

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148995.8 + phase: 0
(325 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
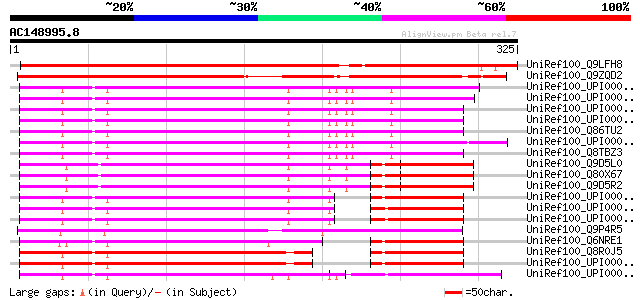
UniRef100_Q9LFH8 Hypothetical protein F4P12_90 [Arabidopsis thal... 508 e-142
UniRef100_Q9ZQD2 Hypothetical protein At2g37160 [Arabidopsis tha... 442 e-123
UniRef100_UPI000019C56F UPI000019C56F UniRef100 entry 213 4e-54
UniRef100_UPI0000369EAE UPI0000369EAE UniRef100 entry 212 1e-53
UniRef100_UPI000019C570 UPI000019C570 UniRef100 entry 211 2e-53
UniRef100_UPI000019C572 UPI000019C572 UniRef100 entry 211 2e-53
UniRef100_Q86TU2 Full-length cDNA clone CS0DC027YE03 of Neurobla... 211 2e-53
UniRef100_UPI00001CFAC8 UPI00001CFAC8 UniRef100 entry 209 6e-53
UniRef100_Q8TBZ3 WD-repeat protein 20 [Homo sapiens] 208 2e-52
UniRef100_Q9D5L0 Mus musculus adult male testis cDNA, RIKEN full... 191 2e-47
UniRef100_Q80X67 WD repeat domain 20 [Mus musculus] 191 2e-47
UniRef100_Q9D5R2 WD-repeat protein 20 [Mus musculus] 191 2e-47
UniRef100_UPI000025D922 UPI000025D922 UniRef100 entry 186 1e-45
UniRef100_UPI000036088C UPI000036088C UniRef100 entry 183 5e-45
UniRef100_UPI000036088A UPI000036088A UniRef100 entry 183 5e-45
UniRef100_Q9P4R5 Catabolite repression protein creC [Emericella ... 183 5e-45
UniRef100_Q6NRE1 Hypothetical protein [Xenopus laevis] 179 1e-43
UniRef100_Q8R0J5 2310040A13Rik protein [Mus musculus] 171 3e-41
UniRef100_UPI000017F7C1 UPI000017F7C1 UniRef100 entry 169 1e-40
UniRef100_UPI00001CE7D9 UPI00001CE7D9 UniRef100 entry 166 8e-40
>UniRef100_Q9LFH8 Hypothetical protein F4P12_90 [Arabidopsis thaliana]
Length = 556
Score = 508 bits (1307), Expect = e-142
Identities = 248/326 (76%), Positives = 279/326 (85%), Gaps = 15/326 (4%)
Query: 8 RCTCISWVPGGDGAFVVAHADGNLYNKDGAGESSFPILKDQTLFSVAHARYSKSNPIARW 67
RCT ISWVPGGDGAFVVAHADGNLYNKDGA ES+FP ++D T FSV A+YSKSNP+ARW
Sbjct: 238 RCTSISWVPGGDGAFVVAHADGNLYNKDGATESTFPAIRDPTQFSVDKAKYSKSNPVARW 297
Query: 68 HICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKY 127
HICQGSINSI+FS DGA+LATVGRDGYLR+FD+ + LVCGGKSYYG LLCC+WSMDGKY
Sbjct: 298 HICQGSINSIAFSNDGAHLATVGRDGYLRIFDFLTQKLVCGGKSYYGALLCCSWSMDGKY 357
Query: 128 ILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVG 187
ILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDS WSSPNS+ +GE + YRFGSVG
Sbjct: 358 ILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSNWSSPNSDGSGEHVMYRFGSVG 417
Query: 188 QDTQLLLWELEMDEIVVPLRRGPPGGSPTFGSPTFSAGSQSSHWDNAVPLGTLQPAPSMR 247
QDTQLLLW+LEMDEIVVPLRR P GSPT+S GS S+HWDN +P+GTLQPAP R
Sbjct: 418 QDTQLLLWDLEMDEIVVPLRRAPG------GSPTYSTGS-SAHWDNVIPMGTLQPAPCKR 470
Query: 248 DVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPGVAESQPSNSE-----T 302
DVPK+SP++AHRVHTEPLS L+FTQESV+TACREGHIKIWTRP +E+Q ++SE
Sbjct: 471 DVPKLSPIIAHRVHTEPLSGLMFTQESVVTACREGHIKIWTRPSESETQTNSSEANPTTA 530
Query: 303 LLATSLKE--KPSLSSK-ISNSIYKQ 325
LL+TS + K SLSSK IS S +KQ
Sbjct: 531 LLSTSFTKDNKASLSSKIISGSTFKQ 556
>UniRef100_Q9ZQD2 Hypothetical protein At2g37160 [Arabidopsis thaliana]
Length = 444
Score = 442 bits (1136), Expect = e-123
Identities = 220/313 (70%), Positives = 251/313 (79%), Gaps = 32/313 (10%)
Query: 6 NSRCTCISWVPGGDGAFVVAHADGNLYNKDGAGESSFPILKDQTLFSVAHARYSKSNPIA 65
NSRCT I+WVPGGDG+FV AHADGNLYNK+GA +SSF ++D T FSV A+YSKSNP+A
Sbjct: 150 NSRCTSIAWVPGGDGSFVAAHADGNLYNKEGATDSSFSAIRDPTQFSVDKAKYSKSNPVA 209
Query: 66 RWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDG 125
RWHI QG+INSI+FS DGAYLATVGRDGYLR+FD++ + LVCG KSYYG LLCCAWSMDG
Sbjct: 210 RWHIGQGAINSIAFSNDGAYLATVGRDGYLRIFDFSTQKLVCGVKSYYGALLCCAWSMDG 269
Query: 126 KYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGS 185
KY+LTGGEDDLVQVWSMEDRKVVAW EG +GE + YRFGS
Sbjct: 270 KYLLTGGEDDLVQVWSMEDRKVVAW-EG---------------------SGENVMYRFGS 307
Query: 186 VGQDTQLLLWELEMDEIVVPLRRGPPGGSPTFGSPTFSAGSQSSHWDNAVPLGTLQPAPS 245
VGQDTQLLLW+LEMDEIVVPLRR PPG GSPT+S GSQS+HWDN VP+GTLQPAP
Sbjct: 308 VGQDTQLLLWDLEMDEIVVPLRR-PPG-----GSPTYSTGSQSAHWDNIVPMGTLQPAPC 361
Query: 246 MRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPGVAESQPSNSETLLA 305
MRDVPK+SP+VAHRVHTEPLS LIFTQES++TACREGH+KIWTRP ++QPS+S
Sbjct: 362 MRDVPKLSPVVAHRVHTEPLSGLIFTQESLITACREGHLKIWTRP---DTQPSSSSE-AT 417
Query: 306 TSLKEKPSLSSKI 318
KPSL+SK+
Sbjct: 418 NPTTSKPSLTSKV 430
>UniRef100_UPI000019C56F UPI000019C56F UniRef100 entry
Length = 581
Score = 213 bits (543), Expect = 4e-54
Identities = 148/424 (34%), Positives = 193/424 (44%), Gaps = 130/424 (30%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLY-----NKDGAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WVPG + F+VAH+ GN+Y + G + +LK F+V H SKS
Sbjct: 150 SRVTCVKWVPGSESLFLVAHSSGNMYLYNVEHTCGTTAPHYQLLKQGESFAV-HTCKSKS 208
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+ +W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC
Sbjct: 209 TRNPLLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDSVELHGTMKSYFGGLLCV 268
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE-- 177
WS DGKYI+TGGEDDLV VWS D +V+A G GH SWVS VAFD Y +S D E
Sbjct: 269 CWSPDGKYIVTGGEDDLVTVWSFVDCRVIARGHGHKSWVSVVAFDPYTTSVEEGDPMEFS 328
Query: 178 -----------------------------------TITYRFGSVGQDTQLLLWELEMDEI 202
++TYRFGSVGQDTQL LW+L D +
Sbjct: 329 GSDEDFQDLLHFGRDRANSTQSRLSKRNSTDSRPVSVTYRFGSVGQDTQLCLWDLTEDIL 388
Query: 203 V--VPLRR-----------GPPGGS-----PTFG---------------SPTFSAGSQSS 229
PL R PP GS T G S +AGS+SS
Sbjct: 389 FPHQPLSRARTHTNVMNATSPPAGSNGNSVTTPGNSVPPPLPRSNSLPHSAVSNAGSKSS 448
Query: 230 HWDNAVPLGTLQPA---------------------------------------------- 243
D A+ G + A
Sbjct: 449 VMDGAIASGVSKFATLSLHDRKERHHEKDHKRNHSMGHISSKSSDKLNLVTKTKTDPAKT 508
Query: 244 ------PSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPGVAESQP 297
P M DVP + PL+ ++ E L+ LIF ++ ++TAC+EG I W RPG S
Sbjct: 509 LGTPLCPRMEDVPLLEPLICKKIAHERLTVLIFLEDCIVTACQEGFICTWGRPGKVGSLS 568
Query: 298 SNSE 301
S S+
Sbjct: 569 SPSQ 572
>UniRef100_UPI0000369EAE UPI0000369EAE UniRef100 entry
Length = 581
Score = 212 bits (539), Expect = 1e-53
Identities = 147/421 (34%), Positives = 191/421 (44%), Gaps = 130/421 (30%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLY-----NKDGAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WVPG + F+VAH+ GN+Y + G + +LK F+V H SKS
Sbjct: 150 SRVTCVKWVPGSESLFLVAHSSGNMYLYNVEHTCGTTAPHYQLLKQGESFAV-HTCKSKS 208
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+ +W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC
Sbjct: 209 TRNPLLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDSVELHGTMKSYFGGLLCV 268
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE-- 177
WS DGKYI+TGGEDDLV VWS D +V+A G GH SWVS VAFD Y +S D E
Sbjct: 269 CWSPDGKYIVTGGEDDLVTVWSFVDCRVIARGHGHKSWVSVVAFDPYTTSVEEGDPMEFS 328
Query: 178 -----------------------------------TITYRFGSVGQDTQLLLWELEMDEI 202
++TYRFGSVGQDTQL LW+L D +
Sbjct: 329 GSDEDFQDLLHFGRDRANSTQSRLSKRNSTDSRPVSVTYRFGSVGQDTQLCLWDLTEDIL 388
Query: 203 V--VPLRR-----------GPPGGS-----PTFG---------------SPTFSAGSQSS 229
PL R PP GS T G S +AGS+SS
Sbjct: 389 FPHQPLSRARTHTNVMNATSPPAGSNGNSVTTPGNSVPPPLPRSNSLPHSAVSNAGSKSS 448
Query: 230 HWDNAVPLGTLQPA---------------------------------------------- 243
D A+ G + A
Sbjct: 449 VMDGAIASGVSKFATLSLHDRKERHHEKDHKRNHSMGHISSKSSDKLNLVTKTKTDPAKT 508
Query: 244 ------PSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPGVAESQP 297
P M DVP + PL+ ++ E L+ LIF ++ ++TAC+EG I W RPG S
Sbjct: 509 LGTPLCPRMEDVPLLEPLICKKIAHERLTVLIFLEDCIVTACQEGFICTWGRPGKVGSLS 568
Query: 298 S 298
S
Sbjct: 569 S 569
>UniRef100_UPI000019C570 UPI000019C570 UniRef100 entry
Length = 445
Score = 211 bits (538), Expect = 2e-53
Identities = 145/414 (35%), Positives = 189/414 (45%), Gaps = 130/414 (31%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLY-----NKDGAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WVPG + F+VAH+ GN+Y + G + +LK F+V H SKS
Sbjct: 26 SRVTCVKWVPGSESLFLVAHSSGNMYLYNVEHTCGTTAPHYQLLKQGESFAV-HTCKSKS 84
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+ +W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC
Sbjct: 85 TRNPLLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDSVELHGTMKSYFGGLLCV 144
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE-- 177
WS DGKYI+TGGEDDLV VWS D +V+A G GH SWVS VAFD Y +S D E
Sbjct: 145 CWSPDGKYIVTGGEDDLVTVWSFVDCRVIARGHGHKSWVSVVAFDPYTTSVEEGDPMEFS 204
Query: 178 -----------------------------------TITYRFGSVGQDTQLLLWELEMDEI 202
++TYRFGSVGQDTQL LW+L D +
Sbjct: 205 GSDEDFQDLLHFGRDRANSTQSRLSKRNSTDSRPVSVTYRFGSVGQDTQLCLWDLTEDIL 264
Query: 203 V--VPLRR-----------GPPGGS-----PTFG---------------SPTFSAGSQSS 229
PL R PP GS T G S +AGS+SS
Sbjct: 265 FPHQPLSRARTHTNVMNATSPPAGSNGNSVTTPGNSVPPPLPRSNSLPHSAVSNAGSKSS 324
Query: 230 HWDNAVPLGTLQPA---------------------------------------------- 243
D A+ G + A
Sbjct: 325 VMDGAIASGVSKFATLSLHDRKERHHEKDHKRNHSMGHISSKSSDKLNLVTKTKTDPAKT 384
Query: 244 ------PSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPG 291
P M DVP + PL+ ++ E L+ LIF ++ ++TAC+EG I W RPG
Sbjct: 385 LGTPLCPRMEDVPLLEPLICKKIAHERLTVLIFLEDCIVTACQEGFICTWGRPG 438
>UniRef100_UPI000019C572 UPI000019C572 UniRef100 entry
Length = 569
Score = 211 bits (538), Expect = 2e-53
Identities = 145/414 (35%), Positives = 189/414 (45%), Gaps = 130/414 (31%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLY-----NKDGAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WVPG + F+VAH+ GN+Y + G + +LK F+V H SKS
Sbjct: 150 SRVTCVKWVPGSESLFLVAHSSGNMYLYNVEHTCGTTAPHYQLLKQGESFAV-HTCKSKS 208
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+ +W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC
Sbjct: 209 TRNPLLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDSVELHGTMKSYFGGLLCV 268
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE-- 177
WS DGKYI+TGGEDDLV VWS D +V+A G GH SWVS VAFD Y +S D E
Sbjct: 269 CWSPDGKYIVTGGEDDLVTVWSFVDCRVIARGHGHKSWVSVVAFDPYTTSVEEGDPMEFS 328
Query: 178 -----------------------------------TITYRFGSVGQDTQLLLWELEMDEI 202
++TYRFGSVGQDTQL LW+L D +
Sbjct: 329 GSDEDFQDLLHFGRDRANSTQSRLSKRNSTDSRPVSVTYRFGSVGQDTQLCLWDLTEDIL 388
Query: 203 V--VPLRR-----------GPPGGS-----PTFG---------------SPTFSAGSQSS 229
PL R PP GS T G S +AGS+SS
Sbjct: 389 FPHQPLSRARTHTNVMNATSPPAGSNGNSVTTPGNSVPPPLPRSNSLPHSAVSNAGSKSS 448
Query: 230 HWDNAVPLGTLQPA---------------------------------------------- 243
D A+ G + A
Sbjct: 449 VMDGAIASGVSKFATLSLHDRKERHHEKDHKRNHSMGHISSKSSDKLNLVTKTKTDPAKT 508
Query: 244 ------PSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPG 291
P M DVP + PL+ ++ E L+ LIF ++ ++TAC+EG I W RPG
Sbjct: 509 LGTPLCPRMEDVPLLEPLICKKIAHERLTVLIFLEDCIVTACQEGFICTWGRPG 562
>UniRef100_Q86TU2 Full-length cDNA clone CS0DC027YE03 of Neuroblastoma of Homo
sapiens [Homo sapiens]
Length = 499
Score = 211 bits (538), Expect = 2e-53
Identities = 145/414 (35%), Positives = 189/414 (45%), Gaps = 130/414 (31%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLY-----NKDGAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WVPG + F+VAH+ GN+Y + G + +LK F+V H SKS
Sbjct: 80 SRVTCVKWVPGSESLFLVAHSSGNMYLYNVEHTCGTTAPHYQLLKQGESFAV-HTCKSKS 138
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+ +W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC
Sbjct: 139 TRNPLLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDSVELHGTMKSYFGGLLCV 198
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE-- 177
WS DGKYI+TGGEDDLV VWS D +V+A G GH SWVS VAFD Y +S D E
Sbjct: 199 CWSPDGKYIVTGGEDDLVTVWSFVDCRVIARGHGHKSWVSVVAFDPYTTSVEEGDPMEFS 258
Query: 178 -----------------------------------TITYRFGSVGQDTQLLLWELEMDEI 202
++TYRFGSVGQDTQL LW+L D +
Sbjct: 259 GSDEDFQDLLHFGRDRANSTQSRLSKRNSTDSRPVSVTYRFGSVGQDTQLCLWDLTEDIL 318
Query: 203 V--VPLRR-----------GPPGGS-----PTFG---------------SPTFSAGSQSS 229
PL R PP GS T G S +AGS+SS
Sbjct: 319 FPHQPLSRARTHTNVMNATSPPAGSNGNSVTTPGNSVPPPLPRSNSLPHSAVSNAGSKSS 378
Query: 230 HWDNAVPLGTLQPA---------------------------------------------- 243
D A+ G + A
Sbjct: 379 VMDGAIASGVSKFATLSLHDRKERHHEKDHKRNHSMGHISSKSSDKLNLVTKTKTDPAKT 438
Query: 244 ------PSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPG 291
P M DVP + PL+ ++ E L+ LIF ++ ++TAC+EG I W RPG
Sbjct: 439 LGTPLCPRMEDVPLLEPLICKKIAHERLTVLIFLEDCIVTACQEGFICTWGRPG 492
>UniRef100_UPI00001CFAC8 UPI00001CFAC8 UniRef100 entry
Length = 595
Score = 209 bits (533), Expect = 6e-53
Identities = 148/442 (33%), Positives = 201/442 (44%), Gaps = 131/442 (29%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLY-----NKDGAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WVPG + F+VAH+ G++Y + G + +LK F+V H SKS
Sbjct: 150 SRVTCVKWVPGSESLFLVAHSSGSMYLYNVEHTCGTTAPHYQLLKQGESFAV-HTCKSKS 208
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+ +W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC
Sbjct: 209 TRNPLLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDSVELHGTMKSYFGGLLCV 268
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE-- 177
WS DGKYI+TGGEDDLV VWS D +V+A G GH SWVS VAFD Y +S +D E
Sbjct: 269 CWSPDGKYIVTGGEDDLVTVWSFLDCRVIARGHGHKSWVSVVAFDPYTTSVEESDPMEFS 328
Query: 178 -----------------------------------TITYRFGSVGQDTQLLLWELEMDEI 202
++TYRFGSVGQDTQL LW+L D +
Sbjct: 329 GSDEDFQDLLHFGRDRANSTQSRLSKRNSTDSRPVSVTYRFGSVGQDTQLCLWDLTEDIL 388
Query: 203 V--VPLRR-----------GPPGGS-----PTFG---------------SPTFSAGSQSS 229
PL R PP GS T G S +AGS+SS
Sbjct: 389 FPHQPLSRARTHTNVMNATSPPAGSNGNSVTTPGDSGPPPLPRSNSLPHSAVSNAGSKSS 448
Query: 230 HWDNAVPLGTLQPA---------------------------------------------- 243
D A+ G + A
Sbjct: 449 VMDGAIASGVSKFATLSLHDRKERHHEKDHKRNHSMGHISSKSSDKLNLVTKAKTDPAKT 508
Query: 244 ------PSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPGVAESQP 297
P M DVP + PL+ ++ E L+ L+F ++ ++TAC+EG I W RPG +P
Sbjct: 509 LGTSLCPRMEDVPLLEPLICKKIAHERLTVLVFLEDCLVTACQEGFICTWARPGKV-CRP 567
Query: 298 SNSETLLATSLKEKPSLSSKIS 319
S L S +++K++
Sbjct: 568 SLGLCPLTNSAVSAARVAAKVA 589
>UniRef100_Q8TBZ3 WD-repeat protein 20 [Homo sapiens]
Length = 569
Score = 208 bits (529), Expect = 2e-52
Identities = 144/414 (34%), Positives = 188/414 (44%), Gaps = 130/414 (31%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLY-----NKDGAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WV G + F+VAH+ GN+Y + G + +LK F+V H SKS
Sbjct: 150 SRVTCVKWVHGSESLFLVAHSSGNMYLYNVEHTCGTTAPHYQLLKQGESFAV-HTCKSKS 208
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+ +W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC
Sbjct: 209 TRNPLLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDSVELHGTMKSYFGGLLCV 268
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE-- 177
WS DGKYI+TGGEDDLV VWS D +V+A G GH SWVS VAFD Y +S D E
Sbjct: 269 CWSPDGKYIVTGGEDDLVTVWSFVDCRVIARGHGHKSWVSVVAFDPYTTSVEEGDPMEFS 328
Query: 178 -----------------------------------TITYRFGSVGQDTQLLLWELEMDEI 202
++TYRFGSVGQDTQL LW+L D +
Sbjct: 329 GSDEDFQDLLHFGRDRANSTQSRLSKRNSTDSRPVSVTYRFGSVGQDTQLCLWDLTEDIL 388
Query: 203 V--VPLRR-----------GPPGGS-----PTFG---------------SPTFSAGSQSS 229
PL R PP GS T G S +AGS+SS
Sbjct: 389 FPHQPLSRARTHTNVMNATSPPAGSNGNSVTTPGNSVPPPLPRSNSLPHSAVSNAGSKSS 448
Query: 230 HWDNAVPLGTLQPA---------------------------------------------- 243
D A+ G + A
Sbjct: 449 VMDGAIASGVSKFATLSLHDRKERHHEKDHKRNHSMGHISSKSSDKLNLVTKTKTDPAKT 508
Query: 244 ------PSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPG 291
P M DVP + PL+ ++ E L+ LIF ++ ++TAC+EG I W RPG
Sbjct: 509 LGTPLCPRMEDVPLLEPLICKKIAHERLTVLIFLEDCIVTACQEGFICTWGRPG 562
>UniRef100_Q9D5L0 Mus musculus adult male testis cDNA, RIKEN full-length enriched
library, clone:4930427E19 product:DMR PROTEIN homolog
[Mus musculus]
Length = 466
Score = 191 bits (485), Expect = 2e-47
Identities = 116/290 (40%), Positives = 152/290 (52%), Gaps = 51/290 (17%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLYNKD-----GAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WVPG D F+VAH+ GN+Y+ + G + +LK F+V + + SKS
Sbjct: 49 SRVTCVKWVPGSDSLFLVAHSSGNMYSYNVEHTYGTTAPHYQLLKQGESFAVHNCK-SKS 107
Query: 62 NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAW 121
+W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC W
Sbjct: 108 TRDLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDSAELHGTMKSYFGGLLCLCW 167
Query: 122 SMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE---- 177
S DGKYI+TGGEDDLV VWS D +V+A G GH SWVS VAFD Y +S +D E
Sbjct: 168 SPDGKYIVTGGEDDLVTVWSFLDCRVIARGRGHKSWVSVVAFDPYTTSVEESDPMEFSDS 227
Query: 178 ---------------------------------TITYRFGSVGQDTQLLLWELEMDEIV- 203
++TYRFGSVGQDTQL LW+L D +
Sbjct: 228 DKNFQDLLHFGRDRANSTQSRLSKQNSTDSRPVSVTYRFGSVGQDTQLCLWDLTEDILFP 287
Query: 204 -VPLRRGPPGGS--PTFGSPTFSAGSQSSHWDNAVPLGTLQPAPSMRDVP 250
PL R + G P S GS + N+VP P P +P
Sbjct: 288 HQPLSRARAHANVMNATGLPAGSNGSAVTTPGNSVP----PPLPRSNSLP 333
Score = 56.2 bits (134), Expect = 1e-06
Identities = 29/67 (43%), Positives = 42/67 (62%), Gaps = 2/67 (2%)
Query: 232 DNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPG 291
D A LGT P M DVP + PL+ ++ E L+ L+F ++ ++TAC+EG I W RPG
Sbjct: 401 DPAKTLGT-SLCPRMEDVPLLEPLICKKIAHERLTVLVFLEDCIVTACQEGFICTWARPG 459
Query: 292 -VAESQP 297
V++ QP
Sbjct: 460 KVSKFQP 466
>UniRef100_Q80X67 WD repeat domain 20 [Mus musculus]
Length = 567
Score = 191 bits (485), Expect = 2e-47
Identities = 116/290 (40%), Positives = 152/290 (52%), Gaps = 51/290 (17%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLYNKD-----GAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WVPG D F+VAH+ GN+Y+ + G + +LK F+V + + SKS
Sbjct: 150 SRVTCVKWVPGSDSLFLVAHSSGNMYSYNVEHTYGTTAPHYQLLKQGESFAVHNCK-SKS 208
Query: 62 NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAW 121
+W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC W
Sbjct: 209 TRDLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDSAELHGTMKSYFGGLLCLCW 268
Query: 122 SMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE---- 177
S DGKYI+TGGEDDLV VWS D +V+A G GH SWVS VAFD Y +S +D E
Sbjct: 269 SPDGKYIVTGGEDDLVTVWSFLDCRVIARGRGHKSWVSVVAFDPYTTSVEESDPMEFSDS 328
Query: 178 ---------------------------------TITYRFGSVGQDTQLLLWELEMDEIV- 203
++TYRFGSVGQDTQL LW+L D +
Sbjct: 329 DKNFQDLLHFGRDRANSTQSRLSKQNSTDSRPVSVTYRFGSVGQDTQLCLWDLTEDILFP 388
Query: 204 -VPLRRGPPGGS--PTFGSPTFSAGSQSSHWDNAVPLGTLQPAPSMRDVP 250
PL R + G P S GS + N+VP P P +P
Sbjct: 389 HQPLSRARAHANVMNATGLPAGSNGSAVTTPGNSVP----PPLPRSNSLP 434
Score = 56.2 bits (134), Expect = 1e-06
Identities = 29/67 (43%), Positives = 42/67 (62%), Gaps = 2/67 (2%)
Query: 232 DNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPG 291
D A LGT P M DVP + PL+ ++ E L+ L+F ++ ++TAC+EG I W RPG
Sbjct: 502 DPAKTLGT-SLCPRMEDVPLLEPLICKKIAHERLTVLVFLEDCIVTACQEGFICTWARPG 560
Query: 292 -VAESQP 297
V++ QP
Sbjct: 561 KVSKFQP 567
>UniRef100_Q9D5R2 WD-repeat protein 20 [Mus musculus]
Length = 567
Score = 191 bits (485), Expect = 2e-47
Identities = 116/290 (40%), Positives = 152/290 (52%), Gaps = 51/290 (17%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLYNKD-----GAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WVPG D F+VAH+ GN+Y+ + G + +LK F+V + + SKS
Sbjct: 150 SRVTCVKWVPGSDSLFLVAHSSGNMYSYNVEHTYGTTAPHYQLLKQGESFAVHNCK-SKS 208
Query: 62 NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAW 121
+W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC W
Sbjct: 209 TRDLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDSAELHGTMKSYFGGLLCLCW 268
Query: 122 SMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE---- 177
S DGKYI+TGGEDDLV VWS D +V+A G GH SWVS VAFD Y +S +D E
Sbjct: 269 SPDGKYIVTGGEDDLVTVWSFLDCRVIARGRGHKSWVSVVAFDPYTTSVEESDPMEFSDS 328
Query: 178 ---------------------------------TITYRFGSVGQDTQLLLWELEMDEIV- 203
++TYRFGSVGQDTQL LW+L D +
Sbjct: 329 DKNFQDLLHFGRDRANSTQSRLSKQNSTDSRPVSVTYRFGSVGQDTQLCLWDLTEDILFP 388
Query: 204 -VPLRRGPPGGS--PTFGSPTFSAGSQSSHWDNAVPLGTLQPAPSMRDVP 250
PL R + G P S GS + N+VP P P +P
Sbjct: 389 HQPLSRARAHANVMNATGLPAGSNGSAVTTPGNSVP----PPLPRSNSLP 434
Score = 56.2 bits (134), Expect = 1e-06
Identities = 29/67 (43%), Positives = 42/67 (62%), Gaps = 2/67 (2%)
Query: 232 DNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPG 291
D A LGT P M DVP + PL+ ++ E L+ L+F ++ ++TAC+EG I W RPG
Sbjct: 502 DPAKTLGT-SLCPRMEDVPLLEPLICKKIAHERLTVLVFLEDCIVTACQEGFICTWARPG 560
Query: 292 -VAESQP 297
V++ QP
Sbjct: 561 KVSKFQP 567
>UniRef100_UPI000025D922 UPI000025D922 UniRef100 entry
Length = 626
Score = 186 bits (471), Expect = 1e-45
Identities = 106/249 (42%), Positives = 138/249 (54%), Gaps = 48/249 (19%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLY-----NKDGAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WVPG + F+V+HA GN+Y + G + +LK +SV H SKS
Sbjct: 150 SRVTCVKWVPGSESLFLVSHASGNMYLYNVEHTCGTTAPHYQLLKQGESYSV-HTCKSKS 208
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+ +W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC
Sbjct: 209 TRNPLLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDAVELHGTMKSYFGGLLCV 268
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE-- 177
WS DGKYI+ GGEDDLV VWS D +V+A G GH SWVS VAFD Y +S +D E
Sbjct: 269 CWSPDGKYIVAGGEDDLVTVWSFLDCRVIARGHGHKSWVSVVAFDHYTTSVEESDPMEFS 328
Query: 178 ------------------------------------TITYRFGSVGQDTQLLLWELEMDE 201
++TYRFGSVGQDTQL LW+L D
Sbjct: 329 GSDEDFQDHVIHFGRDRANSTQSRLSKRNSTDSRPVSVTYRFGSVGQDTQLCLWDLTEDI 388
Query: 202 IV--VPLRR 208
+ +PL R
Sbjct: 389 LFPHLPLSR 397
Score = 56.2 bits (134), Expect = 1e-06
Identities = 28/60 (46%), Positives = 38/60 (62%), Gaps = 1/60 (1%)
Query: 232 DNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPG 291
D A LGT Q P M DVP + PL+ ++ E L+ LIF ++ ++TAC+EG I W RPG
Sbjct: 549 DPAKTLGT-QLCPRMEDVPLLEPLICKKIAHERLTVLIFLEDCLVTACQEGFICTWARPG 607
>UniRef100_UPI000036088C UPI000036088C UniRef100 entry
Length = 519
Score = 183 bits (465), Expect = 5e-45
Identities = 105/249 (42%), Positives = 138/249 (55%), Gaps = 48/249 (19%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLY-----NKDGAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WVPG + F+V+HA G++Y + G + +LK +SV H SKS
Sbjct: 80 SRVTCVKWVPGSESLFLVSHASGSMYLYNVEHTCGTTAPHYQLLKQGENYSV-HTCKSKS 138
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+ +W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC
Sbjct: 139 TRNPLLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDAVELHGTMKSYFGGLLCV 198
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE-- 177
WS DGKYI+ GGEDDLV VWS D +V+A G GH SWVS VAFD Y +S +D E
Sbjct: 199 CWSPDGKYIVAGGEDDLVTVWSFLDCRVIARGHGHKSWVSVVAFDHYTTSVEESDPMEFS 258
Query: 178 ------------------------------------TITYRFGSVGQDTQLLLWELEMDE 201
++TYRFGSVGQDTQL LW+L D
Sbjct: 259 GSDEDFQDQMIHFGRDRANSTQSRLSKRNSTDSRPVSVTYRFGSVGQDTQLCLWDLTEDI 318
Query: 202 IV--VPLRR 208
+ +PL R
Sbjct: 319 LFPHLPLSR 327
Score = 56.2 bits (134), Expect = 1e-06
Identities = 28/60 (46%), Positives = 38/60 (62%), Gaps = 1/60 (1%)
Query: 232 DNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPG 291
D A LGTL P M DVP + PL+ ++ E L+ LIF ++ ++TAC+EG I W RPG
Sbjct: 457 DPAKTLGTLL-CPRMEDVPLLEPLICKKIAHERLTVLIFLEDCLVTACQEGFICTWARPG 515
>UniRef100_UPI000036088A UPI000036088A UniRef100 entry
Length = 589
Score = 183 bits (465), Expect = 5e-45
Identities = 105/249 (42%), Positives = 138/249 (55%), Gaps = 48/249 (19%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLY-----NKDGAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WVPG + F+V+HA G++Y + G + +LK +SV H SKS
Sbjct: 150 SRVTCVKWVPGSESLFLVSHASGSMYLYNVEHTCGTTAPHYQLLKQGENYSV-HTCKSKS 208
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+ +W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC
Sbjct: 209 TRNPLLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDAVELHGTMKSYFGGLLCV 268
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE-- 177
WS DGKYI+ GGEDDLV VWS D +V+A G GH SWVS VAFD Y +S +D E
Sbjct: 269 CWSPDGKYIVAGGEDDLVTVWSFLDCRVIARGHGHKSWVSVVAFDHYTTSVEESDPMEFS 328
Query: 178 ------------------------------------TITYRFGSVGQDTQLLLWELEMDE 201
++TYRFGSVGQDTQL LW+L D
Sbjct: 329 GSDEDFQDQMIHFGRDRANSTQSRLSKRNSTDSRPVSVTYRFGSVGQDTQLCLWDLTEDI 388
Query: 202 IV--VPLRR 208
+ +PL R
Sbjct: 389 LFPHLPLSR 397
Score = 56.2 bits (134), Expect = 1e-06
Identities = 28/60 (46%), Positives = 38/60 (62%), Gaps = 1/60 (1%)
Query: 232 DNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPG 291
D A LGTL P M DVP + PL+ ++ E L+ LIF ++ ++TAC+EG I W RPG
Sbjct: 527 DPAKTLGTLL-CPRMEDVPLLEPLICKKIAHERLTVLIFLEDCLVTACQEGFICTWARPG 585
>UniRef100_Q9P4R5 Catabolite repression protein creC [Emericella nidulans]
Length = 630
Score = 183 bits (465), Expect = 5e-45
Identities = 108/318 (33%), Positives = 163/318 (50%), Gaps = 41/318 (12%)
Query: 6 NSRCTCISWVPGGDGAFVVAHADGNL--YNKDGAGESSFPILKDQTLFSVAHARYS---- 59
+S T I W+PG + F+ AH +G L Y+K+ P L +Q+ SV +R+S
Sbjct: 288 SSPVTHIKWIPGSENFFIAAHENGQLVVYDKEKEDALFIPELPEQSAESVKPSRWSLQVL 347
Query: 60 --------KSNPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKS 111
K+NP+A W + I +FS D +LA V DG LR+ DY +E ++ +S
Sbjct: 348 KSVNSKNQKANPVAVWRLANQKITQFAFSPDHRHLAVVLEDGTLRLMDYLQEEVLDVFRS 407
Query: 112 YYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPN 171
YYGG C WS DGKYI+TGG+DDLV +WS+ +RK++A +GH+SWVS VAFD +
Sbjct: 408 YYGGFTCVCWSPDGKYIVTGGQDDLVTIWSLPERKIIARCQGHDSWVSAVAFDPW----- 462
Query: 172 SNDNGETITYRFGSVGQDTQLLLWELEM-----------------DEIVVPLRRGPPGGS 214
+ TYR GSVG D LLLW+ + ++ P + P
Sbjct: 463 ---RCDERTYRIGSVGDDCNLLLWDFSVGMLHRPKVHHQTSARHRTSLIAPSSQQPNRHR 519
Query: 215 PTFGSPTFSAGSQSSHWDN-AVPLGTLQ-PAPSMRDVPKISPLVAHRVHTEPLSSLIFTQ 272
+ SQ + D+ + P +Q P S + P+++ V +P+ L F +
Sbjct: 520 ADSSGNRMRSDSQRTAADSESAPDQPVQHPVESRARTALLPPIMSKAVGEDPICWLGFQE 579
Query: 273 ESVLTACREGHIKIWTRP 290
++++T+ EGHI+ W RP
Sbjct: 580 DTIMTSSLEGHIRTWDRP 597
>UniRef100_Q6NRE1 Hypothetical protein [Xenopus laevis]
Length = 574
Score = 179 bits (453), Expect = 1e-43
Identities = 104/238 (43%), Positives = 135/238 (56%), Gaps = 45/238 (18%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGN--LYNKD---GAGESSFPILKDQTLFSVAHARYSKS 61
+R T + WVPG + F+VAH+ G+ LYN + G + +LK F+V H SKS
Sbjct: 150 ARVTSVKWVPGSESLFLVAHSSGSMFLYNVEYTCGTTAPHYQLLKQGESFAV-HTCKSKS 208
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+ +W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC
Sbjct: 209 TRNPLLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDSVELHGTMKSYFGGLLCV 268
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFD--------------- 164
WS DGKYI+TGGEDDLV VWS D +V+A G+GH SWVS VAFD
Sbjct: 269 CWSPDGKYIVTGGEDDLVTVWSFVDCRVIARGQGHKSWVSVVAFDPNTTSVEETDPMEFS 328
Query: 165 ---------------------SYWSSPNSNDNGE-TITYRFGSVGQDTQLLLWELEMD 200
S S NS D+ ++TYRFGSVGQDTQL LW+L D
Sbjct: 329 GSDEDFQELMSFGRDRANSTQSRLSKRNSTDSRPVSVTYRFGSVGQDTQLCLWDLTED 386
Score = 55.8 bits (133), Expect = 2e-06
Identities = 27/60 (45%), Positives = 38/60 (63%), Gaps = 1/60 (1%)
Query: 232 DNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPG 291
D+A LGT P M DVP + PL+ ++ E L+ LIF ++ ++TAC+EG I W RPG
Sbjct: 509 DSAKTLGT-PLCPRMEDVPLLEPLICKKIAHERLTVLIFLEDCIVTACQEGFICTWARPG 567
>UniRef100_Q8R0J5 2310040A13Rik protein [Mus musculus]
Length = 539
Score = 171 bits (432), Expect = 3e-41
Identities = 92/195 (47%), Positives = 120/195 (61%), Gaps = 13/195 (6%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLY-----NKDGAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WVPG + F+VAH+ GN+Y + G + +LK F+V H SKS
Sbjct: 150 SRVTCVKWVPGSESLFLVAHSSGNMYLYNVEHTCGTTAPHYQLLKQGESFAV-HTCKSKS 208
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+ +W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC
Sbjct: 209 TRNPLLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDSVELHGTMKSYFGGLLCV 268
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETI 179
WS DGKYI+TGGEDDLV VWS D +V+A G GH SWVS VAFD Y +S +D E
Sbjct: 269 CWSPDGKYIVTGGEDDLVTVWSFLDCRVIARGHGHKSWVSVVAFDPYTTSVEESDPME-- 326
Query: 180 TYRFGSVGQDTQLLL 194
F +D Q LL
Sbjct: 327 ---FSGSDEDFQDLL 338
Score = 54.7 bits (130), Expect = 3e-06
Identities = 26/60 (43%), Positives = 37/60 (61%), Gaps = 1/60 (1%)
Query: 232 DNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPG 291
D A LGT P M DVP + PL+ ++ E L+ L+F ++ ++TAC+EG I W RPG
Sbjct: 474 DPAKTLGT-SLCPRMEDVPLLEPLICKKIAHERLTVLVFLEDCIVTACQEGFICTWARPG 532
>UniRef100_UPI000017F7C1 UPI000017F7C1 UniRef100 entry
Length = 539
Score = 169 bits (427), Expect = 1e-40
Identities = 91/195 (46%), Positives = 120/195 (60%), Gaps = 13/195 (6%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLY-----NKDGAGESSFPILKDQTLFSVAHARYSKS 61
SR TC+ WVPG + F+VAH+ G++Y + G + +LK F+V H SKS
Sbjct: 150 SRVTCVKWVPGSESLFLVAHSSGSMYLYNVEHTCGTTAPHYQLLKQGESFAV-HTCKSKS 208
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+ +W + +G++N +FS DG +LA V +DG+LRVF++ L KSY+GGLLC
Sbjct: 209 TRNPLLKWTVGEGALNEFAFSPDGKFLACVSQDGFLRVFNFDSVELHGTMKSYFGGLLCV 268
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETI 179
WS DGKYI+TGGEDDLV VWS D +V+A G GH SWVS VAFD Y +S +D E
Sbjct: 269 CWSPDGKYIVTGGEDDLVTVWSFLDCRVIARGHGHKSWVSVVAFDPYTTSVEESDPME-- 326
Query: 180 TYRFGSVGQDTQLLL 194
F +D Q LL
Sbjct: 327 ---FSGSDEDFQDLL 338
Score = 53.9 bits (128), Expect = 6e-06
Identities = 26/60 (43%), Positives = 37/60 (61%), Gaps = 1/60 (1%)
Query: 232 DNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPG 291
D A LGT P M DVP + PL+ ++ E L+ L+F ++ ++TAC+EG I W RPG
Sbjct: 474 DPAKTLGT-SLCPRMEDVPLLEPLICKKIAHERLTVLVFLEDCLVTACQEGFICTWARPG 532
>UniRef100_UPI00001CE7D9 UPI00001CE7D9 UniRef100 entry
Length = 1177
Score = 166 bits (420), Expect = 8e-40
Identities = 99/253 (39%), Positives = 139/253 (54%), Gaps = 45/253 (17%)
Query: 7 SRCTCISWVPGGDGAFVVAHADGNLY-----NKDGAGESSFPILKDQTLFSVAHARYSKS 61
++ T + W+P + F+ +HA G+LY + + + +LK F+V +A SK+
Sbjct: 723 TKVTYLKWLPESESLFLASHASGHLYLYNVSHPCASTPPQYSLLKQGEGFAV-YAAKSKA 781
Query: 62 --NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCC 119
NP+A+W + +G +N +FS DG +LA V +DG LRVF + L KSY+GGLLC
Sbjct: 782 PRNPLAKWAVGEGPLNEFAFSPDGRHLACVSQDGCLRVFHFDSMLLRGLMKSYFGGLLCV 841
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYW--------SSPN 171
WS DG+Y++TGGEDDLV VWS + +VVA G GH SWV+ VAFD Y +S +
Sbjct: 842 CWSPDGRYVVTGGEDDLVTVWSFTEGRVVARGHGHKSWVNAVAFDPYTTRAEETASASGD 901
Query: 172 SNDNGE----------------------TITYRFGSVGQDTQLLLWELEMDEIV--VPLR 207
+ +GE ++TYRFGS GQDTQ LW+L D + PL
Sbjct: 902 GDPSGEEEEPEATSSDTGAPVSPLPKAGSVTYRFGSAGQDTQFCLWDLTEDVLSPHPPLA 961
Query: 208 R-----GPPGGSP 215
R G PG +P
Sbjct: 962 RTRTLPGTPGATP 974
Score = 55.8 bits (133), Expect = 2e-06
Identities = 34/110 (30%), Positives = 54/110 (48%), Gaps = 3/110 (2%)
Query: 206 LRRGPPGGSPTFGSPTFSAGSQSSHWDNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPL 265
+ RG GG+ + + + + S D A LGT P + +VP + PLV ++ E L
Sbjct: 1053 ISRGGSGGNSS--NDKLNGPAPRSRLDPAKVLGTAL-CPRIHEVPLLEPLVCKKIAQERL 1109
Query: 266 SSLIFTQESVLTACREGHIKIWTRPGVAESQPSNSETLLATSLKEKPSLS 315
+ L+F ++ ++TAC+EG + W RPG A + S PS S
Sbjct: 1110 TVLLFLEDCIITACQEGLVCTWARPGKAFTDEETETQAGEASWPRSPSKS 1159
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.133 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 645,033,970
Number of Sequences: 2790947
Number of extensions: 30396262
Number of successful extensions: 82856
Number of sequences better than 10.0: 2799
Number of HSP's better than 10.0 without gapping: 1367
Number of HSP's successfully gapped in prelim test: 1446
Number of HSP's that attempted gapping in prelim test: 71055
Number of HSP's gapped (non-prelim): 10219
length of query: 325
length of database: 848,049,833
effective HSP length: 127
effective length of query: 198
effective length of database: 493,599,564
effective search space: 97732713672
effective search space used: 97732713672
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 75 (33.5 bits)
Medicago: description of AC148995.8


