

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149293.2 - phase: 2
(212 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
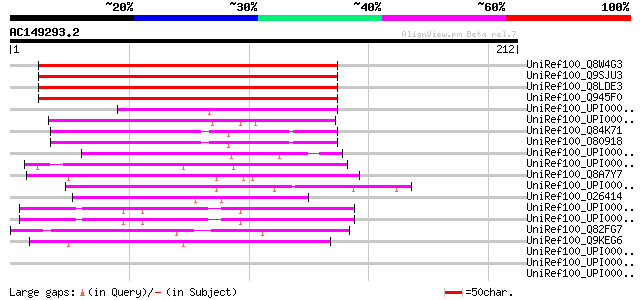
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8W4G3 Hypothetical protein At2g21340; F3K23.10 [Arabi... 204 9e-52
UniRef100_Q9SJU3 Expressed protein [Arabidopsis thaliana] 204 9e-52
UniRef100_Q8LDE3 Enhanced disease susceptibility 5 [Arabidopsis ... 204 9e-52
UniRef100_Q945F0 Enhanced disease susceptibility 5 [Arabidopsis ... 179 6e-44
UniRef100_UPI0000345ADA UPI0000345ADA UniRef100 entry 62 1e-08
UniRef100_UPI00002A7B90 UPI00002A7B90 UniRef100 entry 58 1e-07
UniRef100_Q84K71 Hypothetical protein At2g38330 [Arabidopsis tha... 54 2e-06
UniRef100_O80918 Hypothetical protein At2g38330 [Arabidopsis tha... 54 2e-06
UniRef100_UPI00002F40EF UPI00002F40EF UniRef100 entry 50 3e-05
UniRef100_UPI0000332C91 UPI0000332C91 UniRef100 entry 49 7e-05
UniRef100_Q8A7Y7 Putative Na+-driven multidrug efflux pump [Bact... 49 9e-05
UniRef100_UPI00002E7CB9 UPI00002E7CB9 UniRef100 entry 48 1e-04
UniRef100_O26414 Conserved protein [Methanobacterium thermoautot... 48 1e-04
UniRef100_UPI000033468A UPI000033468A UniRef100 entry 47 3e-04
UniRef100_UPI00002C797B UPI00002C797B UniRef100 entry 47 3e-04
UniRef100_Q82FG7 Putative DNA-damage-inducible protein F [Strept... 47 3e-04
UniRef100_Q9KEG6 BH0886 protein [Bacillus halodurans] 47 3e-04
UniRef100_UPI00002CDB44 UPI00002CDB44 UniRef100 entry 45 0.001
UniRef100_UPI000025ACE3 UPI000025ACE3 UniRef100 entry 45 0.001
UniRef100_UPI0000297230 UPI0000297230 UniRef100 entry 45 0.001
>UniRef100_Q8W4G3 Hypothetical protein At2g21340; F3K23.10 [Arabidopsis thaliana]
Length = 559
Score = 204 bits (520), Expect = 9e-52
Identities = 99/125 (79%), Positives = 110/125 (87%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
L GWVAQSASLGMKDSWGPLKALA AS INGVGD+VLCT+LGYGIAGAAWATM SQVVAA
Sbjct: 252 LIGWVAQSASLGMKDSWGPLKALAVASAINGVGDVVLCTFLGYGIAGAAWATMVSQVVAA 311
Query: 73 YMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHT 132
YMMM LN KGY+AF+ +PS E +TI GLAAPVF+TMMSKV FY+LL+YFATSMGT+
Sbjct: 312 YMMMDALNKKGYSAFSFCVPSPSELLTIFGLAAPVFITMMSKVLFYTLLVYFATSMGTNI 371
Query: 133 MAAHQ 137
+AAHQ
Sbjct: 372 IAAHQ 376
>UniRef100_Q9SJU3 Expressed protein [Arabidopsis thaliana]
Length = 555
Score = 204 bits (520), Expect = 9e-52
Identities = 99/125 (79%), Positives = 110/125 (87%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
L GWVAQSASLGMKDSWGPLKALA AS INGVGD+VLCT+LGYGIAGAAWATM SQVVAA
Sbjct: 248 LIGWVAQSASLGMKDSWGPLKALAVASAINGVGDVVLCTFLGYGIAGAAWATMVSQVVAA 307
Query: 73 YMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHT 132
YMMM LN KGY+AF+ +PS E +TI GLAAPVF+TMMSKV FY+LL+YFATSMGT+
Sbjct: 308 YMMMDALNKKGYSAFSFCVPSPSELLTIFGLAAPVFITMMSKVLFYTLLVYFATSMGTNI 367
Query: 133 MAAHQ 137
+AAHQ
Sbjct: 368 IAAHQ 372
>UniRef100_Q8LDE3 Enhanced disease susceptibility 5 [Arabidopsis thaliana]
Length = 555
Score = 204 bits (520), Expect = 9e-52
Identities = 99/125 (79%), Positives = 110/125 (87%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
L GWVAQSASLGMKDSWGPLKALA AS INGVGD+VLCT+LGYGIAGAAWATM SQVVAA
Sbjct: 248 LIGWVAQSASLGMKDSWGPLKALAVASAINGVGDVVLCTFLGYGIAGAAWATMVSQVVAA 307
Query: 73 YMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHT 132
YMMM LN KGY+AF+ +PS E +TI GLAAPVF+TMMSKV FY+LL+YFATSMGT+
Sbjct: 308 YMMMDALNKKGYSAFSFCVPSPSELLTIFGLAAPVFITMMSKVLFYTLLVYFATSMGTNI 367
Query: 133 MAAHQ 137
+AAHQ
Sbjct: 368 IAAHQ 372
>UniRef100_Q945F0 Enhanced disease susceptibility 5 [Arabidopsis thaliana]
Length = 543
Score = 179 bits (453), Expect = 6e-44
Identities = 86/125 (68%), Positives = 106/125 (84%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
L G VAQSASLGMK+SWGPLKALAAA++ING+GD +LC +LG GIAGAAWAT ASQ+V+A
Sbjct: 235 LVGLVAQSASLGMKNSWGPLKALAAATIINGLGDTILCLFLGQGIAGAAWATTASQIVSA 294
Query: 73 YMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHT 132
YMMM +LN +GYNA++ +IPS +E I LAAPVF+++ SK+AFYS +IY ATSMGTH
Sbjct: 295 YMMMDSLNKEGYNAYSFAIPSPQELWKISALAAPVFISIFSKIAFYSFIIYCATSMGTHV 354
Query: 133 MAAHQ 137
+AAHQ
Sbjct: 355 LAAHQ 359
>UniRef100_UPI0000345ADA UPI0000345ADA UniRef100 entry
Length = 265
Score = 62.0 bits (149), Expect = 1e-08
Identities = 33/96 (34%), Positives = 55/96 (56%), Gaps = 4/96 (4%)
Query: 46 DIVLCTYLGYGIAGAAWATMASQVVAAYMMMRTLNMK----GYNAFALSIPSGREFITIL 101
D++L + LGYGI GAAWAT+ S++ A +++R + K A +P+ T
Sbjct: 6 DVLLVSVLGYGIGGAAWATVVSEIACAGIVVRAVLTKIQSPPSRPRASLLPTREAVATYT 65
Query: 102 GLAAPVFMTMMSKVAFYSLLIYFATSMGTHTMAAHQ 137
A P+ +T++ K+A YS L + AT++ + AAH+
Sbjct: 66 AFAKPLVLTLIGKIATYSSLAHVATTVSVTSTAAHR 101
>UniRef100_UPI00002A7B90 UPI00002A7B90 UniRef100 entry
Length = 422
Score = 58.2 bits (139), Expect = 1e-07
Identities = 46/156 (29%), Positives = 71/156 (45%), Gaps = 36/156 (23%)
Query: 17 VAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAAYMMM 76
V+Q+A L ++D W PL+A++ +V+N V D+ LG+GIAGAA AT SQV+ +++
Sbjct: 76 VSQAAFLAVRDPWTPLRAVSLTTVLNLVLDVWFVCGLGWGIAGAAGATSLSQVITMSLLI 135
Query: 77 RTLNMKG--------------------------------YNAFALSIPSGR---EFITIL 101
R L +G A AL +P R +F++ L
Sbjct: 136 RALVKRGPEIDKVKEMVAEATERSKSTSFAENKSRTLMNVGAPALRLPFQRPRDDFLSRL 195
Query: 102 -GLAAPVFMTMMSKVAFYSLLIYFATSMGTHTMAAH 136
+A PV M K F ++ TS+ AA+
Sbjct: 196 SSIAGPVMMVAAIKCVFVGWVVRAGTSISPEASAAN 231
>UniRef100_Q84K71 Hypothetical protein At2g38330 [Arabidopsis thaliana]
Length = 521
Score = 54.3 bits (129), Expect = 2e-06
Identities = 44/122 (36%), Positives = 62/122 (50%), Gaps = 6/122 (4%)
Query: 18 AQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAAYMMMR 77
AQ A G KD+ PL A+ A +V+N V D +L LG+GI+GAA AT+ S+ + A++++
Sbjct: 228 AQGAFRGFKDTTTPLYAVVAGNVLNAVLDPILIFVLGFGISGAAAATVISEYLIAFILLW 287
Query: 78 TLNMKGYNAFALS--IPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHTMAA 135
LN N LS I GR + + T+ V F +L A G MA
Sbjct: 288 KLN---ENVVLLSPQIKVGRANQYLKSGGLLIGRTVALLVPF-TLATSLAAQNGPTQMAG 343
Query: 136 HQ 137
HQ
Sbjct: 344 HQ 345
>UniRef100_O80918 Hypothetical protein At2g38330 [Arabidopsis thaliana]
Length = 539
Score = 54.3 bits (129), Expect = 2e-06
Identities = 44/122 (36%), Positives = 62/122 (50%), Gaps = 6/122 (4%)
Query: 18 AQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAAYMMMR 77
AQ A G KD+ PL A+ A +V+N V D +L LG+GI+GAA AT+ S+ + A++++
Sbjct: 228 AQGAFRGFKDTTTPLYAVVAGNVLNAVLDPILIFVLGFGISGAAAATVISEYLIAFILLW 287
Query: 78 TLNMKGYNAFALS--IPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHTMAA 135
LN N LS I GR + + T+ V F +L A G MA
Sbjct: 288 KLN---ENVVLLSPQIKVGRANQYLKSGGLLIGRTVALLVPF-TLATSLAAQNGPTQMAG 343
Query: 136 HQ 137
HQ
Sbjct: 344 HQ 345
>UniRef100_UPI00002F40EF UPI00002F40EF UniRef100 entry
Length = 420
Score = 50.4 bits (119), Expect = 3e-05
Identities = 33/117 (28%), Positives = 58/117 (49%), Gaps = 12/117 (10%)
Query: 31 PLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAAYMMMRTLNMKGYNAFALS 90
P+ +V+N + D + YL YG+ GAAWAT SQ++ + + L +K +
Sbjct: 139 PMIVAGLGTVLNTILDPIFIFYLDYGVTGAAWATTISQIIVFCIFIYMLFIKHHTYIRFK 198
Query: 91 I----PSGREFITILGLAAPVFMTM----MSKVAFYSLLIYFATSMGTHTMAAHQYG 139
+ PS I+ + PV M+M + ++ F LL+ ++ T+ +AA+Q G
Sbjct: 199 LKDFSPSSFIIYDIIKVGIPVSMSMVVMAIGQLVFNRLLVNYS----TNAVAAYQIG 251
>UniRef100_UPI0000332C91 UPI0000332C91 UniRef100 entry
Length = 362
Score = 49.3 bits (116), Expect = 7e-05
Identities = 37/141 (26%), Positives = 64/141 (45%), Gaps = 11/141 (7%)
Query: 7 PAGL--LYLCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWAT 64
PA L L L GW+ LG+ GP L + +N V DI +L + +AGAAWA+
Sbjct: 129 PAALCNLVLLGWM-----LGVHYGRGPFYLLLVTNSVNIVLDIYFVVFLDWAVAGAAWAS 183
Query: 65 MASQVVA-AYMMMRTLNMKGYNAFALSIP---SGREFITILGLAAPVFMTMMSKVAFYSL 120
+ + A + + + LS P S ++ +L L +F+ + +S
Sbjct: 184 LIADYTALVFALFLVTKLAKKQGVVLSTPHWFSFKKMAGLLSLNRDIFIRSLILQLCFSF 243
Query: 121 LIYFATSMGTHTMAAHQYGVN 141
+ ++ +G T+AA+ +N
Sbjct: 244 MTFYGARIGETTLAANAVLLN 264
>UniRef100_Q8A7Y7 Putative Na+-driven multidrug efflux pump [Bacteroides
thetaiotaomicron]
Length = 454
Score = 48.9 bits (115), Expect = 9e-05
Identities = 45/170 (26%), Positives = 73/170 (42%), Gaps = 31/170 (18%)
Query: 8 AGLLYLCGWVAQSASL-GMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMA 66
AG+L++ G+ L G+ DS PL +A A VIN V D +L Y G GAA AT+
Sbjct: 141 AGILFIVGYNVVCGILRGLGDSKTPLYFVALACVINIVLDFILVGYFHLGATGAAVATIT 200
Query: 67 SQVVAAYMMMRTLNMKGYN-----------------AFALSIPSGRE---------FITI 100
+Q V+ + + L G++ L P + IT+
Sbjct: 201 AQGVSFMISLWFLYRHGFHFEFTRKDIRLNRNLAKKVMTLGAPIALQDALINISFLIITV 260
Query: 101 ----LGLAAPVFMTMMSKVAFYSLLIYFATSMGTHTMAAHQYGVNLSRKL 146
+G+ A + ++ K+ +++L A S TM A YG L +++
Sbjct: 261 IVNQMGVIASASLGVVEKIIVFAMLPPMAISSAVATMTAQNYGAGLIQRM 310
>UniRef100_UPI00002E7CB9 UPI00002E7CB9 UniRef100 entry
Length = 290
Score = 48.1 bits (113), Expect = 1e-04
Identities = 48/184 (26%), Positives = 81/184 (43%), Gaps = 40/184 (21%)
Query: 24 GMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAAYMMMRTLNMKG 83
G+++ + A+ ++IN +G + G+ GAA++T+ S V +++ TL K
Sbjct: 8 GLQEIKKAMFAIFIMTLINALGSYFSVMHWNLGLKGAAYSTVLSFFVCDVLLILTLLKKS 67
Query: 84 YN--AFALSIPSGREFITILGLAAPVFM-TMMSKVAFYSLLIYFATSMGTHTMAAHQYGV 140
N F S S F+++ A + + T +AF+ LLI F+T +G+ T+AAHQ +
Sbjct: 68 KNIGLFKRSYESVETFLSLGNDALHLTVRTAFLSLAFF-LLIVFSTRLGSQTLAAHQIIL 126
Query: 141 NL------------------------SRKLLNHLCLNCYMELIG------------ICQR 164
L RKL +HL L+ + +G +CQR
Sbjct: 127 QLWLLASFTMDGFAVTATALGAKLIGQRKLKDHLILSRRLVFLGFLMGVFVSLFYFVCQR 186
Query: 165 PGCF 168
P F
Sbjct: 187 PILF 190
>UniRef100_O26414 Conserved protein [Methanobacterium thermoautotrophicum]
Length = 452
Score = 48.1 bits (113), Expect = 1e-04
Identities = 32/116 (27%), Positives = 61/116 (52%), Gaps = 17/116 (14%)
Query: 27 DSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAAYMMM------RTLN 80
D+ P+ A+ ++V+N V D +L G+GIAGAAWAT+ SQ++ + +++ RT
Sbjct: 162 DARRPMYAMGLSAVLNMVLDPILIYTAGWGIAGAAWATVISQLLVSLLIIYWFLAGRTFT 221
Query: 81 MKGYNAF-----------ALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFA 125
G++ F +++IP+ EF+ + + A + + S ++ +Y A
Sbjct: 222 SIGWSHFRADRGVVWSILSVTIPASAEFLVMSMVTALLNWILTSVAGTSAVAVYSA 277
>UniRef100_UPI000033468A UPI000033468A UniRef100 entry
Length = 276
Score = 47.4 bits (111), Expect = 3e-04
Identities = 39/151 (25%), Positives = 73/151 (47%), Gaps = 18/151 (11%)
Query: 5 SIPAGLLYLCGWVAQSASLGMKDSWGPLKALAAASVINGVGD-IVLCTYLG---YGIAGA 60
S+ GL+ + A S G S + ++ ++N +G+ I L + G YG+ G
Sbjct: 23 SLSIGLVMNIAFAAILRSYGFTRS--AMLVTSSTGLMNVLGNYIALYSPFGLPVYGVTGV 80
Query: 61 AWATMASQVVAAYMMMRTLNMKGYNAFALSIPSGR-------EFITILGLAAPVFMTMMS 113
A +T+ SQ++ A +M+ + KG + +P R + +++ + M+S
Sbjct: 81 AISTVTSQIIGALIMLAVIRHKG-----IPLPMPRLKLLPRSTYWSVMRIGLLNAGEMLS 135
Query: 114 KVAFYSLLIYFATSMGTHTMAAHQYGVNLSR 144
+IYF + MGT ++ A+ YG+N+SR
Sbjct: 136 YNVAQMTIIYFISQMGTLSLTAYTYGLNISR 166
>UniRef100_UPI00002C797B UPI00002C797B UniRef100 entry
Length = 455
Score = 47.4 bits (111), Expect = 3e-04
Identities = 39/151 (25%), Positives = 73/151 (47%), Gaps = 18/151 (11%)
Query: 5 SIPAGLLYLCGWVAQSASLGMKDSWGPLKALAAASVINGVGD-IVLCTYLG---YGIAGA 60
S+ GL+ + A S G S + ++ ++N +G+ I L + G YG+ G
Sbjct: 141 SLSIGLVMNIAFAAILRSYGFTRS--AMLVTSSTGLMNVLGNYIALYSPFGLPVYGVTGV 198
Query: 61 AWATMASQVVAAYMMMRTLNMKGYNAFALSIPSGR-------EFITILGLAAPVFMTMMS 113
A +T+ SQ++ A +M+ + KG + +P R + +++ + M+S
Sbjct: 199 AISTVTSQIIGALIMLAVIRHKG-----IPLPMPRLKLLPRSTYWSVMRIGLLNAGEMLS 253
Query: 114 KVAFYSLLIYFATSMGTHTMAAHQYGVNLSR 144
+IYF + MGT ++ A+ YG+N+SR
Sbjct: 254 YNVAQMTIIYFISQMGTLSLTAYTYGLNISR 284
>UniRef100_Q82FG7 Putative DNA-damage-inducible protein F [Streptomyces avermitilis]
Length = 448
Score = 47.4 bits (111), Expect = 3e-04
Identities = 44/149 (29%), Positives = 67/149 (44%), Gaps = 17/149 (11%)
Query: 1 VDSRSIPAGLLYLCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGA 60
+ + IPA L+ L A G++D+ PL A V N V ++VL G GIAG+
Sbjct: 144 ISALGIPAMLVVLA---ATGVLRGLQDTKTPLYVAVAGFVANAVLNVVLVYGAGLGIAGS 200
Query: 61 AWATMASQ----VVAAYMMMRTLNMKGYNAFALSIPSGREFITILGLA---APVFMTMMS 113
AW T+ +Q V Y+++R A L P + I A AP+ + +S
Sbjct: 201 AWGTVIAQYGMAVAYLYVVVR-------GARKLGAPLRPDIAGIRACAQAGAPLLVRTLS 253
Query: 114 KVAFYSLLIYFATSMGTHTMAAHQYGVNL 142
A + A +G +AAHQ ++L
Sbjct: 254 LRAVLMIATAVAARLGDADIAAHQIILSL 282
>UniRef100_Q9KEG6 BH0886 protein [Bacillus halodurans]
Length = 454
Score = 47.0 bits (110), Expect = 3e-04
Identities = 35/128 (27%), Positives = 62/128 (48%), Gaps = 2/128 (1%)
Query: 9 GLLYLCGWVAQSASL-GMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMAS 67
G+L+L G+ L M DS P++ + A ++N + D ++ GI GAA+AT+ S
Sbjct: 144 GILFLFGYNFIGTVLRAMGDSRSPVRFIFIAVILNIILDPLMIAGFNLGIDGAAYATIVS 203
Query: 68 QVVA-AYMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFAT 126
Q A Y ++ ++ G +P R F + L P ++M++ A + ++ T
Sbjct: 204 QGTAFIYGLIYSVRKAGVPFQIPYVPEKRYFFVLFKLGLPAGLSMITISAGVTAILSVVT 263
Query: 127 SMGTHTMA 134
S G +A
Sbjct: 264 SFGEEAVA 271
>UniRef100_UPI00002CDB44 UPI00002CDB44 UniRef100 entry
Length = 291
Score = 45.4 bits (106), Expect = 0.001
Identities = 30/133 (22%), Positives = 66/133 (49%), Gaps = 9/133 (6%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQ---- 68
L GW+ LG +++ GPL + +++N ++ LG+ +AGAAW ++ ++
Sbjct: 149 LVGWL-----LGTQNARGPLLIMLGTNLLNVALTLLFVLGLGWEVAGAAWGSVLAEWSGA 203
Query: 69 VVAAYMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSM 128
++ Y+ +T+ + ++ R + +L + +F+ M+ A + L+ T +
Sbjct: 204 LLGLYLARQTIKRQAGRINWPALRLWRNWRPLLAVNRDIFIRTMALQAAFFLITVQGTRL 263
Query: 129 GTHTMAAHQYGVN 141
G T+AA+ +N
Sbjct: 264 GDATVAANALLIN 276
>UniRef100_UPI000025ACE3 UPI000025ACE3 UniRef100 entry
Length = 295
Score = 45.4 bits (106), Expect = 0.001
Identities = 36/127 (28%), Positives = 58/127 (45%), Gaps = 8/127 (6%)
Query: 19 QSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVA--AYMMM 76
+S G D P+ +++N V D + L +G++GAAWAT SQ+V + M
Sbjct: 159 RSILAGEGDMKFPMYLFGLGTILNIVLDPIFIFLLDFGVSGAAWATTISQIVVFIIFTFM 218
Query: 77 RTLNMKGYNAFALS--IPSGREFITILGLAAPVFMTMMSKVAFYSLLIY--FATSMGTHT 132
+ Y F L PS + + I+ + P ++M+ V LIY + T
Sbjct: 219 LFVKEHAYVQFKLKDFSPSSKIILDIIKVGVPASISMI--VMALGQLIYNRMLATFSTDA 276
Query: 133 MAAHQYG 139
+AA+Q G
Sbjct: 277 VAAYQVG 283
>UniRef100_UPI0000297230 UPI0000297230 UniRef100 entry
Length = 299
Score = 45.1 bits (105), Expect = 0.001
Identities = 39/156 (25%), Positives = 72/156 (46%), Gaps = 18/156 (11%)
Query: 6 IPAGLLY--LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWA 63
+PA L L GW+ LG + + GPL L A++IN D++ L +G+AGAAWA
Sbjct: 106 LPASLATYALIGWL-----LGTQSARGPLAILMTANLINVALDLLFVLGLEWGVAGAAWA 160
Query: 64 TMASQ----VVAAYMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYS 119
++ ++ ++ ++ L + ++ + +L + +F+ ++ +
Sbjct: 161 SVIAEWSGALLGLWLARGALARYPGRLDSSALKRWSNWRPLLAVNRDIFIRTLALQLVFF 220
Query: 120 LLIYFATSMGTHTMAAHQYGVNLSRKLLNHLCLNCY 155
L+ T +G T+AA+ LLN L L Y
Sbjct: 221 LITVKGTRLGDATVAANAL-------LLNGLTLTAY 249
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.338 0.146 0.479
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 311,776,929
Number of Sequences: 2790947
Number of extensions: 10836750
Number of successful extensions: 44095
Number of sequences better than 10.0: 169
Number of HSP's better than 10.0 without gapping: 63
Number of HSP's successfully gapped in prelim test: 106
Number of HSP's that attempted gapping in prelim test: 43991
Number of HSP's gapped (non-prelim): 188
length of query: 212
length of database: 848,049,833
effective HSP length: 122
effective length of query: 90
effective length of database: 507,554,299
effective search space: 45679886910
effective search space used: 45679886910
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 72 (32.3 bits)
Medicago: description of AC149293.2


