

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149205.5 + phase: 0 /pseudo
(738 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
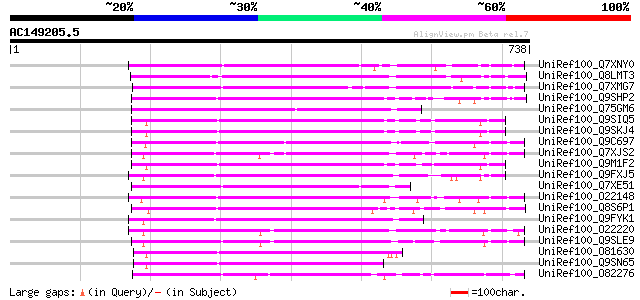
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XNY0 OSJNBb0015N08.11 protein [Oryza sativa] 306 2e-81
UniRef100_Q8LMT3 Putative retroelement [Oryza sativa] 271 5e-71
UniRef100_Q7XMG7 OSJNBa0028I23.15 protein [Oryza sativa] 258 6e-67
UniRef100_Q9SHP2 Putative non-LTR retroelement reverse transcrip... 234 5e-60
UniRef100_Q75GM6 Putative non-LTR retroelement reverse transcrip... 231 8e-59
UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcrip... 229 2e-58
UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcrip... 229 3e-58
UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [A... 228 5e-58
UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcrip... 228 5e-58
UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis tha... 227 1e-57
UniRef100_Q9FXJ5 F5A9.24 [Arabidopsis thaliana] 225 3e-57
UniRef100_Q7XE51 Putative non-LTR retroelement reverse transcrip... 224 5e-57
UniRef100_O22148 Putative non-LTR retroelement reverse transcrip... 224 9e-57
UniRef100_Q8S6P1 Putative reverse transcriptase [Oryza sativa] 223 1e-56
UniRef100_Q9FYK1 F21J9.30 [Arabidopsis thaliana] 223 2e-56
UniRef100_O22220 Putative non-LTR retroelement reverse transcrip... 221 5e-56
UniRef100_Q9SLE9 Putative non-LTR retroelement reverse transcrip... 221 8e-56
UniRef100_O81630 F8M12.22 protein [Arabidopsis thaliana] 218 7e-55
UniRef100_Q9SN65 Hypothetical protein F25I24.40 [Arabidopsis tha... 217 9e-55
UniRef100_O82276 Putative non-LTR retroelement reverse transcrip... 216 2e-54
>UniRef100_Q7XNY0 OSJNBb0015N08.11 protein [Oryza sativa]
Length = 1026
Score = 306 bits (783), Expect = 2e-81
Identities = 187/576 (32%), Positives = 296/576 (50%), Gaps = 40/576 (6%)
Query: 169 VKDMQILDGILIANEVVDEAKKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWR 228
+K+ I++G++I +EV++ + K+ +LFKVDFEKAYD V+ ++ +++ FP W
Sbjct: 352 LKNRFIMEGVVILHEVLNSIHQKKQSGILFKVDFEKAYDKVNWVFIYRMLKAKGFPDQWC 411
Query: 229 KWIKECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNL 288
WI + + ++ VN F H+GLRQGDPLSP LF +AAE L +L++ EN+L
Sbjct: 412 DWIMKVVMGGKVAVKVNDQIGSFFKTHKGLRQGDPLSPLLFNLAAEALTLLVQRAEENSL 471
Query: 289 FKGYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLV 348
+G N + + LQ+ADDT+ L + + L+ +L LF ++SGLK+NFNKS +
Sbjct: 472 IEGLGTNGDNKIAI--LQYADDTIFLINDKLDHAKNLKYILCLFEQLSGLKINFNKSEVF 529
Query: 349 GVNIGESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCF 408
+ +++ CKVG + YLG+ I W+ + ++ +L WQ
Sbjct: 530 CFGEAKEKQDLYSNIFTCKVGSLPLKYLGIPIDQKRILNKDWKLAENKMEHKLGCWQGRL 589
Query: 409 LSFGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSS 468
S GGRL+LL S L+S+P+Y +SF++ P G+ I+ +F W+ + RK + W
Sbjct: 590 QSIGGRLILLNSTLSSVPMYMISFYRLPKGVQERIDYFRKRFLWQEDQGIRKYHLVNWPL 649
Query: 469 VCLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEE----ARRLEVG 524
VC ++ GGLGV L N A++ KW WR L + G+W ++ A+Y + RL+ G
Sbjct: 650 VCSPRDQGGLGVLDLEAMNKAMLGKWIWR-LENEEGWWQEIIYAKYCSDKPLSGLRLKAG 708
Query: 525 GRSVSSWWKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFNRRFARL 584
S +W+ V ++KD F +K+VG+G TLFW D WLG P +F L
Sbjct: 709 S---SHFWQGVMEVKDD--------FFSFCTKIVGNGEKTLFWEDSWLGGKPLAIQFPSL 757
Query: 585 FDLTTNKMCTVADMYRLGWE--------EGGDAWVWRRRLWAWEEECRSLLDNVRLRLNV 636
+ + K T+AD+ R G + G WR+ + +WE + L N
Sbjct: 758 YGIVITKRITIADLNRKGIDCMKFRRDLHGDKLRDWRKIVNSWE--------GLNLVENC 809
Query: 637 LDRWQWDPDTIDVYTVRGVYQILTTSVSNTIVKTDELVWHKQVPLN*VSIVAWRLLKDRL 696
D+ W +TVR Y+ L ++ ++ +W +VPL + I W K+++
Sbjct: 810 KDKLWWTLSKDGKFTVRSFYRALKLQQTSF---PNKKIWKFRVPLK-IRIFIWFFTKNKI 865
Query: 697 PTKTNLQRRDCLQATVNTCVSGCGIAESASHLFLHC 732
TK NL +R + N C C E+ HLF C
Sbjct: 866 LTKDNLLKRGWRKGD-NKC-QFCDKVETVQHLFFDC 899
>UniRef100_Q8LMT3 Putative retroelement [Oryza sativa]
Length = 764
Score = 271 bits (693), Expect = 5e-71
Identities = 173/567 (30%), Positives = 284/567 (49%), Gaps = 31/567 (5%)
Query: 173 QILDGILIANEVVDEAKKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIK 232
Q LDG++I +EV+ E KK KK ++FK+DFEKAYD V +L V+ K F W +W+K
Sbjct: 92 QNLDGVVILHEVLHELKKEKKSGIIFKLDFEKAYDKVQWSFLFDVLHKKGFSDRWIQWVK 151
Query: 233 ECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGY 292
++ +NG D F +RG+RQGDPLSP LF + A+ L+ ++ + +
Sbjct: 152 MATIGGKMAVNINGEVKDFFKTYRGVRQGDPLSPLLFNLVADALSEMLNNAKQ--AVPHL 209
Query: 293 VVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLVGVNI 352
V G ++HLQ+ADDT+L + N+ ++ +L + MSGLK+N+ KS ++ +
Sbjct: 210 VPGG-----LTHLQYADDTILFMTNTEENIVTVKFLLYCYEAMSGLKINYQKSEIMVIGG 264
Query: 353 GESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCFLSFG 412
E A + NC+ GK+ F YLG+ I + A + + I+ RL+ W+ +LS+G
Sbjct: 265 DEMETQRVADLFNCQAGKMPFTYLGIPISMNKLTNADLDIPPNKIEKRLATWKCGYLSYG 324
Query: 413 GRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSVCLK 472
G+ +L+ S L+S+P+Y + + P G+ + ++S+ +FFW G E RK I W ++C
Sbjct: 325 GKAILINSCLSSIPLYMMGVYLLPEGVHNKMDSIRARFFWEGLEKKRKYHMIKWEALCRP 384
Query: 473 KEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARR-LEVGGRSVSSW 531
KEFGGLG R+ N+AL+ KW +RL + +L +Y ++ + S +
Sbjct: 385 KEFGGLGFIDTRKMNIALLCKWIYRLESGKEDPCCVLLRNKYMKDGGGFFQSKAEESSQF 444
Query: 532 WKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFNRRFARLFDLTTNK 591
WK + ++K W S VG+G T FW D W+G+ P ++ ++ + +K
Sbjct: 445 WKGLHEVKK--------WMDLGSSYKVGNGKATNFWSDVWIGETPLKTQYPNIYRMCADK 496
Query: 592 MCTVADMYRLG-WEEGGDAWVWRRRLWAWEEECRSLLDNVRLRLNVLDRWQ---WDPDTI 647
TV+ M G W + R L W + L N +++ + W
Sbjct: 497 EKTVSQMCLEGDWYIELRRSLGERDLNEWND-----LHNTPREIHLKEERDCIIWKLTKN 551
Query: 648 DVYTVRGVYQILTTSVSNTIVKTDELVWHKQVPLN*VSIVAWRLLKDRLPTKTNLQRRDC 707
Y + +YQ L+ V D +W +PL V I+ W +LK R+ L++
Sbjct: 552 GFYKAKTLYQALSFGGVKDKVMQD--LWRSSIPLK-VKILFWLMLKGRIQAAGQLKK--- 605
Query: 708 LQATVNTCVSGCGIAESASHLFLHCEV 734
++ + + CG E HL C +
Sbjct: 606 MKWSGSPNCKLCGQTEDVDHLMFRCHI 632
>UniRef100_Q7XMG7 OSJNBa0028I23.15 protein [Oryza sativa]
Length = 593
Score = 258 bits (658), Expect = 6e-67
Identities = 179/560 (31%), Positives = 276/560 (48%), Gaps = 27/560 (4%)
Query: 175 LDGILIANEVVDEAKKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIKEC 234
++G +I +E + E K KK+ ++ K+DFEKAYD VD +L ++ F W KWI +
Sbjct: 1 MEGAIILHETLHEMHKRKKDGVILKLDFEKAYDKVDWKFLQQTLRMKGFEPRWCKWIDQV 60
Query: 235 ISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGYVV 294
+ + ++ VN + F +GLRQGDPLSP LF + A+ L VL++ E+ G +
Sbjct: 61 VRGGSVAVKVNDEIGNFFQTKKGLRQGDPLSPLLFNLVADMLAVLIQRTSEHGKIHGVIP 120
Query: 295 GASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLVGVNIGE 354
+ + S LQ+ADDT+L E + + L+ VL+ F +SGLK+NF+KS L+
Sbjct: 121 HLVDDGL-SILQYADDTILFMEHDLEDAKNLKLVLSAFERLSGLKINFHKSELLCFGKAI 179
Query: 355 SWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCFLSFGGR 414
E A + CK G YLGL + W+ V + RL W+ LS GGR
Sbjct: 180 EVEREYALLFGCKTGSYTLKYLGLPMHYRKLSNKDWKEVEERFQKRLGSWKGKLLSVGGR 239
Query: 415 LVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSVCLKKE 474
LVL+ SVL+SL +Y LSFF+ P GII ++ ++FFW+ E +K WS +C KE
Sbjct: 240 LVLINSVLSSLAMYMLSFFEVPKGIIKKLDYYRSRFFWQSDEHKKKYRLARWSVLCKPKE 299
Query: 475 FGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLEV-GGRSVSSWWK 533
GGLG++ NL + +K R L W +L +Y + ++V R S +W
Sbjct: 300 CGGLGIQ-----NLEVQNKCLLRGLAKL--VWQNLLRKKYLAKKTIMQVQKQRGDSHFWT 352
Query: 534 EVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFNRRFARLFDLTTNKMC 593
+ +KD F S + DG FW D WLG+ + + L++L NK
Sbjct: 353 GLMGVKD--------TFSSFGSFKLQDGLQIRFWEDCWLGNQSLEKIYPSLYNLVRNKNV 404
Query: 594 TVAD-MYRLGWEEGGDAWVWRRRLWAWEEECRSLLDNVRLRLNVLDRWQWDPDTIDVYTV 652
VA + R+ + L AW E +++D + LR DR+ WD +TV
Sbjct: 405 VVAKVLERVPLNVSFRRAIIGDNLKAWHEIVANVVD-INLR-GQKDRFTWDASKNGTFTV 462
Query: 653 RGVYQILTTSVSNTIVKTDELVWHKQVPLN*VSIVAWRLLKDRLPTKTNLQRRDCLQATV 712
+Y+ L + ++ +W ++PL + I W L + + TK NL +R + ++
Sbjct: 463 NSMYKKL---MFGNVIPRQSFIWKLRLPLK-IKIFLWYLKEGVILTKDNLAKRR-WKGSL 517
Query: 713 NTCVSGCGIAESASHLFLHC 732
C C E+ +LF C
Sbjct: 518 RCCF--CRSHETIQYLFFDC 535
>UniRef100_Q9SHP2 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 773
Score = 234 bits (598), Expect = 5e-60
Identities = 179/591 (30%), Positives = 281/591 (47%), Gaps = 64/591 (10%)
Query: 174 ILDGILIANEVVDE--AKKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWI 231
I D ILIA+E++ KK + + K+D KA+D ++ G+++A+M++M F W WI
Sbjct: 2 ITDNILIAHELIHSLHTKKLVQPFVATKLDITKAFDKIEWGFIEAIMKQMGFSEKWCNWI 61
Query: 232 KECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKG 291
CI+TTT SIL+NG P RG+RQGDP+SP+L+++ EGL+ L+++ ++ G
Sbjct: 62 MTCITTTTYSILINGQPVRRIIPKRGIRQGDPISPYLYLLCTEGLSALIQASIKAKQLHG 121
Query: 292 YVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKS-LLVGV 350
+ + N +SHL FA D+L+ + + L VL L+ + SG VNF KS +L G
Sbjct: 122 FKA-SRNGPAISHLLFAHDSLVFCKATLEECMTLVNVLKLYEKASGQAVNFQKSAILFGK 180
Query: 351 NIGESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCFLS 410
+ + + +L + YLGL K + + T+ ++ W + LS
Sbjct: 181 GLDFRTSEQLSQLLGIYKTEGFGRYLGLPEFVGRNKTNAFSFIAQTMDQKMDNWYNKLLS 240
Query: 411 FGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSVC 470
G+ VL+KS++T++P Y++S F P +I I S F+W ++ KI W+AWS +
Sbjct: 241 PAGKEVLIKSIVTAIPTYSMSCFLLPMRLIHQITSAMRWFWWSNTKVKHKIPWVAWSKLN 300
Query: 471 LKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLEVGGRSVSS 530
K+ GGL +R L++FN+AL++K WR+L RV A+Y + R L+ S SS
Sbjct: 301 DPKKMGGLAIRDLKDFNIALLAKQSWRILQQPFSLMARVFKAKYFPKERLLDAKATSQSS 360
Query: 531 W-WKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFN--RRFARLFDL 587
+ WK I G G + + G+G+N W D WL P N R D
Sbjct: 361 YAWK---SILHGTKLISRG-----LKYIAGNGNNIQLWKDNWL---PLNPPRPPVGTCDS 409
Query: 588 TTNKMCTVADMYRLGWEEGGDAWVWRRRLWAWEEE--CRSLLDNVRLRLNVL-------- 637
+++ V+D+ G W E+ C+ + N + +
Sbjct: 410 IYSQL-KVSDLLIEG---------------RWNEDLLCKLIHQNDIPHIRAIRPSITGAN 453
Query: 638 DRWQWDPDTIDVYTVRGVYQIL-------------TTSVSNTIVKTDELVWHKQVPLN*V 684
D W Y+V+ Y +L VS V T+ +W + P +
Sbjct: 454 DAITWIYTHDGNYSVKSGYHLLRKLSQQQHASLPSPNEVSAQTVFTN--IWKQNAPPK-I 510
Query: 685 SIVAWRLLKDRLPTKTNLQRRDCLQATVNTCVSGCGIA-ESASHLFLHCEV 734
WR + LPT NL+RR + T +TC CG A E +HL C V
Sbjct: 511 KHFWWRSAHNALPTAGNLKRRRLI--TDDTC-QRCGEASEDVNHLLFQCRV 558
>UniRef100_Q75GM6 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 1614
Score = 231 bits (588), Expect = 8e-59
Identities = 134/412 (32%), Positives = 211/412 (50%), Gaps = 10/412 (2%)
Query: 174 ILDGILIANEVVDEAKKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIKE 233
IL+G +I +EV+ E + E ++ K+DFEKAYD V +L VM + FP W WIK
Sbjct: 1176 ILEGCVIIHEVLHEMNRKNLEGIILKIDFEKAYDKVSWDFLIEVMVRKGFPSKWVNWIKT 1235
Query: 234 CISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGYV 293
C+ I +NG TD F RGLRQGDPLSP LF + ++ L ++ S + G V
Sbjct: 1236 CVMGGRVCININGERTDFFRTFRGLRQGDPLSPLLFNLISDALAAMLDSAKREGVLSGLV 1295
Query: 294 VGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLVGVNIG 353
++HLQ+ADDT+L + A + +L F EM+ LKVN++KS + + +
Sbjct: 1296 PDIFPG-GITHLQYADDTVLFVANDDKQIVATKFILYCFEEMAVLKVNYHKSEIFTLGLS 1354
Query: 354 ESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCFLSFGG 413
++ A + NC VG+ YLGL IG D ++ + ++ RL+ W + LS G
Sbjct: 1355 DNDTNRVAMMFNCPVGQFPMKYLGLPIGPDKILNLGFDFLGQKLEKRLNSWGN-NLSHAG 1413
Query: 414 RLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSVCLKK 473
R V + + L+S+P YA+ F++ P G+ S+ +++W + K + W + K
Sbjct: 1414 RAVQINTCLSSIPSYAMCFYQLPEGVHQKFGSIRGRYYWARNRLKGKYHMVKWEDLAFPK 1473
Query: 474 EFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLEVGGRSVSSWWK 533
++GGLG R N AL++KW ++ + +L +Y ++ + R S +WK
Sbjct: 1474 DYGGLGFTETRRMNTALLAKWIMKIESEDDSLCIELLRRKYLQDGGFFQCKERYASQFWK 1533
Query: 534 EVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFNRRFARLF 585
+ I+ + +G V + VGDGS+ FW D W G P F +F
Sbjct: 1534 GLLNIRRWLS-------LGSVWQ-VGDGSHISFWRDVWWGQCPLRTLFPAIF 1577
>UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1524
Score = 229 bits (584), Expect = 2e-58
Identities = 168/549 (30%), Positives = 264/549 (47%), Gaps = 35/549 (6%)
Query: 174 ILDGILIANEVVDEAKKTK---KELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKW 230
I D ++IA+EV+ K K K + K D KAYD V+ +L+ M+ F W W
Sbjct: 702 INDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWIGW 761
Query: 231 IKECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFK 290
I + + S+L+NGSP + RG+RQGDPLSP+LF++ + L+ L+ + +
Sbjct: 762 IMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGDLR 821
Query: 291 GYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLV-G 349
G +G + + ++HLQFADD+L + + N +AL+ V ++ SG K+N KS++ G
Sbjct: 822 GVRIG-NGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITFG 880
Query: 350 VNIGESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCFL 409
+ S + +L YLGL +K +E ++ +K R S W + FL
Sbjct: 881 SRVYGSTQSKLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFL 940
Query: 410 SFGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSV 469
S G+ ++LKSV ++PVYA+S FK P GI+S IESL F+W + + R I W+AW +
Sbjct: 941 SPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRL 1000
Query: 470 CLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLEVGGRSVS 529
K+ GGLG R L +FN AL++K WRL+ + RV+ ARY ++ L+ R
Sbjct: 1001 QYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKDVSILDAKVRKQQ 1060
Query: 530 SW-WKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFNRRFARLFDLT 588
S+ W A + DG+ +G L+GDG N D + P L
Sbjct: 1061 SYGW---ASLLDGIALLKKG-----TRHLIGDGQNIRIGLDNIVDSHPPR----PLNTEE 1108
Query: 589 TNKMCTVADMYRLGWEEGGDAWVW--RRRLWAWEEECRSLLDNVRL-RLNVLDRWQWDPD 645
T K T+ +++ E G + W + ++ + + L + D+ W+ +
Sbjct: 1109 TYKEMTINNLF----ERKGSYYFWDDSKISQFVDQSDHGFIHRIYLAKSKKPDKIIWNYN 1164
Query: 646 TIDVYTVRGVYQILTTSVSNTI---------VKTDELVWHKQVPLN*VSIVAWRLLKDRL 696
T YTVR Y +LT S I + +W+ + + + WR L L
Sbjct: 1165 TTGEYTVRSGYWLLTHDPSTNIPAINPPHGSIDLKTRIWNLPI-MPKLKHFLWRALSQAL 1223
Query: 697 PTKTNLQRR 705
T L R
Sbjct: 1224 ATTERLTTR 1232
>UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1750
Score = 229 bits (583), Expect = 3e-58
Identities = 168/549 (30%), Positives = 263/549 (47%), Gaps = 35/549 (6%)
Query: 174 ILDGILIANEVVDEAKKTK---KELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKW 230
I D ++IA+EV+ K K K + K D KAYD V+ +L+ M+ F W W
Sbjct: 928 INDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWIGW 987
Query: 231 IKECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFK 290
I + + S+L+NGSP + RG+RQGDPLSP+LF++ + L+ L+ + +
Sbjct: 988 IMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGDLR 1047
Query: 291 GYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLV-G 349
G +G + + ++HLQFADD+L + + N +AL+ V ++ SG K+N KS++ G
Sbjct: 1048 GVRIG-NGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITFG 1106
Query: 350 VNIGESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCFL 409
+ S +L YLGL +K +E ++ +K R S W + FL
Sbjct: 1107 SRVYGSTQSRLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFL 1166
Query: 410 SFGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSV 469
S G+ ++LKSV ++PVYA+S FK P GI+S IESL F+W + + R I W+AW +
Sbjct: 1167 SPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRL 1226
Query: 470 CLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLEVGGRSVS 529
K+ GGLG R L +FN AL++K WRL+ + RV+ ARY ++ L+ R
Sbjct: 1227 QYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKDVSILDAKVRKQQ 1286
Query: 530 SW-WKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFNRRFARLFDLT 588
S+ W A + DG+ +G L+GDG N D + P L
Sbjct: 1287 SYGW---ASLLDGIALLKKG-----TRHLIGDGQNIRIGLDNIVDSHPPR----PLNTEE 1334
Query: 589 TNKMCTVADMYRLGWEEGGDAWVW--RRRLWAWEEECRSLLDNVRL-RLNVLDRWQWDPD 645
T K T+ +++ E G + W + ++ + + L + D+ W+ +
Sbjct: 1335 TYKEMTINNLF----ERKGSYYFWDDSKISQFVDQSDHGFIHRIYLAKSKKPDKIIWNYN 1390
Query: 646 TIDVYTVRGVYQILTTSVSNTI---------VKTDELVWHKQVPLN*VSIVAWRLLKDRL 696
T YTVR Y +LT S I + +W+ + + + WR L L
Sbjct: 1391 TTGEYTVRSGYWLLTHDPSTNIPAINPPHGSIDLKTRIWNLPI-MPKLKHFLWRALSQAL 1449
Query: 697 PTKTNLQRR 705
T L R
Sbjct: 1450 ATTERLTTR 1458
>UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [Arabidopsis thaliana]
Length = 1142
Score = 228 bits (581), Expect = 5e-58
Identities = 170/575 (29%), Positives = 271/575 (46%), Gaps = 37/575 (6%)
Query: 174 ILDGILIANEVVDEAKKT---KKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKW 230
I D ILIA E+ + K + + K D KAYD V+ +++A+++KM F W W
Sbjct: 330 ISDNILIAQEMFHGLRTNSSCKDKFMAIKTDMSKAYDQVEWNFIEALLRKMGFCEKWISW 389
Query: 231 IKECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFK 290
I CI+T +L+NG P RGLRQGDPLSP+LF++ E L ++ NL
Sbjct: 390 IMWCITTVQYKVLINGQPKGLIIPERGLRQGDPLSPYLFILCTEVLIANIRKAERQNLIT 449
Query: 291 GYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLL-VG 349
G V A+ S VSHL FADD+L + + + +L + +SG ++NF+KS + G
Sbjct: 450 GIKV-ATPSPAVSHLLFADDSLFFCKANKEQCGIILEILKQYESVSGQQINFSKSSIQFG 508
Query: 350 VNIGESWLLEAASVLNCKVGKILFLYLGL--SIGGDPRKLAFWEPVVSTIKTRLSGWQSC 407
+ +S + +L + YLGL S+GG K+ + V +++R++GW +
Sbjct: 509 HKVEDSIKADIKLILGIHNLGGMGSYLGLPESLGGSKTKVFSF--VRDRLQSRINGWSAK 566
Query: 408 FLSFGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWS 467
FLS GG+ V++KSV +LP Y +S F+ P I S + S KF+W + D+R + W+AW
Sbjct: 567 FLSKGGKEVMIKSVAATLPRYVMSCFRLPKAITSKLTSAVAKFWWSSNGDSRGMHWMAWD 626
Query: 468 SVCLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLE-VGGR 526
+C K GGLG R + +FN AL++K WRL+ + +V RY ++ L+ +
Sbjct: 627 KLCSSKSDGGLGFRNVDDFNSALLAKQLWRLITAPDSLFAKVFKGRYFRKSNPLDSIKSY 686
Query: 527 SVSSWWKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWL-GDVPFNRRFARLF 585
S S W+ + + V + G + K VG G++ W D W+ P ++
Sbjct: 687 SPSYGWRSMISARSLV-------YKGLI-KRVGSGASISVWNDPWIPAQFPRPAKYGGSI 738
Query: 586 DLTTNKMCTVADMYRLGWEEGGDAWVWRRRLWAWEEECRSLLDNVRL-RLNVLDRWQWDP 644
+ K+ ++ D + W ++ E L+ + + N+ D W
Sbjct: 739 VDPSLKVKSLID-------SRSNFWNIDLLKELFDPEDVPLISALPIGNPNMEDTLGWHF 791
Query: 645 DTIDVYTVRGVYQIL-------TTSVSNTIVKTDELVWHKQVPLN*VSIVAWRLLKDRLP 697
YTV+ Y TT + + +W Q P + W++L +P
Sbjct: 792 TKAGNYTVKSGYHTARLDLNEGTTLIGPDLTTLKAYIWKVQCPPK-LRHFLWQILSGCVP 850
Query: 698 TKTNLQRRDCLQATVNTCVSGCGIAESASHLFLHC 732
NL++R L CVS ES +H C
Sbjct: 851 VSENLRKRGIL--CDKGCVSCGASEESINHTLFQC 883
>UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1344
Score = 228 bits (581), Expect = 5e-58
Identities = 165/584 (28%), Positives = 279/584 (47%), Gaps = 53/584 (9%)
Query: 174 ILDGILIANEVVDEAK---KTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKW 230
I D IL+A+E++ + + KE + FK D KAYD V+ +L+ +M + F W W
Sbjct: 524 ISDNILVAHEMIHSLRTNDRISKEHMAFKTDMSKAYDRVEWPFLETMMTALGFNNKWISW 583
Query: 231 IKECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFK 290
I C+++ + S+L+NG P RG+RQGDPLSP LF++ E L ++ +
Sbjct: 584 IMNCVTSVSYSVLINGQPYGHIIPTRGIRQGDPLSPALFVLCTEALIHILNKAEQAGKIT 643
Query: 291 GYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLV-G 349
G + V V+HL FADDTLL+ + + L L+ + ++SG +N NKS + G
Sbjct: 644 G-IQFQDKKVSVNHLLFADDTLLMCKATKQECEELMQCLSQYGQLSGQMINLNKSAITFG 702
Query: 350 VNIG---ESWLLEAASV-LNCKVGKILFLYLGLS--IGGDPRKLAFWEPVVSTIKTRLSG 403
N+ + W+ + + L GK YLGL + G R L + + +++RL+G
Sbjct: 703 KNVDIQIKDWIKSRSGISLEGGTGK----YLGLPECLSGSKRDLFGF--IKEKLQSRLTG 756
Query: 404 WQSCFLSFGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISW 463
W + LS GG+ VLLKS+ +LPVY +S FK P + + ++ F+W + RKI W
Sbjct: 757 WYAKTLSQGGKEVLLKSIALALPVYVMSCFKLPKNLCQKLTTVMMDFWWNSMQQKRKIHW 816
Query: 464 IAWSSVCLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLEV 523
++W + L K+ GG G + L+ FN AL++K WR+L ++G + RV +RY + L
Sbjct: 817 LSWQRLTLPKDQGGFGFKDLQCFNQALLAKQAWRVLQEKGSLFSRVFQSRYFSNSDFLSA 876
Query: 524 GGRSVSSW-WKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGD----VPFN 578
S S+ W+ + ++ + + ++G+G T W D+WL D P N
Sbjct: 877 TRGSRPSYAWRSILFGRE--------LLMQGLRTVIGNGQKTFVWTDKWLHDGSNRRPLN 928
Query: 579 RRFARLFDLTTNKMCTVADMYRLGWEEGGDAWVWRRRLWAWEEECRSLLDNVRLRLNVLD 638
RR DL K+ + D W R L+ W++ ++ R D
Sbjct: 929 RRRFINVDL---KVSQLIDPTSRNWNLN-----MLRDLFPWKDV--EIILKQRPLFFKED 978
Query: 639 RWQWDPDTIDVYTVRGVYQILTTSVSNTIVKTDEL----------VWHKQVPLN*VSIVA 688
+ W +Y+V+ Y+ L+ V + + + ++ +W+ + I
Sbjct: 979 SFCWLHSHNGLYSVKTGYEFLSKQVHHRLYQEAKVKPSVNSLFDKIWNLHTAPK-IRIFL 1037
Query: 689 WRLLKDRLPTKTNLQRRDCLQATVNTCVSGCGIAESASHLFLHC 732
W+ L +P + L+ R + + C+ E+ +H+ C
Sbjct: 1038 WKALHGAIPVEDRLRTRGI--RSDDGCLMCDTENETINHILFEC 1079
>UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis thaliana]
Length = 851
Score = 227 bits (578), Expect = 1e-57
Identities = 167/549 (30%), Positives = 267/549 (48%), Gaps = 35/549 (6%)
Query: 174 ILDGILIANEVVDEAKKTK---KELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKW 230
I D ++IA+E++ K K K + K D KAYD V+ +L+ M+ F W W
Sbjct: 167 INDNVMIAHEIMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCDKWIGW 226
Query: 231 IKECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFK 290
I + + S+L+NGSP S RG+RQGDPLSP+LF++ + L+ L+K + +
Sbjct: 227 IMAAVKSVHYSVLINGSPHGYISPTRGIRQGDPLSPYLFILCGDILSHLIKVKASSGDIR 286
Query: 291 GYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLV-G 349
G +G + + ++HLQFADD+L + + N +AL+ V ++ SG K+N KSL+ G
Sbjct: 287 GVRIG-NGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSLITFG 345
Query: 350 VNIGESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCFL 409
+ S ++LN YLGL +K + ++ +K R + W + FL
Sbjct: 346 SRVYGSTQTRLKTLLNIPNQGGGGKYLGLPEQFGRKKKEMFNYIIDRVKERTASWSAKFL 405
Query: 410 SFGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSV 469
S G+ +LLKSV ++PVYA+S FK P GI+S IESL F+W + + R I W+AW +
Sbjct: 406 SPAGKEILLKSVALAMPVYAMSCFKLPQGIVSEIESLLMNFWWEKASNKRGIPWVAWKRL 465
Query: 470 CLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLEVGGRSVS 529
K+ GGLG R L +FN AL++K WR++ + RV+ ARY ++ ++ RS
Sbjct: 466 QYSKKEGGLGFRDLAKFNDALLAKQAWRIIQYPNSLFARVMKARYFKDNSIIDAKTRSQQ 525
Query: 530 SW-WKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFNRRFARLFDLT 588
S+ W + + G+ +G ++GDG D + P L
Sbjct: 526 SYGW---SSLLSGIALLRKG-----TRYVIGDGKTIRLGIDNVVDSHPPR-------PLL 570
Query: 589 TNKMCTVADMYRLGWEEGGDAWVW-RRRLWAW-EEECRSLLDNVRLRL-NVLDRWQWDPD 645
T++ + L ++ G + W +L + ++ + + L + DR W +
Sbjct: 571 TDEQHNGLSLDNL-FQHRGHSRCWDNAKLQTFVDQSDHDYIKRIYLSTRSKTDRLIWSYN 629
Query: 646 TIDVYTVRGVYQILTTSVSNTI---------VKTDELVWHKQVPLN*VSIVAWRLLKDRL 696
+ YTVR Y + T SNTI V +W+ + + + WR+L L
Sbjct: 630 STGDYTVRSGYWLSTHDPSNTIPTMAKPHGSVDLKTKIWNLPI-MPKLKHFLWRILSKAL 688
Query: 697 PTKTNLQRR 705
PT L R
Sbjct: 689 PTTDRLTTR 697
>UniRef100_Q9FXJ5 F5A9.24 [Arabidopsis thaliana]
Length = 1254
Score = 225 bits (574), Expect = 3e-57
Identities = 167/565 (29%), Positives = 261/565 (45%), Gaps = 57/565 (10%)
Query: 169 VKDMQILDGILIANEVVDEAK---KTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPL 225
V + I D IL+A+E+V K + E + K D KAYD V+ YL +++ + F L
Sbjct: 507 VSERLITDNILVAHELVHSLKVHPRISSEFMAVKSDMSKAYDRVEWSYLRSLLLSLGFHL 566
Query: 226 LWRKWIKECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVE 285
W WI C+S+ T S+L+N P L RGLRQGDPLSPFLF++ EGL L+
Sbjct: 567 KWVNWIMVCVSSVTYSVLINDCPFGLIILQRGLRQGDPLSPFLFVLCTEGLTHLLNKAQW 626
Query: 286 NNLFKGYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKS 345
+G + + N +V HL FADD+L L + S L+ +L ++ +G +N NKS
Sbjct: 627 EGALEG-IQFSENGPMVHHLLFADDSLFLCKASREQSLVLQKILKVYGNATGQTINLNKS 685
Query: 346 LLV-GVNIGESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGW 404
+ G + E + L YLGL K+ + +K +L W
Sbjct: 686 SITFGEKVDEQLKGTIRTCLGIFTEGGAGTYLGLPECFSGSKVDMLHYLKDRLKEKLDVW 745
Query: 405 QSCFLSFGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWI 464
+ LS GG+ VLLKSV ++PV+A+S FK P +ES F+W + +RKI W
Sbjct: 746 FTRCLSQGGKEVLLKSVALAMPVFAMSCFKLPITTCENLESAMASFWWDSCDHSRKIHWQ 805
Query: 465 AWSSVCLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLEVG 524
+W +CL K+ GGLG R ++ FN AL++K WRLL R+L +RY + L+
Sbjct: 806 SWERLCLPKDSGGLGFRDIQSFNQALLAKQAWRLLHFPDCLLSRLLKSRYFDATDFLDAA 865
Query: 525 -GRSVSSWWKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFNRRFAR 583
+ S W+ + ++ + + + K VGDG++ W D W+ D F + +
Sbjct: 866 LSQRPSFGWRSILFGRELLSKG--------LQKRVGDGASLFVWIDPWIDDNGFRAPWRK 917
Query: 584 --LFDLTTNKMCTVADMYRLGWEEGGDAWVWRRRLWAWEEECRSLL----DNVRLR---- 633
++D+T + R W+EE L D +R++
Sbjct: 918 NLIYDVTLKVKALL-----------------NPRTGFWDEEVLHDLFLPEDILRIKAIKP 960
Query: 634 -LNVLDRWQWDPDTIDVYTVRGVYQILTTSVSNTI------------VKTDELVWHKQVP 680
++ D + W + ++V+ Y + + S + +KT VW+ Q
Sbjct: 961 VISQADFFVWKLNKSGDFSVKSAYWLAYQTKSQNLRSEVSMQPSTLGLKTQ--VWNLQTD 1018
Query: 681 LN*VSIVAWRLLKDRLPTKTNLQRR 705
+ I W++L LP NL R
Sbjct: 1019 PK-IKIFLWKVLSGILPVAENLNGR 1042
>UniRef100_Q7XE51 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 1652
Score = 224 bits (572), Expect = 5e-57
Identities = 138/397 (34%), Positives = 209/397 (51%), Gaps = 11/397 (2%)
Query: 174 ILDGILIANEVVDEAKKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIKE 233
I++G++I +E + E K KK ++ K+DFEKAYD VD +L ++ F W WI
Sbjct: 1265 IMEGVVILHETLHELHKKKKNGVILKLDFEKAYDKVDWKFLQQSLRMKGFSSKWCDWIDS 1324
Query: 234 CISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGYV 293
+ + ++ VN F +GLRQGDPLSP LF + + L +L++ + FKG V
Sbjct: 1325 IVRGGSVAVKVNDEIGSYFQTRKGLRQGDPLSPILFNLVVDMLAILIQRAKDQGRFKGVV 1384
Query: 294 VGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLVGVNIG 353
++ + S LQ+ADDT+L + R L+ VL+ F ++S LK+NF KS L
Sbjct: 1385 PHLVDNGL-SILQYADDTILFMDHDLDEARDLKLVLSTFEKLSSLKINFYKSELFCYGKA 1443
Query: 354 ESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCFLSFGG 413
+ E + C F YLG+ + W+ V I+ +LS W+ FLS GG
Sbjct: 1444 KDVEHEYVKLFGCDTEDYPFKYLGIRMHHKRINNKDWQGVEERIQKKLSSWKGKFLSVGG 1503
Query: 414 RLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSVCLKK 473
RLVL+ SVL++L ++ LSFF+ P GI+ ++ ++FFW+ E +K WS +C K
Sbjct: 1504 RLVLINSVLSNLAIFMLSFFEIPKGILKKLDYYRSRFFWQCDEHKKKYRLARWSVLCKPK 1563
Query: 474 EFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLEVGG-RSVSSWW 532
E GGLG++ L N L+SKW ++ LI+ G W +L +Y + +V S +W
Sbjct: 1564 ECGGLGIQNLEIQNKCLLSKWLYK-LINEEGVWQDILRNKYLTKKTITQVEKCPGDSHFW 1622
Query: 533 KEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFD 569
+ ++D +F G K VGDGS FW D
Sbjct: 1623 AGLMGVRD-------VFFKGGAFK-VGDGSQIRFWED 1651
>UniRef100_O22148 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1374
Score = 224 bits (570), Expect = 9e-57
Identities = 167/596 (28%), Positives = 272/596 (45%), Gaps = 59/596 (9%)
Query: 169 VKDMQILDGILIANEVV---DEAKKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPL 225
VK I D ILIA+E++ K +E + K D KAYD V+ +L+ M+ + F
Sbjct: 541 VKGRLISDNILIAHELLHALSSNNKCSEEFIAIKTDISKAYDRVEWPFLEKAMRGLGFAD 600
Query: 226 LWRKWIKECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVE 285
W + I EC+ + +L+NG+P E RGLRQGDPLSP+LF+I E L +++S +
Sbjct: 601 HWIRLIMECVKSVRYQVLINGTPHGEIIPSRGLRQGDPLSPYLFVICTEMLVKMLQSAEQ 660
Query: 286 NNLFKGYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKS 345
N G V A + +SHL FADD++ + + + + ++ ++ SG +VN+ KS
Sbjct: 661 KNQITGLKV-ARGAPPISHLLFADDSMFYCKVNDEALGQIIRIIEEYSLASGQRVNYLKS 719
Query: 346 -LLVGVNIGESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGW 404
+ G +I E L + +YLGL K+A + + ++ GW
Sbjct: 720 SIYFGKHISEERRCLVKRKLGIEREGGEGVYLGLPESFQGSKVATLSYLKDRLGKKVLGW 779
Query: 405 QSCFLSFGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWI 464
QS FLS GG+ +LLK+V +LP Y +S FK P I IES+ +F+W+ ++ R + W
Sbjct: 780 QSNFLSPGGKEILLKAVAMALPTYTMSCFKIPKTICQQIESVMAEFWWKNKKEGRGLHWK 839
Query: 465 AWSSVCLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLE-- 522
AW + K GGLG + + FN+AL+ K WR++ ++ +V +RY ++ L
Sbjct: 840 AWCHLSRPKAVGGLGFKEIEAFNIALLGKQLWRMITEKDSLMAKVFKSRYFSKSDPLNAP 899
Query: 523 VGGRSVSSW---WKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFN- 578
+G R +W ++ IK G + ++G+G W D W+G P
Sbjct: 900 LGSRPSFAWKSIYEAQVLIKQG------------IRAVIGNGETINVWTDPWIGAKPAKA 947
Query: 579 ----RRFARLFDLTTNKMCTVADMYRLGWEEGGDAWVWRRRLWAWEEECRSLLDNVRLRL 634
+R + N + V D+ +G D W W DN + +
Sbjct: 948 AQAVKRSHLVSQYAANSIHVVKDLL---LPDGRD--------WNWNLVSLLFPDNTQENI 996
Query: 635 NVL--------DRWQWDPDTIDVYTVRGVYQILTTSVSN----------TIVKTDELVWH 676
L DR+ W+ Y+V+ Y ++T ++ ++ + +W
Sbjct: 997 LALRPGGKETRDRFTWEYSRSGHYSVKSGYWVMTEIINQRNNPQEVLQPSLDPIFQQIWK 1056
Query: 677 KQVPLN*VSIVAWRLLKDRLPTKTNLQRRDCLQATVNTCVSGCGIAESASHLFLHC 732
VP + WR + + L +NL R A +CV E+ +HL C
Sbjct: 1057 LDVPPK-IHHFLWRCVNNCLSVASNLAYRHL--AREKSCVRCPSHGETVNHLLFKC 1109
>UniRef100_Q8S6P1 Putative reverse transcriptase [Oryza sativa]
Length = 1509
Score = 223 bits (569), Expect = 1e-56
Identities = 173/586 (29%), Positives = 270/586 (45%), Gaps = 52/586 (8%)
Query: 174 ILDGILIANEVVDEAKKTKKELL---LFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKW 230
I D ILIA E+ + + + FK+D KAYD V+ +L +M K+ F W
Sbjct: 747 ISDNILIAYEMTHYMRNKRSGQVGYAAFKLDMSKAYDRVEWSFLHDMMLKLGFHTDWVNL 806
Query: 231 IKECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFK 290
I +C+ST T I VNG ++ FS RGLRQGDPLSP+LF++ AEG + L+ E
Sbjct: 807 IMKCVSTVTYRIRVNGELSESFSPERGLRQGDPLSPYLFLLCAEGFSALLSKTEEEGRLH 866
Query: 291 GYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKS-LLVG 349
G + + VSHL FADD+L+L + + L+ +L ++ E SG +N +KS ++
Sbjct: 867 GIRI-CQGAPSVSHLLFADDSLILCRANGGEAQQLQTILQIYEECSGQVINKDKSAVMFS 925
Query: 350 VNIGESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCFL 409
N + LN + YLGL + + + + I R+ GW+ L
Sbjct: 926 PNTSSLEKGAVMAALNMQRETTNEKYLGLPVFVGRSRTKIFSYLKERIWQRIQGWKEKLL 985
Query: 410 SFGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSV 469
S G+ +L+K+V +P +A+ F+ + I + K++W E + K+ W++W+ +
Sbjct: 986 SRAGKEILIKAVAQVIPTFAMGCFELTKDLCDQISKMIAKYWWSNQEKDNKMHWLSWNKL 1045
Query: 470 CLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARY---GEEARRLEVGGR 526
L K GGLG R + FNLA+++K WRL+ D RVL A+Y G+ R +
Sbjct: 1046 TLPKNMGGLGFRDIYIFNLAMLAKQGWRLIQDPDSLCSRVLRAKYFPLGDCFRPKQTS-- 1103
Query: 527 SVSSWWKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWL----GDVPFNRRFA 582
+VS W+ + K G+ G + VGDGS W D W+ P R A
Sbjct: 1104 NVSYTWRSIQK---GLRVLQNG-----MIWRVGDGSKINIWADPWIPRGWSRKPMTPRGA 1155
Query: 583 RLFDLTTNKMCTVADMYRLGWEEGGDAWVWRRRLWAWEEECRSLLDNVRLRLNVLDRWQW 642
L K+ + D Y W+E + + WEE+ + + ++ + + + D W
Sbjct: 1156 NL----VTKVEELIDPYTGTWDEDLLSQTF------WEEDV-AAIKSIPVHVEMEDVLAW 1204
Query: 643 DPDTIDVYTVRGVYQIL-----------TTSVSNTIVKTDEL---VWHKQVPLN*VSIVA 688
D +TV+ Y++ VSN D+ +W VP +
Sbjct: 1205 HFDARGCFTVKSAYKVQREMERRASRNGCPGVSNWESGDDDFWKKLWKLGVP-GKIKHFL 1263
Query: 689 WRLLKDRLPTKTNLQRRDCLQATVNTCVSGCG-IAESASHLFLHCE 733
WR+ + L + NL R V+T CG E A HLF C+
Sbjct: 1264 WRMCHNTLALRANLHHRG---MDVDTRCVMCGRYNEDAGHLFFKCK 1306
>UniRef100_Q9FYK1 F21J9.30 [Arabidopsis thaliana]
Length = 1270
Score = 223 bits (567), Expect = 2e-56
Identities = 142/427 (33%), Positives = 216/427 (50%), Gaps = 16/427 (3%)
Query: 169 VKDMQILDGILIANEVVDEAK---KTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPL 225
V + I D IL+A+E+V K + E + K D KAYD V+ YL +++ + F L
Sbjct: 504 VSERLITDNILVAHELVHSLKVHPRISSEFMAVKSDMSKAYDRVEWSYLRSLLLSLGFHL 563
Query: 226 LWRKWIKECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVE 285
W WI C+S+ T S+L+N P L RGLRQGDPLSPFLF++ EGL L+
Sbjct: 564 KWVNWIMVCVSSVTYSVLINDCPFGLIILQRGLRQGDPLSPFLFVLCTEGLTHLLNKAQW 623
Query: 286 NNLFKGYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKS 345
+G + + N +V HL FADD+L L + S L+ +L ++ +G +N NKS
Sbjct: 624 EGALEG-IQFSENGPMVHHLLFADDSLFLCKASREQSLVLQKILKVYGNATGQTINLNKS 682
Query: 346 LLV-GVNIGESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGW 404
+ G + E + L YLGL K+ + +K +L W
Sbjct: 683 SITFGEKVDEQLKGTIRTCLGIFTEGGAGTYLGLPECFSGSKVDMLHYLKDRLKEKLDVW 742
Query: 405 QSCFLSFGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWI 464
+ LS GG+ VLLKSV ++PV+A+S FK P +ES F+W + +RKI W
Sbjct: 743 FTRCLSQGGKEVLLKSVALAMPVFAMSCFKLPITTCENLESAMASFWWDSCDHSRKIHWQ 802
Query: 465 AWSSVCLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLEVG 524
+W +CL K+ GGLG R ++ FN AL++K WRLL R+L +RY + L+
Sbjct: 803 SWERLCLPKDSGGLGFRDIQSFNQALLAKQAWRLLHFPDCLLSRLLKSRYFDATDFLDAA 862
Query: 525 -GRSVSSWWKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFNRRFAR 583
+ S W+ + ++ + + + K VGDG++ W D W+ D F + +
Sbjct: 863 LSQRPSFGWRSILFGRELLSKG--------LQKRVGDGASLFVWIDPWIDDNGFRAPWRK 914
Query: 584 --LFDLT 588
++D+T
Sbjct: 915 NLIYDVT 921
>UniRef100_O22220 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1094
Score = 221 bits (564), Expect = 5e-56
Identities = 160/589 (27%), Positives = 269/589 (45%), Gaps = 53/589 (8%)
Query: 169 VKDMQILDGILIANEVVDEAK---KTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPL 225
V + + D I++A+E+V + K K+ ++FK D KAYD V+ +L ++ + F
Sbjct: 270 VAERLVSDNIILAHEIVHNLRTNEKISKDFMVFKTDMSKAYDRVEWPFLKGILLALGFNS 329
Query: 226 LWRKWIKECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVE 285
W W+ C+S+ + S+L+NG P + HRGLRQGDPLSPFLF++ E L ++ +
Sbjct: 330 TWINWMMACVSSVSYSVLINGQPFGHITPHRGLRQGDPLSPFLFVLCTEALIHILNQAEK 389
Query: 286 NNLFKGYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKS 345
G + V+HL FADDTLL+ + S + L+ + +SG +N KS
Sbjct: 390 IGKISGIQFNGTGP-SVNHLLFADDTLLICKASQLECAEIMHCLSQYGHISGQMINSEKS 448
Query: 346 LLV-GVNIGE---SWLLEAASV-LNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTR 400
+ G + E W++ + + GK YLGL K + + +++R
Sbjct: 449 AITFGAKVNEETKQWIMNRSGIQTEGGTGK----YLGLPECFQGSKQVLFGFIKEKLQSR 504
Query: 401 LSGWQSCFLSFGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRK 460
LSGW + LS GG+ +LLKS+ + PVYA++ F+ + + + S+ F+W +D +K
Sbjct: 505 LSGWYAKTLSQGGKDILLKSIAMAFPVYAMTCFRLSKTLCTKLTSVMMDFWWNSVQDKKK 564
Query: 461 ISWIAWSSVCLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARR 520
I WI + L K GG G + L+ FN AL++K RL D ++L +RY +
Sbjct: 565 IHWIGAQKLMLPKFLGGFGFKDLQCFNQALLAKQASRLHTDSDSLLSQILKSRYYMNSDF 624
Query: 521 LE-VGGRSVSSWWKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWL-GDVPFN 578
L G S W+ + ++ V + K++G+G NT W D W+ D P
Sbjct: 625 LSATKGTRPSYAWQSILYGRE--------LLVSGLKKIIGNGENTYVWMDNWIFDDKPRR 676
Query: 579 RRFARLFDLTTNKMCTVADMYRLGWEEGGDAWVWRRRLWAWEEECRSLLDNVRLRLNVLD 638
++ K+ + D + W R L+ W+E ++ R + D
Sbjct: 677 PESLQIMVDIQLKVSQLIDPFSRNWNLN-----MLRDLFPWKE--IQIICQQRPMASRQD 729
Query: 639 RWQWDPDTIDVYTVRGVYQILTTSVSNTIVKTDE----------LVWH-KQVPLN*VSIV 687
+ W +YTV+ Y + + V + K E +W+ P + +
Sbjct: 730 SFCWFGTNHGLYTVKSEYDLCSRQVHKQMFKEAEEQPSLNPLFGKIWNLNSAPK--IKVF 787
Query: 688 AWRLLKDRLPTKTNLQRRDCLQATVNTCVSGCGIA----ESASHLFLHC 732
W++LK + + L+ R L GC + E+ +H+ C
Sbjct: 788 LWKVLKGAVAVEDRLRTRGVL------IEDGCSMCPEKNETLNHILFQC 830
>UniRef100_Q9SLE9 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1319
Score = 221 bits (562), Expect = 8e-56
Identities = 163/581 (28%), Positives = 267/581 (45%), Gaps = 47/581 (8%)
Query: 174 ILDGILIANEVVDEAK---KTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKW 230
I D ILIA+E+V + K KE ++FK D KAYD V+ +L ++ + F W W
Sbjct: 498 ISDNILIAHEIVHSLRTNDKISKEFMVFKTDMSKAYDRVEWSFLQEILVALGFNDKWISW 557
Query: 231 IKECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFK 290
I C+++ T S+L+NG + RG+RQGDP+SPFLF++ E L +++ +
Sbjct: 558 IMGCVTSVTYSVLINGQHFGHITPERGIRQGDPISPFLFVLCTEALIHILQQAENSKKVS 617
Query: 291 GYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLV-G 349
G S V +HL F DDT L+ + ++ + L+ + +SG +N KS + G
Sbjct: 618 GIQFNGSGPSV-NHLLFVDDTQLVCRATKSDCEQMMLCLSQYGHISGQLINVEKSSITFG 676
Query: 350 VNIGES---WLLEAASV-LNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQ 405
V + E W+ + + L GK YLGL K + + +++ LSGW
Sbjct: 677 VKVDEDTKRWIKNRSGIHLEGGTGK----YLGLPENLSGSKQDLFGYIKEKLQSHLSGWY 732
Query: 406 SCFLSFGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIA 465
LS GG+ +LLKS+ +LPVY ++ F+ P G+ + + S+ F+W E + KI WI
Sbjct: 733 DKTLSQGGKEILLKSIALALPVYIMTCFRLPKGLCTKLTSVMMDFWWNSMEFSNKIHWIG 792
Query: 466 WSSVCLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLEV-G 524
+ L K GG G + L+ FN AL++K WRL D ++ +RY L
Sbjct: 793 GKKLTLPKSLGGFGFKDLQCFNQALLAKQAWRLFSDSKSIVSQIFKSRYFMNTDFLNARQ 852
Query: 525 GRSVSSWWKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFNRRFARL 584
G S W+ + ++ + G + +L+G+G T W D+WL D +RR L
Sbjct: 853 GTRPSYTWRSILYGRELLN--------GGLKRLIGNGEQTNVWIDKWLFD-GHSRRPMNL 903
Query: 585 FDLTT--NKMCTVADMYRLGWEEGGDAWVWRRRLWAWEEECRSLLDNVRLRLNVLDRWQW 642
L K+ + D W ++ + E+ L+ + R ++ D + W
Sbjct: 904 HSLMNIHMKVSHLIDPLTRNWN-------LKKLTELFHEKDVQLIMHQRPLISSEDSYCW 956
Query: 643 DPDTIDVYTVRGVYQILTTSVSNTIVKTDEL----------VWH-KQVPLN*VSIVAWRL 691
+YTV+ Y+ + + K ++ VW + VP + + W+
Sbjct: 957 AGTNNGLYTVKSGYERSSRETFKNLFKEADVYPSVNPLFDKVWSLETVPK--IKVFMWKA 1014
Query: 692 LKDRLPTKTNLQRRDCLQATVNTCVSGCGIAESASHLFLHC 732
LK L + L+ R T + C+ E+ +HL C
Sbjct: 1015 LKGALAVEDRLRSRGI--RTADGCLFCKEEIETINHLLFQC 1053
>UniRef100_O81630 F8M12.22 protein [Arabidopsis thaliana]
Length = 1662
Score = 218 bits (554), Expect = 7e-55
Identities = 139/401 (34%), Positives = 214/401 (52%), Gaps = 20/401 (4%)
Query: 176 DGILIANEVVDEAKKTKK---ELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIK 232
D ++IA+E++ K K+ + K D KAYD V+ +L+ M+ F W KWI
Sbjct: 930 DNVMIAHEMMHSLKTRKRVSQSYMAVKTDVSKAYDRVEWNFLETTMRLFGFSETWIKWIM 989
Query: 233 ECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGY 292
+ + S+LVNG P RG+RQGDPLSP+LF++ A+ LN L+K+ V +G
Sbjct: 990 GAVKSVNYSVLVNGIPHGTIQPQRGIRQGDPLSPYLFILCADILNHLIKNRVAEGDIRGI 1049
Query: 293 VVGASNSVV-VSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLV-GV 350
+G N V V+HLQFADD+L + + N +AL+ V ++ SG K+N +KS++ G
Sbjct: 1050 RIG--NGVPGVTHLQFADDSLFFCQSNVRNCQALKDVFDVYEYYSGQKINMSKSMITFGS 1107
Query: 351 NIGESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCFLS 410
+ + ++L + YLGL +K + ++ +K R S W + +LS
Sbjct: 1108 RVHGTTQNRLKNILGIQSHGGGGKYLGLPEQFGRKKRDMFNYIIERVKKRTSSWSAKYLS 1167
Query: 411 FGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSVC 470
G+ ++LKSV S+PVYA+S FK P I+S IE+L F+W + R+I WIAW +
Sbjct: 1168 PAGKEIMLKSVAMSMPVYAMSCFKLPLNIVSEIEALLMNFWWEKNAKKREIPWIAWKRLQ 1227
Query: 471 LKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLEVGGRSVSS 530
K+ GGLG R L +FN AL++K WR++ + + R++ ARY E L+ + S
Sbjct: 1228 YSKKEGGLGFRDLAKFNDALLAKQVWRMINNPNSLFARIMKARYFREDSILDAKRQRYQS 1287
Query: 531 W-WKEVAK----IKDG----VGEDDEG----WFVGKVSKLV 558
+ W + IK G VG+ G W +S+LV
Sbjct: 1288 YGWTSMLAGLDVIKKGSRFIVGDGKTGSYRYWNAHLISQLV 1328
>UniRef100_Q9SN65 Hypothetical protein F25I24.40 [Arabidopsis thaliana]
Length = 1294
Score = 217 bits (553), Expect = 9e-55
Identities = 128/361 (35%), Positives = 199/361 (54%), Gaps = 7/361 (1%)
Query: 176 DGILIANEVVDEAKKTKK---ELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIK 232
D ++IA+E++ K K+ + K D KAYD V+ +L+ M+ F W KWI
Sbjct: 910 DNVMIAHEMMHSLKTRKRVSQSYMAVKTDVSKAYDRVEWNFLETTMRLFGFSETWIKWIM 969
Query: 233 ECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGY 292
+ + S+LVNG P RG+RQGDPLSP+LF++ A+ LN L+K+ V +G
Sbjct: 970 GAVKSVNYSVLVNGIPHGTIQPQRGIRQGDPLSPYLFILCADILNHLIKNRVAEGDIRGI 1029
Query: 293 VVGASNSVV-VSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLV-GV 350
+G N V V+HLQFADD+L + + N +AL+ V ++ SG K+N +KS++ G
Sbjct: 1030 RIG--NGVPGVTHLQFADDSLFFCQSNVRNCQALKDVFDVYEYYSGQKINMSKSMITFGS 1087
Query: 351 NIGESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCFLS 410
+ + ++L + YLGL +K + ++ +K R S W + +LS
Sbjct: 1088 RVHGTTQNRLKNILGIQSHGGGGKYLGLPEQFGRKKRDMFNYIIERVKKRTSSWSAKYLS 1147
Query: 411 FGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSVC 470
G+ ++LKSV S+PVYA+S FK P I+S IE+L F+W + R+I WIAW +
Sbjct: 1148 PAGKEIMLKSVAMSMPVYAMSCFKLPLNIVSEIEALLMNFWWEKNAKKREIPWIAWKRLQ 1207
Query: 471 LKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLEVGGRSVSS 530
K+ GGLG R L +FN AL++K WR++ + + R++ ARY E L+ + S
Sbjct: 1208 YSKKEGGLGFRDLAKFNDALLAKQVWRMINNPNSLFARIMKARYFREDSILDAKRQRYQS 1267
Query: 531 W 531
+
Sbjct: 1268 Y 1268
>UniRef100_O82276 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1231
Score = 216 bits (550), Expect = 2e-54
Identities = 172/575 (29%), Positives = 256/575 (43%), Gaps = 47/575 (8%)
Query: 175 LDGILIANEVVDEA--KKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIK 232
+D I++ E V KK +K +L K+D EKAYD V +L ++ W I
Sbjct: 411 IDNIVLVQEAVHSMRRKKGRKGWMLLKLDLEKAYDRVRWDFLQETLEAAGLSEGWTSRIM 470
Query: 233 ECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGY 292
++ + S+L NG TD F RGLRQGDPLSP+LF++ E L L+++ V +K
Sbjct: 471 AGVTDPSMSVLWNGERTDSFVPARGLRQGDPLSPYLFVLCLERLCHLIEASVGKREWKPI 530
Query: 293 VVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLV---G 349
V S + SH+ FADD +L E S A +R +R VL F E SG KV+ KS +
Sbjct: 531 AVSCGGSKL-SHVCFADDLILFAEASVAQIRIIRRVLERFCEASGQKVSLEKSKIFFSHN 589
Query: 350 VNIGESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCFL 409
V+ L+ S + C K L YLG+ I + V+ + RL+GW+ L
Sbjct: 590 VSREMEQLISEESGIGCT--KELGKYLGMPILQKRMNKETFGEVLERVSARLAGWKGRSL 647
Query: 410 SFGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSV 469
S GR+ L K+VL+S+PV+ +S P + ++ F W + + +K ++W +
Sbjct: 648 SLAGRITLTKAVLSSIPVHVMSAILLPVSTLDTLDRYSRTFLWGSTMEKKKQHLLSWRKI 707
Query: 470 CLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLEVGGRSVS 529
C K GG+G+R R+ N AL++K WRLL D+ W RV+ +Y +VGG +
Sbjct: 708 CKPKAEGGIGLRSARDMNKALVAKVGWRLLQDKESLWARVVRKKY-------KVGGVQDT 760
Query: 530 SW----------WKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFNR 579
SW W+ VA V GW + GDG FW DRWL P
Sbjct: 761 SWLKPQPRWSSTWRSVAVGLREVVVKGVGW-------VPGDGCTIRFWLDRWLLQEPLVE 813
Query: 580 RFARLFDLTTNKMCTVADMYRLGWEEGGDAWVWRRRLWAWEEECRSLLD-NVRLRLNVLD 638
+ + + VA Y W G + L+ E R LL V++ L D
Sbjct: 814 LGTDM--IPEGERIKVAADY---WLPGSGWNLEILGLYLPETVKRRLLSVVVQVFLGNGD 868
Query: 639 RWQWDPDTIDVYTVRGVYQILTTSVSN--TIVKTDELVWHKQVPLN*VSIVAWRLLKDRL 696
W +TVR Y +L V + + +W K + V + W + ++ +
Sbjct: 869 EISWKGTQDGAFTVRSAYSLLQGDVGDRPNMGSFFNRIW-KLITPERVRVFIWLVSQNVI 927
Query: 697 PTKTNLQRRDCLQATVNTCVSGCGIAESASHLFLH 731
T RR + + C + A LH
Sbjct: 928 MTNVERVRRHLSENAI------CSVCNGAEETILH 956
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.335 0.146 0.498
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,224,516,488
Number of Sequences: 2790947
Number of extensions: 51288755
Number of successful extensions: 164119
Number of sequences better than 10.0: 836
Number of HSP's better than 10.0 without gapping: 579
Number of HSP's successfully gapped in prelim test: 257
Number of HSP's that attempted gapping in prelim test: 162220
Number of HSP's gapped (non-prelim): 1255
length of query: 738
length of database: 848,049,833
effective HSP length: 135
effective length of query: 603
effective length of database: 471,271,988
effective search space: 284177008764
effective search space used: 284177008764
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 79 (35.0 bits)
Medicago: description of AC149205.5


