

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
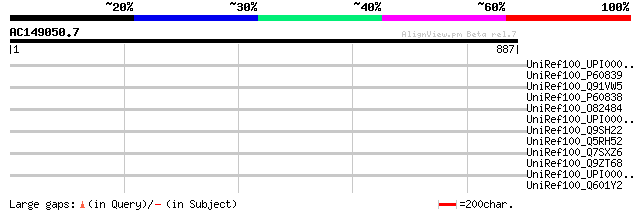
Query= AC149050.7 + phase: 0 /pseudo
(887 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_UPI0000339ED7 UPI0000339ED7 UniRef100 entry 39 0.90
UniRef100_P60839 Probable disease resistance protein At1g12290 [... 38 1.5
UniRef100_Q91VW5 Golgi autoantigen, golgin subfamily A member 4 ... 37 2.6
UniRef100_P60838 Probable disease resistance protein At1g12280 [... 37 3.4
UniRef100_O82484 Putative disease resistance protein At4g10780 [... 37 3.4
UniRef100_UPI00002DA2C0 UPI00002DA2C0 UniRef100 entry 36 4.5
UniRef100_Q9SH22 Putative disease resistance protein At1g63360 [... 36 4.5
UniRef100_Q5RH52 Novel RasGEF domain containing protein [Brachyd... 35 7.6
UniRef100_Q7SXZ6 Hypothetical protein ralgps2 [Brachydanio rerio] 35 7.6
UniRef100_Q9ZT68 Resistance protein candidate RGC2K [Lactuca sat... 35 7.6
UniRef100_UPI00002A1D22 UPI00002A1D22 UniRef100 entry 35 10.0
UniRef100_Q601Y2 Hypothetical protein [Mycoplasma hyopneumoniae] 35 10.0
>UniRef100_UPI0000339ED7 UPI0000339ED7 UniRef100 entry
Length = 314
Score = 38.5 bits (88), Expect = 0.90
Identities = 27/100 (27%), Positives = 50/100 (50%), Gaps = 12/100 (12%)
Query: 7 LGKETVTKLGELVVESTMKHFKYLTQHKKITTNLEEELERLKMIKQALQTRVET------ 60
+ K+ VT ++ V++ ++ F+ KK E+E +++ A+ T ET
Sbjct: 102 ISKKLVTLKKDVPVKNKIEEFELKKIDKKKLYEFLREMEFNRLLSSAISTYGETDFENIK 161
Query: 61 --ERRKGYEIAPNVQKWLYDVTTIENELQKWL--SDDNAQ 96
E+ K Y + N + Y++ T E E++KWL S+DN +
Sbjct: 162 SYEKNKNYNLKVNNKN--YELVTEEKEIKKWLEISEDNGE 199
>UniRef100_P60839 Probable disease resistance protein At1g12290 [Arabidopsis
thaliana]
Length = 884
Score = 37.7 bits (86), Expect = 1.5
Identities = 19/57 (33%), Positives = 32/57 (55%)
Query: 29 YLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIAPNVQKWLYDVTTIENE 85
Y+ K+ T+LEE +E LK ++ L +V+T G + ++ WL V TIE++
Sbjct: 28 YIQNIKENLTSLEEAMEDLKALRDDLLRKVQTAEEGGLQRLHQIKVWLKRVKTIESQ 84
>UniRef100_Q91VW5 Golgi autoantigen, golgin subfamily A member 4 [Mus musculus]
Length = 2238
Score = 37.0 bits (84), Expect = 2.6
Identities = 27/100 (27%), Positives = 48/100 (48%), Gaps = 5/100 (5%)
Query: 18 LVVESTMKHFKYLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIAPNVQKWLY 77
+ +E + + LTQ K+ +L LE L++ K+A+ T E + + E+ + +
Sbjct: 533 IALEKSRSEYLKLTQEKEQQESLA--LEELELQKKAILTESENKLQ---ELGQEAEAYRT 587
Query: 78 DVTTIENELQKWLSDDNAQI*HLIIHWERKPLKALKILQA 117
+ +E L+K L + Q HL +H E + K K L A
Sbjct: 588 RILELETSLEKSLQESKTQSEHLAVHLEAEKNKHNKELTA 627
>UniRef100_P60838 Probable disease resistance protein At1g12280 [Arabidopsis
thaliana]
Length = 894
Score = 36.6 bits (83), Expect = 3.4
Identities = 20/75 (26%), Positives = 43/75 (56%), Gaps = 1/75 (1%)
Query: 29 YLTQHKKITTNLEEELERLKMIKQALQTRVETER-RKGYEIAPNVQKWLYDVTTIENELQ 87
Y+ + K +++++E LK + ++ RV+ E + E VQ WL +V+T+EN+
Sbjct: 28 YICELSKNVVAMKKDMEVLKKKRDDVKRRVDIEEFTRRRERLSQVQGWLTNVSTVENKFN 87
Query: 88 KWLSDDNAQI*HLII 102
+ L+ ++A++ L +
Sbjct: 88 ELLTTNDAELQRLCL 102
>UniRef100_O82484 Putative disease resistance protein At4g10780 [Arabidopsis
thaliana]
Length = 892
Score = 36.6 bits (83), Expect = 3.4
Identities = 24/75 (32%), Positives = 34/75 (45%)
Query: 29 YLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIAPNVQKWLYDVTTIENELQK 88
Y+ + K LE+ +E L + + RV+ E KG E VQ WL V I N+
Sbjct: 28 YIHKLKDNIVALEKAIEDLTATRDDVLRRVQMEEGKGLERLQQVQVWLKRVEIIRNQFYD 87
Query: 89 WLSDDNAQI*HLIIH 103
LS N +I L +
Sbjct: 88 LLSARNIEIQRLCFY 102
>UniRef100_UPI00002DA2C0 UPI00002DA2C0 UniRef100 entry
Length = 589
Score = 36.2 bits (82), Expect = 4.5
Identities = 22/70 (31%), Positives = 42/70 (59%), Gaps = 7/70 (10%)
Query: 14 KLGELVVESTMKHFK-YLTQHKKITTNLEEELERLKMIKQALQTRVETERRK-----GYE 67
K+GE+ V+ ++ FK + HK+ +++ + E LK +++++ TR ER+K YE
Sbjct: 381 KVGEITVDEAVQEFKAWQFDHKQRASSIRYQHENLKQLRESI-TRRHKERQKPGKELDYE 439
Query: 68 IAPNVQKWLY 77
I+ +Q+ LY
Sbjct: 440 ISAPLQRSLY 449
>UniRef100_Q9SH22 Putative disease resistance protein At1g63360 [Arabidopsis
thaliana]
Length = 884
Score = 36.2 bits (82), Expect = 4.5
Identities = 21/74 (28%), Positives = 36/74 (48%)
Query: 29 YLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIAPNVQKWLYDVTTIENELQK 88
Y +K LE+ ++ LK + L+ R++ E +G + Q WL V T+E+ +
Sbjct: 26 YTHNLEKNLAALEKTMKELKAKRDDLERRLKREEARGLQRLSEFQVWLDSVATVEDIIIT 85
Query: 89 WLSDDNAQI*HLII 102
L D N +I L +
Sbjct: 86 LLRDRNVEIQRLCL 99
>UniRef100_Q5RH52 Novel RasGEF domain containing protein [Brachydanio rerio]
Length = 374
Score = 35.4 bits (80), Expect = 7.6
Identities = 24/71 (33%), Positives = 39/71 (54%), Gaps = 3/71 (4%)
Query: 29 YLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIA--PNVQKWLYDVTTIENEL 86
Y+ T ++ E +R ++ L+ + +R YEI P+VQK+L V IE EL
Sbjct: 8 YIDSAYPSTGSILENEQRSNLMNNILRIISDLQRSCEYEIPVLPHVQKYLNSVRYIE-EL 66
Query: 87 QKWLSDDNAQI 97
QK++ DDN ++
Sbjct: 67 QKFVEDDNYKL 77
>UniRef100_Q7SXZ6 Hypothetical protein ralgps2 [Brachydanio rerio]
Length = 510
Score = 35.4 bits (80), Expect = 7.6
Identities = 24/71 (33%), Positives = 39/71 (54%), Gaps = 3/71 (4%)
Query: 29 YLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIA--PNVQKWLYDVTTIENEL 86
Y+ T ++ E +R ++ L+ + +R YEI P+VQK+L V IE EL
Sbjct: 211 YIDSAYPSTGSILENEQRSNLMNNILRIISDLQRSCEYEIPVLPHVQKYLNSVRYIE-EL 269
Query: 87 QKWLSDDNAQI 97
QK++ DDN ++
Sbjct: 270 QKFVEDDNYKL 280
>UniRef100_Q9ZT68 Resistance protein candidate RGC2K [Lactuca sativa]
Length = 1715
Score = 35.4 bits (80), Expect = 7.6
Identities = 19/93 (20%), Positives = 43/93 (45%), Gaps = 3/93 (3%)
Query: 1 MDCLTELGKETVTKLGELVVESTMKHFKYLTQHKKITTNLEEELERLKMIKQALQTRVET 60
M+C+T + + ++ +H L + + ++ + L K ++ R
Sbjct: 1 MECITGIFSNP---FAQCLIAPVKEHLCLLIFYTQYVGDMLTAMTELNAAKDIVEERKNQ 57
Query: 61 ERRKGYEIAPNVQKWLYDVTTIENELQKWLSDD 93
K +E+ +V +WL DV TI ++++ L+D+
Sbjct: 58 NVEKCFEVPNHVNRWLEDVQTINRKVERVLNDN 90
>UniRef100_UPI00002A1D22 UPI00002A1D22 UniRef100 entry
Length = 238
Score = 35.0 bits (79), Expect = 10.0
Identities = 31/124 (25%), Positives = 66/124 (53%), Gaps = 19/124 (15%)
Query: 18 LVVESTMKHFKYLTQHK--KITTN---LEEELERLKMIKQALQTRVETERRKG------- 65
L V+S +K + +++ + K +N +EEE ER K+ + + +E E+RKG
Sbjct: 48 LKVDSELKELRVISKKETEKWKSNKRIIEEEQER-KIQLETYRNELEIEKRKGNWDRAGE 106
Query: 66 --YEIAPNVQKWLYDVTTIENELQKWLSDDNAQI*HLIIHWERKPLKALKILQA*RKKKT 123
Y+I PN++K + + +++ + +++D+ I ++ W P++ K+L+ R K
Sbjct: 107 LSYQIIPNLEKMILESKVLDDLVNTTVTEDD--IAAVVSKWTGIPVE--KMLEFERTKLL 162
Query: 124 NFKL 127
N ++
Sbjct: 163 NIEM 166
>UniRef100_Q601Y2 Hypothetical protein [Mycoplasma hyopneumoniae]
Length = 869
Score = 35.0 bits (79), Expect = 10.0
Identities = 22/78 (28%), Positives = 42/78 (53%), Gaps = 3/78 (3%)
Query: 20 VESTMKHFKYLTQHKKITTNLEEELERLKMIKQALQTRVETERRKGYEIAPNVQKWLYDV 79
++ K+ ++ + KI T +E LER+ I + + + + Y+I P +QK+L D+
Sbjct: 126 LKENRKNTDFVLKVDKIFTRIEANLERIITIIKKINPKAHINLIEYYQIQPKIQKYLQDL 185
Query: 80 TTIENELQKWLSDDNAQI 97
EN L + ++D+N I
Sbjct: 186 A--ENYLIE-INDNNLSI 200
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.362 0.160 0.608
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,219,817,301
Number of Sequences: 2790947
Number of extensions: 42979621
Number of successful extensions: 169963
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 169952
Number of HSP's gapped (non-prelim): 18
length of query: 887
length of database: 848,049,833
effective HSP length: 136
effective length of query: 751
effective length of database: 468,481,041
effective search space: 351829261791
effective search space used: 351829261791
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.9 bits)
S2: 79 (35.0 bits)
Medicago: description of AC149050.7


