

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149205.3 + phase: 0 /pseudo
(523 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
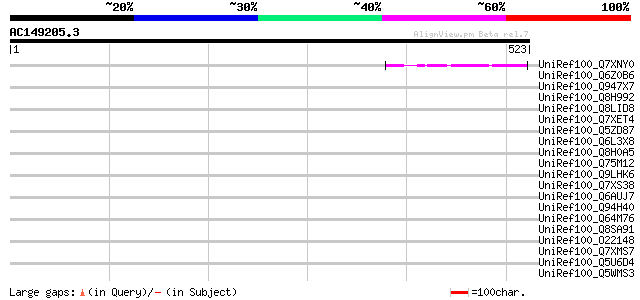
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XNY0 OSJNBb0015N08.11 protein [Oryza sativa] 50 1e-04
UniRef100_Q6Z0B6 Cyst nematode resistance protein-like protein [... 47 0.001
UniRef100_Q947X7 Hypothetical protein OSJNBa0067N01.20 [Oryza sa... 47 0.001
UniRef100_Q8H992 Oryza sativa (japonica cultivar-group) retrotra... 46 0.002
UniRef100_Q8LID8 Cyst nematode resistance protein-like protein [... 44 0.011
UniRef100_Q7XET4 Contains similarity to non-LTR reverse transcri... 44 0.015
UniRef100_Q5ZD87 HcrVf2 protein-like [Oryza sativa] 43 0.020
UniRef100_Q6L3X8 Hypothetical protein [Solanum demissum] 43 0.020
UniRef100_Q8H0A5 Putative reverse transcriptase [Oryza sativa] 42 0.043
UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa] 41 0.097
UniRef100_Q9LHK6 Similarity to non-LTR retroelement reverse tran... 41 0.097
UniRef100_Q7XS38 OSJNBa0081G05.2 protein [Oryza sativa] 40 0.17
UniRef100_Q6AUJ7 Hypothetical protein OSJNBa0052E20.4 [Oryza sat... 39 0.28
UniRef100_Q94H40 Putative reverse transcriptase [Oryza sativa] 39 0.48
UniRef100_Q64M76 Phosphoribosyl pyrophosphate synthase-like [Ory... 38 0.63
UniRef100_Q8SA91 Putative gag-pol polyprotein [Zea mays] 38 0.63
UniRef100_O22148 Putative non-LTR retroelement reverse transcrip... 38 0.82
UniRef100_Q7XMS7 OSJNBa0029L02.10 protein [Oryza sativa] 37 1.8
UniRef100_Q5U6D4 Orf147a protein [Beta vulgaris subsp. vulgaris] 37 1.8
UniRef100_Q5WMS3 Hypothetical protein OJ1333_C12.8 [Oryza sativa] 36 2.4
>UniRef100_Q7XNY0 OSJNBb0015N08.11 protein [Oryza sativa]
Length = 1026
Score = 50.4 bits (119), Expect = 1e-04
Identities = 41/144 (28%), Positives = 66/144 (45%), Gaps = 19/144 (13%)
Query: 379 WRRHLWVWEE-DLVEECGDLLLTVSLQELETDRWMWLSDQIRGYTVRGVYDMLTSQE*PH 437
WR+ + WE +LVE C D L W LS + +TVR Y L Q+
Sbjct: 793 WRKIVNSWEGLNLVENCKDKL------------WWTLSKDGK-FTVRSFYRALKLQQTSF 839
Query: 438 VHQNMELIWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDTTH 497
++ IW +VPLK+ I ++++ TK NL RG +++ C VE H
Sbjct: 840 PNKK---IWKFRVPLKIRIFIWFFTKNKILTKDNLLKRGWRKGDNK--CQFCDKVETVQH 894
Query: 498 MFLSCPIFGALLSMVQTWVGVEGV 521
+F CP+ + +++ + V+ V
Sbjct: 895 LFFDCPLARLIWNIIACALNVKPV 918
>UniRef100_Q6Z0B6 Cyst nematode resistance protein-like protein [Oryza sativa]
Length = 184
Score = 47.0 bits (110), Expect = 0.001
Identities = 32/116 (27%), Positives = 50/116 (42%), Gaps = 6/116 (5%)
Query: 404 QELETDRWMWLSDQIRGYTVRGVYDMLTSQE*PHVHQNMELIWHKQVPLKVSILALRLLR 463
QE +T RW D ++V YD+ V + +L+W + P KV +
Sbjct: 13 QEDDTIRWKLTPDGF--FSVSSAYDLFFMAR--EVSLSGQLVWQTKAPSKVRFFLWLATK 68
Query: 464 DRLPTKSNLADRGITFVEDRICVTGCGHVEDTTHMFLSCPIFGALLSMVQTWVGVE 519
R T NLA RG + CV ED H+F+SC + +++ W+ V+
Sbjct: 69 SRCLTADNLAKRGWPHQDQ--CVLCQRQQEDCLHLFVSCDYTKRVWRLLRDWINVD 122
>UniRef100_Q947X7 Hypothetical protein OSJNBa0067N01.20 [Oryza sativa]
Length = 387
Score = 47.0 bits (110), Expect = 0.001
Identities = 24/69 (34%), Positives = 37/69 (52%), Gaps = 2/69 (2%)
Query: 445 IWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDTTHMFLSCPI 504
IW P +V A + ++RLPT+ NL + T + C ++EDT H+FL CP+
Sbjct: 229 IWRSAAPPRVKFFAWLMSKNRLPTRVNLHKK--TILPTPTCELCNANLEDTYHIFLRCPM 286
Query: 505 FGALLSMVQ 513
A +M+Q
Sbjct: 287 AAAFWNMIQ 295
>UniRef100_Q8H992 Oryza sativa (japonica cultivar-group) retrotransposon Karma DNA,
complete sequence [Oryza sativa]
Length = 1197
Score = 46.2 bits (108), Expect = 0.002
Identities = 28/90 (31%), Positives = 46/90 (51%), Gaps = 4/90 (4%)
Query: 414 LSDQIRGYTVRGVYDMLTSQE*PHVHQNMELIWHKQVPLKVSILALRLLRDRLPTKSNLA 473
L++++ T + +YD+ + P N + IW ++PLK+ + A L+RDRL TK NL
Sbjct: 1007 LNNRLGNLTTKTIYDLRSPPGFPS--PNWKFIWDCRMPLKIKLFAWLLVRDRLSTKLNLL 1064
Query: 474 DRGITFVEDRICVTGCGHVEDTTHMFLSCP 503
+ I V+ C E H+ +CP
Sbjct: 1065 KKKI--VQTATCDICSTTDETADHLSFNCP 1092
>UniRef100_Q8LID8 Cyst nematode resistance protein-like protein [Oryza sativa]
Length = 210
Score = 43.9 bits (102), Expect = 0.011
Identities = 32/137 (23%), Positives = 54/137 (39%), Gaps = 18/137 (13%)
Query: 385 VWEEDLVEECGDLLLTVSLQELETDRWMWLSDQIRGYTVRGVYDM--LTSQE*PHVHQNM 442
+W+E L+ G+ + D+ W ++V YDM + Q P
Sbjct: 7 LWDELLLFNLGN----------DPDKIRWKPGPDGAFSVSSAYDMFFMARQSYPFGQH-- 54
Query: 443 ELIWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDTTHMFLSC 502
IW P +V R R T NLA +G + + C ED H+F++C
Sbjct: 55 --IWQTNAPSRVRFFFWLAARGRCQTADNLAKKG--WPHEDSCALCMREQEDCHHLFVTC 110
Query: 503 PIFGALLSMVQTWVGVE 519
G + +++ W+ V+
Sbjct: 111 EFSGRVWELMRAWISVD 127
>UniRef100_Q7XET4 Contains similarity to non-LTR reverse transcriptase [Oryza sativa]
Length = 379
Score = 43.5 bits (101), Expect = 0.015
Identities = 24/61 (39%), Positives = 32/61 (52%), Gaps = 2/61 (3%)
Query: 443 ELIWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDTTHMFLSC 502
+ IW + PLKV A L++DRL TK NL + T V + IC G E +H+ C
Sbjct: 264 DAIWKCKAPLKVKFFAWLLVKDRLSTKKNLHKK--TIVPNDICDICNGATETASHLCFFC 321
Query: 503 P 503
P
Sbjct: 322 P 322
>UniRef100_Q5ZD87 HcrVf2 protein-like [Oryza sativa]
Length = 1064
Score = 43.1 bits (100), Expect = 0.020
Identities = 26/77 (33%), Positives = 39/77 (49%), Gaps = 2/77 (2%)
Query: 396 DLLLTVSLQELETDRWMWLSDQIRGYTVRGVYDMLTSQE*PHVHQNMELIWHKQVPLKVS 455
D LL+V L E R S Q+ + + +Y+ +Q N + +WH + PLK
Sbjct: 778 DKLLSVHLHECPDFRAAKCSPQLPILSAKHIYE--ATQVWLDACPNWKFVWHNKAPLKAQ 835
Query: 456 ILALRLLRDRLPTKSNL 472
+ A L ++RLPTK NL
Sbjct: 836 LFAWLLAKERLPTKHNL 852
>UniRef100_Q6L3X8 Hypothetical protein [Solanum demissum]
Length = 1155
Score = 43.1 bits (100), Expect = 0.020
Identities = 32/136 (23%), Positives = 57/136 (41%), Gaps = 21/136 (15%)
Query: 375 GAWQWRRHLWVWEEDLVEECGDLL--LTVSLQELETDRWMWLSDQIRGYTVRGVYDMLTS 432
G W W + + L + ++ L ++L E D+ +W +TV +++
Sbjct: 840 GKWNWS----ILQNQLPNQIKVMITGLGLNLNNEEQDKPVWSPTSAGNFTVTSAWNICR- 894
Query: 433 QE*PHVHQNMEL-----IWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVT 487
H+ +E+ IWHK +P K+S L + + +RLPT + GI C T
Sbjct: 895 ------HKGIEITDFNKIWHKDIPFKMSFLTWKAIINRLPTGAKFKRLGIPLSPTCYCCT 948
Query: 488 GCG---HVEDTTHMFL 500
+E H+F+
Sbjct: 949 NNNIPTTLESAEHIFM 964
>UniRef100_Q8H0A5 Putative reverse transcriptase [Oryza sativa]
Length = 1557
Score = 42.0 bits (97), Expect = 0.043
Identities = 30/106 (28%), Positives = 43/106 (40%), Gaps = 11/106 (10%)
Query: 407 ETDRWMWLSDQIRGYTVRGVYDML---------TSQE*PHVHQNMELIWHKQVPLKVSIL 457
E D W D++ ++VR Y + +S ++ + EL+W VP KV I
Sbjct: 1366 EEDFIAWHPDKLGNFSVRTAYRLAENWAKEEASSSSSDVNIRKAWELLWKCNVPSKVKIF 1425
Query: 458 ALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDTTHMFLSCP 503
R + LPT N R + + CV EDT H CP
Sbjct: 1426 TWRATSNCLPTWDNKKKRNLEISD--TCVICGMEKEDTMHALCRCP 1469
>UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa]
Length = 1936
Score = 40.8 bits (94), Expect = 0.097
Identities = 38/162 (23%), Positives = 67/162 (40%), Gaps = 23/162 (14%)
Query: 367 LVGGGGWWGAWQWRRHLWVWEEDLVEECGDLLLTVSLQELETDRWMWLSDQIRGYTVRGV 426
L+ G W + + ++ + D++++ + +S + LE D W D+ ++VR
Sbjct: 1603 LIAEDGTWDSAKINQYFLKIDADIIQK-----ICISAR-LEEDFIAWHPDKTGRFSVRSA 1656
Query: 427 YDML---------TSQE*PHVHQNMELIWHKQVPLKVSILALRLLRDRLPTKSNLADRGI 477
Y + +S ++++ ELIW VP KV I A R+ + L T N R +
Sbjct: 1657 YKLALQLADMNNCSSSSSSRLNKSWELIWKCNVPQKVRIFAWRVASNSLATMENKKKRNL 1716
Query: 478 TFVEDRICVTGCGHVEDTTHMFLSCPIFGALLSMVQTWVGVE 519
+ +C ED H C +L WV +E
Sbjct: 1717 ERFD--VCGICDREKEDAGHALCRCVHANSL------WVNLE 1750
>UniRef100_Q9LHK6 Similarity to non-LTR retroelement reverse transcriptase
[Arabidopsis thaliana]
Length = 332
Score = 40.8 bits (94), Expect = 0.097
Identities = 33/132 (25%), Positives = 52/132 (39%), Gaps = 15/132 (11%)
Query: 382 HLWVWEE---DLVEECGDLLLTVSLQELETDRWMWLSDQIRGYTVRGVYDMLTSQE*PHV 438
H W E+ +L+ D+L + DR WL + Y+ + Y M S P +
Sbjct: 63 HQWDCEDIRRELLHHEEDILSLILSSTKRPDRLWWLPAKAGVYSTKSGYAMAKSNLVPCL 122
Query: 439 HQNMELI---WHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGC---GHV 492
+ W + + + + LP + L+ RGI + TGC G +
Sbjct: 123 EEGFNWKTNNWKVYTSPMIRMFLSKATKKALPIGTALSARGI------VVETGCKRCGEI 176
Query: 493 EDTTHMFLSCPI 504
ED H+F CPI
Sbjct: 177 EDALHIFFRCPI 188
>UniRef100_Q7XS38 OSJNBa0081G05.2 protein [Oryza sativa]
Length = 1711
Score = 40.0 bits (92), Expect = 0.17
Identities = 34/145 (23%), Positives = 61/145 (41%), Gaps = 17/145 (11%)
Query: 367 LVGGGGWWGAWQWRRHLWVWEEDLVEECGDLLLTVSLQELETDRWMWLSDQIRGYTVRGV 426
L+ G W + + ++ + D++++ + +S + LE D W D+ ++VR
Sbjct: 1447 LIAEDGTWDSAKINQYFLKIDADIIQK-----ICISAR-LEEDFIAWHPDKTGRFSVRSA 1500
Query: 427 YDML---------TSQE*PHVHQNMELIWHKQVPLKVSILALRLLRDRLPTKSNLADRGI 477
Y + +S ++++ ELIW VP KV I A R+ + L T N R +
Sbjct: 1501 YKLALQLADMNNCSSSSSSRLNKSWELIWKCNVPQKVRIFAWRVASNSLATMENKKKRNL 1560
Query: 478 TFVEDRICVTGCGHVEDTTHMFLSC 502
+ +C ED H C
Sbjct: 1561 ERFD--VCGICDREKEDAGHALCCC 1583
>UniRef100_Q6AUJ7 Hypothetical protein OSJNBa0052E20.4 [Oryza sativa]
Length = 144
Score = 39.3 bits (90), Expect = 0.28
Identities = 25/78 (32%), Positives = 38/78 (48%), Gaps = 4/78 (5%)
Query: 437 HVHQNMELIWHKQVPLKVSILALRLLRDRLPTKSNLADRG-ITFVEDRICVTGCGHVEDT 495
H+ E IW P +V + +RLPT NL + I + ++C T C EDT
Sbjct: 5 HLWPPWEFIWKSSAPPRVKFFGWLMTMNRLPTAVNLHKKSTIPSLTCQLCNT-C--PEDT 61
Query: 496 THMFLSCPIFGALLSMVQ 513
H+FL+ P+ A ++Q
Sbjct: 62 DHIFLAYPLASAFWGLIQ 79
>UniRef100_Q94H40 Putative reverse transcriptase [Oryza sativa]
Length = 1185
Score = 38.5 bits (88), Expect = 0.48
Identities = 37/157 (23%), Positives = 63/157 (39%), Gaps = 28/157 (17%)
Query: 386 WEEDLVEEC-----GDLLLTVSLQELETDRWMWLSDQIRGYTVRGVY------------- 427
W+EDL++ + +L + + D W + ++VR Y
Sbjct: 788 WDEDLIDTLFWPIDANRILQIPISAGREDFVAWHHTRSGIFSVRSAYHCQWNAKFGSKER 847
Query: 428 --DMLTSQE*PHVHQNMELIWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVED-RI 484
D +S P V N+ W ++P K+ I A R+L +P K LAD+ I +
Sbjct: 848 LPDGASSSSRP-VWDNL---WKLEIPPKIKIFAWRMLHGVIPCKGILADKHIAVQGGCPV 903
Query: 485 CVTGCGHVEDTTHMFLSCPIFGALLSMVQTWVGVEGV 521
C +G VED HM +C + + W ++ +
Sbjct: 904 CSSG---VEDIKHMVFTCRRAKLIWKQLGIWSRIQPI 937
>UniRef100_Q64M76 Phosphoribosyl pyrophosphate synthase-like [Oryza sativa]
Length = 183
Score = 38.1 bits (87), Expect = 0.63
Identities = 21/64 (32%), Positives = 30/64 (46%), Gaps = 4/64 (6%)
Query: 441 NMELIWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDTT-HMF 499
N + +W + P+KV A L +DR P K NL + T V C C ++T H+
Sbjct: 6 NWKFVWENKAPMKVQFFAWLLTKDRFPIKRNLHKK--TIVPAPTCEL-CNSADETALHLC 62
Query: 500 LSCP 503
CP
Sbjct: 63 FQCP 66
>UniRef100_Q8SA91 Putative gag-pol polyprotein [Zea mays]
Length = 2396
Score = 38.1 bits (87), Expect = 0.63
Identities = 22/59 (37%), Positives = 31/59 (52%), Gaps = 4/59 (6%)
Query: 445 IWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHV-EDTTHMFLSC 502
+W Q P +V + R+ + NLA + I V+D +C CG EDTTH+FL C
Sbjct: 2294 VWRNQAPPRVCFFGWLVNNGRIQCRVNLARKQI--VQDTMCEV-CGRAPEDTTHIFLHC 2349
>UniRef100_O22148 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1374
Score = 37.7 bits (86), Expect = 0.82
Identities = 29/105 (27%), Positives = 44/105 (41%), Gaps = 12/105 (11%)
Query: 409 DRWMWLSDQIRGYTVRGVYDMLTS--------QE*--PHVHQNMELIWHKQVPLKVSILA 458
DR+ W + Y+V+ Y ++T QE P + + IW VP K+
Sbjct: 1008 DRFTWEYSRSGHYSVKSGYWVMTEIINQRNNPQEVLQPSLDPIFQQIWKLDVPPKIHHFL 1067
Query: 459 LRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDTTHMFLSCP 503
R + + L SNLA R + ++ CV H E H+ CP
Sbjct: 1068 WRCVNNCLSVASNLAYRHL--AREKSCVRCPSHGETVNHLLFKCP 1110
>UniRef100_Q7XMS7 OSJNBa0029L02.10 protein [Oryza sativa]
Length = 389
Score = 36.6 bits (83), Expect = 1.8
Identities = 30/115 (26%), Positives = 52/115 (45%), Gaps = 8/115 (6%)
Query: 407 ETDRWMWLSDQIRGYTVRGVYDMLTSQE*PHVHQNMELIWHKQVPLKVSILALRLLRDRL 466
+T RW D +TV+ + L Q+ H+ IW +VPLK+ I ++++
Sbjct: 92 DTLRWNLTKDG--KFTVKSFHTALKMQQVIFPHKK---IWGLKVPLKIKIFIWMFTKNKI 146
Query: 467 PTKSNLADRGITFVEDRICVTGCGHVEDTTHMFLSCPIFGALLSMVQTWVGVEGV 521
TK NL RG + + + C E +F CP+ L ++V+ + + V
Sbjct: 147 LTKDNLFKRG--WRKGKANCQFCDQTETLQPLFY-CPLARLLWNIVKCALNINSV 198
>UniRef100_Q5U6D4 Orf147a protein [Beta vulgaris subsp. vulgaris]
Length = 147
Score = 36.6 bits (83), Expect = 1.8
Identities = 23/70 (32%), Positives = 37/70 (52%), Gaps = 4/70 (5%)
Query: 451 PLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDTT-HMFLSCPIFGALL 509
P K + + + DRL T L GI V D++CV CG+V++T H+F C +
Sbjct: 11 PPKCTFITWLTILDRLATCDRLQKFGI--VCDQLCVL-CGNVDETRDHLFFVCEFSYEIW 67
Query: 510 SMVQTWVGVE 519
S + W+G++
Sbjct: 68 SSLLCWLGIQ 77
>UniRef100_Q5WMS3 Hypothetical protein OJ1333_C12.8 [Oryza sativa]
Length = 244
Score = 36.2 bits (82), Expect = 2.4
Identities = 24/71 (33%), Positives = 39/71 (54%), Gaps = 3/71 (4%)
Query: 408 TDRWMWLSDQIRGYTVRGVYDMLTSQE*PHVHQNMELIWHKQVPLKVSILALRLLRDRLP 467
+D + W Q ++VR +Y L + ++ +N +LIW ++PLK+ I LL+ +
Sbjct: 38 SDYFRWNYHQNGQFSVRSMYLALINNG--YIDRN-KLIWKLKMPLKIKIFMWYLLKGVVL 94
Query: 468 TKSNLADRGIT 478
TK NLA R T
Sbjct: 95 TKDNLARRNWT 105
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.364 0.164 0.688
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 730,552,174
Number of Sequences: 2790947
Number of extensions: 25732087
Number of successful extensions: 142294
Number of sequences better than 10.0: 39
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 35
Number of HSP's that attempted gapping in prelim test: 142265
Number of HSP's gapped (non-prelim): 47
length of query: 523
length of database: 848,049,833
effective HSP length: 132
effective length of query: 391
effective length of database: 479,644,829
effective search space: 187541128139
effective search space used: 187541128139
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 77 (34.3 bits)
Medicago: description of AC149205.3


