

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149038.3 + phase: 0
(1557 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
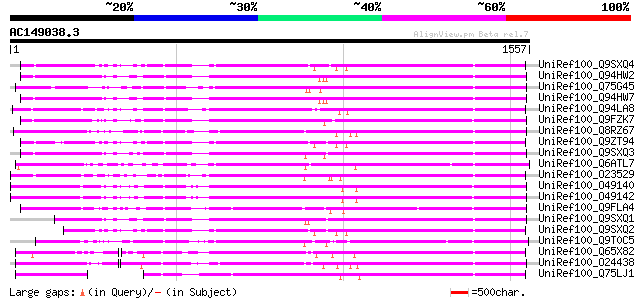
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SXQ4 Polyprotein [Arabidopsis thaliana] 934 0.0
UniRef100_Q94HW2 Polyprotein, putative [Arabidopsis thaliana] 930 0.0
UniRef100_Q75G45 Putative polyprotein [Oryza sativa] 929 0.0
UniRef100_Q94HW7 Polyprotein, putative [Arabidopsis thaliana] 928 0.0
UniRef100_Q94LA8 Polyprotein, putative [Arabidopsis thaliana] 923 0.0
UniRef100_Q9FZK7 F17L21.7 [Arabidopsis thaliana] 922 0.0
UniRef100_Q8RZ67 Putative rice retrotransposon retrofit gag/pol ... 921 0.0
UniRef100_Q9ZT94 Putative polyprotein of LTR transposon [Arabido... 918 0.0
UniRef100_Q9SXQ3 Polyprotein [Arabidopsis thaliana] 918 0.0
UniRef100_Q6ATL7 Putative polyprotein [Oryza sativa] 910 0.0
UniRef100_O23529 RETROTRANSPOSON like protein [Arabidopsis thali... 909 0.0
UniRef100_O49140 Polyprotein [Arabidopsis thaliana] 906 0.0
UniRef100_O49142 Polyprotein [Arabidopsis thaliana] 904 0.0
UniRef100_Q9FLA4 Polyprotein [Arabidopsis thaliana] 897 0.0
UniRef100_Q9SXQ1 Polyprotein [Arabidopsis thaliana] 884 0.0
UniRef100_Q9SXQ2 Polyprotein [Arabidopsis thaliana] 880 0.0
UniRef100_Q9T0C5 Retrotransposon like protein [Arabidopsis thali... 860 0.0
UniRef100_Q65X82 Putative polyprotein [Oryza sativa] 857 0.0
UniRef100_O24438 Retrofit [Oryza longistaminata] 835 0.0
UniRef100_Q75LJ1 Putative copia-like retrotransposon protein [Or... 832 0.0
>UniRef100_Q9SXQ4 Polyprotein [Arabidopsis thaliana]
Length = 1475
Score = 934 bits (2413), Expect = 0.0
Identities = 604/1577 (38%), Positives = 827/1577 (52%), Gaps = 199/1577 (12%)
Query: 31 KLDETNYLQWKQQVEGVLRGTKMVRHV--VSPQIPPVFLNDAAREAGTENPAYTEWEEQD 88
KL TNYL W +QV + G ++ + +P P DA NP YT W QD
Sbjct: 25 KLTSTNYLMWSRQVHALFDGYELAGFLDGSTPMPPATIGTDAVPRV---NPDYTRWRRQD 81
Query: 89 SLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRSELRTITKGTRS 148
L+ + IL IS S+ R + Q+W+ + QLR++L+ TKG ++
Sbjct: 82 KLIYSAILGAISMSVQPAVSRATTAAQIWETLRKIYANPSYGHVTQLRTQLKQWTKGAKT 141
Query: 149 IAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVATVNSQTEVISLDELES 208
I +++ + + L +G P+ H + +E VLE LP+++ P++ + ++ SL E+
Sbjct: 142 IDDYMQGFITRFDQLALLGKPMDHDEQVERVLENLPDDYKPVIDQIAAKDTPPSLTEIHE 201
Query: 209 QLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYPDQQFNISGNPTGNN 268
+L+ QE++ A V + +TH + +N +N+ +Y + NN
Sbjct: 202 RLINQESKLLALNSAEVVPITANVVTHRNTNTNRNQNNRGDNRNYNNN----------NN 251
Query: 269 SSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTGHDASICYHRLSVP 328
S + P+ +GSR + + GR CQIC GH A C
Sbjct: 252 RSNSWQPS---SSGSRSDNRQPKPYLGR-----------CQICSVQGHSAKRC------- 290
Query: 329 PQYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYGAPRAPFPPQFGNSR 388
PQ + Q + +PF P
Sbjct: 291 PQLHQF---------------------------------QSTTNQQQSTSPFTPW----- 312
Query: 389 PPAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLMDAASFSGSDQMYIGNGQGL 448
P+A + N ++N+ W DSGATHH+T D NNL ++G D + I +G +
Sbjct: 313 --QPRANLAVNSPYNANN----WLLDSGATHHITSDFNNLSFHQPYTGGDDVMIADGSTI 366
Query: 449 AINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICYVKS 508
I GS S + + +L LN +L+VP+I KNL+SV + C N V EF VK
Sbjct: 367 PITHTGSASLPTS---SRSLDLNKVLYVPNINKNLISVYRLCNTNRVSVEFFPASFQVKD 423
Query: 509 QDSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVATSSLSLNNCFSPSSLSLSRSQC 568
++ LL+G D+ LY++ +VS AS S AT
Sbjct: 424 LNTGVPLLQGKTKDE-LYEWPIASSQAVSMFASPCSKAT--------------------- 461
Query: 569 NNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQLP--NKSFTD 626
H+S WH+RLGHP ++ SV+ N LP N S
Sbjct: 462 ---------HSS---------------WHSRLGHPSLAILNSVIS--NHSLPVLNPSHKL 495
Query: 627 F-CSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSRYT 685
CS C + KSH++P +S +KP E I+ D+W + + S Y Y++I VD ++RYT
Sbjct: 496 LSCSDCFINKSHKVPFSNSTITSSKPLEYIYSDVWS-SPILSIDNYRYYVIFVDHFTRYT 554
Query: 686 WIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCP 745
W++PLK KS TF FK++VE ++ I ++ +D GGEF +L+ GI+H + P
Sbjct: 555 WLYPLKQKSQVKDTFIIFKSLVENRFQTRIGTLYSDNGGEFVVLRDYLSQHGISHFTSPP 614
Query: 746 HTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFF 805
HT NG ERKHRHIVE GLTLL++A +P YW +AF A YLINRLP+P+L +SPF
Sbjct: 615 HTPEHNGLSERKHRHIVEMGLTLLSHASVPKTYWPYAFSVAVYLINRLPTPLLQLQSPFQ 674
Query: 806 LLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLD-PTGRM 864
L Q P+Y+ LK FGC+C+P+ RPYN HKLE +SK+C F+GYS + Y CL PTGR+
Sbjct: 675 KLFGQPPNYEKLKVFGCACYPWLRPYNRHKLEDKSKQCAFMGYSLTQSAYLCLHIPTGRL 734
Query: 865 FISKDVIFNEYKFPYSE----LFTSGQ--SSSPPTTSSDHTPLPSFLFPLNNKQC----- 913
+ S+ V F+E FP+S + TS + S S P S HT LP+ L C
Sbjct: 735 YTSRHVQFDERCFPFSTTNFGVSTSQEQRSDSAPNWPS-HTTLPTTPLVLPAPPCLGPHL 793
Query: 914 -------------PTTQSSSTPTTTLHTASPHSSFPESNQSNHHHSIQDTHASSHSNHHN 960
TTQ SS+ + +SP SS P + N H + +SN
Sbjct: 794 DTSPRPPSLPSPLCTTQVSSSNLPSSSISSPSSSEPTAPSHNGPQPTAQPHQTQNSN--- 850
Query: 961 ISPGPIF-NPTPISTHPPSP--------SPSSHSHNTYHSISV----EPVTSQPSTQAEP 1007
S PI NP P S P SP SP S H S S+ P +S ST P
Sbjct: 851 -SNSPILNNPNPNSPSPNSPNQNSPLPQSPISSPHIPTPSTSISEPNSPSSSSTSTPPLP 909
Query: 1008 -----------HRIHPNNTHSMATRAKHGI---VQKRKHPTLLLTHIEPTGYRQAMKQPQ 1053
+ P NTHSMATRAK GI QK + T L + EP QAMK +
Sbjct: 910 PVLPAPPIIQVNAQAPVNTHSMATRAKDGIRKPNQKYSYATSLAANSEPRTAIQAMKDDR 969
Query: 1054 WLQAMQLEHEALMKNNTWTLVPLPADR-QAVGCKWVFRTKQNPDGSINKYKARLVAKGFH 1112
W QAM + A + N+TW LVP P VGC+W+F K N DGS+N+YKARLVAKG++
Sbjct: 970 WRQAMGSKINAQIGNHTWDLVPPPPPSVTIVGCRWIFTKKFNSDGSLNRYKARLVAKGYN 1029
Query: 1113 QMPGFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGF 1172
Q PG DY ETFSPV+K ++R VL +AV W I+QLDVNNAFL G L +EVYM+QPPGF
Sbjct: 1030 QRPGLDYAETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDEVYMSQPPGF 1089
Query: 1173 EAVD-PSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFM 1231
D P VC+L KA+YGLKQAPRAW+ L++ LL +GF +S D SLF+L + +M
Sbjct: 1090 VDKDRPDYVCRLRKAIYGLKQAPRAWYVELRTYLLTVGFVNSISDTSLFVLQRGRSIIYM 1149
Query: 1232 LVYVDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYI 1291
LVYVDDILITG+ L++ + L+ FS+K+ L YFLGIE G L LSQ++Y
Sbjct: 1150 LVYVDDILITGNDTVLLKHTLDALSQRFSVKEHEDLHYFLGIEAKRVPQG-LHLSQRRYT 1208
Query: 1292 QDLLVKANMANANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSV 1351
DLL + NM A +A+PMA+S KLT + + DPT +R IVG LQY+ TRP++SY+V
Sbjct: 1209 LDLLARTNMLTAKPVATPMATSPKLTLHSGTKLPDPTEYRGIVGSLQYLAFTRPDLSYAV 1268
Query: 1352 NKVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDD 1411
N++ Q++ P +D+W A+KR+LRYL GT HG+ + LSL A+ DADWA D DD
Sbjct: 1269 NRLSQYMHMPTDDNWNALKRVLRYLAGTPDHGIFLKKG---NTLSLHAYSDADWAGDTDD 1325
Query: 1412 RRSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVPTA 1471
ST+G + LG + ISW +KKQ V RSS EAEYRS+A S+E+ WI SLL EL + +
Sbjct: 1326 YVSTNGYIVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSELQWICSLLTELGIQLS 1385
Query: 1472 IPQ-IFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILT 1530
P I+CDN+ L NPV HSR KH+ LD F+R +V S L V H+ Q+AD LT
Sbjct: 1386 HPPVIYCDNVGATYLCANPVFHSRMKHIALDYHFIRNQVQSGALRVVHVSTHDQLADTLT 1445
Query: 1531 KPLSASRFLELRNKLRV 1547
KPLS F K+ V
Sbjct: 1446 KPLSRVAFQNFSRKIGV 1462
>UniRef100_Q94HW2 Polyprotein, putative [Arabidopsis thaliana]
Length = 1466
Score = 930 bits (2403), Expect = 0.0
Identities = 590/1568 (37%), Positives = 826/1568 (52%), Gaps = 181/1568 (11%)
Query: 31 KLDETNYLQWKQQVEGVLRGTKMVRHVV-SPQIPPVFLNDAAREAGTENPAYTEWEEQDS 89
KL TNYL W +QV + G ++ + S +PP + A A NP YT W+ QD
Sbjct: 25 KLTSTNYLMWSRQVHALFDGYELAGFLDGSTTMPPATIGTDA--APRVNPDYTRWKRQDK 82
Query: 90 LLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRSELRTITKGTRSI 149
L+ + +L IS S+ R + Q+W+ + QLR++L+ TKGT++I
Sbjct: 83 LIYSAVLGAISMSVQPAVSRATTAAQIWETLRKIYANPSYGHVTQLRTQLKQWTKGTKTI 142
Query: 150 AEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVATVNSQTEVISLDELESQ 209
+++ + + + L +G P+ H + +E VLE LPEE+ P++ + ++ +L E+ +
Sbjct: 143 DDYMQGLVTRFDQLALLGKPMDHDEQVERVLENLPEEYKPVIDQIAAKDTPPTLTEIHER 202
Query: 210 LLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYPDQQFNISGNPTGNNS 269
LL E++ A V + ++H + N +N + Y ++ N + P S
Sbjct: 203 LLNHESKILAVSSATVIPITANAVSHRNTTTTNNNNNGNRNNRYDNRNNNNNSKPW-QQS 261
Query: 270 SQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTGHDASICYHRLSVPP 329
S F+PN ++ + + G +CQIC GH A C
Sbjct: 262 STNFHPN-----NNQSKPYLG----------------KCQICGVQGHSAKRC-------- 292
Query: 330 QYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYGAPRAPFPPQFGNSRP 389
+ ++ V + P +P P+ P+
Sbjct: 293 ---------------------------SQLQHFLSSVNSQQPP--SPFTPWQPR------ 317
Query: 390 PAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLMDAASFSGSDQMYIGNGQGLA 449
N + S +N W DSGATHH+T D NNL ++G D + + +G +
Sbjct: 318 --------ANLALGSPYSSNNWLLDSGATHHITSDFNNLSLHQPYTGGDDVMVADGSTIP 369
Query: 450 INSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICYVKSQ 509
I+ GS S S+ P L L+N+L+VP+I KNL+SV + C N V EF VK
Sbjct: 370 ISHTGSTSLSTKSRP---LNLHNILYVPNIHKNLISVYRLCNANGVSVEFFPASFQVKDL 426
Query: 510 DSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVATSSLSLNNCFSPSSLSLSRSQCN 569
++ LL+G D+ LY++ VS AS SS AT
Sbjct: 427 NTGVPLLQGKTKDE-LYEWPIASSQPVSLFASPSSKAT---------------------- 463
Query: 570 NGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQLPNKSFTDF-C 628
H+S WH RLGHP ++ SV+ + + N S C
Sbjct: 464 --------HSS---------------WHARLGHPAPSILNSVISNYSLSVLNPSHKFLSC 500
Query: 629 SACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSRYTWIF 688
S C + KS+++P S +P E I+ D+W + + SH Y Y++I VD ++RYTW++
Sbjct: 501 SDCLINKSNKVPFSQSTINSTRPLEYIYSDVWS-SPILSHDNYRYYVIFVDHFTRYTWLY 559
Query: 689 PLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCPHTH 748
PLK KS TF FK ++E ++ I + +D GGEF ++ + GI+H + PHT
Sbjct: 560 PLKQKSQVKETFITFKNLLENRFQTRIGTFYSDNGGEFVALWEYFSQHGISHLTSPPHTP 619
Query: 749 HQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFFLLH 808
NG ERKHRHIVETGLTLL++A +P YW +AF A YLINRLP+P+L +SPF L
Sbjct: 620 EHNGLSERKHRHIVETGLTLLSHASIPKTYWPYAFAVAVYLINRLPTPLLQLESPFQKLF 679
Query: 809 LQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLD-PTGRMFIS 867
P+Y L+ FGC+C+P+ RPYN HKL+ +S++CVFLGYS + Y CL T R++IS
Sbjct: 680 GTSPNYDKLRVFGCACYPWLRPYNQHKLDDKSRQCVFLGYSLTQSAYLCLHLQTSRLYIS 739
Query: 868 KDVIFNEYKFPYSELFTS-----GQSSSPPTTSSDHTPLPSFLFPLNNKQCPTTQSSSTP 922
+ V F+E FP+S + Q S HT LP+ L C ++TP
Sbjct: 740 RHVRFDENCFPFSNYLATLSPVQEQRRESSCVWSPHTTLPTRTPVLPAPSCSDPHHAATP 799
Query: 923 TTTLHT-------------ASPHSSFPES------NQSNHHHSIQ------DTHAS---S 954
++ +S SSFP S Q+ + Q TH+S S
Sbjct: 800 PSSPSAPFRNSQVSSSNLDSSFSSSFPSSPEPTAPRQNGPQPTTQPTQTQTQTHSSQNTS 859
Query: 955 HSNHHNISPGPIFN--PTPISTHPPSPSP-----SSHSHNTYHSISVEPVTSQPSTQ-AE 1006
+N N SP + TP + SPSP SS + T SI + P P Q
Sbjct: 860 QNNPTNESPSQLAQSLSTPAQSSSSSPSPTTSASSSSTSPTPPSILIHP--PPPLAQIVN 917
Query: 1007 PHRIHPNNTHSMATRAKHGIVQKRKHPTL---LLTHIEPTGYRQAMKQPQWLQAMQLEHE 1063
+ P NTHSM TRAK GI++ +L L EP QA+K +W AM E
Sbjct: 918 NNNQAPLNTHSMGTRAKAGIIKPNPKYSLAVSLAAESEPRTAIQALKDERWRNAMGSEIN 977
Query: 1064 ALMKNNTWTLVPLPADR-QAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKET 1122
A + N+TW LVP P VGC+W+F K N DGS+N+YKARLVAKG++Q PG DY ET
Sbjct: 978 AQIGNHTWDLVPPPPSHVTIVGCRWIFTKKYNSDGSLNRYKARLVAKGYNQRPGLDYAET 1037
Query: 1123 FSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVD-PSLVC 1181
FSPV+K ++R VL +AV W I+QLDVNNAFL G L ++VYM+QPPGF D P+ VC
Sbjct: 1038 FSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDDVYMSQPPGFIDKDRPNYVC 1097
Query: 1182 KLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDILIT 1241
KL KALYGLKQAPRAW+ L++ LL +GF +S D SLF+L + +MLVYVDDILIT
Sbjct: 1098 KLRKALYGLKQAPRAWYVELRNYLLTIGFVNSVSDTSLFVLQRGKSIVYMLVYVDDILIT 1157
Query: 1242 GSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMA 1301
G+ +L+ + L+ FS+KD +L YFLGIE G L LSQ++YI DLL + NM
Sbjct: 1158 GNDPTLLHNTLDNLSQRFSVKDHEELHYFLGIEAKRVPTG-LHLSQRRYILDLLARTNMI 1216
Query: 1302 NANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAP 1361
A + +PMA S KL+ Y ++DPT +R IVG LQY+ TRP+ISY+VN++ QF+ P
Sbjct: 1217 TAKPVTTPMAPSPKLSLYSGTKLTDPTEYRGIVGSLQYLAFTRPDISYAVNRLSQFMHMP 1276
Query: 1362 LEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACIL 1421
E+H +A+KRILRYL GT +HG+ + LSL A+ DADWA D DD ST+G +
Sbjct: 1277 TEEHLQALKRILRYLAGTPNHGIFLKKG---NTLSLHAYSDADWAGDKDDYVSTNGYIVY 1333
Query: 1422 LGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVP-TAIPQIFCDNL 1480
LG + ISW +KKQ V RSS EAEYRS+A S+E+ WI SLL EL + T P I+CDN+
Sbjct: 1334 LGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSEMQWICSLLTELGIRLTRPPVIYCDNV 1393
Query: 1481 STVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASRFLE 1540
L NPV HSR KH+ +D F+R +V S L V H+ Q+AD LTKPLS + F
Sbjct: 1394 GATYLCANPVFHSRMKHIAIDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPLSRTAFQN 1453
Query: 1541 LRNKLRVS 1548
+K+ V+
Sbjct: 1454 FASKIGVT 1461
>UniRef100_Q75G45 Putative polyprotein [Oryza sativa]
Length = 1431
Score = 929 bits (2401), Expect = 0.0
Identities = 588/1581 (37%), Positives = 825/1581 (51%), Gaps = 209/1581 (13%)
Query: 18 AAASKN--FKQIISVKLDETNYLQWKQQVEGVLRGTKMVRHVVSPQIPP---VFLNDAAR 72
AAA N F IS KL + N+ W Q+ LRG ++ H+VS P + D +
Sbjct: 7 AAAISNPLFGLQISDKLTKQNHPLWAAQILTTLRGAQLEEHIVSTTAAPAAEIEKEDGDK 66
Query: 73 EAGTE----NPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQM 128
+ T+ NP Y W QD + +I S++S +L + R + Q W+ I +
Sbjct: 67 DKKTKIVIPNPEYKTWFVQDQQVLGFIFSSLSREVLQQVAGARTAAQAWNMIDDMF---- 122
Query: 129 RTRSRQLRSELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFN 188
+S+ TI KG S++E+IA++RS+++ + + G P+ +L+ ++ L EF+
Sbjct: 123 -----SCKSKAGTINKGPMSMSEYIAKMRSLADKMAATGKPLDEEELVAYIINGLDSEFD 177
Query: 189 PIVATVNSQTEV--ISLDELESQLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHN 246
V + + + +S+ + SQLL+ E R + ++A + T S N
Sbjct: 178 AAVEGLMATARIAPVSISHVYSQLLSYENRI-RIRQAYL--TTSANAA------------ 222
Query: 247 QPQTGSYPDQQFNISGNPTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNI 306
N GG G RG GNR GRGG FGRG
Sbjct: 223 -----------------------------NRGGGRGGRGSS-TGNRGGGRGG--FGRG-- 248
Query: 307 QCQICYKTGHDASICYHRLSVPPQYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGV 366
G+G G G + G G G T + V
Sbjct: 249 --------------------------GHGR-GAPSGASQGRGRGNDT----------RPV 271
Query: 367 GQRNPSYGAPRAPFPPQFGNSRPPAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDAN 426
Q G + ++ +S P + G +T + + WY D+GAT H+T +
Sbjct: 272 CQVCHKRGHVASDCWHRYDDSYVPDEKL---GGAATYAYGVDTNWYVDTGATDHITGQLD 328
Query: 427 NLMDAASFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSV 486
L + G+DQ++ +G+G +I +G +P P L L N+LHVP KNLVSV
Sbjct: 329 KLTTKERYKGTDQIHTASGEGTSIKHVGHAIVPTPSHP---LHLKNVLHVPEAAKNLVSV 385
Query: 487 SQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVA 546
+ DN F E H +K + + + +L+G GLY
Sbjct: 386 HKLVADNYAFLEIHGKYFLIKDKATRRTILEGPCRR-GLYPLP----------------- 427
Query: 547 TSSLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHE 606
+RS V TP WH RLGHP
Sbjct: 428 -----------------ARSSLRQAFVATPSFVR---------------WHGRLGHPSKP 455
Query: 607 VVRSVMKLCNQQLP---NKSFTDFCSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPA 663
+V + L +LP N C AC K H+LP S +V N P ELI D+WGPA
Sbjct: 456 IVLRI--LSQNKLPCLSNSVNESVCDACQQAKCHQLPFPRSTSVSNNPLELIHSDVWGPA 513
Query: 664 SVESHGGYSYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGG 723
S ES G Y++ +D YS++ WI+ LK KS F F+ +VE Q++ I S+QTD G
Sbjct: 514 S-ESVGAKKYYVSFIDDYSKFVWIYFLKHKSEVFQKFHEFQKLVERQFDRKILSMQTDWG 572
Query: 724 GEFRPFTQFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAF 783
GE+ F +GI+H ++CPHTH QNG+VERKHRHIVE GL+LLA+A +PL +WD AF
Sbjct: 573 GEYLKLHTFFNEIGISHHVSCPHTHQQNGAVERKHRHIVEVGLSLLAHASMPLKFWDEAF 632
Query: 784 LTATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKEC 843
L ATYLINRLPS ++ +P L Q PDYK L++FGC+C+P RPYN HKL+ RSK+C
Sbjct: 633 LAATYLINRLPSKVIKFDTPLERLFKQTPDYKSLRTFGCACWPNLRPYNTHKLQFRSKQC 692
Query: 844 VFLGYSPSHKGYKCLD-PTGRMFISKDVIFNEYKFPYSELFTSGQ-------SSSPPTTS 895
VFLG+S HKG+KCLD TGR+++S+DV F+E FP++ L + S PPT
Sbjct: 693 VFLGFSNIHKGFKCLDVSTGRVYVSRDVTFDEQVFPFANLHPNAGARLRAEISLLPPTLI 752
Query: 896 SD----------HTPLPSFLFP----LNNKQCPTTQSSSTPTT--TLHTA-------SPH 932
++ H L + P ++ + P +S P +H A S
Sbjct: 753 NEKTSDQGGEEHHDHLFNISMPNATDISCAESPRNVNSDIPGAFGRVHGANGDLAGESAS 812
Query: 933 SSFPESNQSNHHHSIQDTHASSHSNHHNISPGPIFNPTPISTH-PPSPSPSSHSHNTYHS 991
S Q S T S I P +P T PPSP+ + S +
Sbjct: 813 DSASVQAQLQRQASGSATQGESEQQRSGIQPARATSPAASPTAAPPSPARAVASSSGGAQ 872
Query: 992 ISVEPVTSQPSTQAEPHR-IHPNNTHSMATRAKHGIVQKRKHPTLLLTHIEPTGYRQAMK 1050
S +P +S PS A+ I P R + + + EP +A+
Sbjct: 873 QSHQPGSSAPSESAQLSEVIRPKTRLQSGIRKEKIYTDGTVRYSCFTSSGEPQTLHEALG 932
Query: 1051 QPQWLQAMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKG 1110
W +AM E++ALMKN TW LVP + + CKWV++ K+ DGS+++YKAR+VAKG
Sbjct: 933 DKNWKEAMDSEYQALMKNKTWHLVPSKKGQNIIDCKWVYKVKRKADGSLDRYKARVVAKG 992
Query: 1111 FHQMPGFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPP 1170
F Q G DY++TF+PVVK T+R++L++A++ W ++QLDV NAFL+G LEE+V+M QPP
Sbjct: 993 FKQRYGIDYEDTFNPVVKAATIRTILSIAISRGWTLRQLDVQNAFLHGVLEEDVFMRQPP 1052
Query: 1171 GFEAVDPSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTF 1230
G+E D VCKL+KALYGLKQAPRAW+ RL + L +LGF SSK D SLF + F
Sbjct: 1053 GYEQKD-GYVCKLDKALYGLKQAPRAWYSRLSTKLHELGFKSSKSDTSLFFYSKGDVAMF 1111
Query: 1231 MLVYVDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKY 1290
MLVYVDDI++ SS L+K LN EF+LKDLG+L YFLGIEV NG ++L+Q+KY
Sbjct: 1112 MLVYVDDIIVASSSIDATNALLKNLNQEFALKDLGRLHYFLGIEVKEVNNG-IVLTQEKY 1170
Query: 1291 IQDLLVKANMANANGIASPMASSTKLTKYGSNHV--SDPTFFRSIVGGLQYVTVTRPEIS 1348
D+L + NM++ + +P++ S KL+ + N D T +RS+VG LQY+T+TRP++S
Sbjct: 1171 AMDVLKRVNMSDCKAVNTPLSISEKLSAHEGNPFGPEDSTRYRSLVGALQYLTLTRPDLS 1230
Query: 1349 YSVNKVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASD 1408
+SVNKVCQ+L AP HW A KRILRYLK T+ GL I+ + L ++AF DADWA
Sbjct: 1231 FSVNKVCQYLHAPTTKHWAAAKRILRYLKHTVKLGLKISKS---NSLLVSAFSDADWAGC 1287
Query: 1409 PDDRRSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKV 1468
DDR ST G + +GPNL+SW A+KQ V+RSS EAEY++LA +AE++WIQ+LL EL +
Sbjct: 1288 LDDRHSTGGFAVFIGPNLVSWSARKQATVSRSSTEAEYKALANVTAEIMWIQTLLHELGI 1347
Query: 1469 PT-AIPQIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVAD 1527
I +++CDN+ + NPV H+RTKH+E+D FVRE+V K L V +I + QVAD
Sbjct: 1348 QAPKIAKVWCDNIGAKYMTANPVFHARTKHIEVDYHFVRERVARKFLQVEYISTKDQVAD 1407
Query: 1528 ILTKPLSASRFLELRNKLRVS 1548
TK LS + RN L ++
Sbjct: 1408 GFTKTLSVRQLEMFRNNLNLT 1428
>UniRef100_Q94HW7 Polyprotein, putative [Arabidopsis thaliana]
Length = 1466
Score = 928 bits (2399), Expect = 0.0
Identities = 589/1568 (37%), Positives = 825/1568 (52%), Gaps = 181/1568 (11%)
Query: 31 KLDETNYLQWKQQVEGVLRGTKMVRHVV-SPQIPPVFLNDAAREAGTENPAYTEWEEQDS 89
KL TNYL W +QV + G ++ + S +PP + A A NP YT W+ QD
Sbjct: 25 KLTSTNYLMWSRQVHALFDGYELAGFLDGSTTMPPATIGTDA--APRVNPDYTRWKRQDK 82
Query: 90 LLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRSELRTITKGTRSI 149
L+ + +L IS S+ R + Q+W+ + QLR++L+ TKGT++I
Sbjct: 83 LIYSAVLGAISMSVQPAVSRATTAAQIWETLRKIYANPSYGHVTQLRTQLKQWTKGTKTI 142
Query: 150 AEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVATVNSQTEVISLDELESQ 209
+++ + + + L +G P+ H + +E VLE LPEE+ P++ + ++ +L E+ +
Sbjct: 143 DDYMQGLVTRFDQLALLGKPMDHDEQVERVLENLPEEYKPVIDQIAAKDTPPTLTEIHER 202
Query: 210 LLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYPDQQFNISGNPTGNNS 269
LL E++ A V + ++H + N +N + Y ++ N + P S
Sbjct: 203 LLNHESKILAVSSATVIPITANAVSHRNTTTTNNNNNGNRNNRYDNRNNNNNSKPW-QQS 261
Query: 270 SQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTGHDASICYHRLSVPP 329
S F+PN ++ + + G +CQIC GH A C
Sbjct: 262 STNFHPN-----NNQSKPYLG----------------KCQICGVQGHSAKRC-------- 292
Query: 330 QYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYGAPRAPFPPQFGNSRP 389
+ ++ V + P +P P+ P+
Sbjct: 293 ---------------------------SQLQHFLSSVNSQQPP--SPFTPWQPR------ 317
Query: 390 PAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLMDAASFSGSDQMYIGNGQGLA 449
N + S +N W DSGATHH+T D NNL ++G D + + +G +
Sbjct: 318 --------ANLALGSPYSSNNWLLDSGATHHITSDFNNLSLHQPYTGGDDVMVADGSTIP 369
Query: 450 INSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICYVKSQ 509
I+ GS S S+ P L L+N+L+VP+I KNL+SV + C N V EF VK
Sbjct: 370 ISHTGSTSLSTKSRP---LNLHNILYVPNIHKNLISVYRLCNANGVSVEFFPASFQVKDL 426
Query: 510 DSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVATSSLSLNNCFSPSSLSLSRSQCN 569
++ LL+G D+ LY++ VS AS SS AT
Sbjct: 427 NTGVPLLQGKTKDE-LYEWPIASSQPVSLFASPSSKAT---------------------- 463
Query: 570 NGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQLPNKSFTDF-C 628
H+S WH RLGHP ++ SV+ + + N S C
Sbjct: 464 --------HSS---------------WHARLGHPAPSILNSVISNYSLSVLNPSHKFLSC 500
Query: 629 SACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSRYTWIF 688
S C + KS+++P S +P E I+ D+W + + SH Y Y++I VD ++RYTW++
Sbjct: 501 SDCLINKSNKVPFSQSTINSTRPLEYIYSDVWS-SPILSHDNYRYYVIFVDHFTRYTWLY 559
Query: 689 PLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCPHTH 748
PLK KS TF FK ++E ++ I + +D GGEF ++ + GI+H + PHT
Sbjct: 560 PLKQKSQVKETFITFKNLLENRFQTRIGTFYSDNGGEFVALWEYFSQHGISHLTSPPHTP 619
Query: 749 HQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFFLLH 808
NG ERKHRHIVETGLTLL++A +P YW +AF A YLINRLP+P+L +SPF L
Sbjct: 620 EHNGLSERKHRHIVETGLTLLSHASIPKTYWPYAFAVAVYLINRLPTPLLQLESPFQKLF 679
Query: 809 LQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLD-PTGRMFIS 867
P+Y L+ FGC+C+P+ RPYN HKL+ +S++CVFLGYS + Y CL T R++IS
Sbjct: 680 GTSPNYDKLRVFGCACYPWLRPYNQHKLDDKSRQCVFLGYSLTQSAYLCLHLQTSRLYIS 739
Query: 868 KDVIFNEYKFPYSELFTS-----GQSSSPPTTSSDHTPLPSFLFPLNNKQCPTTQSSSTP 922
+ V F+E FP+S + Q S HT LP+ L C ++TP
Sbjct: 740 RHVRFDENCFPFSNYLATLSPVQEQRRESSCVWSPHTTLPTRTPVLPAPSCSDPHHAATP 799
Query: 923 TTTLHT-------------ASPHSSFPES------NQSNHHHSIQ------DTHAS---S 954
++ +S SSFP S Q+ + Q TH+S S
Sbjct: 800 PSSPSAPFRNSQVSSSNLDSSFSSSFPSSPEPTAPRQNGPQPTTQPTQTQTQTHSSQNTS 859
Query: 955 HSNHHNISPGPIFN--PTPISTHPPSPSP-----SSHSHNTYHSISVEPVTSQPSTQ-AE 1006
+N N SP + TP + SPSP SS + T SI + P P Q
Sbjct: 860 QNNPTNESPSQLAQSLSTPAQSSSSSPSPTTSASSSSTSPTPPSILIHP--PPPLAQIVN 917
Query: 1007 PHRIHPNNTHSMATRAKHGIVQKRKHPTL---LLTHIEPTGYRQAMKQPQWLQAMQLEHE 1063
+ P NTHSM TRAK GI++ +L L EP QA+K +W AM E
Sbjct: 918 NNNQAPLNTHSMGTRAKAGIIKPNPKYSLAVSLAAESEPRTAIQALKDERWRNAMGSEIN 977
Query: 1064 ALMKNNTWTLVPLPADR-QAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKET 1122
A + N+TW LVP P VGC+W+F K N DGS+N+YKAR VAKG++Q PG DY ET
Sbjct: 978 AQIGNHTWDLVPPPPSHVTIVGCRWIFTKKYNSDGSLNRYKARFVAKGYNQRPGLDYAET 1037
Query: 1123 FSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVD-PSLVC 1181
FSPV+K ++R VL +AV W I+QLDVNNAFL G L ++VYM+QPPGF D P+ VC
Sbjct: 1038 FSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDDVYMSQPPGFIDKDRPNYVC 1097
Query: 1182 KLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDILIT 1241
KL KALYGLKQAPRAW+ L++ LL +GF +S D SLF+L + +MLVYVDDILIT
Sbjct: 1098 KLRKALYGLKQAPRAWYVELRNYLLTIGFVNSVSDTSLFVLQRGKSIVYMLVYVDDILIT 1157
Query: 1242 GSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMA 1301
G+ +L+ + L+ FS+KD +L YFLGIE G L LSQ++YI DLL + NM
Sbjct: 1158 GNDPTLLHNTLDNLSQRFSVKDHEELHYFLGIEAKRVPTG-LHLSQRRYILDLLARTNMI 1216
Query: 1302 NANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAP 1361
A + +PMA S KL+ Y ++DPT +R IVG LQY+ TRP+ISY+VN++ QF+ P
Sbjct: 1217 TAKPVTTPMAPSPKLSLYSGTKLTDPTEYRGIVGSLQYLAFTRPDISYAVNRLSQFMHMP 1276
Query: 1362 LEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACIL 1421
E+H +A+KRILRYL GT +HG+ + LSL A+ DADWA D DD ST+G +
Sbjct: 1277 TEEHLQALKRILRYLAGTPNHGIFLKKG---NTLSLHAYSDADWAGDKDDYVSTNGYIVY 1333
Query: 1422 LGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVP-TAIPQIFCDNL 1480
LG + ISW +KKQ V RSS EAEYRS+A S+E+ WI SLL EL + T P I+CDN+
Sbjct: 1334 LGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSEMQWICSLLTELGIRLTRPPVIYCDNV 1393
Query: 1481 STVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASRFLE 1540
L NPV HSR KH+ +D F+R +V S L V H+ Q+AD LTKPLS + F
Sbjct: 1394 GATYLCANPVFHSRMKHIAIDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPLSRTAFQN 1453
Query: 1541 LRNKLRVS 1548
+K+ V+
Sbjct: 1454 FASKIGVT 1461
>UniRef100_Q94LA8 Polyprotein, putative [Arabidopsis thaliana]
Length = 1459
Score = 923 bits (2386), Expect = 0.0
Identities = 579/1588 (36%), Positives = 825/1588 (51%), Gaps = 189/1588 (11%)
Query: 7 PVNNSETQRVTAAASKNFKQIISVKLDETNYLQWKQQVEGVLRGTKMVRHVV-SPQIPPV 65
P E +T N KL TNYL W Q+ +L G + ++ S IPP
Sbjct: 8 PATRDEAIVLTPQTLFNVNTSNVTKLTSTNYLMWSIQIHALLDGYDLAGYLDNSVVIPPE 67
Query: 66 FLNDAAREAGTENPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCY 125
+ NP++T W+ QD L+ + ++ IS ++ S R S Q+W ++N
Sbjct: 68 --TTTINSVVSANPSFTLWKRQDKLIFSALIGAISPAVQSLVSRATNSSQIWSTLNNTYA 125
Query: 126 TQMRTRSRQLRSELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPE 185
+QLR +++ +TKGT++I E++ + + L +G P+ H + +E +L+ LPE
Sbjct: 126 KPSYGHIKQLRQQIQRLTKGTKTIDEYVQSHTTRLDQLAILGKPMEHEEQVEHILKGLPE 185
Query: 186 EFNPIVATVNSQTEVISLDELESQLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGH 245
E+ +V + + ++ E+ +L+ E+ K V ++S ++ A ++N +
Sbjct: 186 EYKTVVDQIEGKDNTPTITEIHERLINHES---KLLSDEVPPSSSFPMS-ANAVQQRNFN 241
Query: 246 NQPQTGSYPDQQFNISGNPTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGN 305
N + ++ GN NN++ P+ ++G R F+ G+
Sbjct: 242 NNCNQNQHKNRY---QGNTHNNNTNTNSQPSTYNKSGQR-------TFKPYLGK------ 285
Query: 306 IQCQICYKTGHDASICYHRLSVPPQYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQG 365
CQIC GH A C PQ +
Sbjct: 286 --CQICSVQGHSARRC-------PQLQA-------------------------------- 304
Query: 366 VGQRNPSYGAPRAPFPPQFGNSRPPAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDA 425
+ P+ + +PF P P+A N + S N W DSGATHH+T D
Sbjct: 305 --MQLPASSSAHSPFTPW-------QPRA----NLAIGSPYAANPWLLDSGATHHITSDL 351
Query: 426 NNLMDAASFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVS 485
N L ++G + + I +G GL I GS S N L L+ +L+VP I KNL+S
Sbjct: 352 NALSLHQPYNGGEYVMIADGTGLTIKQTGSTFLPSQ---NRDLALHKVLYVPDIRKNLIS 408
Query: 486 VSQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSV 545
V + C N V EF VK ++ +LL+G DD LY++ P+
Sbjct: 409 VYRLCNTNQVSVEFFPASFQVKDLNTGTLLLQGRTKDD-LYEWPVTNPPA---------- 457
Query: 546 ATSSLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHH 605
T + TS S + +S WH+RLGHP
Sbjct: 458 -----------------------------TALFTSPSPKTTLSS------WHSRLGHPSA 482
Query: 606 EVVRSVMKLCNQQLPNKSFTDF-CSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPAS 664
++ +++ + + S CS C + KSH+LP +S + P E IF D+W +
Sbjct: 483 SILNTLLSKFSLPVSVASSNKTSCSDCLINKSHKLPFATSSIHSSSPLEYIFTDVW-TSP 541
Query: 665 VESHGGYSYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGG 724
+ SH Y Y+L+ VD Y+RYTW++PL+ KS TF FK +VE ++ I+++ +D GG
Sbjct: 542 IISHDNYKYYLVLVDHYTRYTWLYPLQQKSQVKATFIAFKALVENRFQAKIRTLYSDNGG 601
Query: 725 EFRPFTQFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFL 784
EF FL + GI+H + PHT NG ERKHRHIVETGLTLL A +P YW +AF
Sbjct: 602 EFIALRDFLVSNGISHLTSPPHTPEHNGLSERKHRHIVETGLTLLTQASVPREYWTYAFA 661
Query: 785 TATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECV 844
TA YLINR+P+P+L +SPF L P+Y+ L+ FGC CFP+ RPY +KLE RSK CV
Sbjct: 662 TAVYLINRMPTPVLCLQSPFQKLFGSSPNYQRLRVFGCLCFPWLRPYTRNKLEERSKRCV 721
Query: 845 FLGYSPSHKGYKCLD-PTGRMFISKDVIFNEYKFPYSELFTSGQSSS---PPTTSSDHTP 900
FLGYS + Y CLD R++ S+ V+F+E +P++ SS PP +SS +P
Sbjct: 722 FLGYSLTQTAYLCLDVDNNRLYTSRHVMFDESTYPFAASIREQSQSSLVTPPESSSSSSP 781
Query: 901 LPSFLFPLNNKQCPTTQSSSTPTTTLHTASPHSSFPESNQSNHHHSIQDTHASSHSNHHN 960
S FP C + S P +SP + P Q++ S + T + + S+H
Sbjct: 782 ANSG-FP-----CSVLRLQSPP-----ASSPETPSPPQQQNDSPVSPRQTGSPTPSHHSQ 830
Query: 961 I-----SPGP-IFNPTPISTHPPSPSPSSHSH----------------NTYHSISVEPVT 998
+ SP P + N P + H P P + S+ N SI PV
Sbjct: 831 VRDSTLSPSPSVSNSEPTAPHENGPEPEAQSNPNSPFIGPLPNPNPETNPSSSIEQRPVD 890
Query: 999 SQPSTQAEPHRI-----------HPNNTHSMATRAKHGIVQKRKHPTLLLT----HI-EP 1042
+T P++ P N H M TR+K+ I + + +L + H+ EP
Sbjct: 891 KSTTTALPPNQTTIAATSNSRSQPPKNNHQMKTRSKNNITKPKTKTSLTVALTQPHLSEP 950
Query: 1043 TGYRQAMKQPQWLQAMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKY 1102
QA+K +W AM E +A +N+TW LVP + VGC+WVF+ K P+G I+KY
Sbjct: 951 NTVTQALKDKKWRFAMSDEFDAQQRNHTWDLVPPNPTQHLVGCRWVFKLKYLPNGLIDKY 1010
Query: 1103 KARLVAKGFHQMPGFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEE 1162
KARLVAKGF+Q G DY ETFSPV+K T+R VL +AV W ++QLDVNNAFL G L E
Sbjct: 1011 KARLVAKGFNQQYGVDYAETFSPVIKATTIRVVLDVAVKKNWPLKQLDVNNAFLQGTLTE 1070
Query: 1163 EVYMTQPPGFEAVD-PSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFI 1221
EVYM QPPGF D PS VC+L KA+YGLKQAPRAW+ LK LL +GF +S D SLFI
Sbjct: 1071 EVYMAQPPGFVDKDRPSHVCRLRKAIYGLKQAPRAWYMELKQHLLNIGFVNSLADTSLFI 1130
Query: 1222 LHANQHSTFMLVYVDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENG 1281
++LVYVDDI++TGS + ++ L FS+KD L YFLGIE + G
Sbjct: 1131 YSHGTTLLYLLVYVDDIIVTGSDHKSVSAVLSSLAERFSIKDPTDLHYFLGIEATRTNTG 1190
Query: 1282 SLLLSQKKYIQDLLVKANMANANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVT 1341
L L Q+KY+ DLL K NM +A +A+P+ +S KLT +G ++D + +RS+VG LQY+
Sbjct: 1191 -LHLMQRKYMTDLLAKHNMLDAKPVATPLPTSPKLTLHGGTKLNDASEYRSVVGSLQYLA 1249
Query: 1342 VTRPEISYSVNKVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFC 1401
TRP+I+++VN++ QF+ P DHW+A KR+LRYL GT HG+ +N + P+ L AF
Sbjct: 1250 FTRPDIAFAVNRLSQFMHQPTSDHWQAAKRVLRYLAGTTTHGIFLNSS---SPIHLHAFS 1306
Query: 1402 DADWASDPDDRRSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQS 1461
DADWA D D ST+ I LG N ISW +KKQ V+RSS E+EYR++A A++E+ W+ S
Sbjct: 1307 DADWAGDSADYVSTNAYVIYLGRNPISWSSKKQRGVSRSSTESEYRAVANAASEIRWLCS 1366
Query: 1462 LLKEL--KVPTAIPQIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHI 1519
LL EL ++P P IFCDN+ + NPV HSR KH+ LD FVR + S+ L VSH+
Sbjct: 1367 LLTELHIRLPHG-PTIFCDNIGATYICANPVFHSRMKHIALDYHFVRGMIQSRALRVSHV 1425
Query: 1520 PAQYQVADILTKPLSASRFLELRNKLRV 1547
Q+AD LTK LS FL R+K+ V
Sbjct: 1426 STNDQLADALTKSLSRPHFLSARSKIGV 1453
>UniRef100_Q9FZK7 F17L21.7 [Arabidopsis thaliana]
Length = 1534
Score = 922 bits (2383), Expect = 0.0
Identities = 572/1546 (36%), Positives = 822/1546 (52%), Gaps = 170/1546 (10%)
Query: 31 KLDETNYLQWKQQVEGVLRGTKMVRHVV-SPQIPPVFLNDAAREAGTENPAYTEWEEQDS 89
KL +N+L W++QV+ +L G + ++ S +PP + A A T NPA+ W+ QD
Sbjct: 126 KLTSSNFLMWRRQVQALLNGYDLTGYIDGSIVVPPATIT--ANGAVTVNPAFKHWQRQDQ 183
Query: 90 LLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRSELRTITKGTRSI 149
L+ + +L IS S+ R S ++W ++ + + +QLR +++ K T+SI
Sbjct: 184 LIYSALLGAISISVQPILSRTTTSAEIWTKLMDTYAKPSWSHIQQLRQQIKQWKKDTKSI 243
Query: 150 AEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVATVNSQTEVISLDELESQ 209
EF + + L +G P+ + +E ++E L +++ ++ + + SL E+ +
Sbjct: 244 DEFFQGLVMRFDQLALLGKPMESEEQMEVIVEGLSDDYKQVIDQIQGREVPPSLTEIHEK 303
Query: 210 LLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYPDQQFNISGNPTGNNS 269
LL E + + +L + + N+ N +N Y Q N + N G NS
Sbjct: 304 LLNHEVKLQAAASSLPISANAASYRPPANNKHNNSNN------YRGQ--NRNNNNRGANS 355
Query: 270 SQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTGHDASICYHRLSVPP 329
Y P + RG++G +CQIC GH A C
Sbjct: 356 --YQQPR---NDQPSSRGYQG----------------KCQICGVFGHSARRC-------- 386
Query: 330 QYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYGAPRAPFPPQFGNSRP 389
S + M G A P P Q+ N+
Sbjct: 387 -----------------------------SQLQMSG---------AYSTPSPSQYPNATV 408
Query: 390 P-APQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLMDAASFSGSDQMYIGNGQGL 448
P P+A + ++ S+N W DSGATHH+T D NNL ++G +++ I +G L
Sbjct: 409 PWQPRA------NMAAMSYNP-WLLDSGATHHLTTDLNNLALHQPYNGGEEVTIADGSTL 461
Query: 449 AINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICYVKS 508
I GS + S+ + +L LNN+L+VP++ KNL+SV + C N V EF VK
Sbjct: 462 PITHTGSSTLSTQ---SRSLALNNILYVPNLHKNLISVYKLCNANKVSVEFFPAHFQVKD 518
Query: 509 QDSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVATSSLSLNNCFSPSSLSLSRSQC 568
+ LL+G D+ LY++ P +S AS + T PS
Sbjct: 519 LSTGARLLQGRTKDE-LYEWPVPSNTPISLFASPTPKTTL---------PS--------- 559
Query: 569 NNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQLPNKSFTDF- 627
WH+RLGHP V++S++ + + N S F
Sbjct: 560 ---------------------------WHSRLGHPSPPVLKSLVSQFSLPVSNSSQKHFP 592
Query: 628 CSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSRYTWI 687
CS C + KSH+LP S+ + P E ++ D+W + V S + Y+LI VD Y+RYTW+
Sbjct: 593 CSHCLINKSHKLPFYSNTIISYTPLEYVYSDVW-TSPVTSVDNFKYYLILVDHYTRYTWL 651
Query: 688 FPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCPHT 747
+PLK KS TF FK +VE ++ I+++ +D GGEF QFL T GI+H + PHT
Sbjct: 652 YPLKQKSQVRETFVAFKALVENRFQTKIRTLYSDNGGEFIALRQFLLTHGISHLTSLPHT 711
Query: 748 HHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFFLL 807
NG ERKHRHI+ETGLTLL A +P YW +AF TA YLINRLPS +LNN+SP+ L
Sbjct: 712 PEHNGIAERKHRHILETGLTLLTQASIPTSYWTYAFGTAVYLINRLPSSVLNNESPYSKL 771
Query: 808 HLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLD-PTGRMFI 866
P+Y L+ FGCSCFP+ RPY NHKLE RS+ CVFLGYS + Y CLD +GR++
Sbjct: 772 FKTSPNYLKLRVFGCSCFPWLRPYTNHKLERRSQPCVFLGYSLTQSAYLCLDRSSGRVYT 831
Query: 867 SKDVIFNEYKFPYSELFTSGQSSSPPTTSSDHTPLPSFLFPLNNKQCPTTQ--SSSTPTT 924
S+ V F E +FP+S T S+S P +S P P+ + P Q SS P +
Sbjct: 832 SRHVQFVEDQFPFSISDTHSVSNSSPEEASPSCHQPPSRIPIQSSSPPLVQAPSSLPPLS 891
Query: 925 TLHTASPHSSFPESNQS---------------NHHHSIQDTHASSHSNHHNISPGPIFNP 969
+ P++ S+ S N ++ I T +SS + +N +P
Sbjct: 892 SDSHRRPNAETSSSSSSTNNDVVVSKDNTQVDNRNNFIGPTSSSSAQSQNNSNPSSSIQ- 950
Query: 970 TPISTHPPSPSPSSHSHNTYHSISVEPVTSQPSTQAEPHRIHPNNTHSMATRAKHGIVQK 1029
+ + P+PSPS N+ S TS ST P P N H M TRAK+ I +
Sbjct: 951 ---TQNEPNPSPSPTPQNSSPESSPSSSTSATSTVPNPPPPPPTNNHPMRTRAKNHITKP 1007
Query: 1030 RKHPTLLLTHIE-----PTGYRQAMKQPQWLQAMQLEHEALMKNNTWTLVPLPADRQAVG 1084
+ +LL ++ P QA++ +W AM E A ++NNT+ LVP ++ +
Sbjct: 1008 KTKLSLLAKTVQTRPQIPNTVNQALRDEKWRNAMGEEINAQIRNNTFELVPPKPNQNVIS 1067
Query: 1085 CKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKETFSPVVKPVTVRSVLTLAVTNKW 1144
KW+F K P+G++++YKARLVA+GF Q G Y ETFSPVVK +T+R VL LAV+ W
Sbjct: 1068 TKWIFTLKYLPNGTLDRYKARLVARGFRQQYGLHYSETFSPVVKSLTIRLVLQLAVSRSW 1127
Query: 1145 CIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVD-PSLVCKLNKALYGLKQAPRAWFERLKS 1203
I+QLDVNNAFL G L +EVY+TQPPGF D P VC+L KALYGLKQAPRAW++ L++
Sbjct: 1128 TIKQLDVNNAFLQGTLTDEVYVTQPPGFIDPDRPHHVCRLKKALYGLKQAPRAWYQELRN 1187
Query: 1204 TLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDILITGSSASLIQQLVKKLNAEFSLKD 1263
+ LGF +S D S+F+ + + LVYVDDI++TGSS +L+ + L+ FSLKD
Sbjct: 1188 FVCSLGFTNSLADTSVFVYINDIQIVYCLVYVDDIIVTGSSDALVMAFITALSRRFSLKD 1247
Query: 1264 LGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMANANGIASPMASSTKLTKYGSNH 1323
L YFLGIE + G L L Q KY+ DLL + M +A +++PMA+ KL+ Y
Sbjct: 1248 PTDLVYFLGIEATRTSQG-LHLMQHKYVYDLLSRMKMLDAKPVSTPMATHPKLSLYSGIA 1306
Query: 1324 VSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAPLEDHWKAVKRILRYLKGTIHHG 1383
+ +P +R+++G LQY+ TRP+I+Y+VN++ QF+ P + HW+A KR+LRYL GT HG
Sbjct: 1307 LDEPGEYRTVIGSLQYLAFTRPDIAYAVNRLSQFMHRPTDIHWQAAKRVLRYLAGTATHG 1366
Query: 1384 LLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACILLGPNLISWWAKKQTLVARSSAE 1443
+L+ + PLSL AF DADWA D DD ST+ + LG I+W +KKQ VARSS E
Sbjct: 1367 ILLR---SNSPLSLHAFSDADWAGDNDDFVSTNAYIVYLGSTPIAWSSKKQKGVARSSTE 1423
Query: 1444 AEYRSLAQASAEVLWIQSLLKELKVP-TAIPQIFCDNLSTVSLAHNPVLHSRTKHMELDI 1502
AEYR++A ++E+ W+ SLL EL + +P I+CDN+ L+ NPV HSR KH+ LD
Sbjct: 1424 AEYRAVANTTSEIRWVCSLLTELGITLPKMPVIYCDNVGATYLSANPVFHSRMKHLALDY 1483
Query: 1503 FFVREKVISKDLIVSHIPAQYQVADILTKPLSASRFLELRNKLRVS 1548
F+R+ V + L VSHI Q+AD LTKPL FL+ +K+ VS
Sbjct: 1484 HFIRDNVSAGALRVSHISTHDQLADALTKPLPRQHFLQFSSKIGVS 1529
>UniRef100_Q8RZ67 Putative rice retrotransposon retrofit gag/pol polyprotein [Oryza
sativa]
Length = 1448
Score = 921 bits (2381), Expect = 0.0
Identities = 568/1587 (35%), Positives = 844/1587 (52%), Gaps = 191/1587 (12%)
Query: 10 NSETQRVTAAASKNFKQIISVKLDETNYLQWKQQVEGVLRGTKMVRHVVSPQIPP---VF 66
+S + TAAA+ +S KL + N+ WK QV + G +++ ++ P +
Sbjct: 3 SSSSSSGTAAANLLQGHSVSEKLGKANHALWKAQVSAAVHGARLLGYLNGDIKAPNAEIS 62
Query: 67 LNDAAREAGTENPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYT 126
+ + NPA+ +WE D L+ ++LS++S +L + + + + W I T
Sbjct: 63 VTIDGKTTTKPNPAFEDWEANDQLVLGYLLSSLSRDVLIQVATCKTAAEAWRNIEALYST 122
Query: 127 QMRTRSRQLRSELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEE 186
R R+ R L KGT IAE++A++R++ + + + G P+ L++ ++ L E+
Sbjct: 123 GTRARAVNTRLALTNTKKGTMKIAEYVAKMRALCDEMAAGGRPLDEEGLVQYIIAGLNED 182
Query: 187 FNPIVATVNSQTEVISLDELESQLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHN 246
F+PIV+ + ++++ I++ EL SQL+ E + ++ G A V N G G
Sbjct: 183 FSPIVSNLCNKSDPITVGELYSQLVNFETLLDLYRSTGQGGAAFV-----ANRGRGGG-- 235
Query: 247 QPQTGSYPDQQFNISGNPTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNI 306
G GNN N S G G G GGR RG
Sbjct: 236 --------------GGGGRGNN------------NNSDGGG-------GGGGRGAPRGR- 261
Query: 307 QCQICYKTGHDASICYHRLSVPPQYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGV 366
GG GG G+G TG G Q
Sbjct: 262 -------------------------------GG--GGQGRGGHGRGTG-GQDRRPTCQVC 287
Query: 367 GQRNPSYGAPRAPFPPQFGNSRPPAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDAN 426
+R + F + A + + +T+S + WY D+GAT H+T +
Sbjct: 288 FKRGHTAADCWYRFDEDY-----VADEKLVAA--ATNSYGIDTNWYIDTGATDHITGELE 340
Query: 427 NLMDAASFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSV 486
L ++G +Q++ +G G+ I+ IG + +P+ + LNN+L+VP KNL+S
Sbjct: 341 KLTTKEKYNGGEQIHTASGAGMDISHIGH---TIVHTPSRNIHLNNVLYVPQAKKNLISA 397
Query: 487 SQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVA 546
SQ DN+ F E HS +K Q + +LL+G GLY P S R+ + ++
Sbjct: 398 SQLAADNSAFLELHSKFFSIKDQVTRDVLLEGKCRH-GLY----PIPKSFGRSTNKQALG 452
Query: 547 TSSLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHE 606
+ LSL+ WH+RLGHP
Sbjct: 453 AAKLSLSR-----------------------------------------WHSRLGHPSLP 471
Query: 607 VVRSVMKL----CNQQLPNKSFTDFCSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGP 662
+V+ V+ C+ + N+S C+AC KSH+LP + S +V P EL+F D+WGP
Sbjct: 472 IVKQVISRNNLPCSVESVNQSV---CNACQEAKSHQLPYIRSTSVSQFPLELVFSDVWGP 528
Query: 663 ASVESHGGYSYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDG 722
A ES G Y++ +D +S++TWI+ LK KS F+ F+ +VE ++ I ++QTD
Sbjct: 529 AP-ESVGRNKYYVSFIDDFSKFTWIYLLKYKSEVFEKFKEFQALVERMFDRKIIAMQTDW 587
Query: 723 GGEFRPFTQFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHA 782
GGE++ F +GI H ++CPHTH QNGS ERKH+HI+E GL+LL+ A +PL +WD A
Sbjct: 588 GGEYQKLNSFFAQIGIDHHVSCPHTHQQNGSAERKHKHIIEVGLSLLSYASMPLKFWDEA 647
Query: 783 FLTATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKE 842
F+ ATYLINR+PS + N +P L Q PDY L+ F C+C+P PYN HKL+ RSK+
Sbjct: 648 FVAATYLINRIPSKTIQNSTPLEKLFNQKPDYSSLRVFSCTCWPHLHPYNTHKLQFRSKQ 707
Query: 843 CVFLGYSPSHKGYKCLD-PTGRMFISKDVIFNEYKFPYSELFTSGQSSSPPTTSSDHTPL 901
CVFLG+S HKG+K LD +GR++IS+DV+F+E FP+S L S++ S+ L
Sbjct: 708 CVFLGFSTHHKGFKFLDVSSGRVYISRDVVFDENVFPFSTL----HSNAGARLRSEILLL 763
Query: 902 PSFLFPLNNKQCPTTQ----SSSTPTTTLHTASPHSSFPESNQSNHH----HSIQDTHAS 953
PS L N T ++TP + + S + S H + Q+
Sbjct: 764 PSPLTNYNTVSAGGTHIVAPVANTPLPSDNLISNAADVTSGENSAAHEQEMENEQEIENV 823
Query: 954 SHSN--HHNISPGPIFN-PTPISTHPPS---------PSPSSHSHNTYHSISVEPVTSQP 1001
H N H + + GP+ + PT S+ P +S + ++
Sbjct: 824 MHGNDVHGDAASGPVLDQPTADSSTAPDQGANTSDAVSGAASDAGGDTATLGAGAANGAA 883
Query: 1002 STQAEPHRIHPNNTHSMA----------TRAKHGIVQKRKHPTLLLTH------IEPTGY 1045
+ E + P+ T ++A TR + GI +++ + + + EP
Sbjct: 884 AGGEESQPVPPDVTGTVAATVAPASRPHTRLRSGIRKEKVYTNGTVKYGCFSSTGEPQND 943
Query: 1046 RQAMKQPQWLQAMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKAR 1105
++A+ W AM+ E+ AL+KN+TW LVP + +GCKWV++ K+ DG++++YK R
Sbjct: 944 KEALGDKNWRDAMETEYNALIKNDTWHLVPYEKGQNIIGCKWVYKVKRKADGTLDRYKTR 1003
Query: 1106 LVAKGFHQMPGFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVY 1165
L+AKGF Q G DY++TFSPVVK T+R +L++AV+ W ++QLDV N FL+G+LEEEVY
Sbjct: 1004 LIAKGFKQRYGIDYEDTFSPVVKAATIRIILSIAVSRGWSLRQLDVQNDFLHGFLEEEVY 1063
Query: 1166 MTQPPGFEAVD-PSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHA 1224
M QPPGFE+ P VCKLNKALYGLKQAPRAW+ RL L +LGF +SK D SLF L+
Sbjct: 1064 MQQPPGFESSSKPDYVCKLNKALYGLKQAPRAWYSRLSKKLTELGFEASKADTSLFFLNK 1123
Query: 1225 NQHSTFMLVYVDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLL 1284
F+LVYVDDI++ S+ L+K LN EF+LKDLG L YFLGIEV NG ++
Sbjct: 1124 GGIIMFVLVYVDDIIVASSTEKATTALLKDLNKEFTLKDLGDLHYFLGIEVTKVSNG-VI 1182
Query: 1285 LSQKKYIQDLLVKANMANANGIASPMASSTKLTKYGSNHV--SDPTFFRSIVGGLQYVTV 1342
L+Q+KY D+L + NM+N +++P++ S KLT Y + + +D T +RSIVG LQY+T+
Sbjct: 1183 LTQEKYANDMLKRVNMSNCKPVSTPLSVSEKLTLYEGSPLGPNDATQYRSIVGALQYLTL 1242
Query: 1343 TRPEISYSVNKVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCD 1402
TRP+I+YSVNKVCQFL AP HW AVKRILRYL GL ++ + + + D
Sbjct: 1243 TRPDIAYSVNKVCQFLHAPTTSHWIAVKRILRYLNQCTSLGLHVHKS---ASTLVHGYSD 1299
Query: 1403 ADWASDPDDRRSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSL 1462
AD A DDR+ST G + LG NL+SW A+KQ V+RSS EAEY+++A +AE++W+Q++
Sbjct: 1300 ADGAGSIDDRKSTGGFAVFLGSNLVSWSARKQPTVSRSSTEAEYKAVANTTAELIWVQTV 1359
Query: 1463 LKELKVPT-AIPQIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPA 1521
LKEL + + +I+CDNL L+ NPV H++TKH+ +D FVRE+V K L + +P+
Sbjct: 1360 LKELGIESPKAAKIWCDNLGAKYLSANPVFHAKTKHIHVDYHFVRERVSQKLLEIDFVPS 1419
Query: 1522 QYQVADILTKPLSASRFLELRNKLRVS 1548
QVAD TK LSA ++ L ++
Sbjct: 1420 GDQVADGFTKALSARLLENFKHNLNLT 1446
>UniRef100_Q9ZT94 Putative polyprotein of LTR transposon [Arabidopsis thaliana]
Length = 1456
Score = 918 bits (2372), Expect = 0.0
Identities = 604/1574 (38%), Positives = 817/1574 (51%), Gaps = 212/1574 (13%)
Query: 31 KLDETNYLQWKQQVEGVLRGTKMVRHV--VSPQIPPVFLNDAAREAGTENPAYTEWEEQD 88
KL TNYL W +QV + G ++ + +P P DA NP YT W QD
Sbjct: 25 KLTSTNYLMWSRQVHALFDGYELAGFLDGSTPMPPATIGTDAVPRV---NPDYTRWRRQD 81
Query: 89 SLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRSELRTITKGTRS 148
L+ + IL IS S+ R + Q+W+ + QLR
Sbjct: 82 KLIYSAILGAISMSVQPAVSRATTAAQIWETLRKIYANPSYGHVTQLR------------ 129
Query: 149 IAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVATVNSQTEVISLDELES 208
FI R + L +G P+ H + +E VLE LP+++ P++ + ++ SL E+
Sbjct: 130 ---FITRF----DQLALLGKPMDHDEQVERVLENLPDDYKPVIDQIAAKDTPPSLTEIHE 182
Query: 209 QLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYPDQQFNISGNPTGNN 268
+L+ +E++ A V + +TH + +N +N+ +Y + NN
Sbjct: 183 RLINRESKLLALNSAEVVPITANVVTHRNTNTNRNQNNRGDNRNYNNN----------NN 232
Query: 269 SSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTGHDASICYHRLSVP 328
S + P+ +GSR + + GR CQIC GH A C
Sbjct: 233 RSNSWQPS---SSGSRSDNRQPKPYLGR-----------CQICSVQGHSAKRC------- 271
Query: 329 PQYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYGAPRAPFPPQFGNSR 388
PQ + Q + +PF P
Sbjct: 272 PQLHQF---------------------------------QSTTNQQQSTSPFTPW----- 293
Query: 389 PPAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLMDAASFSGSDQMYIGNGQGL 448
P+A + N ++N+ W DSGATHH+T D NNL ++G D + I +G +
Sbjct: 294 --QPRANLAVNSPYNANN----WLLDSGATHHITSDFNNLSFHQPYTGGDDVMIADGSTI 347
Query: 449 AINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICYVKS 508
I GS S + + +L LN +L+VP+I KNL+SV + C N V EF VK
Sbjct: 348 PITHTGSASLPTS---SRSLDLNKVLYVPNIHKNLISVYRLCNTNRVSVEFFPASFQVKD 404
Query: 509 QDSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVATSSLSLNNCFSPSSLSLSRSQC 568
++ LL+G D+ LY++ +VS AS S AT
Sbjct: 405 LNTGVPLLQGKTKDE-LYEWPIASSQAVSMFASPCSKAT--------------------- 442
Query: 569 NNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQLP--NKSFTD 626
H+S WH+RLGHP ++ SV+ N LP N S
Sbjct: 443 ---------HSS---------------WHSRLGHPSLAILNSVIS--NHSLPVLNPSHKL 476
Query: 627 F-CSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSRYT 685
CS C + KSH++P +S +KP E I+ D+W + + S Y Y++I VD ++RYT
Sbjct: 477 LSCSDCFINKSHKVPFSNSTITSSKPLEYIYSDVWS-SPILSIDNYRYYVIFVDHFTRYT 535
Query: 686 WIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCP 745
W++PLK KS TF FK++VE ++ I ++ +D GGEF +L+ GI+H + P
Sbjct: 536 WLYPLKQKSQVKDTFIIFKSLVENRFQTRIGTLYSDNGGEFVVLRDYLSQHGISHFTSPP 595
Query: 746 HTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFF 805
HT NG ERKHRHIVE GLTLL++A +P YW +AF A YLINRLP+P+L +SPF
Sbjct: 596 HTPEHNGLSERKHRHIVEMGLTLLSHASVPKTYWPYAFSVAVYLINRLPTPLLQLQSPFQ 655
Query: 806 LLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLD-PTGRM 864
L Q P+Y+ LK FGC+C+P+ RPYN HKLE +SK+C F+GYS + Y CL PTGR+
Sbjct: 656 KLFGQPPNYEKLKVFGCACYPWLRPYNRHKLEDKSKQCAFMGYSLTQSAYLCLHIPTGRL 715
Query: 865 FISKDVIFNEYKFPYSE----LFTSGQ--SSSPPTTSSDHTPLPSFLFPLNNKQC----- 913
+ S+ V F+E FP+S + TS + S S P S HT LP+ L C
Sbjct: 716 YTSRHVQFDERCFPFSTTNFGVSTSQEQRSDSAPNWPS-HTTLPTTPLVLPAPPCLGPHL 774
Query: 914 ---PTTQSSSTPTTTLHTAS---PHSSFPESNQSN----HHHSIQDTHASSHSNHHNISP 963
P SS +P T +S P SS + S H+ Q T A H ++ S
Sbjct: 775 DTSPRPPSSPSPLCTTQVSSSNLPSSSISSPSSSEPTAPSHNGPQPT-AQPHQTQNSNSN 833
Query: 964 GPIF-NPTPISTHPPSP--------SPSSHSHNTYHSISV----EPVTSQPSTQAEP--- 1007
PI NP P S P SP SP S H S S+ P +S ST P
Sbjct: 834 SPILNNPNPNSPSPNSPNQNSPLPQSPISSPHIPTPSTSISEPNSPSSSSTSTPPLPPVL 893
Query: 1008 --------HRIHPNNTHSMATRAKHGI---VQKRKHPTLLLTHIEPTGYRQAMKQPQWLQ 1056
+ P NTHSMATRAK GI QK + T L + EP QAMK +W Q
Sbjct: 894 PAPPIIQVNAQAPVNTHSMATRAKDGIRKPNQKYSYATSLAANSEPRTAIQAMKDDRWRQ 953
Query: 1057 AMQLEHEALMKNNTWTLVPLPADR-QAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMP 1115
AM E A + N+TW LVP P VGC+W+F K N DGS+N+YKARLVAKG++Q P
Sbjct: 954 AMGSEINAQIGNHTWDLVPPPPPSVTIVGCRWIFTKKFNSDGSLNRYKARLVAKGYNQRP 1013
Query: 1116 GFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAV 1175
G DY ETFSPV+K ++R VL +AV W I+QLDVNNAFL G L +EVYM+QPPGF
Sbjct: 1014 GLDYAETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDEVYMSQPPGFVDK 1073
Query: 1176 D-PSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVY 1234
D P VC+L KA+YGLKQAPRAW+ L++ LL +GF +S D SLF+L + +MLVY
Sbjct: 1074 DRPDYVCRLRKAIYGLKQAPRAWYVELRTYLLTVGFVNSISDTSLFVLQRGRSIIYMLVY 1133
Query: 1235 VDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDL 1294
VDDILITG+ L++ + L+ FS+K+ L YFLGIE G L LSQ++Y DL
Sbjct: 1134 VDDILITGNDTVLLKHTLDALSQRFSVKEHEDLHYFLGIEAKRVPQG-LHLSQRRYTLDL 1192
Query: 1295 LVKANMANANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKV 1354
L + NM A +A+PMA+S KLT + + DPT +R IVG LQY+ TRP++SY+VN++
Sbjct: 1193 LARTNMLTAKPVATPMATSPKLTLHSGTKLPDPTEYRGIVGSLQYLAFTRPDLSYAVNRL 1252
Query: 1355 CQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRS 1414
Q++ P +DHW A+KR+LRYL GT HG+ + LSL A+ DADWA D DD S
Sbjct: 1253 SQYMHMPTDDHWNALKRVLRYLAGTPDHGIFLKKG---NTLSLHAYSDADWAGDTDDYVS 1309
Query: 1415 TSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVPTAIPQ 1474
T+G + LG + ISW +KKQ V RSS EAEYRS+A S+E+ WI SLL EL + + P
Sbjct: 1310 TNGYIVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSELQWICSLLTELGIQLSHPP 1369
Query: 1475 -IFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPL 1533
I+CDN+ L NPV HSR KH+ LD F+R +V S L V H+ Q+AD LTKPL
Sbjct: 1370 VIYCDNVGATYLCANPVFHSRMKHIALDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPL 1429
Query: 1534 SASRFLELRNKLRV 1547
S F K+ V
Sbjct: 1430 SRVAFQNFSRKIGV 1443
>UniRef100_Q9SXQ3 Polyprotein [Arabidopsis thaliana]
Length = 1447
Score = 918 bits (2372), Expect = 0.0
Identities = 590/1571 (37%), Positives = 827/1571 (52%), Gaps = 187/1571 (11%)
Query: 31 KLDETNYLQWKQQVEGVLRGTKMVRHVV-SPQIPPVFLNDAAREAGTENPAYTEWEEQDS 89
KL TNYL W +QV + G ++ + S +PP + A A NP YT W+ QD
Sbjct: 6 KLTSTNYLMWSRQVHALFDGYELAGFLDGSTTMPPATIGTDA--APRVNPDYTRWKRQDK 63
Query: 90 LLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRSELRTITKGTRSI 149
L+ + +L IS S+ R + Q+W+ + Q R++L+ TKGT++I
Sbjct: 64 LIYSAVLGAISMSVQPAVSRATTAAQIWETLRKIYANPSYGHVTQFRTQLKQWTKGTKTI 123
Query: 150 AEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVATVNSQTEVISLDELESQ 209
+++ + + L +G P+ H + +E VLE LPEE+ P++ + ++ +L E+ +
Sbjct: 124 DDYMQGFVTHFDQLALLGKPMDHDEQVERVLENLPEEYKPVIDQIAAKDTPPTLTEIHER 183
Query: 210 LLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYPDQQFNISGNPTGNNS 269
LL QE++ A V + ++H + N +N +T Y ++ N + P +S
Sbjct: 184 LLNQESKILAVSSATVIPITANAVSHRNTTTTTNNNNGNRTNRYDNRNNNNNSKPWQQSS 243
Query: 270 SQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTGHDASICYHRLSVPP 329
S F PN ++ + + G +CQIC GH A C
Sbjct: 244 SN-FRPN-----NNQSKPYLG----------------KCQICGVQGHSAKRC-------- 273
Query: 330 QYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYGAPRAPFPPQFGNSRP 389
+ ++ V + P P F +P
Sbjct: 274 ---------------------------SQLQHFLSSVNSQQP---------PSPFTLWQP 297
Query: 390 PAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLMDAASFSGSDQMYIGNGQGLA 449
A N + S +N W DSGATHH+T D NNL ++G D + + +G +
Sbjct: 298 RA-------NLALGSPYSSNSWLLDSGATHHITSDFNNLSLHQPYTGGDDVMVVDGSTIP 350
Query: 450 INSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICYVKSQ 509
I+ GS S S+ P L L+N+L+VP+I KNL+SV + C N V EF VK
Sbjct: 351 ISHTGSTSLSTKSRP---LNLHNILYVPNIHKNLISVYRLCNANGVSVEFFLASFQVKDL 407
Query: 510 DSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVATSSLSLNNCFSPSSLSLSRSQCN 569
++ LL+G D+ LY++ VS AS SS AT
Sbjct: 408 NTGVPLLQGKTKDE-LYEWPIASSQPVSLFASPSSKAT---------------------- 444
Query: 570 NGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQLPNKSFTDF-C 628
H+S WH RLGHP ++ SV+ + + N S C
Sbjct: 445 --------HSS---------------WHARLGHPAPSILNSVISNYSLSVLNPSHKFLSC 481
Query: 629 SACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSRYTWIF 688
+ KS+++P S +P E I+ D+W + + SH Y Y++I VD ++RYTW++
Sbjct: 482 LDSLINKSNKVPFSQSTINSTRPLEYIYSDVWS-SPILSHDNYRYYVIFVDHFTRYTWLY 540
Query: 689 PLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCPHTH 748
PLK KS TF FK +VE ++ I + +D GGEF ++ + GI+H + PHT
Sbjct: 541 PLKQKSQVKETFITFKNLVENRFQTRIGTFYSDNGGEFVALREYFSQHGISHLTSPPHTP 600
Query: 749 HQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFFLLH 808
NG ERKHRHIVETGLTLL++A +P YW +AF A YLINRLP+P+L +SP L
Sbjct: 601 EHNGLSERKHRHIVETGLTLLSHASIPKTYWPYAFAVAVYLINRLPTPLLQLESPCQKLF 660
Query: 809 LQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLD-PTGRMFIS 867
P+Y L+ FGC+C+P+ RPYN HKL+ +S++CVFLGYS + Y CL T R++IS
Sbjct: 661 GTSPNYDKLRVFGCACYPWLRPYNQHKLDDKSRQCVFLGYSLTQSAYLCLHLQTSRIYIS 720
Query: 868 KDVIFNEYKFPYSELFTS--------GQSS---SPPTTSSDHTPL---PSFLFPLNNKQC 913
+ V F+E FP+S + +SS SP TT TP+ PS P +
Sbjct: 721 RHVRFDENCFPFSNYLATLSPVQEQRRESSCVWSPHTTLPTRTPVLPAPSCSDPHHAATP 780
Query: 914 PTTQSSSTPTTTLHTASPHSSFPESNQSN----------------------HHHSIQDTH 951
P++ S+ + + + +++ SSF S S+ HS Q+T
Sbjct: 781 PSSPSAPSRNSQVSSSNLDSSFSSSFPSSPEPTAPRQNGPQPTTQPTQTQTQTHSSQNT- 839
Query: 952 ASSHSNHHNISPGPIFN--PTPISTHPPSPSP-----SSHSHNTYHSISVEPVTSQPSTQ 1004
S +N N SP + TP + SPSP SS + T SI + P P Q
Sbjct: 840 --SQNNPTNESPSQLAQSLSTPAQSSSSSPSPTTSASSSSTSPTPPSILIHP--PPPLAQ 895
Query: 1005 -AEPHRIHPNNTHSMATRAKHGIVQKRKHPTL---LLTHIEPTGYRQAMKQPQWLQAMQL 1060
+ P NTHSM TRAK GI++ +L L EP QA+K +W AM
Sbjct: 896 IVNNNNQAPINTHSMGTRAKAGIIKPNLKYSLAVSLAAESEPRTAIQALKDERWRNAMGS 955
Query: 1061 EHEALMKNNTWTLVPLPADR-QAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDY 1119
E A + N+TW LVP P VGC+W+F K N DGS+N+YKARLVAKG++Q PG DY
Sbjct: 956 EINAQIGNHTWDLVPPPPSHVTIVGCRWIFTKKYNSDGSLNRYKARLVAKGYNQRPGLDY 1015
Query: 1120 KETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVD-PS 1178
ETFSPV+K ++R VL +AV W I+QLDVNNAFL G L ++VYM+QPPGF D P+
Sbjct: 1016 VETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDDVYMSQPPGFIDKDRPN 1075
Query: 1179 LVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDI 1238
VCKL KALYGLKQAPRAW+ L++ LL +GF +S D SLF+L + +MLVYVDDI
Sbjct: 1076 YVCKLRKALYGLKQAPRAWYVELRNYLLTIGFVNSVSDTSLFVLQRGKSIVYMLVYVDDI 1135
Query: 1239 LITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKA 1298
LITG+ +L+ + L+ FS+KD +L YFLGIE G L LSQ++YI DLL +
Sbjct: 1136 LITGNDPTLLHNTLDNLSQRFSVKDHEELHYFLGIEAKRVPTG-LHLSQRRYILDLLART 1194
Query: 1299 NMANANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFL 1358
NM A + +PMA S KL+ Y ++DPT +R IVG LQY+ TRP+ISY+VN++ QF+
Sbjct: 1195 NMITAKPVTTPMAPSPKLSLYSGTKLTDPTEYRGIVGSLQYLAFTRPDISYAVNRLSQFM 1254
Query: 1359 SAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGA 1418
P E+H +A+KRILRYL GT +HG+ + LSL A+ DADW D DD ST+G
Sbjct: 1255 HMPTEEHLQALKRILRYLAGTPNHGIFLKKG---NTLSLHAYSDADWTGDKDDYVSTNGY 1311
Query: 1419 CILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVP-TAIPQIFC 1477
+ LG + ISW +KKQ V RSS EAEYRS+A S+E+ WI SLL EL + T P I+C
Sbjct: 1312 IVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSEMQWICSLLTELGIRLTRPPVIYC 1371
Query: 1478 DNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASR 1537
DN+ L NPV HSR KH+ +D F+R +V S L V H+ Q+AD LTKPLS +
Sbjct: 1372 DNVGATYLCANPVFHSRMKHIAIDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPLSRTA 1431
Query: 1538 FLELRNKLRVS 1548
F +K+ V+
Sbjct: 1432 FQNFASKIGVT 1442
>UniRef100_Q6ATL7 Putative polyprotein [Oryza sativa]
Length = 1437
Score = 910 bits (2353), Expect = 0.0
Identities = 575/1587 (36%), Positives = 839/1587 (52%), Gaps = 198/1587 (12%)
Query: 18 AAASKN------FKQIISVKLDETNYLQWKQQVEGVLRGTKMVRHVVSPQIPPVFL---N 68
A++SKN Q +S KL ++N+ WK Q+ +RG ++ H+ PP +
Sbjct: 2 ASSSKNNTGNPLVGQPVSEKLGKSNHAVWKAQILATIRGARLEGHLTGDDQPPAPILRRK 61
Query: 69 DAAREAGTENPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQM 128
+ +E NP Y EW D + ++LS+++ LL + R + W I +
Sbjct: 62 EGEKEVVVSNPEYEEWVATDQQVLAYLLSSMTKDLLVQVATCRTAASAWSMIQGMFGSMT 121
Query: 129 RTRSRQLRSELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFN 188
R R+ R L T+ KG +I ++ ++R++++ LM++G PV +LI + L +EF
Sbjct: 122 RARTINTRLSLSTLQKGDMNITTYVGKMRALADDLMAVGKPVDDDELIGYIFAGLDDEFE 181
Query: 189 PIVATVNSQTEVISLDELESQLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQP 248
P+++T+ + + +++ E +QL++ E R + G+ +SVN QP
Sbjct: 182 PVISTIVGRPDPVTIGETYAQLISFEQRLAHRRS---GDQSSVN-------SASRSRGQP 231
Query: 249 QTGSYPDQQFNISGNPTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGN-IQ 307
Q G G+ +G +S++ G N RGRG RGN GR G N +
Sbjct: 232 QRG----------GSRSGGDSNRGRGAPSNGAN--RGRG-RGNPSGGRANVGGGTDNRPK 278
Query: 308 CQICYKTGHDASICYHRLSVPPQYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVG 367
CQ+CYK GH C++R Y+ F G A +G +N W G
Sbjct: 279 CQLCYKRGHTVCDCWYR------YDENFVPDERFAGT-------AVSYGVDTN-WYLDTG 324
Query: 368 QRNPSYGAPRAPFPPQFGNSRPPAPQAYITG--NESTSSNSFNNGWYPDSGATHHVTPDA 425
+ ++TG ++ T + ++
Sbjct: 325 ATD------------------------HVTGELDKLTVRDKYH----------------G 344
Query: 426 NNLMDAASFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVS 485
N+ + AS +G + +IGN S +P+ L L ++L+VP KNLVS
Sbjct: 345 NDQVHTASGAGMEISHIGN--------------SVVKTPSRNLHLKDVLYVPKANKNLVS 390
Query: 486 VSQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSV 545
+ DN F E + ++K + LL+G GLY P SS
Sbjct: 391 AYKLTSDNLAFIELYRKFFFIKDLAMRRTLLRGRC-HKGLYALPSP-----------SSH 438
Query: 546 ATSSLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHH 605
+ PS ++ WH+RLGHP +
Sbjct: 439 HHQVKQVYGVTKPS---------------------------------FERWHSRLGHPSY 465
Query: 606 EVVRSVMKLCNQQLPNKSFTD---FCSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGP 662
VV V+K +Q LP ++ C AC KSH+L S + P EL+F D+WGP
Sbjct: 466 TVVEKVIK--SQNLPCLDVSEQVSVCDACQKAKSHQLSFPKSTSESKYPLELVFSDVWGP 523
Query: 663 ASVESHGGYSYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDG 722
A +S G Y++ +D YS++TWI+ LK KS F F+++VE +N I ++QTD
Sbjct: 524 AP-QSVGNNKYYVSFIDDYSKFTWIYLLKYKSEVFDKFHEFQSLVERLFNRKIVAMQTDW 582
Query: 723 GGEFRPFTQFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHA 782
GGE++ F +GITH ++CPHTH QNGS ERKHRHIVE GL LLA + +PL +W A
Sbjct: 583 GGEYQKLHSFFNKVGITHHVSCPHTHQQNGSAERKHRHIVEVGLALLAYSSMPLKFWGEA 642
Query: 783 FLTATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKE 842
FL+A YLINR PS +L++ SP L PDY L+ FGC+C+P RPYN HKL+ RS
Sbjct: 643 FLSAVYLINRTPSRVLHDVSPLERLLGHKPDYNALRVFGCACWPNLRPYNKHKLQFRSTT 702
Query: 843 CVFLGYSPSHKGYKCLDP-TGRMFISKDVIFNEYKFPYSELFTS------GQSSSPPTTS 895
C FLGYS HKG+KCLDP TGR++IS+DV+F+E +FP+++L + + + P +
Sbjct: 703 CTFLGYSTLHKGFKCLDPSTGRVYISRDVVFDETQFPFTKLHPNVGAKLRAEIALVPELA 762
Query: 896 SDHTPLPSFLFPLNNKQCPTTQSSSTPTTTLHTASPHSSFPESNQSNHHHSIQDTHASSH 955
+ LP L +++ T ++++ + S + + PE+ ++ +
Sbjct: 763 AS---LPRGLQQISS-VINTPENANVSNENMQQDSTYDNEPETETDGAPDTVSANAPAES 818
Query: 956 SNHHNIS-PGPIFNPTPISTHPPSPSP--SSHSHNTYHSISVEP--VTSQPSTQA----E 1006
S I+ P F + +T P+ +P S+ + S S P TSQ T + +
Sbjct: 819 SGSPPINEPASPFGESDSATASPASAPVNSAPHPDAAASGSSAPRGSTSQGGTPSVAIDD 878
Query: 1007 PH---RIHPNNTHSMATRAKHGIVQKRKHPT------LLLTHIEPTGYRQAMKQPQWLQA 1057
PH + TR + GI +++ + +L + EP + A++ W A
Sbjct: 879 PHPATTVTGQEAQRPRTRLQSGIRKEKVYTDGTVKWGMLTSTGEPENLQDALQNNNWKCA 938
Query: 1058 MQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGF 1117
M E+ AL+KNNTW LVP R + CKWV++ K+ DGS+++YKARLVAKGF Q G
Sbjct: 939 MDAEYMALIKNNTWHLVPPQQGRNVIDCKWVYKIKRKQDGSLDRYKARLVAKGFKQRYGI 998
Query: 1118 DYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFE-AVD 1176
DY++TFSPVVK T+R +L++AV+ WC++QLDV NAFL+G LEEEVYM QPPG+E
Sbjct: 999 DYEDTFSPVVKAATIRIILSIAVSRGWCLRQLDVQNAFLHGVLEEEVYMKQPPGYENPST 1058
Query: 1177 PSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVD 1236
P VCKL+KALYGLKQAPRAW+ RL L LGF SK D SLF + + F+L+YVD
Sbjct: 1059 PDYVCKLDKALYGLKQAPRAWYSRLSGKLHDLGFKGSKADTSLFFYNKGSLTIFLLIYVD 1118
Query: 1237 DILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLV 1296
DI++ S + L++ L EF+LKDLG L YFLGIEV G +L+SQ+KY DLL
Sbjct: 1119 DIIVVSSRKEAVSALLQDLQKEFALKDLGDLHYFLGIEV-TKIPGGILMSQEKYASDLLK 1177
Query: 1297 KANMANANGIASPMASSTKL-----TKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSV 1351
+ NM++ +A+P+++S KL T G N D T +RSIVG LQY+T+TR +I++SV
Sbjct: 1178 RVNMSDCKSVATPLSASEKLIAGKGTILGPN---DATQYRSIVGALQYLTLTRLDIAFSV 1234
Query: 1352 NKVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDD 1411
NKVCQFL P +HW AVKRILRY+K GL I + + ++ + DADWA DD
Sbjct: 1235 NKVCQFLHNPTTEHWAAVKRILRYIKQCTGLGLRICKS---SSMIVSGYSDADWAGCLDD 1291
Query: 1412 RRSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELK-VPT 1470
RRST G + LG NL+SW AKKQ V+RSS EAEY++LA A+AE++W+Q+LL+EL V
Sbjct: 1292 RRSTGGFAVYLGDNLVSWNAKKQATVSRSSTEAEYKALANATAEIMWVQTLLQELNIVSP 1351
Query: 1471 AIPQIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILT 1530
A+ Q++CDN+ L+ NPV H+RTKH+E+D FVRE+V K L V ++ QVAD T
Sbjct: 1352 AMAQLWCDNMGAKYLSFNPVFHARTKHIEVDYHFVRERVARKLLQVDYVSTNDQVADGFT 1411
Query: 1531 KPLSASRFLELRNKLRVSDPMSLRGDC 1557
K L + + L + + + G C
Sbjct: 1412 KALPVKQLENFKYNLNLG-KVVIEGGC 1437
>UniRef100_O23529 RETROTRANSPOSON like protein [Arabidopsis thaliana]
Length = 1474
Score = 909 bits (2350), Expect = 0.0
Identities = 594/1607 (36%), Positives = 826/1607 (50%), Gaps = 203/1607 (12%)
Query: 2 SSPSSPVNNSETQRVTAAASKNFKQIISVKLDETNYLQWKQQVEGVLRGTKMVRHV---V 58
S+ P E T N KL NYL W Q+ +L G ++ H+ +
Sbjct: 4 SANGLPATTDEAIVFTPQTIFNINTSNVTKLTSNNYLMWSLQIHALLDGYELAGHLDGSI 63
Query: 59 SPQIPPVFLNDAAREAGTENPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWD 118
P + N+ + NP YT W+ QD L+ + ++ IS + R + Q+W
Sbjct: 64 ETPAPTLTTNNVV----SANPQYTLWKRQDRLIFSALIGAISPPVQPLVSRATKASQIWK 119
Query: 119 EIHNYCYTQMRTRSRQLRSELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIET 178
+ N +QLR++++ + KGT++I E++ ++ + L +G P+ H + +E
Sbjct: 120 TLTNTYAKSSYDHIKQLRTQIKQLKKGTKTIDEYVLSHTTLLDQLAILGKPMEHEEQVER 179
Query: 179 VLEALPEEFNPIVATVNSQTEVISLDELESQLLTQEARNEKFKKALVGETASVNLTHAEN 238
+LE LPE++ +V + + S+ E+ +L+ EA K +S +L + N
Sbjct: 180 ILEGLPEDYKTVVDQIEGKDNTPSITEIHERLINHEA-----KLLSTAALSSSSLPMSAN 234
Query: 239 SGEKNGHNQPQTGSYPDQQFNISGNPT-GNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRG 297
++ HN + N + N T GN + + P+ ++G R F+
Sbjct: 235 VAQQRHHNNNRNN-------NQNKNRTQGNTYTNNWQPSANNKSGQRP-------FKPYL 280
Query: 298 GRNFGRGNIQCQICYKTGHDASICYHRLSVPPQYEGYGSLGGNFGGNLGSGYGPATGFGT 357
G+ CQIC GH A C ++ P + S + P
Sbjct: 281 GK--------CQICNVQGHSARRCPQLQAMQPS-----------SSSSASTFTP------ 315
Query: 358 HSNVWMQGVGQRNPSYGAPRAPFPPQFGNSRPPAPQAYITGNESTSSNSFNNGWYPDSGA 417
W + N + GAP N W DSGA
Sbjct: 316 ----WQP---RANLAMGAPYTA-----------------------------NNWLLDSGA 339
Query: 418 THHVTPDANNLMDAASFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTT--LTLNNLLH 475
THH+T D N L ++G D M I +G L I GS F P+ LTLN +L+
Sbjct: 340 THHITSDLNALALHQPYNGDDVM-IADGTSLKITKTGST-----FLPSNARDLTLNKVLY 393
Query: 476 VPSITKNLVSVSQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQFDQPYVPS 535
VP I KNLVSV + C N V EF VK ++ +LL+G D+ LY++ P
Sbjct: 394 VPDIQKNLVSVYRLCNTNQVSVEFFPASFQVKDLNTGTLLLQGRTKDE-LYEW-----PV 447
Query: 536 VSRTASSSSVATSSLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKV 595
+ A T + T+ S + +S
Sbjct: 448 TNPKA----------------------------------TALFTTPSPKTTLSS------ 467
Query: 596 WHNRLGHPHHEVVRSVMKLCNQQLPNKSFTDF-CSACCLGKSHRLPSVSSKTVYNKPFEL 654
WH+RLGHP ++ +++ + + + CS C + KSH+LP S P E
Sbjct: 468 WHSRLGHPSSSILNTLISKFSLPVSVSASNKLACSDCFINKSHKLPFSISSIKSTSPLEY 527
Query: 655 IFCDLWGPASVESHGGYSYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLP 714
IF D+W + + S Y Y+L+ VD ++RYTW++PL+ KS TF FK +VE ++
Sbjct: 528 IFSDVW-MSPILSPDNYKYYLVLVDHHTRYTWLYPLQQKSQVKSTFIAFKALVENRFQAK 586
Query: 715 IKSVQTDGGGEFRPFTQFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKL 774
I+++ +D GGEF +FL + GI+H + PHT NG ERKHRHIVETGLTLL A +
Sbjct: 587 IRTLYSDNGGEFIALREFLVSNGISHLTSPPHTPEHNGLSERKHRHIVETGLTLLTQASV 646
Query: 775 PLHYWDHAFLTATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNH 834
P YW +AF A YLINR+P+P+L+ +SPF L P+Y+ L+ FGC CFP+ RPY ++
Sbjct: 647 PREYWPYAFAAAVYLINRMPTPVLSMESPFQKLFGSKPNYERLRVFGCLCFPWLRPYTHN 706
Query: 835 KLELRSKECVFLGYSPSHKGYKCLD-PTGRMFISKDVIFNEYKFPYSEL--------FTS 885
KLE RS+ CVFLGYS + Y C D R++ S+ V+F+E FP+S L T
Sbjct: 707 KLEERSRRCVFLGYSLTQTAYLCFDVEHKRLYTSRHVVFDEASFPFSNLTSQNSLPTVTF 766
Query: 886 GQSSSPPTTS--SDHTPLPSFLFP----LNNKQCPTT----QSSSTPTTTLHTASPHSSF 935
QSSSP T S + LPS L L+ +Q P T SS PTT+ SPH S
Sbjct: 767 EQSSSPLVTPILSSSSVLPSCLSSPCTVLHQQQPPVTTPNSPHSSQPTTSPAPLSPHRST 826
Query: 936 PESNQSNHHHSIQDTHASSHS--------NHHNISP-------GPIFNPTPISTHPPSPS 980
Q S +SS S N + P GP+ NPT + P P+
Sbjct: 827 TMDFQVPQVRSSSPLLSSSSSLNSEPTAPNENGPEPEAQSPPIGPLSNPTHEAFIGPLPN 886
Query: 981 PSSHSHNTY------HSISVEP--VTSQPSTQAEPHRIH-----PNNTHSMATRAKHGIV 1027
P+ + N H V+P T+ P+ H N H+M TRAK+ I
Sbjct: 887 PNRNPTNEIEPTPAPHPKPVKPTTTTTTPNRTTVSDASHQPTAPQQNQHNMKTRAKNNIK 946
Query: 1028 QKRKHPTLLLT-----HIEPTGYRQAMKQPQWLQAMQLEHEALMKNNTWTLVPLPADRQA 1082
+ +L T EPT QA+K +W AM E +A +N+TW LVP +
Sbjct: 947 KPNTKFSLTATLPNRSPSEPTNVTQALKDKKWRFAMSDEFDAQQRNHTWDLVP-HESQLL 1005
Query: 1083 VGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKETFSPVVKPVTVRSVLTLAVTN 1142
VGCKWVF+ K P+G+I+KYKARLVAKGF+Q G DY ETFSPV+K T+R VL +AV
Sbjct: 1006 VGCKWVFKLKYLPNGAIDKYKARLVAKGFNQQYGVDYAETFSPVIKSTTIRLVLDVAVKK 1065
Query: 1143 KWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVD-PSLVCKLNKALYGLKQAPRAWFERL 1201
W I+QLDVNNAFL G L EEVYM QPPGF D P+ VC+L KA+YGLKQAPRAW+ L
Sbjct: 1066 DWEIKQLDVNNAFLQGTLTEEVYMAQPPGFIDKDRPTHVCRLRKAIYGLKQAPRAWYMEL 1125
Query: 1202 KSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDILITGSSASLIQQLVKKLNAEFSL 1261
K L +GF +S D SLFI ++LVYVDDI++TGS S I ++ L FS+
Sbjct: 1126 KQHLFNIGFVNSLSDASLFIYCHGTTFVYVLVYVDDIIVTGSDKSSIDAVLTSLAERFSI 1185
Query: 1262 KDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMANANGIASPMASSTKLTKYGS 1321
KD L YFLGIE ++ G L L Q+KYI+DLL K NMA+A + +P+ +S KLT +G
Sbjct: 1186 KDPTDLHYFLGIEATRTKQG-LHLMQRKYIKDLLAKHNMADAKPVLTPLPTSPKLTLHGG 1244
Query: 1322 NHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAPLEDHWKAVKRILRYLKGTIH 1381
++D + +RS+VG LQY+ TRP+I+Y+VN++ Q + P EDHW+A KR+LRYL GT
Sbjct: 1245 TKLNDASEYRSVVGSLQYLAFTRPDIAYAVNRLSQLMPQPTEDHWQAAKRVLRYLAGTST 1304
Query: 1382 HGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACILLGPNLISWWAKKQTLVARSS 1441
HG+ ++ PL+L AF DADWA D DD ST+ I LG N ISW +KKQ VARSS
Sbjct: 1305 HGIFLDTT---SPLNLHAFSDADWAGDSDDYVSTNAYVIYLGKNPISWSSKKQRGVARSS 1361
Query: 1442 AEAEYRSLAQASAEVLWIQSLLKELKVPTAI-PQIFCDNLSTVSLAHNPVLHSRTKHMEL 1500
E+EYR++A A++EV W+ SLL +L + I P IFCDN+ L NPV HSR KH+ +
Sbjct: 1362 TESEYRAVANAASEVKWLCSLLSKLHIRLPIRPSIFCDNIGATYLCANPVFHSRMKHIAI 1421
Query: 1501 DIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASRFLELRNKLRV 1547
D FVR + S L VSH+ + Q+AD LTKPLS + F R K+ V
Sbjct: 1422 DYHFVRNMIQSGALRVSHVSTRDQLADALTKPLSRAHFQSARFKIGV 1468
>UniRef100_O49140 Polyprotein [Arabidopsis thaliana]
Length = 1451
Score = 906 bits (2341), Expect = 0.0
Identities = 579/1580 (36%), Positives = 822/1580 (51%), Gaps = 166/1580 (10%)
Query: 1 MSSPSSPVNNSETQRVTAAASKNFKQIISVKLDETNYLQWKQQVEGVLRGTKMVRHVVSP 60
MS +SPV+ SET V+ + N +L ++N++ W +QV +L G + +V
Sbjct: 1 MSDSTSPVSVSETIAVSTSTLLNVNMTNVTRLTDSNFVMWSRQVHALLDGYDLAGYVDGS 60
Query: 61 QIPPVFLNDAAREAGTENPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEI 120
P A T N YT W+ QD L+ + +L IS S+ + S ++W+ +
Sbjct: 61 IPIPTPTRTTADGVVTTNNDYTLWKRQDKLIYSALLGAISLSVQPLLSKANTSAEIWETL 120
Query: 121 HNYCYTQMRTRSRQLRSELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVL 180
+ +QLR +L+ TKGT+SI + + + L +G + IE +L
Sbjct: 121 SSTFANPSWAHVQQLRQQLKQWTKGTKSIVTYFQGFTTRFDHLALLGKAPEREEQIELIL 180
Query: 181 EALPEEFNPIVATVNSQTEVISLDELESQLLTQEARNEKFKKALVGETASVNLTHAENSG 240
LPE++ +V + + +L E+ +L+ N + K A E SV +T N+
Sbjct: 181 GGLPEDYKTVVDQIEGRENPPALTEVLEKLI-----NHEVKLAAKAEATSVPVT--ANAV 233
Query: 241 EKNGHNQPQTGSYPDQQFNISGNPTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRN 300
G+N S + + N GN + NS N R R ++G
Sbjct: 234 NYRGNNNNNNNSRSNGRNNSRGNTSWQNSQSTSN-----RQQYTPRPYQG---------- 278
Query: 301 FGRGNIQCQICYKTGHDASICYHRLSVPPQYEGYGSLGGNFGGNLGSGYGPATGFGTHSN 360
+CQIC GH A C PQ + + G++++
Sbjct: 279 ------KCQICSVHGHSARRC-------PQLQQHA--------------------GSYAS 305
Query: 361 VWMQGVGQRNPSYGAPRAPFPPQFGNSRPPAPQAYITGNESTSSNSFNNG-WYPDSGATH 419
N S A AP+ P+ S+ +N+G W DSGATH
Sbjct: 306 ---------NQSSSASYAPWQPRAN---------------MVSATPYNSGNWLLDSGATH 341
Query: 420 HVTPDANNLMDAASFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSI 479
H+T + NNL ++G +++ I +G GL I+ GS +P +L L ++L+VP I
Sbjct: 342 HLTSNLNNLALHQPYNGDEEVTIADGSGLPISHSGSALLPTP---TRSLALKDVLYVPDI 398
Query: 480 TKNLVSVSQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQFDQPYVPSVSRT 539
KNL+SV + C N V EF VK + LL+G ++ LY++
Sbjct: 399 QKNLISVYRMCNTNGVSVEFFPAHFQVKDLSTGARLLQGKTKNE-LYEWPV--------- 448
Query: 540 ASSSSVATSSLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNR 599
+SS+ATS + SP+ + S WH R
Sbjct: 449 --NSSIATSMFA-----SPTPKTDLPS-----------------------------WHAR 472
Query: 600 LGHPHHEVVRSVMKLCNQQLPNKSFTDF-CSACCLGKSHRLPSVSSKTVYNKPFELIFCD 658
LGHP ++++++ + + + CS C + KSH+LP S+ + P E ++ D
Sbjct: 473 LGHPSLPILKALISKFSLPISHSLQNQLLCSDCSINKSHKLPFYSNTIASSHPLEYLYTD 532
Query: 659 LWGPASVESHGGYSYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSV 718
+W + + S Y Y+L+ VD Y+RYTW++PL+ KS F F +VE ++ I ++
Sbjct: 533 VW-TSPITSIDNYKYYLVIVDHYTRYTWLYPLRKKSQVREMFITFTALVENKFKFKIGTL 591
Query: 719 QTDGGGEFRPFTQFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHY 778
+D GGEF FL + GI+H T PHT NG ERKHRHIVETGLTLL+ A +P Y
Sbjct: 592 YSDNGGEFIAMRSFLASHGISHMTTPPHTPELNGISERKHRHIVETGLTLLSTASMPKEY 651
Query: 779 WDHAFLTATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLEL 838
W +AF TA YLINR+ +P+L N+SP+ L Q P+Y L+ FGC CFP+ RPY HKL+
Sbjct: 652 WSYAFATAVYLINRMLTPVLGNESPYVKLFGQPPNYLKLRIFGCLCFPWLRPYTAHKLDN 711
Query: 839 RSKECVFLGYSPSHKGYKCLD-PTGRMFISKDVIFNEYKFPYSELFTSGQSSSPPTTSSD 897
RS CV LGYS S Y CLD TGR++ S+ V F E FP+S S S P S D
Sbjct: 712 RSVPCVLLGYSLSQSAYLCLDRATGRVYTSRHVQFAESSFPFSTTSPSVTPPSDPPLSQD 771
Query: 898 HTP--LPSFLFPLNNKQCPTTQSSSTP---TTTLHTASPHSSFPESNQSNHHHSIQDTHA 952
P +P PL P++ S S P + SP + S + I
Sbjct: 772 TRPVSVPLLARPLTTAP-PSSPSCSAPHRSPSQSENLSPPAPLQPSLSLSPTSPITSPSL 830
Query: 953 SSHS-NHHNISPGPIFNPTPISTHPPSPSPSSHSHNTYHSISVEP--------------- 996
S S HN GP + P+S P P P S + HS S +P
Sbjct: 831 SEESLVGHNSETGPTGSSPPLSPQPQRPQPQSPQSTSPHSSSPQPNSPNPQHSPRSLTPT 890
Query: 997 VTSQPSTQAEPHRIHPNNTHSMATRAKHGIVQKRKHPTLLLTH------IEPTGYRQAMK 1050
+TS PS P+ P H+M TR+K+ IV+ L T I P +A+
Sbjct: 891 LTSSPSPSPPPNPNPPPIQHTMRTRSKNNIVKPNPKFANLATKPTPLKPIIPKTVVEALL 950
Query: 1051 QPQWLQAMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKG 1110
P W QAM E A +N T+ LVP ++ VGCKWVF K +G +++YKARLVAKG
Sbjct: 951 DPNWRQAMCDEINAQTRNGTFDLVPPAPNQNVVGCKWVFTLKYLSNGVLDRYKARLVAKG 1010
Query: 1111 FHQMPGFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPP 1170
FHQ G D+KETFSPV+K TVRSVL +AV+ W I+Q+DVNNAFL G L +EVY+TQPP
Sbjct: 1011 FHQQYGHDFKETFSPVIKSTTVRSVLHIAVSKGWSIRQIDVNNAFLQGTLSDEVYVTQPP 1070
Query: 1171 GFEAVDPS-LVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHST 1229
GF D + VC+L KALYGLKQAPRAW++ L+S LL GF +S D SLF L +
Sbjct: 1071 GFVDKDNAHHVCRLYKALYGLKQAPRAWYQELRSYLLTQGFVNSVADTSLFTLRHERTIL 1130
Query: 1230 FMLVYVDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKK 1289
++LVYVDD+LITGS ++I + + L A FSLKDLG++ YFLGIE + G L L QK+
Sbjct: 1131 YVLVYVDDMLITGSDTNIITRFIANLAARFSLKDLGEMSYFLGIEATRTSKG-LHLMQKR 1189
Query: 1290 YIQDLLVKANMANANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISY 1349
Y+ DLL K NM A+ + +PM+ + KL+ + P+ +R+++G LQY+ TRP+I+Y
Sbjct: 1190 YVLDLLEKTNMLAAHPVLTPMSPTPKLSLTSGKPLDKPSEYRAVLGSLQYLLFTRPDIAY 1249
Query: 1350 SVNKVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDP 1409
+VN++ Q++ P + HW+A KRILRYL GT HG+ I PL+L A+ DADWA D
Sbjct: 1250 AVNRLSQYMHCPTDLHWQAAKRILRYLAGTPSHGIFIR---ADTPLTLHAYSDADWAGDI 1306
Query: 1410 DDRRSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVP 1469
D+ ST+ + LG N ISW +KKQ VARSS EAEYR++A A++E+ W+ SLL EL +
Sbjct: 1307 DNYNSTNAYILYLGSNPISWSSKKQKGVARSSTEAEYRAVANATSEIRWVCSLLTELGIT 1366
Query: 1470 -TAIPQIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADI 1528
++ P ++CDN+ L+ NPV HSR KH+ LD FVRE V + L V+H+ + Q+AD
Sbjct: 1367 LSSPPVVYCDNVGATYLSANPVFHSRMKHIALDFHFVRESVQAGALRVTHVSTKDQLADA 1426
Query: 1529 LTKPLSASRFLELRNKLRVS 1548
LTKPL F L +K+ V+
Sbjct: 1427 LTKPLPRQPFTTLISKIGVA 1446
>UniRef100_O49142 Polyprotein [Arabidopsis thaliana]
Length = 1451
Score = 904 bits (2336), Expect = 0.0
Identities = 578/1580 (36%), Positives = 821/1580 (51%), Gaps = 166/1580 (10%)
Query: 1 MSSPSSPVNNSETQRVTAAASKNFKQIISVKLDETNYLQWKQQVEGVLRGTKMVRHVVSP 60
MS +SPV+ SET V+ + N +L ++N++ W +QV +L G + ++
Sbjct: 1 MSDSTSPVSVSETIAVSTSTLLNVNMTNVTRLTDSNFVMWSRQVHALLDGYDLAGYIDGS 60
Query: 61 QIPPVFLNDAAREAGTENPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEI 120
P A T N YT W+ QD L+ + +L IS S+ + S ++W+ +
Sbjct: 61 IPIPTPTRTTADGVVTTNNDYTLWKRQDKLIYSALLGAISLSVQPLLSKANTSAEIWETL 120
Query: 121 HNYCYTQMRTRSRQLRSELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVL 180
+ +QLR +L+ TKGT+SI + + + L +G + IE +L
Sbjct: 121 SSTFANPSWAHVQQLRQQLKQWTKGTKSIVTYFQGFTTRFDHLALLGKAPEREEQIELIL 180
Query: 181 EALPEEFNPIVATVNSQTEVISLDELESQLLTQEARNEKFKKALVGETASVNLTHAENSG 240
LPE++ +V + + +L E+ +L+ N + K A E SV +T N+
Sbjct: 181 GGLPEDYKTVVDQIEGRENPPALTEVLEKLI-----NHEVKLAAKAEATSVPVT--ANAV 233
Query: 241 EKNGHNQPQTGSYPDQQFNISGNPTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRN 300
G+N S + + N GN + NS N R R ++G
Sbjct: 234 NYRGNNNNNNNSRSNGRNNSRGNTSWQNSQSTSN-----RQQYTPRPYQG---------- 278
Query: 301 FGRGNIQCQICYKTGHDASICYHRLSVPPQYEGYGSLGGNFGGNLGSGYGPATGFGTHSN 360
+CQIC GH A C PQ + + G++++
Sbjct: 279 ------KCQICSVHGHSARRC-------PQLQQHA--------------------GSYAS 305
Query: 361 VWMQGVGQRNPSYGAPRAPFPPQFGNSRPPAPQAYITGNESTSSNSFNNG-WYPDSGATH 419
N S A AP+ P+ S+ +N+G W DSGATH
Sbjct: 306 ---------NQSSSASYAPWQPRAN---------------MVSATPYNSGNWLLDSGATH 341
Query: 420 HVTPDANNLMDAASFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSI 479
H+T D NNL ++G +++ I +G GL I+ GS +P +L L ++L+VP I
Sbjct: 342 HLTSDLNNLALHQPYNGDEEVTIADGSGLPISHSGSALLPTP---TRSLALKDVLYVPDI 398
Query: 480 TKNLVSVSQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQFDQPYVPSVSRT 539
KNL+SV + C N V EF VK + LL+G ++ LY++
Sbjct: 399 QKNLISVYRMCNTNGVSVEFFPAHFQVKDLSTGARLLQGKTKNE-LYEWPV--------- 448
Query: 540 ASSSSVATSSLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNR 599
+SS+ATS + SP+ + S WH R
Sbjct: 449 --NSSIATSMFA-----SPTPKTDLPS-----------------------------WHAR 472
Query: 600 LGHPHHEVVRSVMKLCNQQLPNKSFTDF-CSACCLGKSHRLPSVSSKTVYNKPFELIFCD 658
LGHP ++++++ + + + CS C + KSH+LP S+ + P E ++ D
Sbjct: 473 LGHPSLPILKALISKFSLPISHSLQNQLLCSDCSINKSHKLPFYSNTIASSHPLEYLYTD 532
Query: 659 LWGPASVESHGGYSYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSV 718
+W + + S Y Y+L+ VD Y+RYTW++PL+ KS F F +VE ++ I ++
Sbjct: 533 VW-TSPITSIDNYKYYLVIVDHYTRYTWLYPLRKKSQVREMFITFTALVENKFKFKIGTL 591
Query: 719 QTDGGGEFRPFTQFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHY 778
+D GGEF FL + GI+H T PHT NG ERKHRHIVETGLTLL+ A +P Y
Sbjct: 592 YSDNGGEFIAMRSFLASHGISHMTTPPHTPELNGISERKHRHIVETGLTLLSTASMPKEY 651
Query: 779 WDHAFLTATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLEL 838
W +AF TA YLINR+ +P+L N+SP+ L Q P+Y L+ FGC CFP+ RPY HKL+
Sbjct: 652 WSYAFATAVYLINRMLTPVLGNESPYVKLFGQPPNYLKLRIFGCLCFPWLRPYTAHKLDN 711
Query: 839 RSKECVFLGYSPSHKGYKCLD-PTGRMFISKDVIFNEYKFPYSELFTSGQSSSPPTTSSD 897
RS CV LGYS S Y CLD TGR++ S+ V F E FP+S S S P S D
Sbjct: 712 RSVPCVLLGYSLSQSAYLCLDRATGRVYTSRHVQFAESSFPFSTTSPSVTPPSDPPLSQD 771
Query: 898 HTP--LPSFLFPLNNKQCPTTQSSSTP---TTTLHTASPHSSFPESNQSNHHHSIQDTHA 952
P +P PL P++ S S P + SP + S + I
Sbjct: 772 TRPVSVPLLARPLTTAP-PSSPSCSAPHRSPSQSENLSPPAPLQPSLSLSPTSPITSPSL 830
Query: 953 SSHS-NHHNISPGPIFNPTPISTHPPSPSPSSHSHNTYHSISVEP--------------- 996
S S HN GP + P+S P P P S + HS S +P
Sbjct: 831 SEESLVGHNSETGPTGSSPPLSPQPQRPQPQSPQSTSPHSSSPQPNSPNPQHSPRSLTPT 890
Query: 997 VTSQPSTQAEPHRIHPNNTHSMATRAKHGIVQKRKHPTLLLTH------IEPTGYRQAMK 1050
+TS PS P+ P H+M TR+K+ IV+ L T I P +A+
Sbjct: 891 LTSSPSPSPPPNPNPPPIQHTMRTRSKNNIVKPNPKFANLATKPTPLKPIIPKTVVEALL 950
Query: 1051 QPQWLQAMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKG 1110
P W QAM E A +N T+ LVP ++ VGCKWVF K +G +++YKARLVAKG
Sbjct: 951 DPNWRQAMCDEINAQTRNGTFDLVPPAPNQNVVGCKWVFTLKYLSNGVLDRYKARLVAKG 1010
Query: 1111 FHQMPGFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPP 1170
FHQ G D+KETFSPV+K TVRSVL +AV+ W I+Q+DVNNAFL G L +EVY+TQPP
Sbjct: 1011 FHQQYGHDFKETFSPVIKSTTVRSVLHIAVSKGWSIRQIDVNNAFLQGTLSDEVYVTQPP 1070
Query: 1171 GFEAVDPS-LVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHST 1229
GF D + VC+L KALYGLKQAPRAW++ L+S LL GF +S D SLF L +
Sbjct: 1071 GFVDKDNAHHVCRLYKALYGLKQAPRAWYQELRSYLLTQGFVNSVADTSLFTLRHERTIL 1130
Query: 1230 FMLVYVDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKK 1289
++LVYVDD+LITGS ++I + + L A FSLKDLG++ YFLGIE + G L L QK+
Sbjct: 1131 YVLVYVDDMLITGSDTNIITRFIANLAARFSLKDLGEMSYFLGIEATRTSKG-LHLMQKR 1189
Query: 1290 YIQDLLVKANMANANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISY 1349
Y+ DLL K NM A+ + +PM+ + KL+ + P+ +R+++G LQY+ TRP+I+Y
Sbjct: 1190 YVLDLLEKTNMLAAHPVLTPMSPTPKLSLTSGKPLDKPSEYRAVLGSLQYLLFTRPDIAY 1249
Query: 1350 SVNKVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDP 1409
+VN++ Q++ P + HW+A KRILRYL GT HG+ I PL+L A+ DADWA D
Sbjct: 1250 AVNRLSQYMHCPTDLHWQAAKRILRYLAGTPSHGIFIR---ADTPLTLHAYSDADWAGDI 1306
Query: 1410 DDRRSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVP 1469
D+ ST+ + LG N ISW +KKQ VARSS EAEYR++A A++E+ W+ SLL EL +
Sbjct: 1307 DNYNSTNAYILYLGSNPISWSSKKQKGVARSSTEAEYRAVANATSEIRWVCSLLTELGIT 1366
Query: 1470 -TAIPQIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADI 1528
++ P ++CDN+ L+ NPV SR KH+ LD FVRE V + L V+H+ + Q+AD
Sbjct: 1367 LSSPPVVYCDNVGATYLSANPVFDSRMKHIALDFHFVRESVQAGALRVTHVSTKDQLADA 1426
Query: 1529 LTKPLSASRFLELRNKLRVS 1548
LTKPL F L +K+ V+
Sbjct: 1427 LTKPLPRQPFTTLISKIGVA 1446
>UniRef100_Q9FLA4 Polyprotein [Arabidopsis thaliana]
Length = 1429
Score = 897 bits (2319), Expect = 0.0
Identities = 569/1556 (36%), Positives = 814/1556 (51%), Gaps = 173/1556 (11%)
Query: 31 KLDETNYLQWKQQVEGVLRGTKMVRHVVSPQIPPVFLNDAAREAGTENPAYTEWEEQDSL 90
KL TN+L W++QV +L G + +V I + NP Y W+ QD L
Sbjct: 6 KLTSTNFLMWRRQVHALLDGYDLAGYV-DGSIEEPHTTVTVHGVTSPNPEYKLWKRQDKL 64
Query: 91 LCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRSELRTITKGTRSIA 150
+ + ++ IS ++ + S Q+W ++ + R +Q+R +++ KG+RSI
Sbjct: 65 IYSGLIGAISVAVQPLLSQATTSAQIWRKLVDTYANPSRGHKQQIREQIKQWKKGSRSID 124
Query: 151 EFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVATVNSQTEVISLDELESQL 210
+++ + + + L + + + H D I +L L +++ ++ + + S+ EL +L
Sbjct: 125 DYVLGLTTRFDQLALLEEAIPHEDQIAYILGGLSDDYRRVIDQIEGRDISPSITELHEKL 184
Query: 211 LTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYPDQQFNISGNPTGNNSS 270
+ E K + + + V A + NG N + GN ++
Sbjct: 185 INFEL---KLQAMVPDSSTPVTANAASYNNNNNGRNNRSSSR-------------GNQNN 228
Query: 271 QYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTGHDASICYHRLSVPPQ 330
Q+ N ++ S RG +G ++GR CQIC GH A C Q
Sbjct: 229 QW-QQNQTQQSRSNNRGSQGKGYQGR-----------CQICGVHGHSARRC-------SQ 269
Query: 331 YEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYGAPRAPFPPQFGNSRPP 390
++ YG GG+ ++ SGY P G+ +PS P AP+ P+ + P
Sbjct: 270 FQPYGGSGGS--QSVPSGY-PTNGY--------------SPS---PMAPWQPRANIATAP 309
Query: 391 APQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLMDAASFSGSDQMYIGNGQGLAI 450
N W DSGATHH+T D NL ++G +++ I +G GL I
Sbjct: 310 P----------------FNPWVLDSGATHHLTSDLANLSMHQPYTGGEEVTIADGSGLPI 353
Query: 451 NSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICYVKSQD 510
+ GS +P + +L L ++L+VP+++KNL+SV + C N V EF VK +
Sbjct: 354 SHTGSALLPTP---SRSLALKDILYVPNVSKNLISVYRLCNANQVSVEFFPAHFQVKDLN 410
Query: 511 STKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVATSSLSLNNCFSPSSLSLSRSQCNN 570
+ LL+G ++ LY++ TAS S
Sbjct: 411 TGARLLQGRTRNE-LYEWPVNQKSITILTASPSP-------------------------- 443
Query: 571 GSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQLPNKSFTDF-CS 629
D S+ WH RLGHP +++ V+ + L N CS
Sbjct: 444 ------------KTDLSS-------WHQRLGHPALPILKDVVSHFHLPLSNTIPKQLPCS 484
Query: 630 ACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSRYTWIFP 689
C + KSH+LP ++ V ++P E ++ D+W + S Y Y+L+ VD ++RYTW++P
Sbjct: 485 DCSINKSHKLPFFTNTIVSSQPLEYLYTDVWTSPCI-SVDNYKYYLVIVDHFTRYTWMYP 543
Query: 690 LKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCPHTHH 749
LK KS F FK +VE ++ I+++ +D GGEF FL GI+H + PHT
Sbjct: 544 LKQKSQVKDVFVAFKALVENRFQSRIRTLYSDNGGEFIGLRPFLAAHGISHLTSPPHTPE 603
Query: 750 QNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFFLLHL 809
NG ERKHRHIVETGL LL +A LP +W +AF TA YLINR+P+ +L SP+ L
Sbjct: 604 HNGLAERKHRHIVETGLALLTHASLPKTFWTYAFATAVYLINRMPTEVLQGTSPYVKLFQ 663
Query: 810 QIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLD-PTGRMFISK 868
P+Y L+ FGC C+P+ RPYN +KLE RS CVFLGYS + Y CLD T R++ S+
Sbjct: 664 MSPNYLKLRVFGCLCYPWLRPYNTNKLEARSTMCVFLGYSLTQSAYLCLDIATNRIYTSR 723
Query: 869 DVIFNEYKFPYSELFTSGQSSSPPTTSSDHTPLPSFLFPLNNKQCPTTQSSSTPTTTLHT 928
V F E FP F S ++S +T + P + + PL + ++ P +
Sbjct: 724 HVQFVESSFP----FASPRTSETDSTQTMSQPTTTNVIPLLQRPPHIAPPTALPLCPIFH 779
Query: 929 ASPHS-SFPESNQSNHHHSIQDTHASSHSNHHN---------------ISPGPIFNPTPI 972
+ PHS S P S S H + T ASS SN N S P PT
Sbjct: 780 SPPHSPSSPASPPSEH---VPLTAASSSSNAINDDNISSTGQVSVSGPTSQSPHTTPTNQ 836
Query: 973 STHP--PSPSPSSHSHNTYHSISVEPVTS--------QPSTQAEPHRIHPNNTHSMATRA 1022
+T P SP+P++ + + + P TS P Q P P N H M TRA
Sbjct: 837 NTSPLSKSPNPTNTNQSQNSTPPTSPTTSVHQHSPTPSPLPQNPPLPPPPQNDHPMRTRA 896
Query: 1023 KHGIVQKRK--HPTLLLTHIE---PTGYRQAMKQPQWLQAMQLEHEALMKNNTWTLVPLP 1077
K+ I + + + T LT + PT QA+K P W AM E A MKN+TW LV
Sbjct: 897 KNQITKPKTKFNLTTSLTSSKPTIPTTVAQALKDPNWRNAMSEEINAQMKNHTWDLVSPE 956
Query: 1078 ADRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKETFSPVVKPVTVRSVLT 1137
+ + CKW+F K N DGSI +YKARLVA+GF+Q G DY ETFSPV+K T+R+VL
Sbjct: 957 EAKHVISCKWIFTLKYNVDGSIARYKARLVARGFNQQYGIDYSETFSPVIKSTTIRTVLE 1016
Query: 1138 LAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVD-PSLVCKLNKALYGLKQAPRA 1196
+AV W I Q+D+NNAFL G L EEVY++QPPGF D PS VC+LNKALYGLKQAPRA
Sbjct: 1017 VAVKRNWSIHQVDINNAFLQGTLNEEVYVSQPPGFIDRDRPSHVCRLNKALYGLKQAPRA 1076
Query: 1197 WFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFM--LVYVDDILITGSSASLIQQLVKK 1254
W++ L+ LL+ GF +S D SLFI N+H+TFM LVYVDDI+I G +A L+Q
Sbjct: 1077 WYQELRRFLLQAGFVNSLADASLFIY--NRHNTFMYVLVYVDDIIIAGENA-LVQAFNAS 1133
Query: 1255 LNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMANANGIASPMASST 1314
L + FSLKDLG L YFLGIE + G L L Q+KYI DLL K NM + +++PM+ +
Sbjct: 1134 LASRFSLKDLGPLSYFLGIEATRTSRG-LHLMQRKYITDLLKKHNMLDTKPVSTPMSPTP 1192
Query: 1315 KLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAPLEDHWKAVKRILR 1374
KL+ + D T +R+++G LQY+ TRP+I+++VN++ QF+ P +HW+A KRILR
Sbjct: 1193 KLSLLSGTALDDATEYRTVLGSLQYLAFTRPDIAFAVNRLSQFMHRPTNEHWQAAKRILR 1252
Query: 1375 YLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACILLGPNLISWWAKKQ 1434
YL GT HG+ + PL++ AF DADW D D ST+ + G + +SW +KKQ
Sbjct: 1253 YLAGTKSHGIFLR---SDTPLTIHAFSDADWGCDLDAYLSTNAYIVYFGGSPVSWSSKKQ 1309
Query: 1435 TLVARSSAEAEYRSLAQASAEVLWIQSLLKELKV-PTAIPQIFCDNLSTVSLAHNPVLHS 1493
VARSS EAEYR++A ++E+ W+ SLL E+ + T +P I+CDN+ L NPV HS
Sbjct: 1310 RSVARSSTEAEYRAVANTASELRWLCSLLLEMGISQTTVPVIYCDNIGATYLCANPVFHS 1369
Query: 1494 RTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASRFLELRNKLRVSD 1549
R KH+ LD FVR + S L VSH+ + Q+AD LTKPL RF EL +K+ V +
Sbjct: 1370 RMKHVALDYHFVRGYIQSGALRVSHVSTKDQLADALTKPLPRPRFTELNSKIGVQE 1425
>UniRef100_Q9SXQ1 Polyprotein [Arabidopsis thaliana]
Length = 1431
Score = 884 bits (2284), Expect = 0.0
Identities = 563/1467 (38%), Positives = 781/1467 (52%), Gaps = 184/1467 (12%)
Query: 134 QLRSELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVAT 193
QLR++L+ TKGT++I +++ + + + L +G P+ H + +E VLE LPEE+ P++
Sbjct: 92 QLRTQLKQWTKGTKTIDDYMQGLVTRFDQLALLGKPMDHDEQVERVLENLPEEYKPVIDQ 151
Query: 194 VNSQTEVISLDELESQLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSY 253
+ ++ +L E+ +LL E++ A V + ++H + N +N + Y
Sbjct: 152 IAAKDTPPTLTEIHERLLNHESKILAVSSATVIPITANAVSHRNTTTTTNNNNGNRNNRY 211
Query: 254 PDQQFNISGNPTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYK 313
++ N + P SS F+PN ++ + + G +CQIC
Sbjct: 212 DNRNNNNNSKPW-QQSSTNFHPN-----NNQSKPYLG----------------KCQICGV 249
Query: 314 TGHDASICYHRLSVPPQYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSY 373
GH A C + ++ V + P
Sbjct: 250 QGHSAKRC-----------------------------------SQLQHFLSSVNSQQPP- 273
Query: 374 GAPRAPFPPQFGNSRPPAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLMDAAS 433
+P P+ P+ N + S +N W DSGATHH+T D NNL
Sbjct: 274 -SPFTPWQPR--------------ANLALGSPYSSNNWLLDSGATHHITSDFNNLSLHQP 318
Query: 434 FSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDN 493
++G D + + +G + I+ GS S S+ P L L+N+L+VP+I KNL+SV + C N
Sbjct: 319 YTGGDDVMVADGSTIPISHTGSTSLSTKSRP---LNLHNILYVPNIHKNLISVYRLCNAN 375
Query: 494 NVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVATSSLSLN 553
V EF VK ++ LL+G D+ LY++ VS AS SS AT
Sbjct: 376 GVSVEFFPASFQVKDLNTGVPLLQGKTKDE-LYEWPIASSQPVSLFASPSSKAT------ 428
Query: 554 NCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMK 613
H+S WH RLGHP ++ SV+
Sbjct: 429 ------------------------HSS---------------WHARLGHPAPSILNSVIS 449
Query: 614 LCNQQLPNKSFTDF-CSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYS 672
+ + N S CS C + KS+++P S +P E I+ D+W + + SH Y
Sbjct: 450 NYSLSVLNPSHKFLSCSDCLINKSNKVPFSQSTINSTRPLEYIYSDVWS-SPILSHDNYR 508
Query: 673 YFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQF 732
Y++I VD ++RYTW++PLK KS TF FK ++E ++ I + +D GGEF ++
Sbjct: 509 YYVIFVDHFTRYTWLYPLKQKSQVKETFITFKNLLENRFQTRIGTFYSDNGGEFVALWEY 568
Query: 733 LTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINR 792
+ GI+H + PHT NG ERKHRHIVETGLTLL++A +P YW +AF A YLINR
Sbjct: 569 FSQHGISHLTSPPHTPEHNGLSERKHRHIVETGLTLLSHASIPKTYWPYAFAVAVYLINR 628
Query: 793 LPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSH 852
LP+P+L +SPF L P+Y L+ FGC+C+P+ RPYN HKL+ +S++CVFLGYS +
Sbjct: 629 LPTPLLQLESPFQKLFGTSPNYDKLRVFGCACYPWLRPYNQHKLDDKSRQCVFLGYSLTQ 688
Query: 853 KGYKCLD-PTGRMFISKDVIFNEYKFPYSELFTS--------GQSS---SPPTT------ 894
Y CL T R++IS+ V F+E FP+S + +SS SP TT
Sbjct: 689 SAYLCLHLQTSRLYISRHVRFDENCFPFSNYLATLSPVQEQRRESSCVWSPHTTLPTRTP 748
Query: 895 -------SSDH---TPLPSFLFPLNNKQCPTTQ-----SSSTPTTTLHTA----SPHSSF 935
S H TP S P N Q ++ SSS P++ TA P
Sbjct: 749 GLPAPSGSDPHHAATPPSSPSAPFRNSQVSSSNRESYFSSSFPSSPEPTAPRQNGPQPKT 808
Query: 936 PESNQSNHHHSIQDTHASSHSNHHNISPGPIFN--PTPISTHPPSPSP-----SSHSHNT 988
+ HS Q+T S +N N SP + TP + SPSP SS + T
Sbjct: 809 QPTQTQTQTHSSQNT---SQNNPTNESPSQLAQSLSTPAQSSSSSPSPTTSASSSSTSPT 865
Query: 989 YHSISVEPVTSQPSTQ-AEPHRIHPNNTHSMATRAKHGIVQKRKHPTL---LLTHIEPTG 1044
SI + P P Q + P NTHSM TRAK GI++ +L L EP
Sbjct: 866 PPSILIHP--PPPLAQIVNNNNQAPLNTHSMGTRAKAGIIKPNPKYSLAVSLAAESEPRT 923
Query: 1045 YRQAMKQPQWLQAMQLEHEALMKNNTWTLVPLPADR-QAVGCKWVFRTKQNPDGSINKYK 1103
QA+K +W AM E A + N+TW LVP P VGC+W+F K N DGS+N+YK
Sbjct: 924 AIQALKDERWRNAMGSEINAQIGNHTWDLVPPPPSHVTIVGCRWIFTKKYNSDGSLNRYK 983
Query: 1104 ARLVAKGFHQMPGFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEE 1163
AR VAKG++Q PG DY ETFSPV+K ++R VL +AV W I+QLDVNNAFL G L ++
Sbjct: 984 ARFVAKGYNQRPGLDYAETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDD 1043
Query: 1164 VYMTQPPGFEAVD-PSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFIL 1222
VYM+QPPGF D P+ VCKL KALYGLKQAPRAW+ L++ LL +GF +S D SLF+L
Sbjct: 1044 VYMSQPPGFIDKDRPNYVCKLRKALYGLKQAPRAWYVELRNYLLTIGFVNSVSDTSLFVL 1103
Query: 1223 HANQHSTFMLVYVDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGS 1282
+ +MLVYVDDILITG+ +L+ + L+ FS+KD +L YFLGIE G
Sbjct: 1104 QRGKSIVYMLVYVDDILITGNDPTLLHNTLDNLSQRFSVKDHEELHYFLGIEAKRVPTG- 1162
Query: 1283 LLLSQKKYIQDLLVKANMANANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTV 1342
L LSQ++YI DLL + NM A + +PMA S KL+ Y ++DPT +R IVG LQY+
Sbjct: 1163 LHLSQRRYILDLLARTNMITAKPVTTPMAPSPKLSLYSGTKLTDPTEYRGIVGSLQYLAF 1222
Query: 1343 TRPEISYSVNKVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCD 1402
TRP+ISY+VN++ QF+ P E+H +A+KRILRYL GT +HG+ + LSL A+ D
Sbjct: 1223 TRPDISYAVNRLSQFMHMPTEEHLQALKRILRYLAGTPNHGIFLKKG---NTLSLHAYSD 1279
Query: 1403 ADWASDPDDRRSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSL 1462
ADWA D DD ST+G + LG + ISW +KKQ V RSS EAEYRS+A S+E+ WI SL
Sbjct: 1280 ADWAGDKDDYVSTNGYIVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSEMQWICSL 1339
Query: 1463 LKELKVP-TAIPQIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPA 1521
L EL + T P I+CDN+ L NPV HSR KH+ +D F+R +V S L V H+
Sbjct: 1340 LTELGIRLTRPPVIYCDNVGATYLCANPVFHSRMKHIAIDYHFIRNQVQSGALRVVHVST 1399
Query: 1522 QYQVADILTKPLSASRFLELRNKLRVS 1548
Q+AD LTKPLS + F +K+ V+
Sbjct: 1400 HDQLADTLTKPLSRTAFQNFASKIGVT 1426
>UniRef100_Q9SXQ2 Polyprotein [Arabidopsis thaliana]
Length = 1330
Score = 880 bits (2275), Expect = 0.0
Identities = 568/1442 (39%), Positives = 768/1442 (52%), Gaps = 188/1442 (13%)
Query: 161 ESLMSIGDPVAHRDLIETVLEALPEEFNPIVATVNSQTEVISLDELESQLLTQEARNEKF 220
+ L +G P+ H + +E VLE LP+++ P++ + ++ SL E+ +L+ +E++
Sbjct: 9 DQLALLGKPMDHDEQVERVLENLPDDYKPVIDQIAAKDTPPSLTEIHERLINRESKLLAL 68
Query: 221 KKALVGETASVNLTHAENSGEKNGHNQPQTGSYPDQQFNISGNPTGNNSSQYFNPNFGGR 280
A V + +TH + +N +N+ +Y + NN S + P+
Sbjct: 69 NSAEVVPITANVVTHRNTNTNRNQNNRGDNRNYNNN----------NNRSNSWQPS---S 115
Query: 281 NGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTGHDASICYHRLSVPPQYEGYGSLGGN 340
+GSR + + GR CQIC GH A C PQ +
Sbjct: 116 SGSRSDNRQPKPYLGR-----------CQICSVQGHSAKRC-------PQLHQF------ 151
Query: 341 FGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYGAPRAPFPPQFGNSRPPAPQAYITGNE 400
Q + +PF P P+A + N
Sbjct: 152 ---------------------------QSTTNQQQSTSPFTPW-------QPRANLAVNS 177
Query: 401 STSSNSFNNGWYPDSGATHHVTPDANNLMDAASFSGSDQMYIGNGQGLAINSIGSMSFSS 460
++N+ W DSGATHH+T D NNL ++G D + I +G + I GS S +
Sbjct: 178 PYNANN----WLLDSGATHHITSDFNNLSFHQPYTGGDDVMIADGSTIPITHTGSASLPT 233
Query: 461 PFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHL 520
+ +L LN +L+VP+I KNL+SV + C N V EF VK ++ LL+G
Sbjct: 234 S---SRSLDLNKVLYVPNIHKNLISVYRLCNTNRVSVEFFPASFQVKDLNTGVPLLQGKT 290
Query: 521 GDDGLYQFDQPYVPSVSRTASSSSVATSSLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTS 580
D+ LY++ +VS AS S AT H+S
Sbjct: 291 KDE-LYEWPIASSQAVSMFASPCSKAT------------------------------HSS 319
Query: 581 GSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQLP--NKSFTDF-CSACCLGKSH 637
WH+RLGHP ++ SV+ N LP N S CS C + KSH
Sbjct: 320 ---------------WHSRLGHPSLAILNSVIS--NHSLPVLNPSHKLLSCSDCFINKSH 362
Query: 638 RLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSRYTWIFPLKLKSHTL 697
++P +S +KP E I+ D+W + + S Y Y++I VD ++RYTW++PLK KS
Sbjct: 363 KVPFSNSTITSSKPLEYIYSDVWS-SPILSIDNYRYYVIFVDHFTRYTWLYPLKQKSQVK 421
Query: 698 ITFQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCPHTHHQNGSVERK 757
TF FK++VE ++ I ++ +D GGEF +L+ GI+H + PHT NG ERK
Sbjct: 422 DTFIIFKSLVENRFQTRIGTLYSDNGGEFVVLRDYLSQHGISHFTSPPHTPEHNGLSERK 481
Query: 758 HRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFFLLHLQIPDYKFL 817
HRHIVE GLTLL++A +P YW +AF A YLINRLP+P+L +SPF L Q P+Y+ L
Sbjct: 482 HRHIVEMGLTLLSHASVPKTYWPYAFSVAVYLINRLPTPLLQLQSPFQKLFGQPPNYEKL 541
Query: 818 KSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLD-PTGRMFISKDVIFNEYK 876
K FGC+C+P+ RPYN HKLE +SK+C F+GYS + Y CL PTGR++ S+ V F+E
Sbjct: 542 KVFGCACYPWLRPYNRHKLEDKSKQCAFMGYSLTQSAYLCLHIPTGRLYTSRHVQFDERC 601
Query: 877 FPYSE----LFTSGQ--SSSPPTTSSDHTPLPSFLFPLNNKQC--------PTTQSSSTP 922
FP+S + TS + S S P S HT LP+ L C P SS +P
Sbjct: 602 FPFSTTNFGVSTSQEQRSDSAPNWPS-HTTLPTTPLVLPAPPCLGPHLDTSPRPPSSPSP 660
Query: 923 TTTLHTAS---PHSSFPESNQSN----HHHSIQDTHASSHSNHHNISPGPIF-NPTPIST 974
T +S P SS + S H+ Q T A H ++ S PI NP P S
Sbjct: 661 LCTTQVSSSNLPSSSISSPSSSEPTAPSHNGPQPT-AQPHQTQNSNSNSPILNNPNPNSP 719
Query: 975 HPPSP--------SPSSHSHNTYHSISV----EPVTSQPSTQAEP-----------HRIH 1011
P SP SP S H S S+ P +S ST P +
Sbjct: 720 SPNSPNQNSPLPQSPISSPHIPTPSTSISEPNSPSSSSTSTPPLPPVLPAPPIIQVNAQA 779
Query: 1012 PNNTHSMATRAKHGI---VQKRKHPTLLLTHIEPTGYRQAMKQPQWLQAMQLEHEALMKN 1068
P NTHSMATRAK GI QK + T L + EP QAMK +W QAM E A + N
Sbjct: 780 PVNTHSMATRAKDGIRKPNQKYSYATSLAANSEPRTAIQAMKDDRWRQAMGSEINAQIGN 839
Query: 1069 NTWTLVPLPADR-QAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKETFSPVV 1127
+TW LVP P VGC+W+F K N DGS+N+YKARLVAKG++Q PG DY ETFSPV+
Sbjct: 840 HTWDLVPPPPPSVTIVGCRWIFTKKFNSDGSLNRYKARLVAKGYNQRPGLDYAETFSPVI 899
Query: 1128 KPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVD-PSLVCKLNKA 1186
K ++R VL +AV W I+QLDVNNAFL G L +EVYM+QPPGF D P VC+L KA
Sbjct: 900 KSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDEVYMSQPPGFVDKDRPDYVCRLRKA 959
Query: 1187 LYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDILITGSSAS 1246
+YGLKQAPRAW+ L++ LL +GF +S D SLF+L + +MLVYVDDILITG+
Sbjct: 960 IYGLKQAPRAWYVELRTYLLTVGFVNSISDTSLFVLQRGRSIIYMLVYVDDILITGNDTV 1019
Query: 1247 LIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMANANGI 1306
L++ + L+ FS+K+ L YFLGIE G L LSQ++Y DLL + NM A +
Sbjct: 1020 LLKHTLDALSQRFSVKEHEDLHYFLGIEAKRVPQG-LHLSQRRYTLDLLARTNMLTAKPV 1078
Query: 1307 ASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAPLEDHW 1366
A+PMA+S KLT + + DPT +R IVG LQY+ TRP++SY+VN++ Q++ P +DHW
Sbjct: 1079 ATPMATSPKLTLHSGTKLPDPTEYRGIVGSLQYLAFTRPDLSYAVNRLSQYMHMPTDDHW 1138
Query: 1367 KAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACILLGPNL 1426
A+KR+LRYL GT HG+ + LSL A+ DADWA D DD ST+G + LG +
Sbjct: 1139 NALKRVLRYLAGTPDHGIFLKKG---NTLSLHAYSDADWAGDTDDYVSTNGYIVYLGHHP 1195
Query: 1427 ISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVPTAIPQ-IFCDNLSTVSL 1485
ISW +KKQ V RSS EAEYRS+A S+E+ WI SLL EL + + P I+CDN+ L
Sbjct: 1196 ISWSSKKQKGVVRSSTEAEYRSVANTSSELQWICSLLTELGIQLSHPPVIYCDNVGATYL 1255
Query: 1486 AHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASRFLELRNKL 1545
NPV HSR KH+ LD F+R +V S L V H+ Q+AD LTKPLS F K+
Sbjct: 1256 CANPVFHSRMKHIALDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPLSRVAFQNFSRKI 1315
Query: 1546 RV 1547
V
Sbjct: 1316 GV 1317
>UniRef100_Q9T0C5 Retrotransposon like protein [Arabidopsis thaliana]
Length = 1515
Score = 860 bits (2221), Expect = 0.0
Identities = 556/1524 (36%), Positives = 801/1524 (52%), Gaps = 204/1524 (13%)
Query: 78 NPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRS 137
N + +W D L+ WI ++S L + L + +VW + TR L+
Sbjct: 66 NQEFLKWTRIDQLVKAWIFGSLSEEALKVVIGLNSAQEVWLGLARRFNRFSTTRKYDLQK 125
Query: 138 ELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVATVNSQ 197
L T +K +++ +++ +++I + L SIG PV ++ I VL L +E+ I +
Sbjct: 126 RLGTCSKAGKTMDAYLSEVKNICDQLDSIGFPVTEQEKIFGVLNGLGKEYESIATVIEHS 185
Query: 198 TEVIS---LDELESQLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYP 254
+V D++ +L T F L TA+ +T P Y
Sbjct: 186 LDVYPGPCFDDVVYKLTT-------FDDKLSTYTANSEVT-------------PHLAFYT 225
Query: 255 DQQFNISGNPTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKT 314
D+ ++ GN NNS GGR G+ FRGRG
Sbjct: 226 DKSYSSRGN---NNSR-------GGRYGN---------FRGRGS---------------- 250
Query: 315 GHDASICYHRLSVPPQYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYG 374
Y S G F GSG +G G+ Q YG
Sbjct: 251 -------------------YSSRGRGFHQQFGSGSNNGSGNGSKPTC------QICRKYG 285
Query: 375 APRAPFPPQFGNSRPPA--PQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLMDAA 432
+F + P P A+ S + + ++ W PDS AT H+T + L ++
Sbjct: 286 HSAFKCYTRFEENYLPEDLPNAFAAMRVSDQNQASSHEWLPDSAATAHITNTTDGLQNSQ 345
Query: 433 SFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKD 492
++SG D + +GNG L I IG++ + TL L ++L P ITK+L+SVS+ D
Sbjct: 346 TYSGDDSVIVGNGDFLPITHIGTIPLNIS---QGTLPLEDVLVCPGITKSLLSVSKLTDD 402
Query: 493 NNVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQF-DQPYVPSVSRTASSSSVATSSLS 551
F F S+ +K + + ++L +G+ GLY D P+ S
Sbjct: 403 YPCSFTFDSDSVVIKDKRTQQLLTQGNK-HKGLYVLKDVPFQTYYS-------------- 447
Query: 552 LNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSV 611
+R Q SS+D +VWH RLGHP+ EV++ +
Sbjct: 448 ------------TRQQ--------------SSDD--------EVWHQRLGHPNKEVLQHL 473
Query: 612 MKLCNQQLPNKSFTDFCSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGY 671
+K + NK+ ++ C AC +GK RLP V+S+ V ++P E I CDLWGPA V S G+
Sbjct: 474 IKT-KAIVVNKTSSNMCEACQMGKVCRLPFVASEFVSSRPLERIHCDLWGPAPVTSAQGF 532
Query: 672 SYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEF--RPF 729
Y++I +D YSR+TW +PLKLKS F F+ +VE QY I Q DGGGEF F
Sbjct: 533 QYYVIFIDNYSRFTWFYPLKLKSDFFSVFVLFQQLVENQYQHKIAMFQCDGGGEFVSYKF 592
Query: 730 TQFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYL 789
L + GI ++CPHT QNG ER+HR++ E GL+L+ ++K+P W AF T+ +L
Sbjct: 593 VAHLASCGIKQLISCPHTPQQNGIAERRHRYLTELGLSLMFHSKVPHKLWVEAFFTSNFL 652
Query: 790 INRLPSPILN-NKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGY 848
N LPS L+ NKSP+ +LH P Y L+ FG +C+P+ RPY +K + +S CVFLGY
Sbjct: 653 SNLLPSSTLSDNKSPYEMLHGTPPVYTALRVFGSACYPYLRPYAKNKFDPKSLLCVFLGY 712
Query: 849 SPSHKGYKCLDP-TGRMFISKDVIFNEYKFPYSE------------LFTSGQSSSPPTTS 895
+ +KGY+CL P TG+++I + V+F+E KFPYS+ LFT+ Q T
Sbjct: 713 NNKYKGYRCLHPPTGKVYICRHVLFDERKFPYSDIYSQFQTISGSPLFTAWQKGFSSTAL 772
Query: 896 SDHTP---LPSFLFPLNNKQCPTTQSSSTPTTTLHTASPHSSFPESNQSNHHHSI----- 947
S TP + +FP T SSS PT + ++ P+ + + H +
Sbjct: 773 SRETPSTNVEDIIFP------SATVSSSVPTGCAPNIAETATAPDVDVAAAHDMVVPPSP 826
Query: 948 --------QDTHASSHSNHHNISPGPIFNPTPISTHPPSPSPSSHSHNTYHSISVEPVTS 999
Q ++S NH++ + T IS+ + +P S + + + P+ S
Sbjct: 827 ITSTSLPTQPEESTSDQNHYSTD-----SETAISS---AMTPQSINVSLFEDSDFPPLQS 878
Query: 1000 Q-PSTQAEPHRIHPNNTHSMATRAKHGIVQKRKHPTLLLT---HIEPTGYRQAMKQPQWL 1055
ST A P HP M TRAK GI + L + EP ++A+K W
Sbjct: 879 VISSTTAAPETSHP-----MITRAKSGITKPNPKYALFSVKSNYPEPKSVKEALKDEGWT 933
Query: 1056 QAMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMP 1115
AM E + + +TW LVP + +GCKWVF+TK N DGS+++ KARLVA+G+ Q
Sbjct: 934 NAMGEEMGTMHETDTWDLVPPEMVDRLLGCKWVFKTKLNSDGSLDRLKARLVARGYEQEE 993
Query: 1116 GFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAV 1175
G DY ET+SPVV+ TVRS+L +A NKW ++QLDV NAFL+ L+E V+MTQPPGFE
Sbjct: 994 GVDYVETYSPVVRSATVRSILHVATINKWSLKQLDVKNAFLHDELKETVFMTQPPGFEDP 1053
Query: 1176 D-PSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVY 1234
P VCKL KA+Y LKQAPRAWF++ S LLK GF S DPSLF+ + F+L+Y
Sbjct: 1054 SRPDYVCKLKKAIYDLKQAPRAWFDKFSSYLLKYGFICSFSDPSLFVYLKGRDVMFLLLY 1113
Query: 1235 VDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDL 1294
VDD+++TG++ L+QQL+ L+ EF +KD+G L YFLGI+ HY +G L LSQ+KY DL
Sbjct: 1114 VDDMILTGNNDVLLQQLLNILSTEFRMKDMGALHYFLGIQAHYHNDG-LFLSQEKYTSDL 1172
Query: 1295 LVKANMANANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKV 1354
LV A M++ + + +P+ L + + +PT+FR + G LQY+T+TRP+I ++VN V
Sbjct: 1173 LVNAGMSDCSSMPTPL--QLDLLQGNNKPFPEPTYFRRLAGKLQYLTLTRPDIQFAVNFV 1230
Query: 1355 CQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRS 1414
CQ + AP + +KRIL YLKGT+ G+ ++ + L + D+DWA D RRS
Sbjct: 1231 CQKMHAPTMSDFHLLKRILHYLKGTMTMGINLS---SNTDSVLRCYSDSDWAGCKDTRRS 1287
Query: 1415 TSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVP-TAIP 1473
T G C LG N+ISW AK+ V++SS EAEYR+L+ A++EV WI LL+E+ +P IP
Sbjct: 1288 TGGFCTFLGYNIISWSAKRHPTVSKSSTEAEYRTLSFAASEVSWIGFLLQEIGLPQQQIP 1347
Query: 1474 QIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPL 1533
+++CDNLS V L+ NP LHSR+KH ++D ++VRE+V L V HIPA Q+ADI TK L
Sbjct: 1348 EMYCDNLSAVYLSANPALHSRSKHFQVDYYYVRERVALGALTVKHIPASQQLADIFTKSL 1407
Query: 1534 SASRFLELRNKLRVSDP--MSLRG 1555
+ F +LR KL V P SLRG
Sbjct: 1408 PQAPFCDLRFKLGVVLPPDTSLRG 1431
>UniRef100_Q65X82 Putative polyprotein [Oryza sativa]
Length = 1447
Score = 857 bits (2213), Expect = 0.0
Identities = 519/1283 (40%), Positives = 715/1283 (55%), Gaps = 139/1283 (10%)
Query: 335 GSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYGAPRAPFPPQFGNSRPPAPQA 394
G G GGN G+G G G ++N +G G+ N G R PP ++RP
Sbjct: 231 GGGGQQRGGNTGNG---GRGRGGNNNGANRGRGRGNN--GGAR---PPGGVDNRPKCQLC 282
Query: 395 YITGNE--------------------STSSNSFNNGWYPDSGATHHVTPDANNLMDAASF 434
Y G+ S +S + WY D+ AT HVT + + L +
Sbjct: 283 YKRGHTVINCWYRYDEDFVPDEKYAGSATSYGIDTNWYVDTSATDHVTGELDKLTVRDRY 342
Query: 435 SGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNN 494
G DQ++ +G G+ I+ IG S+ +PN + L N+L+VP+ KNLVS ++ DN+
Sbjct: 343 KGQDQVHTASGAGMEISHIGH---STVRTPNRDIHLRNILYVPNANKNLVSANRLVSDNS 399
Query: 495 VFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQFDQPYVPSVSRTAS--SSSVATSSLSL 552
+ E +S +K + K+L +G P R + SSS L
Sbjct: 400 AYMELYSKYFNLKDLATKKLLFRG---------------PCRGRLYALPSSSPHERPRPL 444
Query: 553 NNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVM 612
F S R WH+RLGHP +V V+
Sbjct: 445 KEAFGAIKPSFER------------------------------WHSRLGHPASPIVEKVI 474
Query: 613 KLCNQQ-LPNKSFTDFCSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGY 671
N L + C AC GKSH+LP S ++ + P ELI+ D+WGPA + S GG
Sbjct: 475 SKNNLPCLAESNKQSVCDACQQGKSHQLPYSRSSSMSSHPLELIYSDVWGPA-LTSVGGK 533
Query: 672 SYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQ 731
Y++ + YS++TW++ +K KS + F F+ +VE +N I ++Q+D GGE+
Sbjct: 534 QYYVSFIGDYSKFTWLYLIKHKSEVIQKFHEFQALVERLFNRKIIAMQSDWGGEYEKLHS 593
Query: 732 FLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLIN 791
F T +GITH ++CPHTH QNGS ERKHRHIVE GLTLLA + +PL +WD AF A YLIN
Sbjct: 594 FFTKIGITHHVSCPHTHQQNGSAERKHRHIVEVGLTLLAYSSMPLKFWDEAFQAAVYLIN 653
Query: 792 RLPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPS 851
R P+ +L +P L Q PDY L+ FGC+C+P RPYN HKL+ RSK+C FLGYS
Sbjct: 654 RTPTKLLQFLTPLEHLFNQTPDYSSLRVFGCACWPHLRPYNTHKLQFRSKQCTFLGYSTL 713
Query: 852 HKGYKCLDP-TGRMFISKDVIFNEYKFPYSELFTSGQSSSPPTTSSDHTPLPSFLFPLNN 910
HKG+KCLDP TGR++IS+DVIF+E FP+++L T+ + S+ LPS L LN
Sbjct: 714 HKGFKCLDPSTGRVYISRDVIFDETNFPFAKLHTNAGA----RLRSEILLLPSHL--LNP 767
Query: 911 KQCPTTQ-------------------SSSTPTTTLHTASPHSSFPESNQSNHHHSIQDTH 951
P Q S+ P + + + SS N + H QDT
Sbjct: 768 TSNPGEQQLDDNMANIPVNPANQIFGSNVQPVSAENDEADDSSGATENLAEQIH--QDTA 825
Query: 952 AS-SHSNHHNISPGPI-------FNPTP-ISTH--PPSPSPSSHSHNTYHSISVEPVTSQ 1000
AS S S+ PG F +P + TH S SPS SH T S + V S
Sbjct: 826 ASPSASDTAASQPGAATSLDVVHFPASPDVMTHHSADSSSPSQPSHATATPASNDDVPSP 885
Query: 1001 PSTQA-EPHRIHPNNTHSMA---TRAKHGIVQKRKHP--------TLLLTHIEPTGYRQA 1048
A EP T + + TR + GI +++ + T L EP A
Sbjct: 886 SHISASEPFATGEEVTQAPSRPTTRLQRGIRKEKVYTDGTVKYKNTFLTVTGEPKNLTDA 945
Query: 1049 MKQPQWLQAMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKARLVA 1108
++ W +AM +E+EALM N TW LVP R + CKWV++ K+ DGS+++YKARLVA
Sbjct: 946 LQNTNWKKAMDIEYEALMNNKTWHLVPPKQGRNVIDCKWVYKIKRKQDGSLDRYKARLVA 1005
Query: 1109 KGFHQMPGFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQ 1168
KGF Q G DY++TFSPVVK T+R VL++AV+ WC++QLDV NAFL+G+LEEEVYM Q
Sbjct: 1006 KGFKQRYGIDYEDTFSPVVKAATIRIVLSIAVSRGWCMRQLDVQNAFLHGFLEEEVYMKQ 1065
Query: 1169 PPGFEAVD-PSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQH 1227
PPG+E P VCKL+KALYGLKQAPRAW+ RL L LGF SK D SLF +
Sbjct: 1066 PPGYEDESFPGYVCKLDKALYGLKQAPRAWYSRLSKKLYDLGFQGSKGDTSLFFYNKGGL 1125
Query: 1228 STFMLVYVDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQ 1287
F+L+YVDDI++T S + L++ L EF+LKDLG L YFLGIEV+ ++L+Q
Sbjct: 1126 IIFVLIYVDDIIVTSSRQEAVSALLQDLKKEFALKDLGDLHYFLGIEVN-KVTDEIILTQ 1184
Query: 1288 KKYIQDLLVKANMANANGIASPMASSTKLTKYGSNHVS--DPTFFRSIVGGLQYVTVTRP 1345
KY DLL + NM + +++P+++S KL+ + + + D T +RS+VG LQY+T+TRP
Sbjct: 1185 DKYACDLLRRVNMFDCKPVSTPLSTSEKLSAHEGDLLGPLDATNYRSVVGALQYLTLTRP 1244
Query: 1346 EISYSVNKVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADW 1405
+I++ VNKVCQFL AP HW AVKRILRYLK GL + + + + ++A+ DADW
Sbjct: 1245 DIAFPVNKVCQFLHAPTTVHWAAVKRILRYLKQCTKLGLKLCKS---KSMLVSAYSDADW 1301
Query: 1406 ASDPDDRRSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKE 1465
A DDRRST G + LG NL+SW A+KQ V+RSS E+EY++LA A+AE++W+Q+LL E
Sbjct: 1302 AGSLDDRRSTGGFAVFLGDNLVSWCARKQATVSRSSTESEYKALANATAEIMWVQTLLTE 1361
Query: 1466 LKVPT-AIPQIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQ 1524
L+V + + +++CDNL L+ NPV H+RTKH+E+D FVRE+V K L + +P Q
Sbjct: 1362 LQVQSPPMAKLWCDNLGAKYLSSNPVFHARTKHIEVDYHFVRERVSQKLLEIDFVPTGDQ 1421
Query: 1525 VADILTKPLSASRFLELRNKLRV 1547
VAD TK L + ++ L +
Sbjct: 1422 VADGFTKALPVRQLENFKHNLNL 1444
Score = 113 bits (282), Expect = 5e-23
Identities = 88/320 (27%), Positives = 144/320 (44%), Gaps = 43/320 (13%)
Query: 17 TAAASKNFKQIISVKLDETNYLQWKQQVEGVLRGTKMVRHVVSPQIPPVFL------NDA 70
TAA + IS KL ++N+ WK QV +RG ++ H+ P L +
Sbjct: 7 TAAGNPLGGNAISEKLSKSNHALWKAQVMAAVRGARLEGHLTGATKTPNALITTTAGDKG 66
Query: 71 AREAGTENPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRT 130
+E NP + +W D + ++LST++ +L++ + W + + R
Sbjct: 67 EKEVTVRNPEFDDWVATDQQVLGFLLSTLARDVLAQVATCGTAAAAWQMLEEMYSSVTRA 126
Query: 131 RSRQLRSELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPI 190
R R L KGT SI E++++++++++ + + G V DLI ++ L + + P+
Sbjct: 127 RFINTRIALSNTKKGTLSINEYVSKMKALADEMTAAGKIVDDDDLISYIIAGLDDTYEPV 186
Query: 191 VATVNSQTEVISLDELESQLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQT 250
++T+ + + ++L E SQLL+ E R G +SVNL N G G Q +
Sbjct: 187 ISTIVGK-DTMTLGEAYSQLLSFEQR----LALRHGGDSSVNLA---NRGRGGGGGQQRG 238
Query: 251 GSYPDQQFNISGNPTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGR--GNI-- 306
G+ TGN GGR G G NR RGRG R G +
Sbjct: 239 GN------------TGN----------GGR-GRGGNNNGANRGRGRGNNGGARPPGGVDN 275
Query: 307 --QCQICYKTGHDASICYHR 324
+CQ+CYK GH C++R
Sbjct: 276 RPKCQLCYKRGHTVINCWYR 295
>UniRef100_O24438 Retrofit [Oryza longistaminata]
Length = 1445
Score = 835 bits (2158), Expect = 0.0
Identities = 500/1277 (39%), Positives = 712/1277 (55%), Gaps = 125/1277 (9%)
Query: 333 GYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYGAP--RAPFPPQFGNSRPP 390
G G GG G N SG G QG G R G R F
Sbjct: 231 GRGGGGGGRGNNNNSGGGGGRSAPGGRGSGSQGRGGRGRGTGGQDRRPTCQVCFKRGHTA 290
Query: 391 AP------QAYITGNE----STSSNSFNNGWYPDSGATHHVTPDANNLMDAASFSGSDQM 440
A + Y+ + +T+S + WY D+GAT H+T + L ++G +Q+
Sbjct: 291 ADCWYRFDEDYVADEKLVAAATNSYGIDTNWYIDTGATDHITGELEKLTTKEKYNGGEQI 350
Query: 441 YIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFH 500
+ +G G+ I+ IG + +P+ + LNN+L+VP KNL+S SQ DN+ F E H
Sbjct: 351 HTASGAGMDISHIGH---TIVHTPSRNIHLNNVLYVPQAKKNLISASQLAADNSAFLELH 407
Query: 501 SNICYVKSQDSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVATSSLSLNNCFSPSS 560
S +K Q + +LL+G GLY + + S ++ A ++
Sbjct: 408 SKFFSIKDQVTRDVLLEGKCRH-GLYPIPKFFGRSTNKQALGAA---------------K 451
Query: 561 LSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKL----CN 616
LSLSR WH+RLGHP +V+ V+ C+
Sbjct: 452 LSLSR------------------------------WHSRLGHPSLPIVKQVISRNNLPCS 481
Query: 617 QQLPNKSFTDFCSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLI 676
+ N+S C+AC KSH+LP + S +V P EL+F D+WGPA ES G Y++
Sbjct: 482 VESVNQSV---CNACQEAKSHQLPYIRSTSVSQFPLELVFSDVWGPAP-ESVGRNKYYVS 537
Query: 677 CVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDG-GGEFRPFTQFLTT 735
+D +S++TWI+ LK KS F+ F+ +VE ++ I ++QTD GG ++ F
Sbjct: 538 FIDDFSKFTWIYLLKYKSEVFEKFKEFQALVERMFDRKIIAMQTDWRGGRYQKLNSFFAQ 597
Query: 736 LGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPS 795
+G+ QNGS ERKHRHIVE GL+LL+ A +PL +WD AF+ ATYLINR+PS
Sbjct: 598 IGLIIMCHVLTLIRQNGSAERKHRHIVEVGLSLLSYASMPLKFWDEAFVAATYLINRIPS 657
Query: 796 PILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGY 855
+ N +P L Q PDY L+ FGC+C+P RPYN HKL+ RSK+CVFLG+S HKG+
Sbjct: 658 KTIQNSTPLEKLFNQKPDYSSLRVFGCACWPHLRPYNTHKLQFRSKQCVFLGFSTHHKGF 717
Query: 856 KCLD-PTGRMFISKDVIFNEYKFPYSELFTSGQSSSPPTTSSDHTPLPSFLFPLNNKQCP 914
KCLD +GR++IS+DV+F+E FP+S L S++ S+ LPS PL N
Sbjct: 718 KCLDVSSGRVYISRDVVFDENVFPFSTL----HSNAGARLRSEILLLPS---PLTNYNTA 770
Query: 915 TTQSSSTPTTTLHTASPHSSFPES--------NQSNHHHSI---QDTHASSHSN--HHNI 961
+ + +T P + + N + H + Q+ H N H +
Sbjct: 771 SAGGTHVVAPVANTPLPSDNLISNAADVTSGENSAAHEQEMENEQEIENVMHGNDVHGDA 830
Query: 962 SPGPIFN-PTPISTHPPSPSP---------SSHSHNTYHSISVEPVTSQPSTQAEPHRIH 1011
+ GP+ + PT S+ P +S + ++ S + E +
Sbjct: 831 ASGPVLDQPTADSSTAPDQGADTSDAVSGAASDAGGDTATLGAGAANSAAAGGEESQPVQ 890
Query: 1012 PNNTHSMA----------TRAKHGIVQKRKHPTLLLTHI------EPTGYRQAMKQPQWL 1055
P+ T ++ TR + GI +++ + + + EP ++A+ W
Sbjct: 891 PDVTGTVLATVAPASRPHTRLRSGIRKEKVYTDGTVKYGCFSSTGEPQNDKEALGDKNWR 950
Query: 1056 QAMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMP 1115
AM+ E+ AL+KN+TW LVP + +GCKWV++ K+ DG++++YKARLVAKGF Q
Sbjct: 951 DAMETEYNALIKNDTWHLVPYEKGQNIIGCKWVYKIKRKADGTLDRYKARLVAKGFKQRY 1010
Query: 1116 GFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAV 1175
G DY++TFSPVVK T+R +L++AV+ W ++QLDV NAFL+G+LEEEVYM QPPGFE+
Sbjct: 1011 GIDYEDTFSPVVKAATIRIILSIAVSRGWSLRQLDVQNAFLHGFLEEEVYMQQPPGFESS 1070
Query: 1176 D-PSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVY 1234
P VCKL+KALYGLKQAPRAW+ RL L++LGF +SK D SLF L+ F+LVY
Sbjct: 1071 SKPDYVCKLDKALYGLKQAPRAWYSRLSKKLVELGFEASKADTSLFFLNKGGILMFVLVY 1130
Query: 1235 VDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDL 1294
VDDI++ S+ L+K LN EF+LKDLG L YFLGIEV NG ++L+Q+KY DL
Sbjct: 1131 VDDIIVASSTEKATTALLKDLNKEFALKDLGDLHYFLGIEVTKVSNG-VILTQEKYANDL 1189
Query: 1295 LVKANMANANGIASPMASSTKLTKYGSNHV--SDPTFFRSIVGGLQYVTVTRPEISYSVN 1352
L + NM+N +++P++ S KLT Y + + +D +RSIVG LQY+T+TRP+I+YSVN
Sbjct: 1190 LKRVNMSNCKPVSTPLSVSEKLTLYEGSPLGPNDAIQYRSIVGALQYLTLTRPDIAYSVN 1249
Query: 1353 KVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDR 1412
KVCQFL AP HW AVKRILRYL GL I+ + + + DADWA DDR
Sbjct: 1250 KVCQFLHAPTTSHWIAVKRILRYLNQCTSLGLHIHKS---ASTLVHGYSDADWAGSIDDR 1306
Query: 1413 RSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVPT-A 1471
+ST G + LG NL+SW A+KQ V+RSS EAEY+++A +AE++W+Q+LLKEL + +
Sbjct: 1307 KSTGGFAVFLGSNLVSWSARKQPTVSRSSTEAEYKAVANTTAELIWVQTLLKELGIESPK 1366
Query: 1472 IPQIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTK 1531
+I+CDNL L+ NPV H+RTKH+E+D FVRE+V K L + +P+ QVAD TK
Sbjct: 1367 AAKIWCDNLGAKYLSANPVFHARTKHIEVDYHFVRERVSQKLLEIDFVPSGDQVADGFTK 1426
Query: 1532 PLSASRFLELRNKLRVS 1548
LSA ++ L ++
Sbjct: 1427 ALSACLLENFKHNLNLA 1443
Score = 139 bits (349), Expect = 9e-31
Identities = 90/317 (28%), Positives = 154/317 (48%), Gaps = 37/317 (11%)
Query: 17 TAAASKNFKQ--IISVKLDETNYLQWKQQVEGVLRGTKMVRHV---VSPQIPPVFLNDAA 71
+ AA+ N Q +S KL + N+ WK QV +RG +++ ++ + + +
Sbjct: 8 SGAAAANLLQGHSVSEKLGKANHALWKAQVSAAVRGARLLGYLNGDIKAPDAELSVTIDG 67
Query: 72 REAGTENPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTR 131
+ NPA+ +WE D L+ ++LS++S +L + + + + W I T R R
Sbjct: 68 KTTTKPNPAFEDWEANDQLVLGYLLSSLSRDVLIQVATCKTAAEAWRSIEALYSTGTRAR 127
Query: 132 SRQLRSELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIV 191
+ R L KGT IAE++A++R++ + + + G P+ DL++ ++ L E+F+PIV
Sbjct: 128 AVNTRLALTNTKKGTMKIAEYVAKMRALGDEMAAGGHPLDEEDLVQYIIAGLNEDFSPIV 187
Query: 192 ATVNSQTEVISLDELESQLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTG 251
+ + ++++ I++ EL SQL+ E + ++ G A V N G G
Sbjct: 188 SNLCNKSDPITVGELYSQLVNFETLLDLYRSTGQGGAAFV-----ANRGRGGG------- 235
Query: 252 SYPDQQFNISGNPTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQ---- 307
G GNN+ N GG G G RG+ +GRGGR G G
Sbjct: 236 ----------GGGRGNNN------NSGGGGGRSAPGGRGSGSQGRGGRGRGTGGQDRRPT 279
Query: 308 CQICYKTGHDASICYHR 324
CQ+C+K GH A+ C++R
Sbjct: 280 CQVCFKRGHTAADCWYR 296
>UniRef100_Q75LJ1 Putative copia-like retrotransposon protein [Oryza sativa]
Length = 1399
Score = 832 bits (2150), Expect = 0.0
Identities = 466/1173 (39%), Positives = 671/1173 (56%), Gaps = 86/1173 (7%)
Query: 401 STSSNSFNNGWYPDSGATHHVTPDANNLMDAASFSGSDQMYIGNGQGLAINSIGSMSFSS 460
+ ++ + WY D+GAT H+T + + L + G D+++ +G G+ I IG S
Sbjct: 278 AAAAYGIDTNWYLDTGATDHITNELDKLDVREKYKGGDKIHTASGAGMEIKHIGD---SV 334
Query: 461 PFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHL 520
+P L L N+LHVP KNL+S + DN F E HSN +K + + +LKG
Sbjct: 335 IHTPTRELHLKNILHVPQAKKNLISAHRLAMDNFAFLEVHSNYFLIKDRATRNTILKG-- 392
Query: 521 GDDGLYQFDQPYVPSVSRTASSSSVATSSLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTS 580
+C P TS
Sbjct: 393 ----------------------------------------------RCRRRLYSLP--TS 404
Query: 581 GSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQLPNKSFTD-FCSACCLGKSHRL 639
+ + + + WH+RLGHP +V V+ N + D C AC GKSH+L
Sbjct: 405 PAKQVHAATTPSFSRWHSRLGHPAVPIVTRVLSKNNLPCSTVANKDSICDACQKGKSHQL 464
Query: 640 PSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSRYTWIFPLKLKSHTLIT 699
P S +V ++P ELIF ++WGPA + S GG +++ +D YS++TW++ LK KS
Sbjct: 465 PYPKSSSVSSQPLELIFSNVWGPAPI-SVGGKKFYVSFIDDYSKFTWVYLLKHKSEVFQK 523
Query: 700 FQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCPHTHHQNGSVERKHR 759
FQ F+T+VE + I +VQTD GGE++ F +GI+H ++CP+ H QNGS ERKHR
Sbjct: 524 FQEFQTLVERFFGHKILAVQTDWGGEYQKLNTFFAKIGISHHVSCPYAHQQNGSAERKHR 583
Query: 760 HIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKS 819
H++E LTLLA+A +P+ +WD A L A YLINR PS ++N P L + P+Y L+
Sbjct: 584 HLIEVALTLLAHASMPIKFWDEAVLAAAYLINRTPSKVINFACPLEQLFKEKPNYTALRI 643
Query: 820 FGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLD-PTGRMFISKDVIFNEYKFP 878
FGC+ +P RPYN HKL RSK CVFLGYS HKG+KCL+ TGR+++S+DV F+E FP
Sbjct: 644 FGCAVWPNLRPYNKHKLAFRSKRCVFLGYSNLHKGFKCLEIATGRVYVSRDVTFDESIFP 703
Query: 879 YSELFTSGQSSSPPTTSSDHTPLPSFLFPLNNKQCPTTQSSSTPTTTLHTASPHSSFPES 938
+SEL ++ + S L L L +Q + P T ++ E
Sbjct: 704 FSELHSNAGARLRAEISLLPPSLVPHLSSLGGEQ-NNHVLNYPPNVTDQFGEENAEIGEE 762
Query: 939 NQSNHHHSI-----QDTHASSHSNHHNISPGPIFNPTPISTHPPSPSPSSHSHNTY---- 989
+N + ++ A+++ + G ++ +P + P + + + +
Sbjct: 763 IVANGEENAAAAADENAAAAANGGAQDDVHGVAYDASPEHSSPVTDDAMASAAEQHGNPI 822
Query: 990 ---HSISVEPVTSQPSTQ--AEPHRIHPNNTHSMATRAKHGIVQKRKHPTLLLTHIEPTG 1044
H + P T+ ++ A +H + T + + + + P T ++ +G
Sbjct: 823 QEEHLVQASPQTASSTSPSVASSAGVHDDVTTDQSDQTDQAMPEAAVAPIRPKTRLQ-SG 881
Query: 1045 YR------QAMKQPQWLQAMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGS 1098
R +A+ W +AM E+ AL++N TW LVP R + CKWV++ K+ DGS
Sbjct: 882 IRKEKSLEEAVNNKHWKEAMDAEYMALIENKTWHLVPPQKGRNVIDCKWVYKVKRKADGS 941
Query: 1099 INKYKARLVAKGFHQMPGFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNG 1158
+++YKARLVAKGF Q G DY++TFSPVVK T+R VL+LAV+ W ++QLDV NAFL+G
Sbjct: 942 LDRYKARLVAKGFKQRYGIDYEDTFSPVVKAATIRIVLSLAVSRGWSLRQLDVKNAFLHG 1001
Query: 1159 YLEEEVYMTQPPGFEAVD-PSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDP 1217
LEEEVYM QPPG+E P+ VCKL+KALYGLKQAPRAW+ RL + L +LGF SK D
Sbjct: 1002 VLEEEVYMKQPPGYEKKSMPNYVCKLDKALYGLKQAPRAWYSRLSTKLSELGFVPSKADT 1061
Query: 1218 SLFILHANQHSTFMLVYVDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHY 1277
SLF Q S F+L+YVDDI++ S L+++L+ +F+LKDLG L YFLGIEVH
Sbjct: 1062 SLFFYKKGQVSIFLLIYVDDIIVASSVPDATSTLLQELSKDFALKDLGDLHYFLGIEVHK 1121
Query: 1278 SENGSLLLSQKKYIQDLLVKANMANANGIASPMASSTKLTKYGSNHV--SDPTFFRSIVG 1335
++G L+LSQ+KY DLL + M +++P+++S KL+ + D T +RS+VG
Sbjct: 1122 VKDG-LMLSQEKYASDLLRRVGMYECKPVSTPLSTSEKLSVNEGTLLGPQDSTQYRSVVG 1180
Query: 1336 GLQYVTVTRPEISYSVNKVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPL 1395
LQY+T+TRP+IS+S+NKVCQFL AP HW AVKRILRY+K T+ GL P L
Sbjct: 1181 ALQYLTLTRPDISFSINKVCQFLHAPTTTHWAAVKRILRYVKYTVDTGLKFCRNP---SL 1237
Query: 1396 SLTAFCDADWASDPDDRRSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAE 1455
++ F DADWA PDDRRST G + LGPNL+SW A+KQ V+RSS EAEY++LA A+AE
Sbjct: 1238 LVSGFSDADWAGSPDDRRSTGGFAVFLGPNLVSWSARKQATVSRSSIEAEYKALANATAE 1297
Query: 1456 VLWIQSLLKELKVPT-AIPQIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDL 1514
++W+Q+LL+EL V + +++CDNL L+ NP+ H+RTKH+E+D FVRE+V K L
Sbjct: 1298 IMWVQTLLQELGVESPRAAKLWCDNLGAKYLSANPIFHARTKHIEVDFHFVRERVARKLL 1357
Query: 1515 IVSHIPAQYQVADILTKPLSASRFLELRNKLRV 1547
+++I + QVAD TK + + +N L +
Sbjct: 1358 EIAYISTKDQVADGFTKAIPVRQMEMFKNNLNL 1390
Score = 109 bits (272), Expect = 8e-22
Identities = 64/219 (29%), Positives = 114/219 (51%), Gaps = 5/219 (2%)
Query: 17 TAAASKNFKQIISVKLDETNYLQWKQQVEGVLRGTKMVRHVV-SPQIPPVFLNDAAREAG 75
TA+ + F +S KL + NY W QV +RG ++ H+ + P + + A +
Sbjct: 8 TASTNPLFGVQVSEKLTKGNYALWSAQVLAAIRGARLDGHITGATAAPSMEIEKTASDKT 67
Query: 76 TE---NPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRS 132
TE NPAY EW D + ++LST+S +L++ + Q W ++ Q + RS
Sbjct: 68 TEKIVNPAYQEWFASDQQVLGFLLSTLSRDILTQVATASTAAQAWQQVCAMFTAQTKARS 127
Query: 133 RQLRSELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVA 192
+R L KG SI+E+ +++++++ + S G P+ DL+ VL L ++F P+V+
Sbjct: 128 LNVRLTLTNTQKGNMSISEYCGKMKALADEIASSGKPLDEEDLVAYVLNGLDDDFEPVVS 187
Query: 193 TVNSQTEVISLDELESQLLTQEARNEKFKKALVGETASV 231
+ ++ E ++ E+ SQLL E R + ++A A+V
Sbjct: 188 AIVARNESTTMAEVYSQLLNFENR-QALRQAHASANAAV 225
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.132 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,760,775,631
Number of Sequences: 2790947
Number of extensions: 126048998
Number of successful extensions: 490591
Number of sequences better than 10.0: 4293
Number of HSP's better than 10.0 without gapping: 1598
Number of HSP's successfully gapped in prelim test: 2886
Number of HSP's that attempted gapping in prelim test: 454951
Number of HSP's gapped (non-prelim): 18512
length of query: 1557
length of database: 848,049,833
effective HSP length: 141
effective length of query: 1416
effective length of database: 454,526,306
effective search space: 643609249296
effective search space used: 643609249296
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 82 (36.2 bits)
Medicago: description of AC149038.3


