

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149134.13 - phase: 0
(163 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
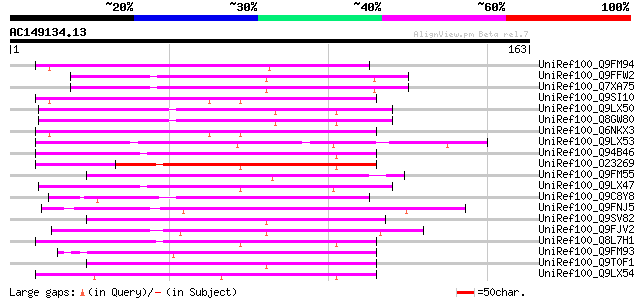
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FM94 Emb|CAB62440.1 [Arabidopsis thaliana] 65 6e-10
UniRef100_Q9FFW2 Similarity to heat shock transcription factor H... 61 9e-09
UniRef100_Q7XA75 At5g38590/MBB18_14 [Arabidopsis thaliana] 61 9e-09
UniRef100_Q9SI10 Hypothetical protein At2g04230 [Arabidopsis tha... 60 2e-08
UniRef100_Q9LX50 Hypothetical protein F25L23_70 [Arabidopsis tha... 60 2e-08
UniRef100_Q8GW80 Hypothetical protein At3g59210/F25L23_70 [Arabi... 60 2e-08
UniRef100_Q6NKX3 Hypothetical protein At2g04230 [Arabidopsis tha... 60 2e-08
UniRef100_Q9LX53 Hypothetical protein F25L23_40 [Arabidopsis tha... 59 6e-08
UniRef100_Q94B46 Hypothetical protein At4g14100 [Arabidopsis tha... 58 7e-08
UniRef100_O23269 Hypothetical protein dl3085w [Arabidopsis thali... 58 7e-08
UniRef100_Q9FM55 Arabidopsis thaliana genomic DNA, chromosome 5,... 58 1e-07
UniRef100_Q9LX47 Hypothetical protein F25L23_100 [Arabidopsis th... 57 2e-07
UniRef100_Q9C8Y8 Hypothetical protein T27F4.4 [Arabidopsis thali... 56 3e-07
UniRef100_Q9FNJ5 Arabidopsis thaliana genomic DNA, chromosome 5,... 55 8e-07
UniRef100_Q9SV82 Hypothetical protein F24G24.200 [Arabidopsis th... 54 1e-06
UniRef100_Q9FJV2 Emb|CAB89229.1 [Arabidopsis thaliana] 54 1e-06
UniRef100_Q8L7H1 Hypothetical protein At4g14103 [Arabidopsis tha... 54 1e-06
UniRef100_Q9FM93 Emb|CAB62440.1 [Arabidopsis thaliana] 54 2e-06
UniRef100_Q9T0F1 Hypothetical protein T5L19.50 [Arabidopsis thal... 53 2e-06
UniRef100_Q9LX54 Hypothetical protein F25L23_30 [Arabidopsis tha... 53 2e-06
>UniRef100_Q9FM94 Emb|CAB62440.1 [Arabidopsis thaliana]
Length = 421
Score = 65.1 bits (157), Expect = 6e-10
Identities = 41/107 (38%), Positives = 61/107 (56%), Gaps = 2/107 (1%)
Query: 9 PIL-SFHLQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSI 67
P+L S HL+ DI ++ IA+++G+ L+ D+ H LP +F+ TL++
Sbjct: 72 PVLESLHLKLGRQCSEVDIGFWVRIAVEKGLCELDFDYEHYKTEPCRLPQSLFTCGTLTV 131
Query: 68 LKLKQITLNEVPF-VNLPSLKALYLDVVTFTYYELILKLLSGCPILQ 113
LKLK ++L +V F V LK L+L+ V F E KLLS CPIL+
Sbjct: 132 LKLKNVSLKDVQFPVCFKLLKTLHLEYVIFLDKETPQKLLSSCPILE 178
>UniRef100_Q9FFW2 Similarity to heat shock transcription factor HSF30 [Arabidopsis
thaliana]
Length = 410
Score = 61.2 bits (147), Expect = 9e-09
Identities = 42/115 (36%), Positives = 63/115 (54%), Gaps = 11/115 (9%)
Query: 20 DYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNEVP 79
D DI ++ IA+ R V +L+ID H ++ S F+ K+L LKL+ +TL ++P
Sbjct: 86 DIKPEDIRRWIEIAVSRHVHDLDID--HFSENENIFLSSFFACKSLVTLKLRSVTLRDIP 143
Query: 80 -FVNLPSLKALYLDVVTFTYYELILKLLSGCPILQ--------YLGTNNLVVELP 125
V LPSLK L LD V+F + + +LLS CP+L+ Y T L + +P
Sbjct: 144 SMVCLPSLKTLLLDNVSFVEGKSLQELLSICPVLEDLSVYCDDYENTKELTIVVP 198
>UniRef100_Q7XA75 At5g38590/MBB18_14 [Arabidopsis thaliana]
Length = 289
Score = 61.2 bits (147), Expect = 9e-09
Identities = 42/115 (36%), Positives = 63/115 (54%), Gaps = 11/115 (9%)
Query: 20 DYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNEVP 79
D DI ++ IA+ R V +L+ID H ++ S F+ K+L LKL+ +TL ++P
Sbjct: 86 DIKPEDIRRWIEIAVSRHVHDLDID--HFSENENIFLSSFFACKSLVTLKLRSVTLRDIP 143
Query: 80 -FVNLPSLKALYLDVVTFTYYELILKLLSGCPILQ--------YLGTNNLVVELP 125
V LPSLK L LD V+F + + +LLS CP+L+ Y T L + +P
Sbjct: 144 SMVCLPSLKTLLLDNVSFVEGKSLQELLSICPVLEDLSVYCDDYENTKELTIVVP 198
>UniRef100_Q9SI10 Hypothetical protein At2g04230 [Arabidopsis thaliana]
Length = 454
Score = 60.1 bits (144), Expect = 2e-08
Identities = 46/110 (41%), Positives = 59/110 (52%), Gaps = 3/110 (2%)
Query: 9 PIL-SFHLQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFS-SKTLS 66
P+L S HL DC + ++ A RGV L +D + +TLPS +FS + +L
Sbjct: 80 PVLESLHLSFEGRTDCLHVGIWIATAFARGVRKLVLDSFYQEDQTVTLPSVLFSYNDSLE 139
Query: 67 ILKLK-QITLNEVPFVNLPSLKALYLDVVTFTYYELILKLLSGCPILQYL 115
ILKLK I L+ V L SL+ LYLD V F E + LL GCP LQ L
Sbjct: 140 ILKLKCAIDLDFPSRVCLKSLRKLYLDQVHFKDEESVCNLLCGCPSLQDL 189
>UniRef100_Q9LX50 Hypothetical protein F25L23_70 [Arabidopsis thaliana]
Length = 491
Score = 60.1 bits (144), Expect = 2e-08
Identities = 44/114 (38%), Positives = 62/114 (53%), Gaps = 5/114 (4%)
Query: 10 ILSFHLQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILK 69
I F L+C + + D++ ++ ++ GV +L++ S SL LPS VF+SKTL LK
Sbjct: 87 INKFSLKCGDGIEDEDVFPWILNTLRHGVSDLSLHVSPSLV--YWLPSKVFASKTLVRLK 144
Query: 70 LKQITLNEVPFVN--LPSLKALYLDVVTFTYYEL-ILKLLSGCPILQYLGTNNL 120
+ V N LP LK L LD V F ++ KLLSGCP+L+ L NL
Sbjct: 145 IGPKDGPRVKLRNVCLPKLKTLNLDSVVFEEGKIGFAKLLSGCPVLEELSLLNL 198
>UniRef100_Q8GW80 Hypothetical protein At3g59210/F25L23_70 [Arabidopsis thaliana]
Length = 484
Score = 60.1 bits (144), Expect = 2e-08
Identities = 44/114 (38%), Positives = 62/114 (53%), Gaps = 5/114 (4%)
Query: 10 ILSFHLQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILK 69
I F L+C + + D++ ++ ++ GV +L++ S SL LPS VF+SKTL LK
Sbjct: 87 INKFSLKCGDGIEDEDVFPWILNTLRHGVSDLSLHVSPSLV--YWLPSKVFASKTLVRLK 144
Query: 70 LKQITLNEVPFVN--LPSLKALYLDVVTFTYYEL-ILKLLSGCPILQYLGTNNL 120
+ V N LP LK L LD V F ++ KLLSGCP+L+ L NL
Sbjct: 145 IGPKDGPRVKLRNVCLPKLKTLNLDSVVFEEGKIGFAKLLSGCPVLEELSLLNL 198
>UniRef100_Q6NKX3 Hypothetical protein At2g04230 [Arabidopsis thaliana]
Length = 448
Score = 60.1 bits (144), Expect = 2e-08
Identities = 46/110 (41%), Positives = 59/110 (52%), Gaps = 3/110 (2%)
Query: 9 PIL-SFHLQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFS-SKTLS 66
P+L S HL DC + ++ A RGV L +D + +TLPS +FS + +L
Sbjct: 89 PVLESLHLSFEGRTDCLHVGIWIATAFARGVRKLVLDSFYQEDQTVTLPSVLFSYNDSLE 148
Query: 67 ILKLK-QITLNEVPFVNLPSLKALYLDVVTFTYYELILKLLSGCPILQYL 115
ILKLK I L+ V L SL+ LYLD V F E + LL GCP LQ L
Sbjct: 149 ILKLKCAIDLDFPSRVCLKSLRKLYLDQVHFKDEESVCNLLCGCPSLQDL 198
>UniRef100_Q9LX53 Hypothetical protein F25L23_40 [Arabidopsis thaliana]
Length = 475
Score = 58.5 bits (140), Expect = 6e-08
Identities = 53/150 (35%), Positives = 77/150 (51%), Gaps = 16/150 (10%)
Query: 9 PILSFHLQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSIL 68
P+ F L+C D + ++ + RGV L++D + S S LPS VF +KTL L
Sbjct: 53 PLNRFSLKCKVRCDSACVDGWIINVLDRGV--LDLDLNISSESDYKLPSEVFLNKTLVRL 110
Query: 69 KL--KQITLNEVPFVNLPSLKALYLDVVTFTYYE----LILKLLSGCPILQYLGTNNLVV 122
KL K + +V V LP LK LYL F +E KL+SGCP+L+ L +++
Sbjct: 111 KLGGKDEFIIDVEEVFLPKLKTLYLH--GFVCFEDSGVACAKLISGCPVLEEL----VMI 164
Query: 123 ELPYSERPVISLSN--LIRANICDIHIEFD 150
L + V S+SN L R IC I+++
Sbjct: 165 NLMWGLMDVCSVSNPTLKRVTICSEIIDYN 194
>UniRef100_Q94B46 Hypothetical protein At4g14100 [Arabidopsis thaliana]
Length = 468
Score = 58.2 bits (139), Expect = 7e-08
Identities = 41/108 (37%), Positives = 58/108 (52%), Gaps = 3/108 (2%)
Query: 9 PILSFHLQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSIL 68
P+ F L+ + + I ++ ++RGV +L D + ++ PS +F SKTL L
Sbjct: 85 PLHKFSLKIGDGVEPDRIIPWINNVLERGVSDL--DLHVYMETEFVFPSEMFLSKTLVRL 142
Query: 69 KLKQITLNEVPFVNLPSLKALYLDVVTFTYYEL-ILKLLSGCPILQYL 115
KL L E V LP LK LY+D F Y + + KLLSGCPIL+ L
Sbjct: 143 KLMLYPLLEFEDVYLPKLKTLYIDSCYFEKYGIGLTKLLSGCPILEDL 190
>UniRef100_O23269 Hypothetical protein dl3085w [Arabidopsis thaliana]
Length = 1047
Score = 58.2 bits (139), Expect = 7e-08
Identities = 41/108 (37%), Positives = 58/108 (52%), Gaps = 3/108 (2%)
Query: 9 PILSFHLQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSIL 68
P+ F L+ + + I ++ ++RGV +L D + ++ PS +F SKTL L
Sbjct: 340 PLHKFSLKIGDGVEPDRIIPWINNVLERGVSDL--DLHVYMETEFVFPSEMFLSKTLVRL 397
Query: 69 KLKQITLNEVPFVNLPSLKALYLDVVTFTYYEL-ILKLLSGCPILQYL 115
KL L E V LP LK LY+D F Y + + KLLSGCPIL+ L
Sbjct: 398 KLMLYPLLEFEDVYLPKLKTLYIDSCYFEKYGIGLTKLLSGCPILEDL 445
Score = 50.4 bits (119), Expect = 2e-05
Identities = 34/85 (40%), Positives = 52/85 (61%), Gaps = 5/85 (5%)
Query: 34 IQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLK--QITLNEVPFVNLPSLKALYL 91
++RGV +L++ + L S+ LPS V+ KTL LKL+ +V V+LP LK LY+
Sbjct: 778 LERGVSDLDLHLN--LESEFLLPSQVYLCKTLVWLKLRFGLYPTIDVEDVHLPKLKTLYI 835
Query: 92 DVVTFTYYEL-ILKLLSGCPILQYL 115
+ F + + + KLLSGCP+L+ L
Sbjct: 836 EATHFEEHGVGLTKLLSGCPMLEDL 860
>UniRef100_Q9FM55 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MJH22
[Arabidopsis thaliana]
Length = 449
Score = 57.8 bits (138), Expect = 1e-07
Identities = 39/101 (38%), Positives = 60/101 (58%), Gaps = 6/101 (5%)
Query: 25 DIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNEVPFV-NL 83
DI ++ IA+ R + L++DF LP +++ K+L LKL+ L +VP V +L
Sbjct: 116 DIKLWVEIAVSRCAQELSVDFFPKEKHNALLPRNLYTCKSLVTLKLRNNILVDVPHVFSL 175
Query: 84 PSLKALYLDVVTFTYYELILKLLSGCPILQYLGTNNLVVEL 124
PSLK L+L+ VT+ E + +LLS C +L+ +LVVEL
Sbjct: 176 PSLKILHLERVTYGDGESLQRLLSNCSVLE-----DLVVEL 211
>UniRef100_Q9LX47 Hypothetical protein F25L23_100 [Arabidopsis thaliana]
Length = 504
Score = 56.6 bits (135), Expect = 2e-07
Identities = 39/114 (34%), Positives = 64/114 (55%), Gaps = 5/114 (4%)
Query: 10 ILSFHLQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILK 69
++ F L C N D I+ ++ I+RGV +L D + S ++PS VF SK+L L+
Sbjct: 88 LVRFSLNCRNGIDRECIFRWISNVIERGVSDL--DLGGNFVSNRSMPSSVFVSKSLVKLR 145
Query: 70 LK--QITLNEVPFVNLPSLKALYLDVVTFTYYE-LILKLLSGCPILQYLGTNNL 120
++ T+ ++ V LP LK L L + F + +LKL+SGC +L+ L ++L
Sbjct: 146 IRTENCTIIDLEDVFLPKLKTLDLSSIWFRDGDTCLLKLISGCQVLEDLTMSDL 199
>UniRef100_Q9C8Y8 Hypothetical protein T27F4.4 [Arabidopsis thaliana]
Length = 453
Score = 56.2 bits (134), Expect = 3e-07
Identities = 35/107 (32%), Positives = 63/107 (58%), Gaps = 12/107 (11%)
Query: 13 FHLQCWNDYDCRDI------YNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLS 66
F L+C +D D D+ + ++Y+AI+R V+++++ + + + +++ ++L
Sbjct: 101 FKLRCGSDLD-GDVDLATCSWKWIYMAIKRKVQHIDVTWPG-----VRIYPEIYNCESLV 154
Query: 67 ILKLKQITLNEVPFVNLPSLKALYLDVVTFTYYELILKLLSGCPILQ 113
LKL ++TL + FV+LPSLK L LD V F L+SGCP+L+
Sbjct: 155 SLKLSEVTLTKPEFVSLPSLKVLVLDWVEFYNEFAFDMLMSGCPVLE 201
>UniRef100_Q9FNJ5 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MDJ22
[Arabidopsis thaliana]
Length = 450
Score = 54.7 bits (130), Expect = 8e-07
Identities = 42/140 (30%), Positives = 75/140 (53%), Gaps = 13/140 (9%)
Query: 11 LSFHLQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQM--TLPSFVFSSKTLSIL 68
+SF ++ D I ++ A++R +++L +D S M TLP V+ S++L L
Sbjct: 89 ISFQMEA---VDMWTIIPWIEDAVKRRIQHLEVD---SRIDHMIDTLPLTVYLSESLVSL 142
Query: 69 KLKQITLNEVPFVNLPSLKALYLDVVTFTYYELILKLLSGCPILQYLGTNNLVVE----- 123
+L + L+ FV+LP+LK ++L+ ++Y E + K +S CP+L+ L V E
Sbjct: 143 RLHLVMLHRFVFVSLPNLKVMHLEENIYSYAETMEKFISSCPVLEDLTVVRNVDEATEKV 202
Query: 124 LPYSERPVISLSNLIRANIC 143
L S + + SL +I ++ C
Sbjct: 203 LRVSSQSLNSLKLVIDSSKC 222
>UniRef100_Q9SV82 Hypothetical protein F24G24.200 [Arabidopsis thaliana]
Length = 326
Score = 54.3 bits (129), Expect = 1e-06
Identities = 37/95 (38%), Positives = 52/95 (53%), Gaps = 1/95 (1%)
Query: 25 DIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNEVP-FVNL 83
DI ++ +A+ R + L I SH Q LPS +++ K+L ILKL L +VP V L
Sbjct: 91 DIRMWVVVAVSRYIRELKIYSSHYGEKQNILPSSLYTCKSLVILKLDGGVLLDVPRMVCL 150
Query: 84 PSLKALYLDVVTFTYYELILKLLSGCPILQYLGTN 118
PSLK L L V + + +LL CP+L+ L N
Sbjct: 151 PSLKTLELKGVRYFKQGSLQRLLCNCPVLEDLVVN 185
>UniRef100_Q9FJV2 Emb|CAB89229.1 [Arabidopsis thaliana]
Length = 477
Score = 54.3 bits (129), Expect = 1e-06
Identities = 41/122 (33%), Positives = 62/122 (50%), Gaps = 9/122 (7%)
Query: 14 HLQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQ-MTLPSFVFSSKTLSILKLKQ 72
HL+ Y DI ++ +A+ R V L ID LF + + LP + S TL L L
Sbjct: 83 HLKLNQKYSASDINFWVQVAVNRSVRELRID----LFGKTLELPCCLCSCITLKELTLHD 138
Query: 73 ITLNEVP-FVNLPSLKALYLDVVTFTYYELILKLLSGCPILQYL---GTNNLVVELPYSE 128
+ + VP + LPSLK L+L V F+ + +L CP+L+ L GT + V++P +
Sbjct: 139 LCIKVVPAWFRLPSLKTLHLLSVKFSSDGFVASILRICPVLERLVVDGTKGVNVKIPNMD 198
Query: 129 RP 130
P
Sbjct: 199 VP 200
>UniRef100_Q8L7H1 Hypothetical protein At4g14103 [Arabidopsis thaliana]
Length = 381
Score = 53.9 bits (128), Expect = 1e-06
Identities = 39/110 (35%), Positives = 62/110 (55%), Gaps = 5/110 (4%)
Query: 9 PILSFHLQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSIL 68
P+ F L+ + D I ++ ++RGV +L++ + L S+ LPS V+ KTL L
Sbjct: 85 PLHKFSLKIGDGIDPVRIIPWINNVLERGVSDLDLHLN--LESEFLLPSQVYLCKTLVWL 142
Query: 69 KLK--QITLNEVPFVNLPSLKALYLDVVTFTYYEL-ILKLLSGCPILQYL 115
KL+ +V V+LP LK LY++ F + + + KLLSGCP+L+ L
Sbjct: 143 KLRFGLYPTIDVEDVHLPKLKTLYIEATHFEEHGVGLTKLLSGCPMLEDL 192
>UniRef100_Q9FM93 Emb|CAB62440.1 [Arabidopsis thaliana]
Length = 412
Score = 53.5 bits (127), Expect = 2e-06
Identities = 35/101 (34%), Positives = 61/101 (59%), Gaps = 5/101 (4%)
Query: 16 QCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLF-SQMTLPSFVFSSKTLSILKLKQIT 74
QC YD DI ++ IA++RG+ L + ++ S + + +LP +++ +TL +LKLK+
Sbjct: 65 QC--SYD--DIAIWVGIAVKRGLMELKLKYTDSYYPKRSSLPRSLYTCETLVVLKLKKGY 120
Query: 75 LNEVPFVNLPSLKALYLDVVTFTYYELILKLLSGCPILQYL 115
L+ V L SLK L L + ++ +L+LL+ CP+L+ L
Sbjct: 121 LDVPDLVCLRSLKTLSLRDMNYSNASCLLRLLASCPVLEEL 161
>UniRef100_Q9T0F1 Hypothetical protein T5L19.50 [Arabidopsis thaliana]
Length = 414
Score = 53.1 bits (126), Expect = 2e-06
Identities = 36/92 (39%), Positives = 51/92 (55%), Gaps = 1/92 (1%)
Query: 25 DIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNEVP-FVNL 83
DI ++ +A+ R + L I SH Q LPS +++ K+L ILKL L +VP V L
Sbjct: 91 DIRMWVVVAVSRYIRELKIYSSHYGEKQNILPSSLYTCKSLVILKLDGGVLLDVPRMVCL 150
Query: 84 PSLKALYLDVVTFTYYELILKLLSGCPILQYL 115
PSLK L L V + + +LL CP+L+ L
Sbjct: 151 PSLKTLELKGVRYFKQGSLQRLLCNCPVLEDL 182
>UniRef100_Q9LX54 Hypothetical protein F25L23_30 [Arabidopsis thaliana]
Length = 473
Score = 53.1 bits (126), Expect = 2e-06
Identities = 37/113 (32%), Positives = 63/113 (55%), Gaps = 6/113 (5%)
Query: 9 PILSFHLQCWNDYDCRD---IYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTL 65
P+ F L+ + +D D + +++ + RGV +L++ + ++ LPS +F S+TL
Sbjct: 105 PLDKFSLKMVDGHDPVDPDSVVPWIHKVLVRGVSDLHLVVDMNEWTSDPLPSRIFLSETL 164
Query: 66 --SILKLKQITLNEVPFVNLPSLKALYLDVVTFTYYEL-ILKLLSGCPILQYL 115
+K++ +V V+LP LK LYL V F ++ KLLSGCP+L+ L
Sbjct: 165 VKLTIKIRDGPFIDVKHVHLPKLKTLYLQSVMFDENDIGFRKLLSGCPVLEEL 217
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.327 0.142 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 255,792,754
Number of Sequences: 2790947
Number of extensions: 9995049
Number of successful extensions: 31670
Number of sequences better than 10.0: 223
Number of HSP's better than 10.0 without gapping: 76
Number of HSP's successfully gapped in prelim test: 147
Number of HSP's that attempted gapping in prelim test: 31494
Number of HSP's gapped (non-prelim): 246
length of query: 163
length of database: 848,049,833
effective HSP length: 117
effective length of query: 46
effective length of database: 521,509,034
effective search space: 23989415564
effective search space used: 23989415564
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 69 (31.2 bits)
Medicago: description of AC149134.13


