0 KIAA0674 protein [Homo sapiens]
Length = 1234
Score = 33.5 bits (75), Expect = 1.0
Identities = 26/101 (25%), Positives = 40/101 (38%), Gaps = 10/101 (9%)
Query: 37 EPP--KPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIK 94
+PP K DV S + ++ + L + +++L D D A +K
Sbjct: 1134 QPPQLKKDDVTSSTGPHKELSSTEAGSTVAGAALRPSHHSQRSSLSGDEEDELFKGATLK 1193
Query: 95 RKRPTQTPRKVRKSGPPMVG--------GDNDEDDDEDWAG 127
RP P + + M G GD+D+DDD DW G
Sbjct: 1194 ALRPKAQPEEEDEDEVSMKGRPPPTPLFGDDDDDDDIDWLG 1234
>UniRef100_Q6DD85 KIAA0674 protein [Homo sapiens]
Length = 662
Score = 33.5 bits (75), Expect = 1.0
Identities = 26/101 (25%), Positives = 40/101 (38%), Gaps = 10/101 (9%)
Query: 37 EPP--KPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIK 94
+PP K DV S + ++ + L + +++L D D A +K
Sbjct: 562 QPPQLKKDDVTSSTGPHKELSSTEAGSTVAGAALRPSHHSQRSSLSGDEEDELFKGATLK 621
Query: 95 RKRPTQTPRKVRKSGPPMVG--------GDNDEDDDEDWAG 127
RP P + + M G GD+D+DDD DW G
Sbjct: 622 ALRPKAQPEEEDEDEVSMKGRPPPTPLFGDDDDDDDIDWLG 662
>UniRef100_UPI000031C155 UPI000031C155 UniRef100 entry
Length = 284
Score = 33.1 bits (74), Expect = 1.3
Identities = 14/39 (35%), Positives = 23/39 (58%)
Query: 41 PGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNAL 79
P D SRQ PF +G+ + + Q+++QRY E K+ +
Sbjct: 205 PLDSTSRQLDPFVVGDDHYQVAQGVQKILQRYNELKDII 243
>UniRef100_UPI00002FD103 UPI00002FD103 UniRef100 entry
Length = 465
Score = 33.1 bits (74), Expect = 1.3
Identities = 14/39 (35%), Positives = 23/39 (58%)
Query: 41 PGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNAL 79
P D SRQ PF +G+ + + Q+++QRY E K+ +
Sbjct: 342 PLDSTSRQLDPFVVGDEHYEVAQGVQKILQRYNELKDII 380
>UniRef100_UPI00002E354D UPI00002E354D UniRef100 entry
Length = 303
Score = 33.1 bits (74), Expect = 1.3
Identities = 14/39 (35%), Positives = 23/39 (58%)
Query: 41 PGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNAL 79
P D SRQ PF +G+ + + Q+++QRY E K+ +
Sbjct: 180 PLDSTSRQLDPFVVGDEHYEVAQGVQKILQRYNELKDII 218
>UniRef100_UPI00002DECA4 UPI00002DECA4 UniRef100 entry
Length = 323
Score = 33.1 bits (74), Expect = 1.3
Identities = 14/39 (35%), Positives = 23/39 (58%)
Query: 41 PGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNAL 79
P D SRQ PF +G+ + + Q+++QRY E K+ +
Sbjct: 200 PLDSTSRQLDPFVVGDDHYEVAQGVQKILQRYNELKDII 238
>UniRef100_UPI00002D43C1 UPI00002D43C1 UniRef100 entry
Length = 461
Score = 33.1 bits (74), Expect = 1.3
Identities = 14/39 (35%), Positives = 23/39 (58%)
Query: 41 PGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNAL 79
P D SRQ PF +G+ + + Q+++QRY E K+ +
Sbjct: 338 PLDSTSRQLDPFVVGDDHYEVAQGVQKILQRYNELKDII 376
>UniRef100_UPI00002BDDFB UPI00002BDDFB UniRef100 entry
Length = 364
Score = 33.1 bits (74), Expect = 1.3
Identities = 14/39 (35%), Positives = 23/39 (58%)
Query: 41 PGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNAL 79
P D SRQ PF +G+ + + Q+++QRY E K+ +
Sbjct: 283 PLDSTSRQLDPFVVGDDHYEVAQGVQKILQRYNELKDII 321
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.329 0.143 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 255,646,955
Number of Sequences: 2790947
Number of extensions: 10545052
Number of successful extensions: 56813
Number of sequences better than 10.0: 122
Number of HSP's better than 10.0 without gapping: 66
Number of HSP's successfully gapped in prelim test: 59
Number of HSP's that attempted gapping in prelim test: 56680
Number of HSP's gapped (non-prelim): 146
length of query: 146
length of database: 848,049,833
effective HSP length: 122
effective length of query: 24
effective length of database: 507,554,299
effective search space: 12181303176
effective search space used: 12181303176
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC149305.1


