

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
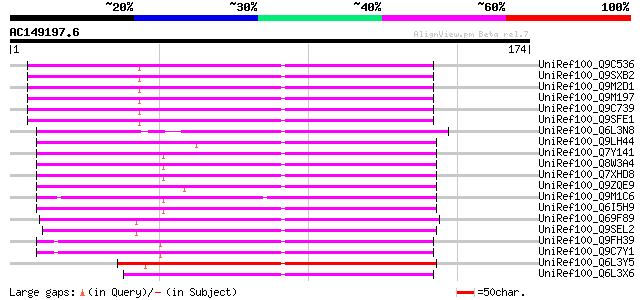
Query= AC149197.6 + phase: 0 /pseudo
(174 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9C536 Copia-type polyprotein, putative [Arabidopsis t... 100 3e-20
UniRef100_Q9SXB2 T28P6.8 protein [Arabidopsis thaliana] 100 3e-20
UniRef100_Q9M2D1 Copia-type polyprotein [Arabidopsis thaliana] 100 3e-20
UniRef100_Q9M197 Copia-type reverse transcriptase-like protein [... 100 3e-20
UniRef100_Q9C739 Copia-type polyprotein, putative [Arabidopsis t... 100 3e-20
UniRef100_Q9SFE1 T26F17.17 [Arabidopsis thaliana] 99 5e-20
UniRef100_Q6L3N8 Putative gag-pol polyprotein [Solanum demissum] 95 9e-19
UniRef100_Q9LH44 Copia-like retrotransposable element [Arabidops... 93 2e-18
UniRef100_Q7Y141 Putative polyprotein [Oryza sativa] 87 2e-16
UniRef100_Q8W3A4 Putative gag-pol polyprotein [Oryza sativa] 87 2e-16
UniRef100_Q7XHD8 Putative copia-type polyprotein [Oryza sativa] 86 3e-16
UniRef100_Q9ZQE9 Putative retroelement pol polyprotein [Arabidop... 81 1e-14
UniRef100_Q9M1C6 Hypothetical protein T2O9.150 [Arabidopsis thal... 81 1e-14
UniRef100_Q6I5H9 Putative polyprotein [Oryza sativa] 80 2e-14
UniRef100_Q69F89 Gag-pol polyprotein [Phaseolus vulgaris] 80 2e-14
UniRef100_Q9SEL2 Gag-pol polyprotein [Vitis vinifera] 76 4e-13
UniRef100_Q9FH39 Copia-type polyprotein [Arabidopsis thaliana] 73 3e-12
UniRef100_Q9C7Y1 Copia-type polyprotein, putative; 28768-32772 [... 73 3e-12
UniRef100_Q6L3Y5 Putative gag-pol polyprotein [Solanum demissum] 69 7e-11
UniRef100_Q6L3X6 Putative polyprotein [Solanum demissum] 67 2e-10
>UniRef100_Q9C536 Copia-type polyprotein, putative [Arabidopsis thaliana]
Length = 1320
Score = 99.8 bits (247), Expect = 3e-20
Identities = 53/138 (38%), Positives = 78/138 (56%), Gaps = 3/138 (2%)
Query: 7 PTHLPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSV--PALASNATDMQ*TANRDAKKKD 64
P +PV NYD W ++MK I DV E+V P + + Q RD++K+D
Sbjct: 7 PFQVPVLTKSNYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRD 66
Query: 65 GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTM 124
KA+ ++Q + + FEK+V SAKE WE L Y G ++KKVRLQ LR ++E L M
Sbjct: 67 KKALCLIYQGLDEDTFEKVVEATSAKE-AWEKLRTSYKGADQVKKVRLQTLRGEFEALQM 125
Query: 125 EEEETISQYFDKFVNLSN 142
+E E +S YF + + ++N
Sbjct: 126 KEGELVSDYFSRVLTVTN 143
>UniRef100_Q9SXB2 T28P6.8 protein [Arabidopsis thaliana]
Length = 1352
Score = 99.8 bits (247), Expect = 3e-20
Identities = 53/138 (38%), Positives = 78/138 (56%), Gaps = 3/138 (2%)
Query: 7 PTHLPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSV--PALASNATDMQ*TANRDAKKKD 64
P +PV NYD W ++MK I DV E+V P + + Q RD++K+D
Sbjct: 7 PFQVPVLTKSNYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRD 66
Query: 65 GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTM 124
KA+ ++Q + + FEK+V SAKE WE L Y G ++KKVRLQ LR ++E L M
Sbjct: 67 KKALCLIYQGLDEDTFEKVVEATSAKE-AWEKLRTSYKGADQVKKVRLQTLRGEFEALQM 125
Query: 125 EEEETISQYFDKFVNLSN 142
+E E +S YF + + ++N
Sbjct: 126 KEGELVSDYFSRVLTVTN 143
>UniRef100_Q9M2D1 Copia-type polyprotein [Arabidopsis thaliana]
Length = 1352
Score = 99.8 bits (247), Expect = 3e-20
Identities = 53/138 (38%), Positives = 78/138 (56%), Gaps = 3/138 (2%)
Query: 7 PTHLPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSV--PALASNATDMQ*TANRDAKKKD 64
P +PV NYD W ++MK I DV E+V P + + Q RD++K+D
Sbjct: 7 PFQVPVLTKSNYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRD 66
Query: 65 GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTM 124
KA+ ++Q + + FEK+V SAKE WE L Y G ++KKVRLQ LR ++E L M
Sbjct: 67 KKALCLIYQGLDEDTFEKVVEATSAKE-AWEKLRTSYKGADQVKKVRLQTLRGEFEALQM 125
Query: 125 EEEETISQYFDKFVNLSN 142
+E E +S YF + + ++N
Sbjct: 126 KEGELVSDYFSRVLTVTN 143
>UniRef100_Q9M197 Copia-type reverse transcriptase-like protein [Arabidopsis
thaliana]
Length = 1272
Score = 99.8 bits (247), Expect = 3e-20
Identities = 53/138 (38%), Positives = 78/138 (56%), Gaps = 3/138 (2%)
Query: 7 PTHLPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSV--PALASNATDMQ*TANRDAKKKD 64
P +PV NYD W ++MK I DV E+V P + + Q RD++K+D
Sbjct: 7 PFQVPVLTKSNYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRD 66
Query: 65 GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTM 124
KA+ ++Q + + FEK+V SAKE WE L Y G ++KKVRLQ LR ++E L M
Sbjct: 67 KKALCLIYQGLDEDTFEKVVEATSAKE-AWEKLRTSYKGADQVKKVRLQTLRGEFEALQM 125
Query: 125 EEEETISQYFDKFVNLSN 142
+E E +S YF + + ++N
Sbjct: 126 KEGELVSDYFSRVLTVTN 143
>UniRef100_Q9C739 Copia-type polyprotein, putative [Arabidopsis thaliana]
Length = 1352
Score = 99.8 bits (247), Expect = 3e-20
Identities = 53/138 (38%), Positives = 78/138 (56%), Gaps = 3/138 (2%)
Query: 7 PTHLPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSV--PALASNATDMQ*TANRDAKKKD 64
P +PV NYD W ++MK I DV E+V P + + Q RD++K+D
Sbjct: 7 PFQVPVLTKSNYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRD 66
Query: 65 GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTM 124
KA+ ++Q + + FEK+V SAKE WE L Y G ++KKVRLQ LR ++E L M
Sbjct: 67 KKALCLIYQGLDEDTFEKVVEATSAKE-AWEKLRTSYKGADQVKKVRLQTLRGEFEALQM 125
Query: 125 EEEETISQYFDKFVNLSN 142
+E E +S YF + + ++N
Sbjct: 126 KEGELVSDYFSRVLTVTN 143
>UniRef100_Q9SFE1 T26F17.17 [Arabidopsis thaliana]
Length = 1291
Score = 99.0 bits (245), Expect = 5e-20
Identities = 53/138 (38%), Positives = 78/138 (56%), Gaps = 3/138 (2%)
Query: 7 PTHLPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSV--PALASNATDMQ*TANRDAKKKD 64
P +PV NYD W ++MK I DV E+V P + + Q RD++K+D
Sbjct: 7 PFQVPVLTKSNYDNWSLQMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRD 66
Query: 65 GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTM 124
KA+ ++Q + + FEK+V SAKE WE L Y G ++KKVRLQ LR ++E L M
Sbjct: 67 KKALCLIYQGLDEDTFEKVVEATSAKE-AWEKLRTSYKGVDQVKKVRLQTLRGEFEALQM 125
Query: 125 EEEETISQYFDKFVNLSN 142
+E E +S YF + + ++N
Sbjct: 126 KEGELVSDYFSRVLTVTN 143
>UniRef100_Q6L3N8 Putative gag-pol polyprotein [Solanum demissum]
Length = 1333
Score = 94.7 bits (234), Expect = 9e-19
Identities = 48/138 (34%), Positives = 83/138 (59%), Gaps = 8/138 (5%)
Query: 10 LPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNATDMQ*TANRDAKKKDGKAMF 69
+P+F G+NY W +KMK + + Q++ ++V +P NA M R+ +K+D KA+F
Sbjct: 14 IPIFRGENYQFWSLKMKTLFKSQELWDIVETGIPE--GNANQM-----REHRKRDSKALF 66
Query: 70 FMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEEET 129
+ Q++ +EIF +I E++K+ WE L++ Y GD K+ V+LQ LRR +E L M E E+
Sbjct: 67 TIQQALDDEIFPRISAVETSKQ-AWEILKQEYFGDDKVITVKLQTLRRDFETLFMNENES 125
Query: 130 ISQYFDKFVNLSNQSQEW 147
+ Y + + N+ + +
Sbjct: 126 VQGYLSRTSAIVNRMRSY 143
>UniRef100_Q9LH44 Copia-like retrotransposable element [Arabidopsis thaliana]
Length = 1499
Score = 93.2 bits (230), Expect = 2e-18
Identities = 52/135 (38%), Positives = 79/135 (58%), Gaps = 2/135 (1%)
Query: 10 LPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNATDMQ*TANRDAK-KKDGKAM 68
+P+F G++Y W +KM I + + + +V+ + V + +S T T RD + KD A+
Sbjct: 9 IPIFNGESYGFWKIKMITILKTRKLWDVIENGVTSNSSPETSPALTRERDDQVMKDMMAL 68
Query: 69 FFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEEE 128
+ +VS+ IF +I SA E W ALE + G ++K + LQ LRR+YE L MEE E
Sbjct: 69 QILQSAVSDSIFPRIAPASSATE-AWNALEMEFQGSSQVKMINLQTLRREYENLKMEEGE 127
Query: 129 TISQYFDKFVNLSNQ 143
TI+ + K +NLSNQ
Sbjct: 128 TINDFTTKLINLSNQ 142
>UniRef100_Q7Y141 Putative polyprotein [Oryza sativa]
Length = 1335
Score = 86.7 bits (213), Expect = 2e-16
Identities = 43/136 (31%), Positives = 78/136 (56%), Gaps = 3/136 (2%)
Query: 10 LPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNAT--DMQ*TANRDAKKKDGKA 67
+PVF G+NYD W +KM+ + Q + ++V + ++ T Q + + + D KA
Sbjct: 6 VPVFAGENYDIWSIKMRTLLLSQGLWDIVENGYQEYSAGETLTAEQKKSLAEDRMSDAKA 65
Query: 68 MFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEE 127
+F + Q V+ +F +I+ + +KE W+ L++ + G K+ V+LQ LRRQ++ L M+E
Sbjct: 66 LFLIQQGVAESLFPRIIGAKKSKE-AWDKLKEEFQGSQKVLAVKLQTLRRQFQNLLMKES 124
Query: 128 ETISQYFDKFVNLSNQ 143
E + YF + + + NQ
Sbjct: 125 EKVKDYFSRVIEIVNQ 140
>UniRef100_Q8W3A4 Putative gag-pol polyprotein [Oryza sativa]
Length = 1167
Score = 86.7 bits (213), Expect = 2e-16
Identities = 43/136 (31%), Positives = 78/136 (56%), Gaps = 3/136 (2%)
Query: 10 LPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNAT--DMQ*TANRDAKKKDGKA 67
+PVF G+NYD W +KM+ + Q + ++V + ++ T Q + + + D KA
Sbjct: 6 VPVFAGENYDIWSIKMRTLLLSQGLWDIVENGYQEYSAGETLTAEQKKSLAEDRMSDAKA 65
Query: 68 MFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEE 127
+F + Q V+ +F +I+ + +KE W+ L++ + G K+ V+LQ LRRQ++ L M+E
Sbjct: 66 LFLIQQGVAESLFPRIIGAKKSKE-AWDKLKEEFQGSQKVLAVKLQTLRRQFQNLLMKES 124
Query: 128 ETISQYFDKFVNLSNQ 143
E + YF + + + NQ
Sbjct: 125 EKVKDYFSRVIEIVNQ 140
>UniRef100_Q7XHD8 Putative copia-type polyprotein [Oryza sativa]
Length = 1350
Score = 86.3 bits (212), Expect = 3e-16
Identities = 43/136 (31%), Positives = 78/136 (56%), Gaps = 3/136 (2%)
Query: 10 LPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNAT--DMQ*TANRDAKKKDGKA 67
+PVF G+NYD W +KM+ + Q + ++V + ++ T Q + + + D KA
Sbjct: 6 VPVFAGENYDIWSIKMRTLLLSQGLWDIVENGYQEYSAGETLTAEQKKSLAEDRMSDAKA 65
Query: 68 MFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEE 127
+F + Q V+ +F +I+ + +KE W+ L++ + G K+ V+LQ LRRQ++ L M+E
Sbjct: 66 LFLIQQGVAESLFPRIIGAKKSKE-AWDKLKEEFQGSQKVLAVKLQTLRRQFQNLLMKES 124
Query: 128 ETISQYFDKFVNLSNQ 143
E + YF + + + NQ
Sbjct: 125 EKVKDYFPRVIEIVNQ 140
>UniRef100_Q9ZQE9 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1347
Score = 81.3 bits (199), Expect = 1e-14
Identities = 46/139 (33%), Positives = 75/139 (53%), Gaps = 6/139 (4%)
Query: 10 LPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNATDMQ*TAN-----RDAKKKD 64
+P+F+G+ YD W +KM I R + + VV + VP A + TA +A D
Sbjct: 9 IPIFDGEKYDFWSIKMATIFRTRKLWSVVEEGVPVEPVQAEETPETARAKTLREEAVTND 68
Query: 65 GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTM 124
A+ + +V+++IF +I S+KE W+ L+ Y G +++ V+LQ LRR+YE L M
Sbjct: 69 TMALQILQTAVTDQIFSRIAAASSSKE-AWDVLKDEYQGSPQVRLVKLQSLRREYENLKM 127
Query: 125 EEEETISQYFDKFVNLSNQ 143
+ + I + DK + L Q
Sbjct: 128 YDNDNIKTFTDKLIVLEIQ 146
>UniRef100_Q9M1C6 Hypothetical protein T2O9.150 [Arabidopsis thaliana]
Length = 1339
Score = 80.9 bits (198), Expect = 1e-14
Identities = 44/137 (32%), Positives = 80/137 (58%), Gaps = 5/137 (3%)
Query: 10 LPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNAT---DMQ*TANRDAKKKDGK 66
+P F+G YD W + M+ R +++ +V + +PA+ T + Q +A +AK KD K
Sbjct: 12 IPRFDGY-YDFWSMTMENFLRSRELWRLVEEGIPAIVVGTTPVSEAQRSAVEEAKLKDLK 70
Query: 67 AMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEE 126
F+ Q++ EI E I+ +S + WE+++K Y G K+K+ +LQ LR+++E+L M+E
Sbjct: 71 VKNFLFQAIDREILETILD-KSTSKAIWESMKKKYQGSTKVKRAQLQALRKEFELLAMKE 129
Query: 127 EETISQYFDKFVNLSNQ 143
E I + + + + N+
Sbjct: 130 GEKIDTFLGRTLTVVNK 146
>UniRef100_Q6I5H9 Putative polyprotein [Oryza sativa]
Length = 1136
Score = 80.5 bits (197), Expect = 2e-14
Identities = 41/136 (30%), Positives = 76/136 (55%), Gaps = 3/136 (2%)
Query: 10 LPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNAT--DMQ*TANRDAKKKDGKA 67
+PVF +NYD W +KM+ + Q + ++V + ++ T Q + + + D KA
Sbjct: 6 VPVFARENYDIWSIKMRTLLLSQGLWDIVENGYQEYSAGETLTAEQKKSLVEDRMSDAKA 65
Query: 68 MFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEE 127
+F + Q V+ +F +I+ + +KE W+ L++ + K+ V+LQ LRRQ++ L M+E
Sbjct: 66 LFLIQQGVAESLFPRIIGAKKSKE-AWDKLKEEFQASQKVLAVKLQTLRRQFQNLQMKES 124
Query: 128 ETISQYFDKFVNLSNQ 143
E + YF + + + NQ
Sbjct: 125 EKVKDYFSRVIEIVNQ 140
>UniRef100_Q69F89 Gag-pol polyprotein [Phaseolus vulgaris]
Length = 529
Score = 80.5 bits (197), Expect = 2e-14
Identities = 43/136 (31%), Positives = 78/136 (56%), Gaps = 3/136 (2%)
Query: 11 PVFEGKNYDQWIVKMKVICRFQDVLEVVNDS--VPALASNATDMQ*TANRDAKKKDGKAM 68
PVF+G +Y W V+M+ D+ E V + + +L N T Q + +D K K KA
Sbjct: 4 PVFDGDSYQMWAVRMETYLEALDLWEAVEEDYEIQSLPENPTVAQIKSQKDKKMKRSKAK 63
Query: 69 FFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEEE 128
+ +VS +F +I+ +SAKE W+ L+ Y GD +++ ++ L R++E+ M+E E
Sbjct: 64 ACLFAAVSPMVFTRIMSLKSAKE-IWDYLKAEYEGDERIRGMQALNLIREFELQKMKESE 122
Query: 129 TISQYFDKFVNLSNQS 144
+I +Y D+ + ++N++
Sbjct: 123 SIKEYSDRLLTIANKN 138
>UniRef100_Q9SEL2 Gag-pol polyprotein [Vitis vinifera]
Length = 581
Score = 75.9 bits (185), Expect = 4e-13
Identities = 42/134 (31%), Positives = 77/134 (57%), Gaps = 3/134 (2%)
Query: 12 VFEGKNYDQWIVKMKVICRFQDVLEVVNDS--VPALASNATDMQ*TANRDAKKKDGKAMF 69
V +G NY+ W V+M V + DV E V ++ VP L ++ T Q +++ + + KA
Sbjct: 12 VLDGDNYETWAVRMTVHLQALDVWEAVEENYEVPPLGADPTVAQMKLHKERRTRKAKAKA 71
Query: 70 FMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEEET 129
+ +VS IF KI+ +SA E WE L++ Y GD ++K +++ L R++E+ M E +
Sbjct: 72 CLFAAVSPSIFIKIMKIDSAAE-IWEYLKEEYKGDERIKNMQVMNLIREFEMKKMRESDA 130
Query: 130 ISQYFDKFVNLSNQ 143
+ Y + ++++N+
Sbjct: 131 VKDYAAQLLSIANK 144
>UniRef100_Q9FH39 Copia-type polyprotein [Arabidopsis thaliana]
Length = 1334
Score = 72.8 bits (177), Expect = 3e-12
Identities = 39/135 (28%), Positives = 77/135 (56%), Gaps = 4/135 (2%)
Query: 10 LPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNA--TDMQ*TANRDAKKKDGKA 67
+P F+G +Y+ W + M+ + R ++ +++ +P N T Q T + KD K
Sbjct: 9 IPKFDG-DYEHWAMLMENLIRSKEWWDIIETGIPRPERNVILTGAQRTELAEKTVKDHKV 67
Query: 68 MFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEE 127
++ S+ I + I+ E++K+ WE++++ Y G+ +++ +LQ LRR +EVL M+
Sbjct: 68 KNYLFASIDKTILKTILQKETSKD-LWESMKRKYQGNDRVQSAQLQRLRRSFEVLEMKIG 126
Query: 128 ETISQYFDKFVNLSN 142
ETI+ YF + + ++N
Sbjct: 127 ETITGYFSRVMEITN 141
>UniRef100_Q9C7Y1 Copia-type polyprotein, putative; 28768-32772 [Arabidopsis
thaliana]
Length = 1334
Score = 72.8 bits (177), Expect = 3e-12
Identities = 39/135 (28%), Positives = 77/135 (56%), Gaps = 4/135 (2%)
Query: 10 LPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNA--TDMQ*TANRDAKKKDGKA 67
+P F+G +Y+ W + M+ + R ++ +++ +P N T Q T + KD K
Sbjct: 9 IPKFDG-DYEHWAMLMENLIRSKEWWDIIETGIPRPERNVILTGAQRTELAEKTVKDHKV 67
Query: 68 MFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEE 127
++ S+ I + I+ E++K+ WE++++ Y G+ +++ +LQ LRR +EVL M+
Sbjct: 68 KNYLFASIDKTILKTILQKETSKD-LWESMKRKYQGNDRVQSAQLQRLRRSFEVLEMKIG 126
Query: 128 ETISQYFDKFVNLSN 142
ETI+ YF + + ++N
Sbjct: 127 ETITGYFSRVMEITN 141
>UniRef100_Q6L3Y5 Putative gag-pol polyprotein [Solanum demissum]
Length = 1133
Score = 68.6 bits (166), Expect = 7e-11
Identities = 36/108 (33%), Positives = 67/108 (61%), Gaps = 2/108 (1%)
Query: 37 VVNDSVPA-LASNATDMQ*TANRDAKKKDGKAMFFMHQSVSNEIFEKIVHYESAKEETWE 95
V D+ P L +N T Q ++ + K KA+ F+H +VS+ IF +I+ ESAKE +W+
Sbjct: 14 VEEDNAPEPLRANPTLQQIRSHEEEIAKGPKALSFIHAAVSDSIFTRIMTCESAKE-SWD 72
Query: 96 ALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEEETISQYFDKFVNLSNQ 143
L++ + G + +++++ LR+++E+L M+E E + Y D+ + + NQ
Sbjct: 73 MLKQEFQGSDRTRQMQILNLRKEFEMLRMKETEKVKDYIDRIMKIFNQ 120
>UniRef100_Q6L3X6 Putative polyprotein [Solanum demissum]
Length = 1758
Score = 66.6 bits (161), Expect = 2e-10
Identities = 34/104 (32%), Positives = 63/104 (59%), Gaps = 1/104 (0%)
Query: 39 NDSVPALASNATDMQ*TANRDAKKKDGKAMFFMHQSVSNEIFEKIVHYESAKEETWEALE 98
N + AL N T Q +N + K K KA M +V++ +F +I+ ++AKE W+ L+
Sbjct: 16 NKPLAALPENPTLAQIKSNNEEKTKKSKAKSLMQNAVADSVFYRIMACKTAKE-AWDRLK 74
Query: 99 KLYSGDGKLKKVRLQVLRRQYEVLTMEEEETISQYFDKFVNLSN 142
+ Y G + +++++ L+R++E L M+++ETIS+Y D+ + N
Sbjct: 75 EEYQGSDRTRQMQVLNLKREFECLNMQDDETISKYADRISLIVN 118
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.332 0.140 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 258,605,571
Number of Sequences: 2790947
Number of extensions: 9399964
Number of successful extensions: 50194
Number of sequences better than 10.0: 266
Number of HSP's better than 10.0 without gapping: 132
Number of HSP's successfully gapped in prelim test: 134
Number of HSP's that attempted gapping in prelim test: 49897
Number of HSP's gapped (non-prelim): 294
length of query: 174
length of database: 848,049,833
effective HSP length: 119
effective length of query: 55
effective length of database: 515,927,140
effective search space: 28375992700
effective search space used: 28375992700
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 70 (31.6 bits)
Medicago: description of AC149197.6


