

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149050.9 - phase: 0 /pseudo
(157 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
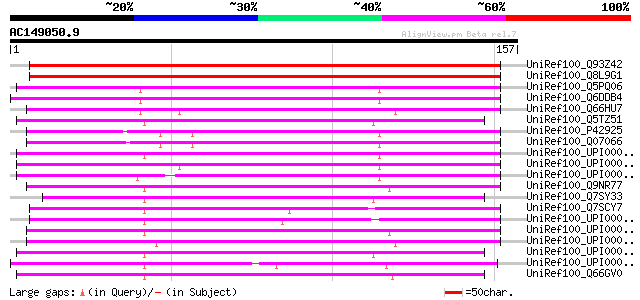
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q93Z42 AT5g19750/T29J13_170 [Arabidopsis thaliana] 220 1e-56
UniRef100_Q8L9G1 Hypothetical protein [Arabidopsis thaliana] 220 1e-56
UniRef100_Q5PQ06 Hypothetical protein [Xenopus laevis] 108 3e-23
UniRef100_Q6DDB4 Peroxisomal membrane protein 2, 22kDa [Xenopus ... 108 6e-23
UniRef100_Q66HU7 Zgc:92599 [Brachydanio rerio] 100 2e-20
UniRef100_Q5TZ51 Novel protein similar to human and mouse MpV17 ... 97 2e-19
UniRef100_P42925 Peroxisomal membrane protein 2 [Mus musculus] 96 3e-19
UniRef100_Q07066 Peroxisomal membrane protein 2 [Rattus norvegicus] 94 8e-19
UniRef100_UPI0000361A71 UPI0000361A71 UniRef100 entry 93 2e-18
UniRef100_UPI00003611E8 UPI00003611E8 UniRef100 entry 93 2e-18
UniRef100_UPI00002DAC6A UPI00002DAC6A UniRef100 entry 92 4e-18
UniRef100_Q9NR77 Peroxisomal membrane protein 2 [Homo sapiens] 92 4e-18
UniRef100_Q7SY33 Similar to MpV17 transgene, murine homolog, glo... 91 7e-18
UniRef100_Q7SCY7 Hypothetical protein [Neurospora crassa] 91 9e-18
UniRef100_UPI000023F32C UPI000023F32C UniRef100 entry 90 2e-17
UniRef100_UPI0000368C60 UPI0000368C60 UniRef100 entry 90 2e-17
UniRef100_UPI000042F6D3 UPI000042F6D3 UniRef100 entry 87 1e-16
UniRef100_UPI00002DA7F6 UPI00002DA7F6 UniRef100 entry 87 1e-16
UniRef100_UPI000021F5C5 UPI000021F5C5 UniRef100 entry 86 2e-16
UniRef100_Q66GV0 LOC446961 protein [Xenopus laevis] 86 3e-16
>UniRef100_Q93Z42 AT5g19750/T29J13_170 [Arabidopsis thaliana]
Length = 288
Score = 220 bits (560), Expect = 1e-56
Identities = 102/146 (69%), Positives = 125/146 (84%)
Query: 7 YMALLAKYPVPVKALTSAILNLIGDLICQLVIDKVQTPDLKRTFLFSFLGLVLVGPTLHF 66
Y ALL+ PV KA+T+A+LNL+GDLICQL I+K + D KRT F+FLGL LVGPTLHF
Sbjct: 118 YQALLSNSPVLTKAVTAALLNLVGDLICQLTINKTSSLDKKRTLTFTFLGLGLVGPTLHF 177
Query: 67 WYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQAVPKLKQEWFS 126
WYLYLS++VT G SGA++RL+LDQFVF+PIF+GVFLS++VTLEG+PS +PKL+QEW
Sbjct: 178 WYLYLSKVVTASGLSGAVIRLLLDQFVFAPIFVGVFLSAVVTLEGKPSNVIPKLQQEWTG 237
Query: 127 AVLANWQLWIPFQFLNFRFVPQQFQV 152
A++ANWQLWIPFQFLNFRFVPQ +QV
Sbjct: 238 AMIANWQLWIPFQFLNFRFVPQNYQV 263
>UniRef100_Q8L9G1 Hypothetical protein [Arabidopsis thaliana]
Length = 289
Score = 220 bits (560), Expect = 1e-56
Identities = 102/146 (69%), Positives = 125/146 (84%)
Query: 7 YMALLAKYPVPVKALTSAILNLIGDLICQLVIDKVQTPDLKRTFLFSFLGLVLVGPTLHF 66
Y ALL+ PV KA+T+A+LNL+GDLICQL I+K + D KRT F+FLGL LVGPTLHF
Sbjct: 119 YQALLSNSPVLTKAVTAALLNLVGDLICQLTINKTSSLDKKRTLTFTFLGLGLVGPTLHF 178
Query: 67 WYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQAVPKLKQEWFS 126
WYLYLS++VT G SGA++RL+LDQFVF+PIF+GVFLS++VTLEG+PS +PKL+QEW
Sbjct: 179 WYLYLSKVVTASGLSGAVIRLLLDQFVFAPIFVGVFLSAVVTLEGKPSNVIPKLQQEWTG 238
Query: 127 AVLANWQLWIPFQFLNFRFVPQQFQV 152
A++ANWQLWIPFQFLNFRFVPQ +QV
Sbjct: 239 AMIANWQLWIPFQFLNFRFVPQNYQV 264
>UniRef100_Q5PQ06 Hypothetical protein [Xenopus laevis]
Length = 193
Score = 108 bits (271), Expect = 3e-23
Identities = 59/157 (37%), Positives = 92/157 (58%), Gaps = 7/157 (4%)
Query: 3 LYYRYMALLAKYPVPVKALTSAILNLIGDLICQLVID------KVQTPDLKRTFLFSFLG 56
L RY+ LL PV KALTSAIL+ +G+++ Q + Q DL+ F F+ G
Sbjct: 18 LLLRYLQLLHSRPVLTKALTSAILSALGNILSQTIQKWRKEQKAPQNVDLRGPFRFAVYG 77
Query: 57 LVLVGPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRP-SQ 115
L+ GP H++YL L QLV + RL++++ + +P FL +F + LEG+ ++
Sbjct: 78 LLFTGPLSHYFYLLLEQLVPSSAPLAGLQRLLIERLMIAPAFLLLFFLVMNLLEGKNLAK 137
Query: 116 AVPKLKQEWFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
KLK ++SA+ NW++W PFQF+N ++P QF+V
Sbjct: 138 LNKKLKDHYWSALKLNWKVWTPFQFININYIPVQFRV 174
>UniRef100_Q6DDB4 Peroxisomal membrane protein 2, 22kDa [Xenopus tropicalis]
Length = 193
Score = 108 bits (269), Expect = 6e-23
Identities = 59/159 (37%), Positives = 91/159 (57%), Gaps = 7/159 (4%)
Query: 1 MTLYYRYMALLAKYPVPVKALTSAILNLIGDLICQLVID------KVQTPDLKRTFLFSF 54
M L RY+ LL PV KALTSAIL+ +G+++ Q + Q DL+ F+
Sbjct: 16 MVLLLRYLQLLHSRPVLTKALTSAILSALGNILSQTIQKWRKEQKHPQNVDLRGPLRFAV 75
Query: 55 LGLVLVGPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRP- 113
GL+ GP H++YL L QLV + RL++++ + +P FL +F + LEG+
Sbjct: 76 YGLLFTGPLSHYFYLLLEQLVPSSAPLAGLQRLLIERLIIAPAFLLLFFLVMNLLEGKNF 135
Query: 114 SQAVPKLKQEWFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
++ KLK ++ A+ NW++W PFQF+N +VP QF+V
Sbjct: 136 TKLNQKLKSSYWQALKLNWKVWTPFQFININYVPVQFRV 174
>UniRef100_Q66HU7 Zgc:92599 [Brachydanio rerio]
Length = 194
Score = 100 bits (248), Expect = 2e-20
Identities = 57/156 (36%), Positives = 92/156 (58%), Gaps = 9/156 (5%)
Query: 6 RYMALLAKYPVPVKALTSAILNLIGDLICQLVID----KVQTPDLKRTFL----FSFLGL 57
+Y++LL KYP+ K++TS IL+ +G+L+ Q++ K +P K + L F+ GL
Sbjct: 20 QYLSLLKKYPIITKSVTSGILSALGNLLSQVLEYQKNVKENSPKKKISILGPVHFAIYGL 79
Query: 58 VLVGPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQAV 117
+ GP H++Y L L+ I RL+L++ +F+P FL +F + LEG+ V
Sbjct: 80 FITGPVSHYFYHLLEVLLPTTVPYCLIKRLLLERLIFAPAFLLLFYVVMNALEGKTLADV 139
Query: 118 P-KLKQEWFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
KLK ++ A+ NW++W PFQF+N +VP QF+V
Sbjct: 140 QNKLKTSYWPAMKMNWKVWTPFQFININYVPVQFRV 175
>UniRef100_Q5TZ51 Novel protein similar to human and mouse MpV17 transgene, murine
homolog, glomerulosclerosis [Brachydanio rerio]
Length = 177
Score = 96.7 bits (239), Expect = 2e-19
Identities = 54/148 (36%), Positives = 87/148 (58%), Gaps = 3/148 (2%)
Query: 3 LYYRYMALLAKYPVPVKALTSAILNLIGDLICQLVIDK--VQTPDLKRTFLFSFLGLVLV 60
L+ Y AL+AK+P V+ +T+ L +GD+I Q +I++ + + +RT +G V
Sbjct: 4 LWRSYQALMAKHPWKVQIITAGSLVGVGDVISQQLIERRGLANHNARRTAKMMSIGFFFV 63
Query: 61 GPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEG-RPSQAVPK 119
GP + WY L +LVT S A+ ++++DQ F+P FLG FL TL G + V K
Sbjct: 64 GPVVGGWYKVLDKLVTGGTKSAALKKMLVDQVGFAPCFLGAFLGITGTLNGLTVEENVAK 123
Query: 120 LKQEWFSAVLANWQLWIPFQFLNFRFVP 147
L++++ A+++N+ LW P Q NF F+P
Sbjct: 124 LQRDYTDALISNYYLWPPVQIANFYFIP 151
>UniRef100_P42925 Peroxisomal membrane protein 2 [Mus musculus]
Length = 193
Score = 95.9 bits (237), Expect = 3e-19
Identities = 57/153 (37%), Positives = 91/153 (59%), Gaps = 7/153 (4%)
Query: 6 RYMALLAKYPVPVKALTSAILNLIGDLICQLVIDKVQTPD---LKRTFLFSFL--GLVLV 60
+Y+ LL YPV KA++S IL+ +G+L+ Q I+K Q D L+ + L +L GL +
Sbjct: 23 QYLLLLKLYPVLTKAVSSGILSALGNLLAQ-TIEKKQRKDSRLLEVSGLLRYLVYGLFVT 81
Query: 61 GPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRP-SQAVPK 119
GP H+ YL++ V ++ RL+LD+ F+P FL +F + LEG+ S V K
Sbjct: 82 GPLSHYLYLFMEYSVPPEVPWASVKRLLLDRLFFAPTFLLLFFFVMNLLEGKNVSVFVAK 141
Query: 120 LKQEWFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
++ ++ A+ NW++W P QF+N +VP QF+V
Sbjct: 142 MRSGFWPALQMNWRMWTPLQFININYVPLQFRV 174
>UniRef100_Q07066 Peroxisomal membrane protein 2 [Rattus norvegicus]
Length = 193
Score = 94.4 bits (233), Expect = 8e-19
Identities = 56/153 (36%), Positives = 90/153 (58%), Gaps = 7/153 (4%)
Query: 6 RYMALLAKYPVPVKALTSAILNLIGDLICQLVIDKVQTPD---LKRTFLFSFL--GLVLV 60
+Y+ L YPV KA++S IL+ +G+L+ Q+ I+K Q D L+ + L +L GL +
Sbjct: 23 QYLLFLKFYPVVTKAVSSGILSALGNLLAQM-IEKKQKKDSRSLEVSGLLRYLVYGLFVT 81
Query: 61 GPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRP-SQAVPK 119
GP H+ YL++ V + RL+LD+ F+P FL +F + LEG+ S V K
Sbjct: 82 GPLSHYLYLFMEYWVPPEVPWARVKRLLLDRLFFAPTFLLLFFFVMNLLEGKNISVFVAK 141
Query: 120 LKQEWFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
++ ++ A+ NW++W P QF+N +VP QF+V
Sbjct: 142 MRSGFWPALQMNWRMWTPLQFININYVPLQFRV 174
>UniRef100_UPI0000361A71 UPI0000361A71 UniRef100 entry
Length = 174
Score = 93.2 bits (230), Expect = 2e-18
Identities = 50/153 (32%), Positives = 89/153 (57%), Gaps = 3/153 (1%)
Query: 3 LYYRYMALLAKYPVPVKALTSAILNLIGDLICQLVIDK--VQTPDLKRTFLFSFLGLVLV 60
++ Y +L+++YP V+ +T+ L +GD+I Q +I++ + +++RT +G V
Sbjct: 1 IWRAYQSLMSRYPWTVQIVTAGSLVGVGDVISQQLIERRGLAHHNMQRTAKMMSIGFFFV 60
Query: 61 GPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRP-SQAVPK 119
GP + WY L +LV G S A+ ++++DQ F+P FL F +L G + V K
Sbjct: 61 GPVIGSWYKVLDRLVVGGGKSAAMKKMLVDQLCFAPCFLAAFFCVSGSLNGLTLEENVRK 120
Query: 120 LKQEWFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
LK+++ A+++N+ LW P Q NF FVP +++
Sbjct: 121 LKRDYTDALISNYYLWPPVQIANFYFVPLHYRL 153
>UniRef100_UPI00003611E8 UPI00003611E8 UniRef100 entry
Length = 173
Score = 93.2 bits (230), Expect = 2e-18
Identities = 52/153 (33%), Positives = 88/153 (56%), Gaps = 3/153 (1%)
Query: 3 LYYRYMALLAKYPVPVKALTSAILNLIGDLICQLVIDKVQTPDLKRTFL--FSFLGLVLV 60
L +Y+ LL +YP+ K++TS IL +G+L+ Q + + + + T + ++ GL +
Sbjct: 2 LLQQYLFLLKRYPIITKSVTSGILTALGNLLSQNLEARKKAGAIDGTGVARYAVYGLFIT 61
Query: 61 GPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRP-SQAVPK 119
GP H +Y + L+ I RL+LD+ +F+P FL +F + LE + + K
Sbjct: 62 GPVSHCFYQLMEALIPTTDPHCIIKRLLLDRLIFAPGFLLIFYFVMNILEFKGWEEFEKK 121
Query: 120 LKQEWFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
LK +++A+ NW++W PFQF+N FVP QF+V
Sbjct: 122 LKGSFWTALKMNWKVWTPFQFVNINFVPVQFRV 154
>UniRef100_UPI00002DAC6A UPI00002DAC6A UniRef100 entry
Length = 175
Score = 92.0 bits (227), Expect = 4e-18
Identities = 55/158 (34%), Positives = 87/158 (54%), Gaps = 11/158 (6%)
Query: 3 LYYRYMALLAKYPVPVKALTSAILNLIGDLICQLVI-------DKVQTPDLKRTFLFSFL 55
L +Y+ LL KYP+ K++TS IL +G+L+ Q + D + P + R ++
Sbjct: 2 LLQQYLFLLRKYPILTKSVTSGILTALGNLLSQSLEARKKASNDAICGPAVAR---YAAY 58
Query: 56 GLVLVGPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRP-S 114
GL + GP H +Y + L+ I RL+LD+ F+P FL +F + LE +
Sbjct: 59 GLFITGPVSHCFYQLMEALIPATDPHCIIKRLLLDRLFFAPGFLLIFYLVMNVLELKGWK 118
Query: 115 QAVPKLKQEWFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
+ KLK +++A+ NW++W PFQF+N FVP QF+V
Sbjct: 119 ELEAKLKGSFWTALKMNWKVWTPFQFVNINFVPVQFRV 156
>UniRef100_Q9NR77 Peroxisomal membrane protein 2 [Homo sapiens]
Length = 194
Score = 92.0 bits (227), Expect = 4e-18
Identities = 48/153 (31%), Positives = 84/153 (54%), Gaps = 6/153 (3%)
Query: 6 RYMALLAKYPVPVKALTSAILNLIGDLICQLVIDK-----VQTPDLKRTFLFSFLGLVLV 60
+Y+ L YPV KA TS IL+ +G+ + Q++ K ++ D+ ++ G
Sbjct: 23 QYLLFLRLYPVLTKAATSGILSALGNFLAQMIEKKRKKENSRSLDVGGPLRYAVYGFFFT 82
Query: 61 GPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQA-VPK 119
GP HF+Y ++ + + RL+LD+ VF+P FL +F + LEG+ + A K
Sbjct: 83 GPLSHFFYFFMEHWIPPEVPLAGLRRLLLDRLVFAPAFLMLFFLIMNFLEGKDASAFAAK 142
Query: 120 LKQEWFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
++ ++ A+ NW++W P QF+N +VP +F+V
Sbjct: 143 MRGGFWPALRMNWRVWTPLQFININYVPLKFRV 175
>UniRef100_Q7SY33 Similar to MpV17 transgene, murine homolog, glomerulosclerosis
[Brachydanio rerio]
Length = 166
Score = 91.3 bits (225), Expect = 7e-18
Identities = 51/140 (36%), Positives = 83/140 (58%), Gaps = 3/140 (2%)
Query: 11 LAKYPVPVKALTSAILNLIGDLICQLVIDK--VQTPDLKRTFLFSFLGLVLVGPTLHFWY 68
+AK+P V+ LT+ L +GD+I Q +I++ + + +RT +G + VGP + WY
Sbjct: 1 MAKHPWKVQILTAGSLVGVGDVISQQLIERRGLANHNARRTAKMMSIGFLFVGPVVGGWY 60
Query: 69 LYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEG-RPSQAVPKLKQEWFSA 127
L +LVT S A+ ++++DQ F+P FLG FL TL G + V KL++++ A
Sbjct: 61 KVLDKLVTGGTKSAALKKMLVDQVGFAPCFLGAFLGITGTLNGLTVEENVAKLQRDYTDA 120
Query: 128 VLANWQLWIPFQFLNFRFVP 147
+++N+ LW P Q NF F+P
Sbjct: 121 LISNYYLWPPVQIANFYFIP 140
>UniRef100_Q7SCY7 Hypothetical protein [Neurospora crassa]
Length = 190
Score = 90.9 bits (224), Expect = 9e-18
Identities = 50/150 (33%), Positives = 85/150 (56%), Gaps = 6/150 (4%)
Query: 7 YMALLAKYPVPVKALTSAILNLIGDLICQLVIDK--VQTPDLKRTFLFSFLGLVLVGPTL 64
Y A LA P+ +A+T++IL +GD+ Q ++D+ + DL RT G + GP
Sbjct: 5 YKAQLAARPLLTQAVTTSILFGVGDVAAQQLVDRRGLSNHDLTRTGRMVLYGGAVFGPAA 64
Query: 65 HFWYLYLSQLVTLPGTSGAIL--RLVLDQFVFSPIFLGVFLSSLVTLEGRPSQAVPKLKQ 122
W+ +L + V +PG++ + R+ DQ +F+P F+G+FL S+ LEG + KL++
Sbjct: 65 TTWFRFLQKRVVVPGSTNKTILARVAADQGLFAPTFIGIFLGSMAVLEG--TDVKEKLQK 122
Query: 123 EWFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
++ A+ NW +W Q +NF+ VP +V
Sbjct: 123 NYWEALSTNWMVWPFVQMVNFKVVPLDHRV 152
>UniRef100_UPI000023F32C UPI000023F32C UniRef100 entry
Length = 175
Score = 90.1 bits (222), Expect = 2e-17
Identities = 48/150 (32%), Positives = 84/150 (56%), Gaps = 6/150 (4%)
Query: 7 YMALLAKYPVPVKALTSAILNLIGDLICQLVIDK--VQTPDLKRTFLFSFLGLVLVGPTL 64
Y + LA P+ +++T+A L GD+ Q +++K Q DL RT + G + GP
Sbjct: 8 YNSRLAARPLLTQSVTTAFLFATGDVTAQQLVEKRGAQKHDLVRTGRMALYGGFVFGPVA 67
Query: 65 HFWYLYLSQLVTLPGTSGA--ILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQAVPKLKQ 122
W+ +L++ V + A + R+ DQ F+P+ +GVFLSS+ T+EG+ ++ +
Sbjct: 68 TTWFAFLARRVNVRNNKKAEVLARVACDQLGFAPVMIGVFLSSMATMEGK--SVKERIDK 125
Query: 123 EWFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
W+ A+ ANW +W Q +NF +P Q+++
Sbjct: 126 TWWPALKANWMVWPAVQVINFSLIPLQYRL 155
>UniRef100_UPI0000368C60 UPI0000368C60 UniRef100 entry
Length = 195
Score = 89.7 bits (221), Expect = 2e-17
Identities = 47/153 (30%), Positives = 83/153 (53%), Gaps = 6/153 (3%)
Query: 6 RYMALLAKYPVPVKALTSAILNLIGDLICQLVIDK-----VQTPDLKRTFLFSFLGLVLV 60
+Y+ L YPV KA TS IL+ +G+ + Q++ K ++ D+ ++ G
Sbjct: 24 QYLLFLRLYPVLTKAATSGILSALGNFLAQMIEKKRKKENSRSLDVGGPLRYAVYGFFFT 83
Query: 61 GPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQA-VPK 119
GP HF+Y ++ + + RL+LD+ V +P FL +F + LEG+ + A K
Sbjct: 84 GPLSHFFYFFMEHWIPPEVPLAGLRRLLLDRLVLAPAFLMLFFLIMNFLEGKDASAFAAK 143
Query: 120 LKQEWFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
++ ++ A+ NW++W P QF+N +VP +F+V
Sbjct: 144 MRGGFWPALRMNWRVWTPLQFININYVPLKFRV 176
>UniRef100_UPI000042F6D3 UPI000042F6D3 UniRef100 entry
Length = 190
Score = 87.4 bits (215), Expect = 1e-16
Identities = 51/149 (34%), Positives = 80/149 (53%), Gaps = 2/149 (1%)
Query: 6 RYMALLAKYPVPVKALTSAILNLIGDLICQLVIDKVQTP-DLKRTFLFSFLGLVLVGPTL 64
+Y A L + PV ++SA+L GD+I Q +I+K DL RT G +L PT+
Sbjct: 7 KYAAFLTRRPVLGNMISSAVLFGTGDVIAQQLIEKKGADHDLPRTARIVTWGGILFAPTV 66
Query: 65 HFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQAVP-KLKQE 123
+ W+ L ++ R+ LDQF F+P+ L F +++ +EG+ A K +
Sbjct: 67 NLWFRTLERIPIRSRWPATFARVGLDQFGFAPVILSGFFTAMTFMEGKDFNAAKVKWHES 126
Query: 124 WFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
+F + ANW L+IPFQ LN VP Q+++
Sbjct: 127 FFPTLQANWMLFIPFQILNMGLVPLQYRL 155
>UniRef100_UPI00002DA7F6 UPI00002DA7F6 UniRef100 entry
Length = 177
Score = 87.0 bits (214), Expect = 1e-16
Identities = 48/148 (32%), Positives = 83/148 (55%), Gaps = 3/148 (2%)
Query: 3 LYYRYMALLAKYPVPVKALTSAILNLIGDLICQLVIDK--VQTPDLKRTFLFSFLGLVLV 60
L+ Y +L+++YP V+ +T+ L +GD+I Q +I++ V +++RT +G V
Sbjct: 4 LWRAYQSLMSRYPWTVQIVTAGSLVGVGDVISQQLIERRGVAHHNMRRTAKMMSIGFFFV 63
Query: 61 GPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEG-RPSQAVPK 119
GP + WY L +LV S A+ ++++DQ F+P FL F + G + K
Sbjct: 64 GPVIGSWYKVLDRLVVGGSRSAAMKKMLVDQLCFAPCFLAAFFCVSGAVNGLTVEDNLGK 123
Query: 120 LKQEWFSAVLANWQLWIPFQFLNFRFVP 147
L++++ A+++N+ LW P Q NF FVP
Sbjct: 124 LQRDYADALISNYYLWPPVQIANFYFVP 151
>UniRef100_UPI000021F5C5 UPI000021F5C5 UniRef100 entry
Length = 154
Score = 86.3 bits (212), Expect = 2e-16
Identities = 55/156 (35%), Positives = 85/156 (54%), Gaps = 7/156 (4%)
Query: 1 MTLYYRYMALLAKYPVPVKALTSAILNLIGDLICQLVIDK--VQTPDLKRTFLFSFLGLV 58
M L+ Y LA +P V+ LT+ L +GD+I Q ++++ +Q RT + LG
Sbjct: 1 MALWRAYQRALAAHPWKVQVLTAGSLMGLGDIISQQLVERRGLQQHQTGRTLTMASLGCG 60
Query: 59 LVGPTLHFWYLYLSQLVTLPGTS--GAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQ- 115
VGP + WY L L+ PGT+ A+ +++LDQ F+P FLG FL + L G +Q
Sbjct: 61 FVGPVVGGWYRVLDHLI--PGTTKVNALKKMLLDQGGFAPCFLGCFLPLVGVLNGMSAQD 118
Query: 116 AVPKLKQEWFSAVLANWQLWIPFQFLNFRFVPQQFQ 151
KLK+++ A++ N+ LW Q NF VP ++
Sbjct: 119 NWAKLKRDYPDALITNYYLWPAVQLANFYLVPLHYR 154
>UniRef100_Q66GV0 LOC446961 protein [Xenopus laevis]
Length = 182
Score = 85.9 bits (211), Expect = 3e-16
Identities = 47/148 (31%), Positives = 82/148 (54%), Gaps = 3/148 (2%)
Query: 3 LYYRYMALLAKYPVPVKALTSAILNLIGDLICQLVIDK--VQTPDLKRTFLFSFLGLVLV 60
L+ Y LL +P V+ +T+ L +GD+I Q ++++ ++ ++RT +G V
Sbjct: 9 LWRAYQRLLGAHPWKVQIVTAGSLVGVGDVISQQLLERKGLKGHSIERTVKMMGIGFCFV 68
Query: 61 GPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQAV-PK 119
GP + WY L +++ G A+ +++LDQ F+P FLG FLS L G + + K
Sbjct: 69 GPVVGGWYKILDRIIPGSGKPVALKKMLLDQVAFAPCFLGCFLSIASALNGLSGEQIWGK 128
Query: 120 LKQEWFSAVLANWQLWIPFQFLNFRFVP 147
LK+++ A++ N+ +W Q NF F+P
Sbjct: 129 LKRDYKDALITNYYIWPAVQVANFYFIP 156
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.332 0.146 0.456
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 250,493,140
Number of Sequences: 2790947
Number of extensions: 9662910
Number of successful extensions: 30858
Number of sequences better than 10.0: 212
Number of HSP's better than 10.0 without gapping: 113
Number of HSP's successfully gapped in prelim test: 99
Number of HSP's that attempted gapping in prelim test: 30581
Number of HSP's gapped (non-prelim): 224
length of query: 157
length of database: 848,049,833
effective HSP length: 116
effective length of query: 41
effective length of database: 524,299,981
effective search space: 21496299221
effective search space used: 21496299221
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 69 (31.2 bits)
Medicago: description of AC149050.9


