

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149205.17 + phase: 0 /pseudo
(626 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
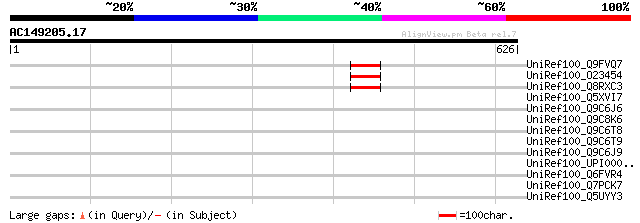
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FVQ7 Hypothetical protein F3C3.9 [Arabidopsis thaliana] 51 9e-05
UniRef100_O23454 Hypothetical protein dl4065w [Arabidopsis thali... 48 8e-04
UniRef100_Q8RXC3 Hypothetical protein At4g16050 [Arabidopsis tha... 48 8e-04
UniRef100_Q5XVI7 Hypothetical protein [Arabidopsis thaliana] 46 0.004
UniRef100_Q9C6J6 Hypothetical protein F8A12.7 [Arabidopsis thali... 46 0.004
UniRef100_Q9C8K6 Hypothetical protein F5D21.24 [Arabidopsis thal... 42 0.071
UniRef100_Q9C6T8 Hypothetical protein F4M15.2 [Arabidopsis thali... 39 0.35
UniRef100_Q9C6T9 Hypothetical protein F4M15.1 [Arabidopsis thali... 39 0.60
UniRef100_Q9C6J9 Hypothetical protein F8A12.4 [Arabidopsis thali... 38 1.0
UniRef100_UPI000036B834 UPI000036B834 UniRef100 entry 36 3.9
UniRef100_Q6FVR4 Candida glabrata strain CBS138 chromosome D com... 36 3.9
UniRef100_Q7PCK7 X-ray radiation resistance associated 1 protein... 35 8.6
UniRef100_Q5UYY3 Hypothetical protein [Haloarcula marismortui] 35 8.6
>UniRef100_Q9FVQ7 Hypothetical protein F3C3.9 [Arabidopsis thaliana]
Length = 1206
Score = 51.2 bits (121), Expect = 9e-05
Identities = 19/38 (50%), Positives = 27/38 (71%)
Query: 421 IAWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKSV 458
+AWK+Y RP++D LYFP++ EA +T Y +WW SV
Sbjct: 427 LAWKDYIRPIADGMLYFPARLHEADVTVGYIRWWKLSV 464
>UniRef100_O23454 Hypothetical protein dl4065w [Arabidopsis thaliana]
Length = 900
Score = 48.1 bits (113), Expect = 8e-04
Identities = 19/37 (51%), Positives = 24/37 (64%)
Query: 422 AWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKSV 458
AW +Y++PL LYFPS+ A +T RY WW KSV
Sbjct: 644 AWDDYNKPLVGLKLYFPSRVATASVTTRYRDWWAKSV 680
>UniRef100_Q8RXC3 Hypothetical protein At4g16050 [Arabidopsis thaliana]
Length = 666
Score = 48.1 bits (113), Expect = 8e-04
Identities = 19/37 (51%), Positives = 24/37 (64%)
Query: 422 AWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKSV 458
AW +Y++PL LYFPS+ A +T RY WW KSV
Sbjct: 410 AWDDYNKPLVGLKLYFPSRVATASVTTRYRDWWAKSV 446
>UniRef100_Q5XVI7 Hypothetical protein [Arabidopsis thaliana]
Length = 768
Score = 45.8 bits (107), Expect = 0.004
Identities = 19/55 (34%), Positives = 31/55 (55%)
Query: 404 NSDLIKMFLVNQTDPKAIAWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKSV 458
+ DL + + + AW +Y++ L +LY PS+ + +TARY WW+KSV
Sbjct: 400 DQDLPGLATCQRNSTEKEAWNDYNKSLIGLNLYMPSRLDQGSVTARYRVWWLKSV 454
>UniRef100_Q9C6J6 Hypothetical protein F8A12.7 [Arabidopsis thaliana]
Length = 768
Score = 45.8 bits (107), Expect = 0.004
Identities = 19/55 (34%), Positives = 31/55 (55%)
Query: 404 NSDLIKMFLVNQTDPKAIAWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKSV 458
+ DL + + + AW +Y++ L +LY PS+ + +TARY WW+KSV
Sbjct: 400 DQDLPGLATCQRNSTEKEAWNDYNKSLIGLNLYMPSRLDQGSVTARYRVWWLKSV 454
>UniRef100_Q9C8K6 Hypothetical protein F5D21.24 [Arabidopsis thaliana]
Length = 1036
Score = 41.6 bits (96), Expect = 0.071
Identities = 15/37 (40%), Positives = 21/37 (56%)
Query: 422 AWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKSV 458
AW Y++ L LY PS+ +T RY WW+KS+
Sbjct: 326 AWDGYNKSLDGLMLYIPSRVATTSVTERYRDWWLKSI 362
>UniRef100_Q9C6T8 Hypothetical protein F4M15.2 [Arabidopsis thaliana]
Length = 816
Score = 39.3 bits (90), Expect = 0.35
Identities = 15/46 (32%), Positives = 29/46 (62%), Gaps = 1/46 (2%)
Query: 413 VNQTD-PKAIAWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKS 457
VNQ + + AW +Y++P++D +L+ PS+ +T + +WW +S
Sbjct: 321 VNQNNLSQEAAWNDYNKPINDLALFIPSRSAIPRVTPTFCEWWRRS 366
>UniRef100_Q9C6T9 Hypothetical protein F4M15.1 [Arabidopsis thaliana]
Length = 649
Score = 38.5 bits (88), Expect = 0.60
Identities = 12/36 (33%), Positives = 24/36 (66%)
Query: 422 AWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKS 457
AW +Y++P+ D +L+ PS+ +T+ + +WW K+
Sbjct: 393 AWNDYNKPIDDLALHIPSRSAIPRVTSTFCEWWRKT 428
>UniRef100_Q9C6J9 Hypothetical protein F8A12.4 [Arabidopsis thaliana]
Length = 812
Score = 37.7 bits (86), Expect = 1.0
Identities = 34/129 (26%), Positives = 50/129 (38%), Gaps = 11/129 (8%)
Query: 422 AWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKSVLRSQ-------GFVTNFEPRKRS 474
AW +Y +PL D LY PS+ + T + W K V+ S G T+F P S
Sbjct: 405 AWNDYYKPLDDLELYIPSRSAISCDTPMFCDSWRKRVVGSAKTLRTSIGDDTSFVP-SGS 463
Query: 475 ASSSKCEPSAKIPPEFPSHKLVGCTVTIGKSSDDGSNTSQGDKIVDDDAPSGSIPKLLKT 534
+ + E KI + +L +DGS Q I DD S+ T
Sbjct: 464 KMNRRSEDGRKIAEDSTKKRL---KFMKSARKNDGSTMGQKQVISDDADDDDSLTVAQIT 520
Query: 535 MSFEKSVED 543
++K +
Sbjct: 521 TLYKKKYSE 529
>UniRef100_UPI000036B834 UPI000036B834 UniRef100 entry
Length = 522
Score = 35.8 bits (81), Expect = 3.9
Identities = 36/137 (26%), Positives = 56/137 (40%), Gaps = 24/137 (17%)
Query: 505 SSDDGSNTSQGDKIVDDDAPSGSIPKLLKTMSFEKSVEDGLLA----EKYVDGGAPLPAK 560
S + SNT D IVD DA S I K+ ++ SFE+++++ + EK D L +K
Sbjct: 154 SKAETSNTDF-DNIVDPDAYSSDIEKIEESASFERNLKEKNIGLESNEKSDDSCVSLESK 212
Query: 561 D--------NTPTPLIFVDDCKHVL-----------EDGNESEEARLSTDTICQSENQTE 601
D P D K L ED ++ + L + C+S+ T+
Sbjct: 213 DTLLGIDLEKAPIEEKLSQDIKESLEFSNLHKRPSFEDSKTTKSSLLLQEIACRSKPITK 272
Query: 602 SYSYLSELIVEELNREL 618
Y L + + N L
Sbjct: 273 QYQGLERFFIFDTNERL 289
>UniRef100_Q6FVR4 Candida glabrata strain CBS138 chromosome D complete sequence
[Candida glabrata]
Length = 990
Score = 35.8 bits (81), Expect = 3.9
Identities = 31/135 (22%), Positives = 59/135 (42%), Gaps = 25/135 (18%)
Query: 503 GKSSDDGSNTSQGDKIVDDDAPSGSIPKLLKTMSFEKSVEDGLLAEKYVDGGAPLPAKDN 562
G+S++ + DK V ++ PS ++ S K E+ + + D + KDN
Sbjct: 280 GESTEASEASDSTDKAVGNNEPSTD-----ESDSNNKDAENSEDQDTFFDSKDTITEKDN 334
Query: 563 ---------TPTPLIFVDDCK--------HVLEDGNESEEARLSTDTICQSENQTESYSY 605
+P PL F + K H ED N+ E +++ DT +EN ++ +
Sbjct: 335 ELESGSKDSSPRPLTFAEKLKLKKKQTEEHKEEDSNKEEHEQVNIDT---TENSGDNNQH 391
Query: 606 LSELIVEELNRELAD 620
++E+ +E + +D
Sbjct: 392 VNEVELESKEKNESD 406
>UniRef100_Q7PCK7 X-ray radiation resistance associated 1 protein [Bos taurus]
Length = 508
Score = 34.7 bits (78), Expect = 8.6
Identities = 26/99 (26%), Positives = 47/99 (47%), Gaps = 9/99 (9%)
Query: 513 SQGDKIVDDDAPSGSIPKLLKTMSFEKSVE-DGLLAEKYVDGGAPLPAKDNTPTPLIFVD 571
S ++++ +A + P KT+S E+ + +GL GG P+P + P P I D
Sbjct: 207 SPSKEMLESEAELTTEPPPTKTISVEREMPIEGL-------GGPPMPHRTFVPLPPICSD 259
Query: 572 DCKHVLEDGNESEEA-RLSTDTICQSENQTESYSYLSEL 609
H E S+ A RLS D + + ++ +L+++
Sbjct: 260 STVHSEETSPRSDAAGRLSADQLSDEDTKSTESIFLTQV 298
>UniRef100_Q5UYY3 Hypothetical protein [Haloarcula marismortui]
Length = 729
Score = 34.7 bits (78), Expect = 8.6
Identities = 32/114 (28%), Positives = 46/114 (40%), Gaps = 7/114 (6%)
Query: 508 DGSNTSQGDKIVDDDAPSGSIPKLLKTMSFEKSVEDGLLAEKYVDGGAPLPAKDNTPTPL 567
D S+T ++ D+ PS P L T + + + AE VDG PL P L
Sbjct: 14 DDSSTDGDERTAKDNGPSDDGPDRLDTAALDPTTFREY-AENVVDGADPLTFD---PDAL 69
Query: 568 IFVDDCKHVLEDGNESEEARLSTDTICQSENQTESYSYLSELIVEELNRELADS 621
+D +E+ E + T +EN SYL EL+V + E S
Sbjct: 70 ARLDTA---IEESYGGETVGDAAGTTTYTENTVRFGSYLGELLVRAYDGEWVQS 120
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.350 0.153 0.557
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 936,562,535
Number of Sequences: 2790947
Number of extensions: 35780003
Number of successful extensions: 148332
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 148317
Number of HSP's gapped (non-prelim): 19
length of query: 626
length of database: 848,049,833
effective HSP length: 134
effective length of query: 492
effective length of database: 474,062,935
effective search space: 233238964020
effective search space used: 233238964020
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 78 (34.7 bits)
Medicago: description of AC149205.17


