

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149038.5 + phase: 0 /partial
(345 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
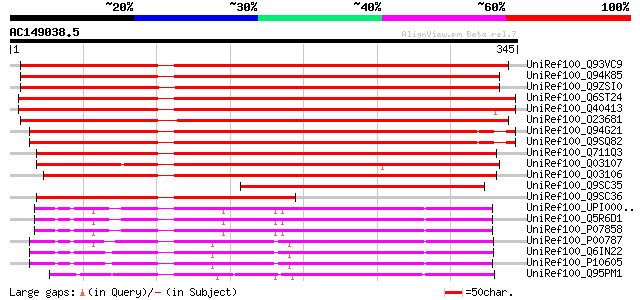
Sequences producing significant alignments: (bits) Value
UniRef100_Q93VC9 At1g02300/T6A9_10 [Arabidopsis thaliana] 538 e-152
UniRef100_Q94K85 Putative cathepsin B cysteine protease [Arabido... 524 e-147
UniRef100_Q9ZSI0 T15B16.17a protein [Arabidopsis thaliana] 524 e-147
UniRef100_Q6ST24 Cathepsin B-like cysteine proteinase [Solanum t... 517 e-145
UniRef100_Q40413 Cathepsin B-like cysteine proteinase [Nicotiana... 514 e-144
UniRef100_O23681 Cathepsin B-like cysteine proteinase [Arabidops... 506 e-142
UniRef100_Q94G21 Cathepsin B-like cysteine proteinase [Ipomoea b... 502 e-141
UniRef100_Q9SQ82 Cathepsin B-like cysteine proteinase [Ipomoea b... 498 e-140
UniRef100_Q711Q3 Cathepsin B [Hordeum vulgare] 485 e-136
UniRef100_Q03107 Cathepsin B [Triticum aestivum] 477 e-133
UniRef100_Q03106 Cathepsin B [Triticum aestivum] 471 e-131
UniRef100_Q9SC35 Putative cathepsin B-like protease [Pisum sativum] 342 1e-92
UniRef100_Q9SC36 Putative cathepsin B-like protease [Pisum sativum] 304 2e-81
UniRef100_UPI000036E1D7 UPI000036E1D7 UniRef100 entry 277 3e-73
UniRef100_Q5R6D1 Hypothetical protein DKFZp459D187 [Pongo pygmaeus] 277 3e-73
UniRef100_P07858 Cathepsin B precursor [Homo sapiens] 275 1e-72
UniRef100_P00787 Cathepsin B precursor [Rattus norvegicus] 273 7e-72
UniRef100_Q6IN22 Cathepsin B, preproprotein [Rattus norvegicus] 271 3e-71
UniRef100_P10605 Cathepsin B precursor [Mus musculus] 267 4e-70
UniRef100_Q95PM1 Cathepsin B endopeptidase precursor [Schistosom... 265 1e-69
>UniRef100_Q93VC9 At1g02300/T6A9_10 [Arabidopsis thaliana]
Length = 362
Score = 538 bits (1386), Expect = e-152
Identities = 244/332 (73%), Positives = 274/332 (82%), Gaps = 10/332 (3%)
Query: 8 EKLNGLKLNSHILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRLLGVKQAPKKELL 67
E L+ KL S ILQ I K++NENP AGW+A+ N RF+N TV +FKRLLGVK PK E L
Sbjct: 34 ENLSKQKLTSWILQNEIVKEVNENPNAGWKASFNDRFANATVAEFKRLLGVKPTPKTEFL 93
Query: 68 STPVVTHPKSLKLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESL 127
P+V+H SLKLPKEFDARTAWSQC++IG+IL QGHCGSCWAFGAVESL
Sbjct: 94 GVPIVSHDISLKLPKEFDARTAWSQCTSIGRILD----------QGHCGSCWAFGAVESL 143
Query: 128 QDRFCIHFDMNISLSVNDLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEECDPYFDQI 187
DRFCI ++MN+SLSVNDLLACCGFLCG GC+GG PI AWRY HHGVVTEECDPYFD
Sbjct: 144 SDRFCIKYNMNVSLSVNDLLACCGFLCGQGCNGGYPIAAWRYFKHHGVVTEECDPYFDNT 203
Query: 188 GCSHPGCEPAYQTPKCVRKCVKGNQIWKRSKHYSVKAYRVKSDPQDIMAEVYKNGPVEVA 247
GCSHPGCEPAY TPKC RKCV GNQ+W+ SKHY V AY+V+S P DIMAEVYKNGPVEVA
Sbjct: 204 GCSHPGCEPAYPTPKCARKCVSGNQLWRESKHYGVSAYKVRSHPDDIMAEVYKNGPVEVA 263
Query: 248 FTVFEDFAHYKSGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLLANQWNTNWGDDGYFK 307
FTV+EDFAHYKSGVYKHITG+ +GGHAVKLIGWGTSD+GEDYWLLANQWN +WGDDGYFK
Sbjct: 264 FTVYEDFAHYKSGVYKHITGTNIGGHAVKLIGWGTSDDGEDYWLLANQWNRSWGDDGYFK 323
Query: 308 IKRGTNECGIEDDVTAGLPSTKNIVREVTDMD 339
I+RGTNECGIE V AGLPS +N+V+ +T D
Sbjct: 324 IRRGTNECGIEHGVVAGLPSDRNVVKGITTSD 355
>UniRef100_Q94K85 Putative cathepsin B cysteine protease [Arabidopsis thaliana]
Length = 359
Score = 524 bits (1349), Expect = e-147
Identities = 241/326 (73%), Positives = 265/326 (80%), Gaps = 10/326 (3%)
Query: 8 EKLNGLKLNSHILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRLLGVKQAPKKELL 67
E L KL+S ILQ+ I K++NENP AGW+AAIN RFSN TV +FKRLLGVK PKK L
Sbjct: 31 ESLTKQKLDSKILQDEIVKKVNENPNAGWKAAINDRFSNATVAEFKRLLGVKPTPKKHFL 90
Query: 68 STPVVTHPKSLKLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESL 127
P+V+H SLKLPK FDARTAW QC++IG IL QGHCGSCWAFGAVESL
Sbjct: 91 GVPIVSHDPSLKLPKAFDARTAWPQCTSIGNILD----------QGHCGSCWAFGAVESL 140
Query: 128 QDRFCIHFDMNISLSVNDLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEECDPYFDQI 187
DRFCI F MNISLSVNDLLACCGF CG GCDGG PI AW+Y ++ GVVTEECDPYFD
Sbjct: 141 SDRFCIQFGMNISLSVNDLLACCGFRCGDGCDGGYPIAAWQYFSYSGVVTEECDPYFDNT 200
Query: 188 GCSHPGCEPAYQTPKCVRKCVKGNQIWKRSKHYSVKAYRVKSDPQDIMAEVYKNGPVEVA 247
GCSHPGCEPAY TPKC RKCV N++W SKHYSV Y VKS+PQDIMAEVYKNGPVEV+
Sbjct: 201 GCSHPGCEPAYPTPKCSRKCVSDNKLWSESKHYSVSTYTVKSNPQDIMAEVYKNGPVEVS 260
Query: 248 FTVFEDFAHYKSGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLLANQWNTNWGDDGYFK 307
FTV+EDFAHYKSGVYKHITGS +GGHAVKLIGWGTS EGEDYWL+ANQWN WGDDGYF
Sbjct: 261 FTVYEDFAHYKSGVYKHITGSNIGGHAVKLIGWGTSSEGEDYWLMANQWNRGWGDDGYFM 320
Query: 308 IKRGTNECGIEDDVTAGLPSTKNIVR 333
I+RGTNECGIED+ AGLPS+KN+ R
Sbjct: 321 IRRGTNECGIEDEPVAGLPSSKNVFR 346
>UniRef100_Q9ZSI0 T15B16.17a protein [Arabidopsis thaliana]
Length = 359
Score = 524 bits (1349), Expect = e-147
Identities = 241/326 (73%), Positives = 265/326 (80%), Gaps = 10/326 (3%)
Query: 8 EKLNGLKLNSHILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRLLGVKQAPKKELL 67
E L KL+S ILQ+ I K++NENP AGW+AAIN RFSN TV +FKRLLGVK PKK L
Sbjct: 31 ESLTKQKLDSKILQDEIVKKVNENPNAGWKAAINDRFSNATVAEFKRLLGVKPTPKKHFL 90
Query: 68 STPVVTHPKSLKLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESL 127
P+V+H SLKLPK FDARTAW QC++IG ILG GHCGSCWAFGAVESL
Sbjct: 91 GVPIVSHDPSLKLPKAFDARTAWPQCTSIGNILGL----------GHCGSCWAFGAVESL 140
Query: 128 QDRFCIHFDMNISLSVNDLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEECDPYFDQI 187
DRFCI F MNISLSVNDLLACCGF CG GCDGG PI AW+Y ++ GVVTEECDPYFD
Sbjct: 141 SDRFCIQFGMNISLSVNDLLACCGFRCGDGCDGGYPIAAWQYFSYSGVVTEECDPYFDNT 200
Query: 188 GCSHPGCEPAYQTPKCVRKCVKGNQIWKRSKHYSVKAYRVKSDPQDIMAEVYKNGPVEVA 247
GCSHPGCEPAY TPKC RKCV N++W SKHYSV Y VKS+PQDIMAEVYKNGPVEV+
Sbjct: 201 GCSHPGCEPAYPTPKCSRKCVSDNKLWSESKHYSVSTYTVKSNPQDIMAEVYKNGPVEVS 260
Query: 248 FTVFEDFAHYKSGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLLANQWNTNWGDDGYFK 307
FTV+EDFAHYKSGVYKHITGS +GGHAVKLIGWGTS EGEDYWL+ANQWN WGDDGYF
Sbjct: 261 FTVYEDFAHYKSGVYKHITGSNIGGHAVKLIGWGTSSEGEDYWLMANQWNRGWGDDGYFM 320
Query: 308 IKRGTNECGIEDDVTAGLPSTKNIVR 333
I+RGTNECGIED+ AGLPS+KN+ R
Sbjct: 321 IRRGTNECGIEDEPVAGLPSSKNVFR 346
>UniRef100_Q6ST24 Cathepsin B-like cysteine proteinase [Solanum tuberosum]
Length = 354
Score = 517 bits (1331), Expect = e-145
Identities = 236/338 (69%), Positives = 272/338 (79%), Gaps = 10/338 (2%)
Query: 7 DEKLNGLKLNSHILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRLLGVKQAPKKEL 66
++ ++ KL S ILQ+SI K++NEN EAGW+AA NP+ SNFTV QFKRLLGVK A + +L
Sbjct: 27 EKPISEAKLESAILQDSIVKRVNENAEAGWKAAFNPQLSNFTVSQFKRLLGVKPAREGDL 86
Query: 67 LSTPVVTHPKSLKLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVES 126
PV+THP+ +LPKEFDAR AW QCSTIGKIL QGHCGSCWAFGAVES
Sbjct: 87 EGIPVLTHPRLKELPKEFDARKAWPQCSTIGKILD----------QGHCGSCWAFGAVES 136
Query: 127 LQDRFCIHFDMNISLSVNDLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEECDPYFDQ 186
L DRFCIH++++ISLSVNDLLACC FLCG+GCDGG PI AWRY GVVTEECDPYFD
Sbjct: 137 LSDRFCIHYNLSISLSVNDLLACCSFLCGSGCDGGYPIAAWRYFKRSGVVTEECDPYFDT 196
Query: 187 IGCSHPGCEPAYQTPKCVRKCVKGNQIWKRSKHYSVKAYRVKSDPQDIMAEVYKNGPVEV 246
GCSHPGCEP Y TPKC RKCVKGN +W++SKHY V AYRV DPQ IMAEVYKNGPVEV
Sbjct: 197 TGCSHPGCEPLYPTPKCHRKCVKGNVLWRKSKHYGVNAYRVSHDPQSIMAEVYKNGPVEV 256
Query: 247 AFTVFEDFAHYKSGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLLANQWNTNWGDDGYF 306
+FTV+EDFAHYKSGVYKH+TG +GGHAVKLIGWGTS++GEDYWL+ N WN WG+DGYF
Sbjct: 257 SFTVYEDFAHYKSGVYKHVTGGNMGGHAVKLIGWGTSEQGEDYWLIVNSWNRGWGEDGYF 316
Query: 307 KIKRGTNECGIEDDVTAGLPSTKNIVREVTDMDVDAGV 344
KI+RGTNECGIE V AGLPS +N+ E+ D +DA +
Sbjct: 317 KIRRGTNECGIEHSVVAGLPSARNLNVELGDAVLDASM 354
>UniRef100_Q40413 Cathepsin B-like cysteine proteinase [Nicotiana rustica]
Length = 356
Score = 514 bits (1325), Expect = e-144
Identities = 234/340 (68%), Positives = 271/340 (78%), Gaps = 12/340 (3%)
Query: 7 DEKLNGLKLNSHILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRLLGVKQAPKKEL 66
++ ++ K S ILQ+SI KQ+NEN +AGW+AA+NPRFSNFTV QFKRLLGVK K +L
Sbjct: 27 EQPISQAKAESAILQDSIVKQVNENEKAGWKAALNPRFSNFTVSQFKRLLGVKPTRKGDL 86
Query: 67 LSTPVVTHPKSLKLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVES 126
P++THPK L+LP+EFDAR AWS CSTIG+IL QGHCGSCWAFGAVES
Sbjct: 87 KGIPILTHPKLLELPQEFDARVAWSNCSTIGRILD----------QGHCGSCWAFGAVES 136
Query: 127 LQDRFCIHFDMNISLSVNDLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEECDPYFDQ 186
L DRFCIH+ +NISLS NDL ACCGFLCG GCDGG P+ AW+Y GVVT+ECDPYFD
Sbjct: 137 LSDRFCIHYGLNISLSANDLYACCGFLCGDGCDGGYPLQAWKYFVRKGVVTDECDPYFDN 196
Query: 187 IGCSHPGCEPAYQTPKCVRKCVKGNQIWKRSKHYSVKAYRVKSDPQDIMAEVYKNGPVEV 246
GCSHPGCEPAY TPKC RKCVK N +W RSKH+ V AY + SDP IM EVYKNGPVEV
Sbjct: 197 EGCSHPGCEPAYPTPKCHRKCVKQNLLWSRSKHFGVNAYMISSDPHSIMTEVYKNGPVEV 256
Query: 247 AFTVFEDFAHYKSGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLLANQWNTNWGDDGYF 306
+FTV+EDFAHYKSGVYKH+TG +GGHAVKLIGWGTS++GEDYWLLANQWN WGDDGYF
Sbjct: 257 SFTVYEDFAHYKSGVYKHVTGDIMGGHAVKLIGWGTSEDGEDYWLLANQWNRGWGDDGYF 316
Query: 307 KIKRGTNECGIEDDVTAGLPSTK--NIVREVTDMDVDAGV 344
KI+RGTNEC IED+V AGLPS + N+ +V+D +DA +
Sbjct: 317 KIRRGTNECEIEDEVVAGLPSARNLNVELDVSDAFLDAAM 356
>UniRef100_O23681 Cathepsin B-like cysteine proteinase [Arabidopsis thaliana]
Length = 357
Score = 506 bits (1302), Expect = e-142
Identities = 230/332 (69%), Positives = 262/332 (78%), Gaps = 12/332 (3%)
Query: 8 EKLNGLKLNSHILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRLLGVKQAPKKELL 67
E L+ KL S ILQ I K++NENP AGW+AA N RF+N TV +FKRLLGV Q PK L
Sbjct: 31 ENLSKQKLTSLILQNEIVKEVNENPNAGWKAAFNDRFANATVAEFKRLLGVIQTPKTAYL 90
Query: 68 STPVVTHPKSLKLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESL 127
P+V H SLKLPKEFDARTAWS C++I +ILG HCGSCWAFGAVESL
Sbjct: 91 GVPIVRHDLSLKLPKEFDARTAWSHCTSIRRILG------------HCGSCWAFGAVESL 138
Query: 128 QDRFCIHFDMNISLSVNDLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEECDPYFDQI 187
DRFCI +++N+SLS ND++ACCG LCG GC+GG P+ AW Y +HGVVT+ECDPYFD
Sbjct: 139 SDRFCIKYNLNVSLSANDVIACCGLLCGFGCNGGFPMGAWLYFKYHGVVTQECDPYFDNT 198
Query: 188 GCSHPGCEPAYQTPKCVRKCVKGNQIWKRSKHYSVKAYRVKSDPQDIMAEVYKNGPVEVA 247
GCSHPGCEP Y TPKC RKCV NQ+W SKHY V AYR+ DPQDIMAEVYKNGPVEVA
Sbjct: 199 GCSHPGCEPTYPTPKCERKCVSRNQLWGESKHYGVGAYRINPDPQDIMAEVYKNGPVEVA 258
Query: 248 FTVFEDFAHYKSGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLLANQWNTNWGDDGYFK 307
FTV+EDFAHYKSGVYK+ITG+ +GGHAVKLIGWGTSD+GEDYWLLANQWN +WGDDGYFK
Sbjct: 259 FTVYEDFAHYKSGVYKYITGTKIGGHAVKLIGWGTSDDGEDYWLLANQWNRSWGDDGYFK 318
Query: 308 IKRGTNECGIEDDVTAGLPSTKNIVREVTDMD 339
I+RGTNECGIE V AGLPS KN+ + +T D
Sbjct: 319 IRRGTNECGIEQSVVAGLPSEKNVFKGITTSD 350
>UniRef100_Q94G21 Cathepsin B-like cysteine proteinase [Ipomoea batatas]
Length = 352
Score = 502 bits (1293), Expect = e-141
Identities = 233/331 (70%), Positives = 264/331 (79%), Gaps = 18/331 (5%)
Query: 14 KLNSHILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRLLGVKQAPKKELLSTPVVT 73
+++ ILQ+ I K +NENPEAGW+A +NPRFS+FTV QFKRLLGVK+APK L TPVVT
Sbjct: 30 EVDPKILQDEIVKTVNENPEAGWKADMNPRFSDFTVSQFKRLLGVKKAPKSLLKRTPVVT 89
Query: 74 HPKSLKLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESLQDRFCI 133
H K ++LPK FDARTAW QC +I IL QGHCGSCWAFGAVESL DRFCI
Sbjct: 90 HSKEIELPKTFDARTAWPQCLSIADILD----------QGHCGSCWAFGAVESLTDRFCI 139
Query: 134 HFDMNISLSVNDLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEECDPYFDQIGCSHPG 193
H+ N++LSVNDLLACCGFLCG GCDGG PI AW+Y GVVT ECDPYFDQ GCSHPG
Sbjct: 140 HYGTNVTLSVNDLLACCGFLCGEGCDGGYPIAAWQYFKRTGVVTSECDPYFDQTGCSHPG 199
Query: 194 CEPAYQTPKCVRKCVKGNQIWKRSKHYSVKAYRVKSDPQDIMAEVYKNGPVEVAFTVFED 253
CEPAY TP C +KCVK N +W SKH+SV AYRV SD IM EVY NGP EV+FTV+ED
Sbjct: 200 CEPAYPTPACEKKCVKKNLLWSESKHFSVNAYRVNSDQHSIMTEVYTNGPAEVSFTVYED 259
Query: 254 FAHYKSGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLLANQWNTNWGDDGYFKIKRGTN 313
FAHYKSGVYKH+TGS +GGHAVKLIGWGTS++GEDYWLLANQWN +WGDDGYFKI RGTN
Sbjct: 260 FAHYKSGVYKHVTGSEMGGHAVKLIGWGTSEDGEDYWLLANQWNRSWGDDGYFKIIRGTN 319
Query: 314 ECGIEDDVTAGLPSTKNIVREVTDMDVDAGV 344
ECGIE DVTAG+PSTKN +D+++GV
Sbjct: 320 ECGIE-DVTAGMPSTKN-------LDIESGV 342
>UniRef100_Q9SQ82 Cathepsin B-like cysteine proteinase [Ipomoea batatas]
Length = 352
Score = 498 bits (1283), Expect = e-140
Identities = 232/331 (70%), Positives = 262/331 (79%), Gaps = 18/331 (5%)
Query: 14 KLNSHILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRLLGVKQAPKKELLSTPVVT 73
+++ ILQ+ I K +NENPEAGW+A +NPRFS+FTV QFKRLLGVK+APK L TPVVT
Sbjct: 30 EVDPKILQDEIVKTVNENPEAGWKADMNPRFSDFTVSQFKRLLGVKKAPKSLLKRTPVVT 89
Query: 74 HPKSLKLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESLQDRFCI 133
H K ++LPK FDARTAW QC +I IL QGHCGSCWAFGAVESL DRFCI
Sbjct: 90 HSKEIELPKTFDARTAWPQCLSIADILD----------QGHCGSCWAFGAVESLTDRFCI 139
Query: 134 HFDMNISLSVNDLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEECDPYFDQIGCSHPG 193
H+ N++LSVNDLLACCGFLCG GCDGG PI AW+Y GVVT ECDPYFDQ GCSHPG
Sbjct: 140 HYGTNVTLSVNDLLACCGFLCGEGCDGGYPIAAWQYFKRTGVVTSECDPYFDQTGCSHPG 199
Query: 194 CEPAYQTPKCVRKCVKGNQIWKRSKHYSVKAYRVKSDPQDIMAEVYKNGPVEVAFTVFED 253
CEPAY TP C +KCVK N +W SKH+SV AYRV SD IM EVY NGP EV+FTV+ED
Sbjct: 200 CEPAYPTPACEKKCVKKNLLWSESKHFSVNAYRVNSDQHSIMTEVYTNGPAEVSFTVYED 259
Query: 254 FAHYKSGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLLANQWNTNWGDDGYFKIKRGTN 313
FAHYKSGVYKH+TGS +GGHAVKLIGWGTS++GEDYWLLANQWN +WG DGYFKI RGTN
Sbjct: 260 FAHYKSGVYKHVTGSEMGGHAVKLIGWGTSEDGEDYWLLANQWNRSWGGDGYFKIIRGTN 319
Query: 314 ECGIEDDVTAGLPSTKNIVREVTDMDVDAGV 344
ECGIE DVTAG PSTKN +D+++GV
Sbjct: 320 ECGIE-DVTAGTPSTKN-------LDIESGV 342
>UniRef100_Q711Q3 Cathepsin B [Hordeum vulgare]
Length = 344
Score = 485 bits (1248), Expect = e-136
Identities = 219/313 (69%), Positives = 247/313 (77%), Gaps = 10/313 (3%)
Query: 19 ILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRLLGVKQAPKKELLSTPVVTHPKSL 78
I+Q+ I + +N +P AGW A NP +N+T+ QFK +LGVK P L THP+S
Sbjct: 35 IIQKGIIQTVNNHPNAGWTAGHNPYLANYTIEQFKHMLGVKPTPPGLLAGVRTKTHPRSE 94
Query: 79 KLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESLQDRFCIHFDMN 138
+LPKEFDAR+ WS CSTIGKIL QGHCGSCWAFGAVE LQDRFCIH +MN
Sbjct: 95 QLPKEFDARSKWSGCSTIGKILD----------QGHCGSCWAFGAVECLQDRFCIHHNMN 144
Query: 139 ISLSVNDLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEECDPYFDQIGCSHPGCEPAY 198
ISLS NDL+ACCGF+CG GCDGG PI AW+Y +GVVTEECDPYFDQ+GC HPGCEPAY
Sbjct: 145 ISLSANDLVACCGFMCGDGCDGGYPISAWQYFVQNGVVTEECDPYFDQVGCKHPGCEPAY 204
Query: 199 QTPKCVRKCVKGNQIWKRSKHYSVKAYRVKSDPQDIMAEVYKNGPVEVAFTVFEDFAHYK 258
TP C +KC NQ+W+ KH+S+ AY+V SDP DIMAEVYKNGPVEVAFTV+EDFAHYK
Sbjct: 205 PTPVCEKKCKVQNQVWQEKKHFSIDAYQVNSDPHDIMAEVYKNGPVEVAFTVYEDFAHYK 264
Query: 259 SGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLLANQWNTNWGDDGYFKIKRGTNECGIE 318
SGVYKHITG +GGHAVKLIGWGTSD GEDYWLLANQWN WGDDGYFKI RG NECGIE
Sbjct: 265 SGVYKHITGGVMGGHAVKLIGWGTSDAGEDYWLLANQWNRGWGDDGYFKIIRGKNECGIE 324
Query: 319 DDVTAGLPSTKNI 331
+DVTAG+PS KNI
Sbjct: 325 EDVTAGMPSMKNI 337
>UniRef100_Q03107 Cathepsin B [Triticum aestivum]
Length = 353
Score = 477 bits (1228), Expect = e-133
Identities = 221/317 (69%), Positives = 247/317 (77%), Gaps = 13/317 (4%)
Query: 19 ILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRLLGVKQAPKKELLSTPVVTHPKSL 78
I+Q+ I + +N++P AGW A NP F+N+T+ QFK +LGVK P L P+ HP+ +
Sbjct: 37 IIQKDIIQTVNKHPNAGWTAGHNPYFANYTIEQFKHILGVKPTPPGLLAGVPIKIHPE-M 95
Query: 79 KLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESLQDRFCIHFDMN 138
LPKEFDART WS CSTIG IL QGHCG+CWAF AVE+LQDRFCIH +M+
Sbjct: 96 DLPKEFDARTQWSSCSTIGNILD----------QGHCGACWAFAAVEALQDRFCIHLNMS 145
Query: 139 ISLSVNDLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEECDPYFDQIGCSHPGCEPAY 198
+SLSVNDLLACCGFLCG+GC+GG PI AWRY GVVTEECDPYFDQ GC HPGCEPAY
Sbjct: 146 VSLSVNDLLACCGFLCGSGCNGGYPISAWRYFRRSGVVTEECDPYFDQTGCQHPGCEPAY 205
Query: 199 QTPKCVRKCVKGNQIWKRSKHYSVKAYRVKSDPQDIMAEVYKNGPVEVAFTVFE--DFAH 256
TPKC RKC NQ WK +KH+SV AYRV S+P DIMAEVYKNGPVEVAFT + DFAH
Sbjct: 206 PTPKCQRKCKVENQAWKENKHFSVNAYRVHSNPHDIMAEVYKNGPVEVAFTYCQILDFAH 265
Query: 257 YKSGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLLANQWNTNWGDDGYFKIKRGTNECG 316
YKSGVYKHITG +GGHAVKLIGWGTSD GEDYWLLANQWN WGDDGYFKI RG NECG
Sbjct: 266 YKSGVYKHITGGVMGGHAVKLIGWGTSDAGEDYWLLANQWNRGWGDDGYFKIIRGENECG 325
Query: 317 IEDDVTAGLPSTKNIVR 333
IE DVTAG+PSTKN R
Sbjct: 326 IEGDVTAGMPSTKNTAR 342
>UniRef100_Q03106 Cathepsin B [Triticum aestivum]
Length = 305
Score = 471 bits (1212), Expect = e-131
Identities = 212/308 (68%), Positives = 241/308 (77%), Gaps = 10/308 (3%)
Query: 24 IAKQINENPEAGWEAAINPRFSNFTVGQFKRLLGVKQAPKKELLSTPVVTHPKSLKLPKE 83
I + +N +P AGW A NP +N+T+ QFK +LGVK P + TH +S +LPK
Sbjct: 1 IIQTVNNHPNAGWTAGHNPYLANYTIEQFKHMLGVKPTPPGLRAAVRTKTHSRSEQLPKV 60
Query: 84 FDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESLQDRFCIHFDMNISLSV 143
FDAR+ WS CSTIGKIL QGHCGSCWAFGAVE LQDRFCIH +MNI+LS
Sbjct: 61 FDARSKWSGCSTIGKILD----------QGHCGSCWAFGAVECLQDRFCIHHNMNITLSA 110
Query: 144 NDLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEECDPYFDQIGCSHPGCEPAYQTPKC 203
NDL+ACCGF+CG GCDGG PI AW+Y +GVVT+ECDPYFDQ+GC HPGCEPAY TP C
Sbjct: 111 NDLVACCGFMCGDGCDGGYPISAWQYFVQNGVVTDECDPYFDQVGCKHPGCEPAYPTPVC 170
Query: 204 VRKCVKGNQIWKRSKHYSVKAYRVKSDPQDIMAEVYKNGPVEVAFTVFEDFAHYKSGVYK 263
+KC NQ+W+ KH+S+ AY+V SDP DIMAEVY NGPVEVAFTV+EDFAHYKSGVYK
Sbjct: 171 EKKCKVQNQVWEEKKHFSINAYQVNSDPHDIMAEVYNNGPVEVAFTVYEDFAHYKSGVYK 230
Query: 264 HITGSALGGHAVKLIGWGTSDEGEDYWLLANQWNTNWGDDGYFKIKRGTNECGIEDDVTA 323
HITG +GGHAVKLIGWGTSD GEDYWLLANQWN WGDDGYFKI RG NECGIE+DVTA
Sbjct: 231 HITGGVMGGHAVKLIGWGTSDAGEDYWLLANQWNRGWGDDGYFKIIRGKNECGIEEDVTA 290
Query: 324 GLPSTKNI 331
G+PSTKNI
Sbjct: 291 GMPSTKNI 298
>UniRef100_Q9SC35 Putative cathepsin B-like protease [Pisum sativum]
Length = 174
Score = 342 bits (876), Expect = 1e-92
Identities = 150/166 (90%), Positives = 158/166 (94%)
Query: 158 CDGGTPIYAWRYLAHHGVVTEECDPYFDQIGCSHPGCEPAYQTPKCVRKCVKGNQIWKRS 217
CDGG PI AW+Y AHHGVVTEECDPYFDQIGCSHPGCEP YQTPKCVRKCVKGNQ+WK+S
Sbjct: 1 CDGGYPISAWKYFAHHGVVTEECDPYFDQIGCSHPGCEPGYQTPKCVRKCVKGNQVWKKS 60
Query: 218 KHYSVKAYRVKSDPQDIMAEVYKNGPVEVAFTVFEDFAHYKSGVYKHITGSALGGHAVKL 277
KHYSVK Y+V SDPQ+IM EVYKNGPVEVAF+V+EDFAHYKSGVYKHITGSALGGHAVKL
Sbjct: 61 KHYSVKPYKVNSDPQNIMEEVYKNGPVEVAFSVYEDFAHYKSGVYKHITGSALGGHAVKL 120
Query: 278 IGWGTSDEGEDYWLLANQWNTNWGDDGYFKIKRGTNECGIEDDVTA 323
GWGTSDEGEDYWLLANQWNTNWGDDGYFKIKRGTNECGIE+DVTA
Sbjct: 121 NGWGTSDEGEDYWLLANQWNTNWGDDGYFKIKRGTNECGIEEDVTA 166
>UniRef100_Q9SC36 Putative cathepsin B-like protease [Pisum sativum]
Length = 206
Score = 304 bits (779), Expect = 2e-81
Identities = 140/176 (79%), Positives = 151/176 (85%), Gaps = 10/176 (5%)
Query: 19 ILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRLLGVKQAPKKELLSTPVVTHPKSL 78
+LQESIAK++NENP AGW+AAINPRFSN TVGQFKRLLGVKQ P+ EL S PVVTHPKSL
Sbjct: 41 LLQESIAKEVNENPGAGWKAAINPRFSNSTVGQFKRLLGVKQTPRNELSSIPVVTHPKSL 100
Query: 79 KLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESLQDRFCIHFDMN 138
LPKEFDARTAW QCSTIG+IL QGHCGSCWAFGAVESL DRFCIHF ++
Sbjct: 101 NLPKEFDARTAWPQCSTIGRILD----------QGHCGSCWAFGAVESLSDRFCIHFGVD 150
Query: 139 ISLSVNDLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEECDPYFDQIGCSHPGC 194
+ LSVNDLLACCGFLCG+GCDGG PI AW+Y AHHGVVTEECDPYFDQIGCSHPGC
Sbjct: 151 VPLSVNDLLACCGFLCGSGCDGGYPISAWKYFAHHGVVTEECDPYFDQIGCSHPGC 206
>UniRef100_UPI000036E1D7 UPI000036E1D7 UniRef100 entry
Length = 339
Score = 277 bits (708), Expect = 3e-73
Identities = 143/330 (43%), Positives = 195/330 (58%), Gaps = 40/330 (12%)
Query: 18 HILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRL----LGVKQAPKKELLSTPVVT 73
H L + + +N+ W+A N F N + KRL LG + P++ + +
Sbjct: 24 HPLSDELVNYVNKR-NTTWQAGHN--FYNVDMSYLKRLCGAFLGGPKPPQRVMFT----- 75
Query: 74 HPKSLKLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESLQDRFCI 133
+ LKLP+ FDAR W QC TI +I QG CGSCWAFGAVE++ DR CI
Sbjct: 76 --EDLKLPESFDAREQWPQCPTIKEIRD----------QGSCGSCWAFGAVEAISDRICI 123
Query: 134 HFDMNISLSVN--DLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEE-------CDPYF 184
H + ++S+ V+ DLL CCG +CG GC+GG P AW + G+V+ C PY
Sbjct: 124 HTNAHVSVEVSAEDLLTCCGSMCGDGCNGGYPAEAWNFWTRKGLVSGGLYESHVGCRPYS 183
Query: 185 -----DQIGCSHPGCEPAYQTPKCVRKCVKG-NQIWKRSKHYSVKAYRVKSDPQDIMAEV 238
+ S P C TPKC + C G + +K+ KHY +Y V + +DIMAE+
Sbjct: 184 IPPCEHHVNGSRPPCTGEGDTPKCSKICEPGYSPTYKQDKHYGYNSYSVSNSEKDIMAEI 243
Query: 239 YKNGPVEVAFTVFEDFAHYKSGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLLANQWNT 298
YKNGPVE AF+V+ DF YKSGVY+H+TG +GGHA++++GWG + G YWL+AN WNT
Sbjct: 244 YKNGPVEGAFSVYSDFLLYKSGVYQHVTGEMMGGHAIRILGWGV-ENGTPYWLVANSWNT 302
Query: 299 NWGDDGYFKIKRGTNECGIEDDVTAGLPST 328
+WGD+G+FKI RG + CGIE +V AG+P T
Sbjct: 303 DWGDNGFFKILRGQDHCGIESEVVAGIPRT 332
>UniRef100_Q5R6D1 Hypothetical protein DKFZp459D187 [Pongo pygmaeus]
Length = 339
Score = 277 bits (708), Expect = 3e-73
Identities = 143/330 (43%), Positives = 195/330 (58%), Gaps = 40/330 (12%)
Query: 18 HILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRL----LGVKQAPKKELLSTPVVT 73
H L + + +N+ W+A N F N V K+L LG + P++ + +
Sbjct: 24 HPLSDELVNYVNKR-NTTWQAGHN--FYNVDVSYLKKLCGTFLGGPKPPQRVMFT----- 75
Query: 74 HPKSLKLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESLQDRFCI 133
+ LKLP+ FDAR W QC TI +I QG CGSCWAFGAVE++ DR CI
Sbjct: 76 --EDLKLPESFDAREQWPQCPTIKEIRD----------QGSCGSCWAFGAVEAISDRICI 123
Query: 134 HFDMNISLSVN--DLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEE-------CDPYF 184
H + ++S+ V+ DLL CCG +CG GC+GG P AW + G+V+ C PY
Sbjct: 124 HTNAHVSVEVSAEDLLTCCGSMCGDGCNGGYPAEAWNFWTRKGLVSGGLYESHVGCRPYS 183
Query: 185 -----DQIGCSHPGCEPAYQTPKCVRKCVKG-NQIWKRSKHYSVKAYRVKSDPQDIMAEV 238
+ S P C TPKC + C G + +K+ KHY +Y V + +DIMAE+
Sbjct: 184 IPPCEHHVNGSRPPCTGEGDTPKCSKICEPGYSPTYKQDKHYGYNSYSVSNSERDIMAEI 243
Query: 239 YKNGPVEVAFTVFEDFAHYKSGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLLANQWNT 298
YKNGPVE AF+V+ DF YKSGVY+H+TG +GGHA++++GWG + G YWL+AN WNT
Sbjct: 244 YKNGPVEGAFSVYSDFLLYKSGVYQHVTGEMMGGHAIRILGWGV-ENGTPYWLVANSWNT 302
Query: 299 NWGDDGYFKIKRGTNECGIEDDVTAGLPST 328
+WGD+G+FKI RG + CGIE +V AG+P T
Sbjct: 303 DWGDNGFFKILRGQDHCGIESEVVAGIPRT 332
>UniRef100_P07858 Cathepsin B precursor [Homo sapiens]
Length = 339
Score = 275 bits (703), Expect = 1e-72
Identities = 142/330 (43%), Positives = 194/330 (58%), Gaps = 40/330 (12%)
Query: 18 HILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRL----LGVKQAPKKELLSTPVVT 73
H + + + +N+ W+A N F N + KRL LG + P++ + +
Sbjct: 24 HPVSDELVNYVNKR-NTTWQAGHN--FYNVDMSYLKRLCGTFLGGPKPPQRVMFT----- 75
Query: 74 HPKSLKLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESLQDRFCI 133
+ LKLP FDAR W QC TI +I QG CGSCWAFGAVE++ DR CI
Sbjct: 76 --EDLKLPASFDAREQWPQCPTIKEIRD----------QGSCGSCWAFGAVEAISDRICI 123
Query: 134 HFDMNISLSVN--DLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEE-------CDPYF 184
H + ++S+ V+ DLL CCG +CG GC+GG P AW + G+V+ C PY
Sbjct: 124 HTNAHVSVEVSAEDLLTCCGSMCGDGCNGGYPAEAWNFWTRKGLVSGGLYESHVGCRPYS 183
Query: 185 -----DQIGCSHPGCEPAYQTPKCVRKCVKG-NQIWKRSKHYSVKAYRVKSDPQDIMAEV 238
+ S P C TPKC + C G + +K+ KHY +Y V + +DIMAE+
Sbjct: 184 IPPCEHHVNGSRPPCTGEGDTPKCSKICEPGYSPTYKQDKHYGYNSYSVSNSEKDIMAEI 243
Query: 239 YKNGPVEVAFTVFEDFAHYKSGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLLANQWNT 298
YKNGPVE AF+V+ DF YKSGVY+H+TG +GGHA++++GWG + G YWL+AN WNT
Sbjct: 244 YKNGPVEGAFSVYSDFLLYKSGVYQHVTGEMMGGHAIRILGWGV-ENGTPYWLVANSWNT 302
Query: 299 NWGDDGYFKIKRGTNECGIEDDVTAGLPST 328
+WGD+G+FKI RG + CGIE +V AG+P T
Sbjct: 303 DWGDNGFFKILRGQDHCGIESEVVAGIPRT 332
>UniRef100_P00787 Cathepsin B precursor [Rattus norvegicus]
Length = 339
Score = 273 bits (697), Expect = 7e-72
Identities = 145/337 (43%), Positives = 189/337 (56%), Gaps = 44/337 (13%)
Query: 14 KLNSHILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRL----LGVKQAPKKELLST 69
K +SH L + + IN+ W+A N F N + K+L LG P++
Sbjct: 20 KPSSHPLSDDMINYINKQ-NTTWQAGRN--FYNVDISYLKKLCGTVLGGPNLPER----- 71
Query: 70 PVVTHPKSLKLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESLQD 129
V + + LP+ FDAR WS C TI +I QG CGSCWAFGAVE++ D
Sbjct: 72 --VGFSEDINLPESFDAREQWSNCPTIAQIRD----------QGSCGSCWAFGAVEAMSD 119
Query: 130 RFCIHFD--MNISLSVNDLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEECDPYFDQI 187
R CIH + +N+ +S DLL CCG CG GC+GG P AW + G+V+ Y I
Sbjct: 120 RICIHTNGRVNVEVSAEDLLTCCGIQCGDGCNGGYPSGAWNFWTRKGLVSGGV--YNSHI 177
Query: 188 GC--------------SHPGCEPAYQTPKCVRKCVKG-NQIWKRSKHYSVKAYRVKSDPQ 232
GC S P C TPKC + C G + +K KHY +Y V +
Sbjct: 178 GCLPYTIPPCEHHVNGSRPPCTGEGDTPKCNKMCEAGYSTSYKEDKHYGYTSYSVSDSEK 237
Query: 233 DIMAEVYKNGPVEVAFTVFEDFAHYKSGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLL 292
+IMAE+YKNGPVE AFTVF DF YKSGVYKH G +GGHA++++GWG + G YWL+
Sbjct: 238 EIMAEIYKNGPVEGAFTVFSDFLTYKSGVYKHEAGDVMGGHAIRILGWGI-ENGVPYWLV 296
Query: 293 ANQWNTNWGDDGYFKIKRGTNECGIEDDVTAGLPSTK 329
AN WN +WGD+G+FKI RG N CGIE ++ AG+P T+
Sbjct: 297 ANSWNVDWGDNGFFKILRGENHCGIESEIVAGIPRTQ 333
>UniRef100_Q6IN22 Cathepsin B, preproprotein [Rattus norvegicus]
Length = 339
Score = 271 bits (692), Expect = 3e-71
Identities = 145/334 (43%), Positives = 187/334 (55%), Gaps = 38/334 (11%)
Query: 14 KLNSHILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRLLG-VKQAPKKELLSTPVV 72
K + H L + + IN+ W+A N F N + K+L G V PK V
Sbjct: 20 KPSFHPLSDDMINYINKQ-NTTWQAGRN--FYNVDISYLKKLCGTVLGGPKLP----ERV 72
Query: 73 THPKSLKLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESLQDRFC 132
+ + LP+ FDAR WS C TI +I QG CGSCWAFGAVE++ DR C
Sbjct: 73 GFSEDINLPESFDAREQWSNCPTIAQIRD----------QGSCGSCWAFGAVEAMSDRIC 122
Query: 133 IHFD--MNISLSVNDLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEECDPYFDQIGC- 189
IH + +N+ +S DLL CCG CG GC+GG P AW + G+V+ Y IGC
Sbjct: 123 IHTNGRVNVEVSAEDLLTCCGIQCGDGCNGGYPSGAWNFWTRKGLVSGGV--YNSHIGCL 180
Query: 190 -------------SHPGCEPAYQTPKCVRKCVKG-NQIWKRSKHYSVKAYRVKSDPQDIM 235
S P C TPKC + C G + +K KHY +Y V ++IM
Sbjct: 181 PYTIPPCEHHVNGSRPPCTGEGDTPKCNKMCEAGYSTSYKEDKHYGYTSYSVSDSEKEIM 240
Query: 236 AEVYKNGPVEVAFTVFEDFAHYKSGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLLANQ 295
AE+YKNGPVE AFTVF DF YKSGVYKH G +GGHA++++GWG + G YWL+AN
Sbjct: 241 AEIYKNGPVEGAFTVFSDFLTYKSGVYKHEAGDVMGGHAIRILGWGI-ENGVPYWLVANS 299
Query: 296 WNTNWGDDGYFKIKRGTNECGIEDDVTAGLPSTK 329
WN +WGD+G+FKI RG N CGIE ++ AG+P T+
Sbjct: 300 WNVDWGDNGFFKILRGENHCGIESEIVAGIPRTQ 333
>UniRef100_P10605 Cathepsin B precursor [Mus musculus]
Length = 339
Score = 267 bits (682), Expect = 4e-70
Identities = 143/333 (42%), Positives = 187/333 (55%), Gaps = 38/333 (11%)
Query: 14 KLNSHILQESIAKQINENPEAGWEAAINPRFSNFTVGQFKRLLG-VKQAPKKELLSTPVV 72
K + H L + + IN+ W+A N F N + K+L G V PK V
Sbjct: 20 KPSFHPLSDDLINYINKQ-NTTWQAGRN--FYNVDISYLKKLCGTVLGGPKLP----GRV 72
Query: 73 THPKSLKLPKEFDARTAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESLQDRFC 132
+ + LP+ FDAR WS C TIG+I QG CGSCWAFGAVE++ DR C
Sbjct: 73 AFGEDIDLPETFDAREQWSNCPTIGQIRD----------QGSCGSCWAFGAVEAISDRTC 122
Query: 133 IHFD--MNISLSVNDLLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEECDPYFDQIGC- 189
IH + +N+ +S DLL CCG CG GC+GG P AW + G+V+ Y +GC
Sbjct: 123 IHTNGRVNVEVSAEDLLTCCGIQCGDGCNGGYPSGAWSFWTKKGLVSGGV--YNSHVGCL 180
Query: 190 -------------SHPGCEPAYQTPKCVRKCVKG-NQIWKRSKHYSVKAYRVKSDPQDIM 235
S P C TP+C + C G + +K KH+ +Y V + ++IM
Sbjct: 181 PYTIPPCEHHVNGSRPPCTGEGDTPRCNKSCEAGYSPSYKEDKHFGYTSYSVSNSVKEIM 240
Query: 236 AEVYKNGPVEVAFTVFEDFAHYKSGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLLANQ 295
AE+YKNGPVE AFTVF DF YKSGVYKH G +GGHA++++GWG + G YWL AN
Sbjct: 241 AEIYKNGPVEGAFTVFSDFLTYKSGVYKHEAGDMMGGHAIRILGWGV-ENGVPYWLAANS 299
Query: 296 WNTNWGDDGYFKIKRGTNECGIEDDVTAGLPST 328
WN +WGD+G+FKI RG N CGIE ++ AG+P T
Sbjct: 300 WNLDWGDNGFFKILRGENHCGIESEIVAGIPRT 332
>UniRef100_Q95PM1 Cathepsin B endopeptidase precursor [Schistosoma mansoni]
Length = 347
Score = 265 bits (678), Expect = 1e-69
Identities = 144/319 (45%), Positives = 184/319 (57%), Gaps = 32/319 (10%)
Query: 28 INENPEAGWEAAINPRFSNFTVGQFKRLLGVKQAPKKELLSTPVVTHPKSLKLPKEFDAR 87
IN W+AA RF TV +R+LG P E L T + T S +LPK FDAR
Sbjct: 45 INYEANTTWKAAPTTRFR--TVSDIRRMLGALPDPNGEQLET-LCTGYISDELPKSFDAR 101
Query: 88 TAWSQCSTIGKILGSNLILMLMMIQGHCGSCWAFGAVESLQDRFCIHFDMNIS--LSVND 145
W C +I +I Q CGSCWAFGAVE++ DR CI LS +
Sbjct: 102 VEWPHCPSISEIRD----------QSSCGSCWAFGAVEAMSDRICIKSKGKHKPFLSAEN 151
Query: 146 LLACCGFLCGAGCDGGTPIYAWRYLAHHGVVTEE-------CDPYFDQIGCSH------P 192
L++CC CG GC+GG P AW Y + G+VT + C PY + C H P
Sbjct: 152 LVSCCSS-CGMGCNGGFPHSAWLYWKNQGIVTGDLYNTTNGCQPY-EFPPCEHHVIGPLP 209
Query: 193 GCEPAYQTPKCVRKCVKGNQI-WKRSKHYSVKAYRVKSDPQDIMAEVYKNGPVEVAFTVF 251
C+ +TP C C G I +++ K Y K YR+ S+P+ IM E+ +NGPVEV F V+
Sbjct: 210 SCDGDVETPSCKTNCQPGYNIPYEKDKWYGEKVYRIHSNPEAIMLELMRNGPVEVDFEVY 269
Query: 252 EDFAHYKSGVYKHITGSALGGHAVKLIGWGTSDEGEDYWLLANQWNTNWGDDGYFKIKRG 311
DF +YKSGVY+H++G+ LGGHAV+L+GWG + YWL+AN WN++WGD GYFKI RG
Sbjct: 270 ADFPNYKSGVYQHVSGALLGGHAVRLLGWG-EENNVPYWLIANSWNSDWGDKGYFKIVRG 328
Query: 312 TNECGIEDDVTAGLPSTKN 330
NECGIE DV AG+P KN
Sbjct: 329 KNECGIESDVNAGIPKIKN 347
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.137 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 643,976,089
Number of Sequences: 2790947
Number of extensions: 28875419
Number of successful extensions: 61440
Number of sequences better than 10.0: 1469
Number of HSP's better than 10.0 without gapping: 1332
Number of HSP's successfully gapped in prelim test: 137
Number of HSP's that attempted gapping in prelim test: 57274
Number of HSP's gapped (non-prelim): 1826
length of query: 345
length of database: 848,049,833
effective HSP length: 128
effective length of query: 217
effective length of database: 490,808,617
effective search space: 106505469889
effective search space used: 106505469889
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 75 (33.5 bits)
Medicago: description of AC149038.5


