

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148996.15 + phase: 0
(153 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
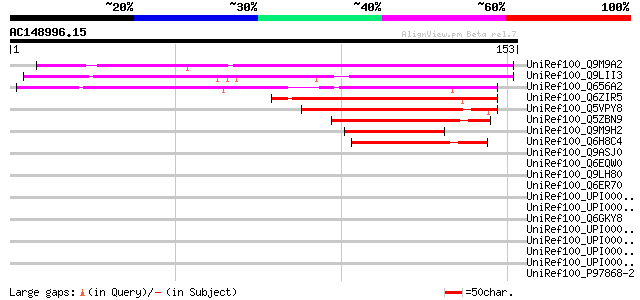
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9M9A2 F27J15.21 [Arabidopsis thaliana] 112 1e-24
UniRef100_Q9LII3 Gb|AAF43227.1 [Arabidopsis thaliana] 97 9e-20
UniRef100_Q656A2 Hypothetical protein P0596H06.16 [Oryza sativa] 71 5e-12
UniRef100_Q6ZIR5 Hypothetical protein OJ1038_A06.25 [Oryza sativa] 67 1e-10
UniRef100_Q5VPY8 Hypothetical protein B1096D03.33 [Oryza sativa] 50 8e-06
UniRef100_Q5ZBN9 Hypothetical protein P0704D04.8 [Oryza sativa] 47 8e-05
UniRef100_Q9M9H2 F14O23.12 [Arabidopsis thaliana] 47 8e-05
UniRef100_Q6H8C4 Hypothetical protein OJ1006_A02.44 [Oryza sativa] 44 5e-04
UniRef100_Q9ASJ0 Hypothetical protein P0439B06.27 [Oryza sativa] 42 0.003
UniRef100_Q6EQW0 Hypothetical protein OSJNBa0086N11.28 [Oryza sa... 39 0.030
UniRef100_Q9LH80 Gb|AAF43227.1 [Arabidopsis thaliana] 37 0.087
UniRef100_Q6ER70 Hypothetical protein OSJNBa0073A18.24 [Oryza sa... 37 0.11
UniRef100_UPI00002FEC1A UPI00002FEC1A UniRef100 entry 35 0.25
UniRef100_UPI000043923D UPI000043923D UniRef100 entry 35 0.33
UniRef100_Q6GKY8 AT12465p [Drosophila melanogaster] 35 0.33
UniRef100_UPI00001CE9FC UPI00001CE9FC UniRef100 entry 34 0.57
UniRef100_UPI00001931EA UPI00001931EA UniRef100 entry 34 0.57
UniRef100_UPI000017E863 UPI000017E863 UniRef100 entry 34 0.57
UniRef100_UPI00000235D8 UPI00000235D8 UniRef100 entry 34 0.57
UniRef100_P97868-2 Splice isoform 2 of P97868 [Mus musculus] 34 0.57
>UniRef100_Q9M9A2 F27J15.21 [Arabidopsis thaliana]
Length = 156
Score = 112 bits (281), Expect = 1e-24
Identities = 64/149 (42%), Positives = 89/149 (58%), Gaps = 9/149 (6%)
Query: 9 SPNRHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRD-----EEWRFDLKTPP 63
SP+ + F V++ + L +R ANRL +K K S + E +R+++ +
Sbjct: 11 SPSHKRLSVSFLVSM---MVLCARHANRLSKKLKLKSKKRTCSGGEGGGERFRWNMISSS 67
Query: 64 KSPMAKPKKLLSNISNKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDFSGV 123
S +PK+L + +SNKA++ +K EE+ G+WQ+EILMGGKCEPLDFSGV
Sbjct: 68 MSS-PRPKELFTTLSNKAMTMVRRKNPPEEKATAMEEEHGLWQREILMGGKCEPLDFSGV 126
Query: 124 IYYDINGKQTREVPIRSPRASPLPGYLTR 152
IYYD NG+ EVP RSPR +PLP Y TR
Sbjct: 127 IYYDSNGRLLNEVPPRSPRGTPLPSYPTR 155
>UniRef100_Q9LII3 Gb|AAF43227.1 [Arabidopsis thaliana]
Length = 195
Score = 96.7 bits (239), Expect = 9e-20
Identities = 67/183 (36%), Positives = 96/183 (51%), Gaps = 40/183 (21%)
Query: 5 TSPESPNRHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRDEEWRFDLKT--- 61
++PE + H +S ++ L +R A+R+ +K K T K E++ K+
Sbjct: 4 STPEHVSSAHKRISVSFLVS-LMVLCARHASRVSKKLKPKKTRKQTHLEDYLESPKSNGN 62
Query: 62 --------------PPK--SPM-AKPKKLLSNISNKALSQFGKKKQR------------- 91
P + SPM +PK+L + +SNKA++ G+K +
Sbjct: 63 GSEDGRGGGRFGWSPARTFSPMRVRPKELYTTLSNKAMTMVGRKNKAYDGGPTKKTAVEM 122
Query: 92 --EEREKEGWGNGGVWQKEILMGGKCEPLDFSGVIYYDINGKQTREVPIRSPRASPLPGY 149
EE E+E GVWQ+EILMGGKCEPLD+SGVIYYD +G Q ++VP RSPRAS +P
Sbjct: 123 VMEEDEEEY----GVWQREILMGGKCEPLDYSGVIYYDCSGHQLKQVPPRSPRASLVPER 178
Query: 150 LTR 152
TR
Sbjct: 179 PTR 181
>UniRef100_Q656A2 Hypothetical protein P0596H06.16 [Oryza sativa]
Length = 171
Score = 70.9 bits (172), Expect = 5e-12
Identities = 49/147 (33%), Positives = 76/147 (51%), Gaps = 13/147 (8%)
Query: 3 KITSPESPNRHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRDEEWRFDLKTP 62
K T+ + +R GG + + A+ AL+S RL+ A+ S+ ++
Sbjct: 23 KATTAAASDRKVMAGGASL-LHAVAALMSTCTRRLQRAARRVSSAAAGAGKQGASSRAVV 81
Query: 63 P-KSPMAKPKKLLSNISNKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDFS 121
P + ++ P + + A + R +EG +GG+W+KEILMG +C+PLDFS
Sbjct: 82 PWRKALSLPAAATAKVKAAAAAA---------RREEG-DSGGLWRKEILMGERCQPLDFS 131
Query: 122 GVIYYDINGKQ-TREVPIRSPRASPLP 147
GVIYYD +G++ P RSP SPLP
Sbjct: 132 GVIYYDADGRRLAHPPPPRSPMRSPLP 158
>UniRef100_Q6ZIR5 Hypothetical protein OJ1038_A06.25 [Oryza sativa]
Length = 138
Score = 66.6 bits (161), Expect = 1e-10
Identities = 34/69 (49%), Positives = 46/69 (66%), Gaps = 2/69 (2%)
Query: 80 KALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDFSGVIYYDINGKQTRE-VPI 138
KA+S +++++ E GVW+KEILMG +C+PLDFSGVIYYD G++ + P
Sbjct: 56 KAISS-SRRRRKAGAELSFRAEDGVWRKEILMGERCQPLDFSGVIYYDAEGRRLEQPPPP 114
Query: 139 RSPRASPLP 147
RSP SPLP
Sbjct: 115 RSPLRSPLP 123
>UniRef100_Q5VPY8 Hypothetical protein B1096D03.33 [Oryza sativa]
Length = 138
Score = 50.4 bits (119), Expect = 8e-06
Identities = 28/62 (45%), Positives = 39/62 (62%), Gaps = 5/62 (8%)
Query: 89 KQREEREKEGWGNGGVWQKEILMGGKCEPLDFSGVIYYDINGKQTREVPIRSPRA---SP 145
++R +E GV +KEILM +C+ LDFSG+IYYD+ G++ + P PRA SP
Sbjct: 66 RRRRRQELSFRVEDGVCRKEILMEERCQSLDFSGMIYYDVAGRRLEQPP--PPRALLHSP 123
Query: 146 LP 147
LP
Sbjct: 124 LP 125
>UniRef100_Q5ZBN9 Hypothetical protein P0704D04.8 [Oryza sativa]
Length = 505
Score = 47.0 bits (110), Expect = 8e-05
Identities = 21/48 (43%), Positives = 30/48 (61%), Gaps = 2/48 (4%)
Query: 98 GWGNGGVWQKEILMGGKCEPLDFSGVIYYDINGKQTREVPIRSPRASP 145
G G G+W++ ILMG +CEPL F G I+YD G++ + R +A P
Sbjct: 422 GGGGEGLWRRAILMGERCEPLSFPGAIHYDSRGRRLSQP--RRAKAKP 467
>UniRef100_Q9M9H2 F14O23.12 [Arabidopsis thaliana]
Length = 130
Score = 47.0 bits (110), Expect = 8e-05
Identities = 20/30 (66%), Positives = 24/30 (79%)
Query: 102 GGVWQKEILMGGKCEPLDFSGVIYYDINGK 131
G +WQK ILMGGKC+ DFSGVI YD +G+
Sbjct: 86 GPLWQKNILMGGKCQLPDFSGVILYDADGQ 115
>UniRef100_Q6H8C4 Hypothetical protein OJ1006_A02.44 [Oryza sativa]
Length = 299
Score = 44.3 bits (103), Expect = 5e-04
Identities = 23/41 (56%), Positives = 26/41 (63%), Gaps = 2/41 (4%)
Query: 104 VWQKEILMGGKCEPLDFSGVIYYDINGKQTREVPIRSPRAS 144
VWQK ILMGGKC+ +FSGVI YD G P PRA+
Sbjct: 98 VWQKNILMGGKCQLPEFSGVINYDAAGNIV--APSGRPRAA 136
>UniRef100_Q9ASJ0 Hypothetical protein P0439B06.27 [Oryza sativa]
Length = 175
Score = 42.0 bits (97), Expect = 0.003
Identities = 19/41 (46%), Positives = 24/41 (58%)
Query: 104 VWQKEILMGGKCEPLDFSGVIYYDINGKQTREVPIRSPRAS 144
+W+K I+MG KC PL FSG I YD +G Q I A+
Sbjct: 126 LWRKTIMMGDKCRPLQFSGHIAYDSDGNQLPATTISKEAAN 166
>UniRef100_Q6EQW0 Hypothetical protein OSJNBa0086N11.28 [Oryza sativa]
Length = 202
Score = 38.5 bits (88), Expect = 0.030
Identities = 17/31 (54%), Positives = 22/31 (70%)
Query: 104 VWQKEILMGGKCEPLDFSGVIYYDINGKQTR 134
VWQ+ ILMG +CE FSG+I YD +G+ R
Sbjct: 138 VWQRRILMGMRCELPRFSGLILYDEHGRPIR 168
>UniRef100_Q9LH80 Gb|AAF43227.1 [Arabidopsis thaliana]
Length = 168
Score = 37.0 bits (84), Expect = 0.087
Identities = 31/98 (31%), Positives = 41/98 (41%), Gaps = 14/98 (14%)
Query: 44 SSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNISNKALSQF-----GKKKQREERE--- 95
+S K + D EW + ++ K K S + + F K + E R
Sbjct: 42 NSRGKQVADAEWASETAFGSRTMEEKRKCKWSKVKRTLMGSFCWSSAAKWMEMETRRQPP 101
Query: 96 ----KEGWGNG--GVWQKEILMGGKCEPLDFSGVIYYD 127
KE N VWQ+ ILMG KCE FSG+I YD
Sbjct: 102 LLAVKERSLNAVDPVWQRPILMGEKCELPRFSGLILYD 139
>UniRef100_Q6ER70 Hypothetical protein OSJNBa0073A18.24 [Oryza sativa]
Length = 143
Score = 36.6 bits (83), Expect = 0.11
Identities = 14/28 (50%), Positives = 21/28 (75%)
Query: 104 VWQKEILMGGKCEPLDFSGVIYYDINGK 131
+WQ+++LMG KC+ FSG+I YD G+
Sbjct: 89 IWQRKVLMGVKCQLPRFSGMILYDERGR 116
>UniRef100_UPI00002FEC1A UPI00002FEC1A UniRef100 entry
Length = 227
Score = 35.4 bits (80), Expect = 0.25
Identities = 21/81 (25%), Positives = 37/81 (44%)
Query: 34 ANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNISNKALSQFGKKKQREE 93
+N + K + + + E +F K P+ K KK++S+ ++L + GK KQ +
Sbjct: 43 SNNFRFGNKRQGDVQSLIESEKKFHFKVIKPKPLIKYKKIVSSSLIRSLLEKGKLKQANK 102
Query: 94 REKEGWGNGGVWQKEILMGGK 114
W G+ QK +G K
Sbjct: 103 LLNRNWTINGIVQKGRQLGKK 123
>UniRef100_UPI000043923D UPI000043923D UniRef100 entry
Length = 264
Score = 35.0 bits (79), Expect = 0.33
Identities = 21/66 (31%), Positives = 34/66 (50%), Gaps = 2/66 (3%)
Query: 32 RKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNISNKALSQFGKKKQR 91
+KAN+ +EK K + K + E R ++ K K +K + K + KK+Q+
Sbjct: 173 KKANKKEEKKKEKKSKKERKKE--RATKRSKRKRKQIKQQKEKKRVKKKIKTNRKKKRQK 230
Query: 92 EEREKE 97
EER+KE
Sbjct: 231 EERKKE 236
>UniRef100_Q6GKY8 AT12465p [Drosophila melanogaster]
Length = 742
Score = 35.0 bits (79), Expect = 0.33
Identities = 29/138 (21%), Positives = 57/138 (41%), Gaps = 4/138 (2%)
Query: 20 FVTISAILALLSRKANRLK-EKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNIS 78
FV +L + R+ RLK + + D E FD P + + +P++L S+
Sbjct: 500 FVDKKRLLYHVRRECERLKIDSCEDELVLTDTSDSEDEFDFGVTPPAGLMRPERLKSSHV 559
Query: 79 NKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDF---SGVIYYDINGKQTRE 135
+Q+ + + V + +MG + +P ++ GVI + + R
Sbjct: 560 KHTETQYNESDWVNPSLYPHMPDATVQYESCVMGEREKPFNWIYGKGVIQEEPKLPKMRN 619
Query: 136 VPIRSPRASPLPGYLTRG 153
+P + + P+PG + G
Sbjct: 620 IPKKPKKKKPVPGRVAGG 637
>UniRef100_UPI00001CE9FC UPI00001CE9FC UniRef100 entry
Length = 1659
Score = 34.3 bits (77), Expect = 0.57
Identities = 22/67 (32%), Positives = 35/67 (51%), Gaps = 4/67 (5%)
Query: 32 RKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKS--PMAKPKKLLSNISNKALSQFGKKK 89
RKA+ + AK KP+++E+ + D KS P +K +K NK L G+K+
Sbjct: 959 RKAH--SKSAKEHQEAKPVKEEKAKKDCSKDIKSEKPASKDEKAKKPEKNKLLESKGEKR 1016
Query: 90 QREEREK 96
+R+ EK
Sbjct: 1017 KRKTEEK 1023
>UniRef100_UPI00001931EA UPI00001931EA UniRef100 entry
Length = 1556
Score = 34.3 bits (77), Expect = 0.57
Identities = 23/67 (34%), Positives = 34/67 (50%), Gaps = 4/67 (5%)
Query: 32 RKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKS--PMAKPKKLLSNISNKALSQFGKKK 89
RKA+ + AK KP +DE+ + D KS P +K +K NK L G+K+
Sbjct: 857 RKAH--SKSAKEHQEAKPAKDEKVKKDCSKDIKSEKPASKDEKAKKPEKNKLLDSKGEKR 914
Query: 90 QREEREK 96
+R+ EK
Sbjct: 915 KRKTEEK 921
>UniRef100_UPI000017E863 UPI000017E863 UniRef100 entry
Length = 1590
Score = 34.3 bits (77), Expect = 0.57
Identities = 22/67 (32%), Positives = 35/67 (51%), Gaps = 4/67 (5%)
Query: 32 RKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKS--PMAKPKKLLSNISNKALSQFGKKK 89
RKA+ + AK KP+++E+ + D KS P +K +K NK L G+K+
Sbjct: 890 RKAH--SKSAKEHQEAKPVKEEKAKKDCSKDIKSEKPASKDEKAKKPEKNKLLESKGEKR 947
Query: 90 QREEREK 96
+R+ EK
Sbjct: 948 KRKTEEK 954
>UniRef100_UPI00000235D8 UPI00000235D8 UniRef100 entry
Length = 1590
Score = 34.3 bits (77), Expect = 0.57
Identities = 23/67 (34%), Positives = 34/67 (50%), Gaps = 4/67 (5%)
Query: 32 RKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKS--PMAKPKKLLSNISNKALSQFGKKK 89
RKA+ + AK KP +DE+ + D KS P +K +K NK L G+K+
Sbjct: 891 RKAH--SKSAKEHQEAKPAKDEKVKKDCSKDIKSEKPASKDEKAKKPEKNKLLDSKGEKR 948
Query: 90 QREEREK 96
+R+ EK
Sbjct: 949 KRKTEEK 955
>UniRef100_P97868-2 Splice isoform 2 of P97868 [Mus musculus]
Length = 1557
Score = 34.3 bits (77), Expect = 0.57
Identities = 23/67 (34%), Positives = 34/67 (50%), Gaps = 4/67 (5%)
Query: 32 RKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKS--PMAKPKKLLSNISNKALSQFGKKK 89
RKA+ + AK KP +DE+ + D KS P +K +K NK L G+K+
Sbjct: 857 RKAH--SKSAKEHQEAKPAKDEKVKKDCSKDIKSEKPASKDEKAKKPEKNKLLDSKGEKR 914
Query: 90 QREEREK 96
+R+ EK
Sbjct: 915 KRKTEEK 921
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.133 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 277,359,703
Number of Sequences: 2790947
Number of extensions: 11264211
Number of successful extensions: 28121
Number of sequences better than 10.0: 132
Number of HSP's better than 10.0 without gapping: 35
Number of HSP's successfully gapped in prelim test: 98
Number of HSP's that attempted gapping in prelim test: 28041
Number of HSP's gapped (non-prelim): 164
length of query: 153
length of database: 848,049,833
effective HSP length: 129
effective length of query: 24
effective length of database: 488,017,670
effective search space: 11712424080
effective search space used: 11712424080
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC148996.15


