

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148994.8 - phase: 0 /pseudo
(569 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
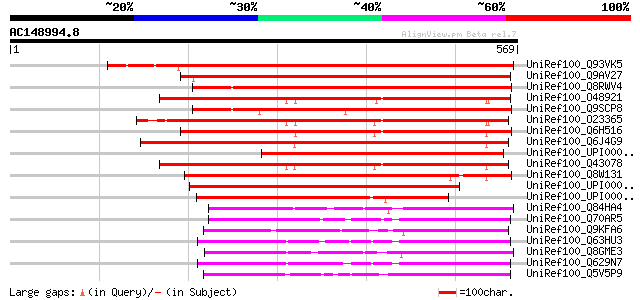
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q93VK5 At1g31800/68069_m00159 [Arabidopsis thaliana] 736 0.0
UniRef100_Q9AV27 Putative cytochrome P450 monooxygenase [Oryza s... 387 e-106
UniRef100_Q8RWV4 Putative cytochrome P450 [Arabidopsis thaliana] 370 e-101
UniRef100_O48921 Cytochrome P450 97B2 [Glycine max] 353 1e-95
UniRef100_Q9SCP8 Cytochrom P450-like protein [Arabidopsis thaliana] 352 2e-95
UniRef100_O23365 Cytochrome P450 97B3 [Arabidopsis thaliana] 352 2e-95
UniRef100_Q6H516 Putative cytochrome P450 [Oryza sativa] 349 1e-94
UniRef100_Q6J4G9 Putative 97B2-like cytochrome P450 [Ginkgo biloba] 347 5e-94
UniRef100_UPI000028DB9A UPI000028DB9A UniRef100 entry 346 1e-93
UniRef100_Q43078 Cytochrome P450 97B1 [Pisum sativum] 343 6e-93
UniRef100_Q8W131 Cytochrome P450 [Skeletonema costatum] 335 2e-90
UniRef100_UPI000028DAE2 UPI000028DAE2 UniRef100 entry 269 1e-70
UniRef100_UPI00002B0E7E UPI00002B0E7E UniRef100 entry 245 2e-63
UniRef100_Q84HA4 Cytochrome P-450 [Streptomyces carzinostaticus ... 170 9e-41
UniRef100_Q70AR5 Putative cytochrome P450 [Streptomyces peucetius] 165 4e-39
UniRef100_Q9KFA6 Cytochrome P450 hydroxylase [Bacillus halodurans] 161 4e-38
UniRef100_Q63HU3 Cytochrome P450 family protein [Burkholderia ps... 159 3e-37
UniRef100_Q8GME3 P450 hydroxylase [Streptomyces globisporus] 158 4e-37
UniRef100_Q629N7 Cytochrome P450-related protein [Burkholderia m... 157 8e-37
UniRef100_Q5V5P9 Cytochrome P450 [Haloarcula marismortui] 153 1e-35
>UniRef100_Q93VK5 At1g31800/68069_m00159 [Arabidopsis thaliana]
Length = 595
Score = 736 bits (1901), Expect = 0.0
Identities = 370/465 (79%), Positives = 420/465 (89%), Gaps = 11/465 (2%)
Query: 110 SSFIA-CSSSNGRSP--NDSVDDGVVKSADQLLEEKRRAELSAKIASGEFTVKQESGLPS 166
S+F+ SSSNGR P +SV +GV KS ++L EEKRRAELSA+IASG FTV++ S PS
Sbjct: 24 SNFVVFSSSSNGRDPLEENSVPNGV-KSLEKLQEEKRRAELSARIASGAFTVRKSS-FPS 81
Query: 167 ILKKSLSNLGVSNEILEFLFGL------YPKIPEAKGSISAIRSEAFFIPLYELYITYGG 220
+K LS +G+ + +L+F+F YPK+PEAKGSI A+R+EAFFIPLYEL++TYGG
Sbjct: 82 TVKNGLSKIGIPSNVLDFMFDWTGSDQDYPKVPEAKGSIQAVRNEAFFIPLYELFLTYGG 141
Query: 221 IFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILDFVMGKGLIPADGEIWRVRR 280
IFRL FGPKSFLIVSDP+IAKHILKDN+KAYSKGILAEILDFVMGKGLIPADGEIWR RR
Sbjct: 142 IFRLTFGPKSFLIVSDPSIAKHILKDNAKAYSKGILAEILDFVMGKGLIPADGEIWRRRR 201
Query: 281 RTIVPALHLKFVAAMIGLFGQATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIGKAVFN 340
R IVPALH K+VAAMI LFG+A+DRLCQKLD AA GE+VEMESLFSRLTLD+IGKAVFN
Sbjct: 202 RAIVPALHQKYVAAMISLFGEASDRLCQKLDAAALKGEEVEMESLFSRLTLDIIGKAVFN 261
Query: 341 YDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIWKDISPRQRKVTAALKLVNDTL 400
YDFDSL+NDTG+IEAVYTVLREAEDRS+SPIPVWD+PIWKDISPRQRKV +LKL+NDTL
Sbjct: 262 YDFDSLTNDTGVIEAVYTVLREAEDRSVSPIPVWDIPIWKDISPRQRKVATSLKLINDTL 321
Query: 401 NNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLASGDDVTSKQLRDDLMTMLIAGHET 460
++LIA CKRMV+EEELQFHEEYMNE+DPSILHFLLASGDDV+SKQLRDDLMTMLIAGHET
Sbjct: 322 DDLIATCKRMVEEEELQFHEEYMNERDPSILHFLLASGDDVSSKQLRDDLMTMLIAGHET 381
Query: 461 SAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMKKLKYTTRVINESLRLYPQP 520
SAAVLTWTFYLL+ EPSV++KLQEEVDSV+GDRFPTI+DMKKLKYTTRV+NESLRLYPQP
Sbjct: 382 SAAVLTWTFYLLTTEPSVVAKLQEEVDSVIGDRFPTIQDMKKLKYTTRVMNESLRLYPQP 441
Query: 521 PVLIRRSIEDDVLGEYPIKRGEDIFISVWNLHRSPTLWNDADKLN 565
PVLIRRSI++D+LGEYPIKRGEDIFISVWNLHRSP W+DA+K N
Sbjct: 442 PVLIRRSIDNDILGEYPIKRGEDIFISVWNLHRSPLHWDDAEKFN 486
>UniRef100_Q9AV27 Putative cytochrome P450 monooxygenase [Oryza sativa]
Length = 584
Score = 387 bits (994), Expect = e-106
Identities = 192/377 (50%), Positives = 272/377 (71%), Gaps = 6/377 (1%)
Query: 192 IPEAKGSISAIRS---EAFFIPLYELYITYGGIFRLNFGPKSFLIVSDPAIAKHILKDNS 248
IP A + +R A F+PL++ + G ++RL GP+ ++VSDPA+A+H+L+
Sbjct: 86 IPVASAKLDDVRDLLGGALFLPLFKWFREEGPVYRLAAGPRDLVVVSDPAVARHVLRGYG 145
Query: 249 KAYSKGILAEILDFVMGKGLIPADGEIWRVRRRTIVPALHLKFVAAMIG-LFGQATDRLC 307
Y KG++AE+ +F+ G G A+G +W VRRR++VP+LH +F++ M+ +F + +RL
Sbjct: 146 SRYEKGLVAEVSEFLFGSGFAIAEGALWTVRRRSVVPSLHKRFLSVMVDRVFCKCAERLV 205
Query: 308 QKLDTAASDGEDVEMESLFSRLTLDVIGKAVFNYDFDSLSNDTGIIEAVYTVLREAEDRS 367
+KL+T+A G+ V ME+ FS++TLDVIG ++FNY+FDSL++D+ +I+AVYT L+EAE RS
Sbjct: 206 EKLETSALSGKPVNMEARFSQMTLDVIGLSLFNYNFDSLTSDSPVIDAVYTALKEAELRS 265
Query: 368 ISPIPVWDLPIWKDISPRQRKVTAALKLVNDTLNNLIAICKRMVDEEELQFH-EEYMNEQ 426
+P W + + I PRQ K A+ ++ +T+ +LI CK++VD E Q EEY+NE
Sbjct: 266 TDLLPYWKIDLLCKIVPRQIKAEKAVNIIRNTVEDLITKCKKIVDAENEQIEGEEYVNEA 325
Query: 427 DPSILHFLLASGDDVTSKQLRDDLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEV 486
DPSIL FLLAS ++VTS QLRDDL++ML+AGHET+ +VLTWT YLLSK+P+ + + Q EV
Sbjct: 326 DPSILRFLLASREEVTSVQLRDDLLSMLVAGHETTGSVLTWTIYLLSKDPAALRRAQAEV 385
Query: 487 DSVLGDRFPTIEDMKKLKYTTRVINESLRLYPQPPVLIRRSIEDDVL-GEYPIKRGEDIF 545
D VL R P ED+K+LKY R INES+RLYP PPVLIRR+I DDVL G Y IK G+DI
Sbjct: 386 DRVLQGRLPRYEDLKELKYLMRCINESMRLYPHPPVLIRRAIVDDVLPGNYKIKAGQDIM 445
Query: 546 ISVWNLHRSPTLWNDAD 562
ISV+N+HRSP +W+ AD
Sbjct: 446 ISVYNIHRSPEVWDRAD 462
>UniRef100_Q8RWV4 Putative cytochrome P450 [Arabidopsis thaliana]
Length = 552
Score = 370 bits (951), Expect = e-101
Identities = 188/361 (52%), Positives = 261/361 (72%), Gaps = 4/361 (1%)
Query: 206 AFFIPLYELYITYGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILDFVMG 265
A F+PLY+ YG I+RL GP++F+IVSDPAIAKH+L++ K Y+KG++AE+ +F+ G
Sbjct: 109 ALFLPLYKWMNEYGPIYRLAAGPRNFVIVSDPAIAKHVLRNYPK-YAKGLVAEVSEFLFG 167
Query: 266 KGLIPADGEIWRVRRRTIVPALHLKFVAAMIG-LFGQATDRLCQKLDTAASDGEDVEMES 324
G A+G +W RRR +VP+LH ++++ ++ +F + +RL +KL A DG V ME+
Sbjct: 168 SGFAIAEGPLWTARRRAVVPSLHRRYLSVIVERVFCKCAERLVEKLQPYAEDGSAVNMEA 227
Query: 325 LFSRLTLDVIGKAVFNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIWKDISP 384
FS++TLDVIG ++FNY+FDSL+ D+ +IEAVYT L+EAE RS +P W + I P
Sbjct: 228 KFSQMTLDVIGLSLFNYNFDSLTTDSPVIEAVYTALKEAELRSTDLLPYWKIDALCKIVP 287
Query: 385 RQRKVTAALKLVNDTLNNLIAICKRMVDEEELQFH-EEYMNEQDPSILHFLLASGDDVTS 443
RQ K A+ L+ +T+ +LIA CK +V+ E + + EEY+N+ DPSIL FLLAS ++V+S
Sbjct: 288 RQVKAEKAVTLIRETVEDLIAKCKEIVEREGERINDEEYVNDADPSILRFLLASREEVSS 347
Query: 444 KQLRDDLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMKKL 503
QLRDDL++ML+AGHET+ +VLTWT YLLSK S + K QEEVD VL R P ED+K+L
Sbjct: 348 VQLRDDLLSMLVAGHETTGSVLTWTLYLLSKNSSALRKAQEEVDRVLEGRNPAFEDIKEL 407
Query: 504 KYTTRVINESLRLYPQPPVLIRRSIEDDVL-GEYPIKRGEDIFISVWNLHRSPTLWNDAD 562
KY TR INES+RLYP PPVLIRR+ D+L G Y + G+DI ISV+N+HRS +W A+
Sbjct: 408 KYITRCINESMRLYPHPPVLIRRAQVPDILPGNYKVNTGQDIMISVYNIHRSSEVWEKAE 467
Query: 563 K 563
+
Sbjct: 468 E 468
>UniRef100_O48921 Cytochrome P450 97B2 [Glycine max]
Length = 576
Score = 353 bits (905), Expect = 1e-95
Identities = 197/416 (47%), Positives = 273/416 (65%), Gaps = 23/416 (5%)
Query: 169 KKSLSNL--GVSNEILEFLFG-LYPKIPEAKGSISAIRSEAFFIPLYELYITYGGIFRLN 225
KKS NL SN + + L G +P A+G++S + F LY+ ++ +G +++L
Sbjct: 53 KKSSRNLLGNASNLLTDLLSGGSIGSMPIAEGAVSDLLGRPLFFSLYDWFLEHGAVYKLA 112
Query: 226 FGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILDFVMGKGLIPADGEIWRVRRRTIVP 285
FGPK+F++VSDP +A+HIL++N+ +Y KG+LA+IL+ +MGKGLIPAD + W+ RRR I P
Sbjct: 113 FGPKAFVVVSDPIVARHILRENAFSYDKGVLADILEPIMGKGLIPADLDTWKQRRRVIAP 172
Query: 286 ALHLKFVAAMIGLFGQATDRLCQK----LDTAASDGED---VEMESLFSRLTLDVIGKAV 338
A H ++ AM+ +F ++R K L+ DG D +++E+ FS L LD+IG V
Sbjct: 173 AFHNSYLEAMVKIFTTCSERTILKFNKLLEGEGYDGPDSIELDLEAEFSSLALDIIGLGV 232
Query: 339 FNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIWKDISPRQRKVTAALKLVND 398
FNYDF S++ ++ +I+AVY L EAE RS IP W +P+ + I PRQRK LK++N
Sbjct: 233 FNYDFGSVTKESPVIKAVYGTLFEAEHRSTFYIPYWKIPLARWIVPRQRKFQDDLKVINT 292
Query: 399 TLNNLIAICKRM---VDEEELQFHEEYMNEQDPSILHFLL-ASGDDVTSKQLRDDLMTML 454
L+ LI K D E+LQ +Y+N +D S+L FL+ G DV +QLRDDLMTML
Sbjct: 293 CLDGLIRNAKESRQETDVEKLQ-QRDYLNLKDASLLRFLVDMRGADVDDRQLRDDLMTML 351
Query: 455 IAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMKKLKYTTRVINESL 514
IAGHET+AAVLTW +LL++ PS M K Q EVD VLG PT E +K+L+Y ++ E+L
Sbjct: 352 IAGHETTAAVLTWAVFLLAQNPSKMKKAQAEVDLVLGTGRPTFESLKELQYIRLIVVEAL 411
Query: 515 RLYPQPPVLIRRSIEDDVL-----GE---YPIKRGEDIFISVWNLHRSPTLWNDAD 562
RLYPQPP+LIRRS++ DVL GE Y I G D+FISV+NLHRSP W+ D
Sbjct: 412 RLYPQPPLLIRRSLKSDVLPGGHKGEKDGYAIPAGTDVFISVYNLHRSPYFWDRPD 467
>UniRef100_Q9SCP8 Cytochrom P450-like protein [Arabidopsis thaliana]
Length = 566
Score = 352 bits (903), Expect = 2e-95
Identities = 188/388 (48%), Positives = 262/388 (67%), Gaps = 31/388 (7%)
Query: 206 AFFIPLYELYITYGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILDFVMG 265
A F+PLY+ YG I+RL GP++F+IVSDPAIAKH+L++ K Y+KG++AE+ +F+ G
Sbjct: 96 ALFLPLYKWMNEYGPIYRLAAGPRNFVIVSDPAIAKHVLRNYPK-YAKGLVAEVSEFLFG 154
Query: 266 KGLIPADGEIWRV---------------RRRTIVPALHLKFVAAMIG-LFGQATDRLCQK 309
G A+G +W V +RR +VP+LH ++++ ++ +F + +RL +K
Sbjct: 155 SGFAIAEGPLWTVISSPPISILKFLELWKRRAVVPSLHRRYLSVIVERVFCKCAERLVEK 214
Query: 310 LDTAASDGEDVEMESLFSRLTLDVIGKAVFNYDFDSLSNDTGIIEAVYTVLREAEDRSIS 369
L A DG V ME+ FS++TLDVIG ++FNY+FDSL+ D+ +IEAVYT L+EAE RS
Sbjct: 215 LQPYAEDGSAVNMEAKFSQMTLDVIGLSLFNYNFDSLTTDSPVIEAVYTALKEAELRSTD 274
Query: 370 PIPVWD------------LPIWKDISPRQRKVTAALKLVNDTLNNLIAICKRMVDEEELQ 417
+P W + I PRQ K A+ L+ +T+ +LIA CK +V+ E +
Sbjct: 275 LLPYWKASFLCFFCGLLIIDALCKIVPRQVKAEKAVTLIRETVEDLIAKCKEIVEREGER 334
Query: 418 FH-EEYMNEQDPSILHFLLASGDDVTSKQLRDDLMTMLIAGHETSAAVLTWTFYLLSKEP 476
+ EEY+N+ DPSIL FLLAS ++V+S QLRDDL++ML+AGHET+ +VLTWT YLLSK
Sbjct: 335 INDEEYVNDADPSILRFLLASREEVSSVQLRDDLLSMLVAGHETTGSVLTWTLYLLSKNS 394
Query: 477 SVMSKLQEEVDSVLGDRFPTIEDMKKLKYTTRVINESLRLYPQPPVLIRRSIEDDVL-GE 535
S + K QEEVD VL R P ED+K+LKY TR INES+RLYP PPVLIRR+ D+L G
Sbjct: 395 SALRKAQEEVDRVLEGRNPAFEDIKELKYITRCINESMRLYPHPPVLIRRAQVPDILPGN 454
Query: 536 YPIKRGEDIFISVWNLHRSPTLWNDADK 563
Y + G+DI ISV+N+HRS +W A++
Sbjct: 455 YKVNTGQDIMISVYNIHRSSEVWEKAEE 482
>UniRef100_O23365 Cytochrome P450 97B3 [Arabidopsis thaliana]
Length = 580
Score = 352 bits (902), Expect = 2e-95
Identities = 198/439 (45%), Positives = 284/439 (64%), Gaps = 33/439 (7%)
Query: 143 RRAELSAKIASGEFTVKQESGLPSILKKSLSNLGVSNEILEFLFG-LYPKIPEAKGSISA 201
RRA +S K S E P L N SN + FL G +P A+GS+S
Sbjct: 45 RRASVSIKCQSTE---------PKTNGNILDN--ASNLLTNFLSGGSLGSMPTAEGSVSD 93
Query: 202 IRSEAFFIPLYELYITYGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILD 261
+ + F+ LY+ ++ +GGI++L FGPK+F+++SDP IA+H+L++N+ +Y KG+LAEIL+
Sbjct: 94 LFGKPLFLSLYDWFLEHGGIYKLAFGPKAFVVISDPIIARHVLRENAFSYDKGVLAEILE 153
Query: 262 FVMGKGLIPADGEIWRVRRRTIVPALHLKFVAAMIGLFGQATDRLCQK-----LDTAASD 316
+MGKGLIPAD + W++RRR I PA H ++ AM+ +F ++++ K + S
Sbjct: 154 PIMGKGLIPADLDTWKLRRRAITPAFHKLYLEAMVKVFSDCSEKMILKSEKLIREKETSS 213
Query: 317 GED---VEMESLFSRLTLDVIGKAVFNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPV 373
GED +++E+ FS L LD+IG +VFNYDF S++ ++ +I+AVY L EAE RS P
Sbjct: 214 GEDTIELDLEAEFSSLALDIIGLSVFNYDFGSVTKESPVIKAVYGTLFEAEHRSTFYFPY 273
Query: 374 WDLPIWKDISPRQRKVTAALKLVNDTLNNLIAICK---RMVDEEELQFHEEYMNEQDPSI 430
W+ P + I PRQRK + LK++ND L+ LI K + D E+LQ +Y N +D S+
Sbjct: 274 WNFPPARWIVPRQRKFQSDLKIINDCLDGLIQNAKETRQETDVEKLQ-ERDYTNLKDASL 332
Query: 431 LHFLL-ASGDDVTSKQLRDDLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSV 489
L FL+ G D+ +QLRDDLMTMLIAGHET+AAVLTW +LLS+ P + K Q E+D+V
Sbjct: 333 LRFLVDMRGVDIDDRQLRDDLMTMLIAGHETTAAVLTWAVFLLSQNPEKIRKAQAEIDAV 392
Query: 490 LGDRFPTIEDMKKLKYTTRVINESLRLYPQPPVLIRRSIEDDVL-----GE---YPIKRG 541
LG PT E MKKL+Y ++ E LRL+PQPP+LIRR+++ + L GE + + +G
Sbjct: 393 LGQGPPTYESMKKLEYIRLIVVEVLRLFPQPPLLIRRTLKPETLPGGHKGEKEGHKVPKG 452
Query: 542 EDIFISVWNLHRSPTLWND 560
DIFISV+NLHRSP W++
Sbjct: 453 TDIFISVYNLHRSPYFWDN 471
>UniRef100_Q6H516 Putative cytochrome P450 [Oryza sativa]
Length = 571
Score = 349 bits (895), Expect = 1e-94
Identities = 183/390 (46%), Positives = 263/390 (66%), Gaps = 19/390 (4%)
Query: 192 IPEAKGSISAIRSEAFFIPLYELYITYGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAY 251
+P A+G+++ + F LY+ ++ +G +++L FGPK+F++VSDP +A+HIL++N+ Y
Sbjct: 78 MPVAEGAVTDLFGRPLFFSLYDWFLEHGSVYKLAFGPKAFVVVSDPIVARHILRENAFCY 137
Query: 252 SKGILAEILDFVMGKGLIPADGEIWRVRRRTIVPALHLKFVAAMIGLFGQATDRLCQKLD 311
KG+LAEIL +MGKGLIPAD + W+ RR+ I P H F+ AM+G+F + ++R KL+
Sbjct: 138 DKGVLAEILKPIMGKGLIPADLDTWKQRRKVITPGFHALFIDAMVGVFTKCSERTIFKLE 197
Query: 312 TAASDGED------VEMESLFSRLTLDVIGKAVFNYDFDSLSNDTGIIEAVYTVLREAED 365
GE V++E+ FS L LD+IG VFN+DFDS++ ++ +I+AVY L EAE
Sbjct: 198 ELIERGEHGEKYTIVDLEAEFSNLALDIIGLGVFNFDFDSVTKESPVIKAVYGTLFEAEH 257
Query: 366 RSISPIPVWDLPIWKDISPRQRKVTAALKLVNDTLNNLIAICK---RMVDEEELQFHEEY 422
RS IP W+LP+ + I PRQRK + LK++ND L++LI K + D E+LQ +Y
Sbjct: 258 RSTFYIPYWNLPLTRWIVPRQRKFHSDLKVINDCLDSLIKNAKETRQEADVEKLQ-QRDY 316
Query: 423 MNEQDPSILHFLL-ASGDDVTSKQLRDDLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSK 481
+ +D S+L FL+ G DV +QLRDDLMTMLIAGHET+AAVLTW+ +LL++ PS M K
Sbjct: 317 SSLKDASLLRFLVDMRGADVDDRQLRDDLMTMLIAGHETTAAVLTWSVFLLAQNPSKMRK 376
Query: 482 LQEEVDSVLGDRFPTIEDMKKLKYTTRVINESLRLYPQPPVLIRRSIEDDVL-------- 533
Q EVDSVL + ++ +KKL+Y +I E+LRLYPQPP+LIRR++ D L
Sbjct: 377 AQAEVDSVLSNETINVDQLKKLEYIRLIIVEALRLYPQPPLLIRRALRPDKLPGGYNGAK 436
Query: 534 GEYPIKRGEDIFISVWNLHRSPTLWNDADK 563
Y I G DIF+S++NLHRSP W+ D+
Sbjct: 437 EGYEIPAGTDIFLSIYNLHRSPYFWDRPDE 466
>UniRef100_Q6J4G9 Putative 97B2-like cytochrome P450 [Ginkgo biloba]
Length = 586
Score = 347 bits (890), Expect = 5e-94
Identities = 190/433 (43%), Positives = 274/433 (62%), Gaps = 21/433 (4%)
Query: 148 SAKIASGEFTVKQESGLPSILKKSLSNL--GVSNEILEFLFG-LYPKIPEAKGSISAIRS 204
S ++ +F + ES KS L SN + L G +P A+G++S +
Sbjct: 38 SFRLPGSKFFPRCESSSTERATKSNRTLLDNASNFLTNLLSGGQLGTMPIAEGAVSDLFG 97
Query: 205 EAFFIPLYELYITYGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILDFVM 264
+ F LY+ +I +G +++L FGPKSF++VSDP + +HIL++N+ AY KG+LA+IL+ +M
Sbjct: 98 KPLFFSLYDWFIEHGSVYKLAFGPKSFVVVSDPIVVRHILRENAFAYDKGVLADILEPIM 157
Query: 265 GKGLIPADGEIWRVRRRTIVPALHLKFVAAMIGLFGQATDRLCQKLDTAASDGE------ 318
GKGLIPAD W+ RR+ IVP H F+ AM+ +FG ++R KL + E
Sbjct: 158 GKGLIPADLGTWKPRRKAIVPGFHAAFLEAMVKVFGDCSERTINKLQSLLLAAEADKTMH 217
Query: 319 -DVEMESLFSRLTLDVIGKAVFNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLP 377
DV+ME+ FS L LD+IG +VFNYDF S++ ++ +I+AVY L EAE RS IP W P
Sbjct: 218 IDVDMEAEFSNLALDIIGLSVFNYDFGSVTRESPVIKAVYGTLFEAEHRSTFYIPYWKFP 277
Query: 378 IWKDISPRQRKVTAALKLVNDTLNNLIAICKRMVDEEELQFHE--EYMNEQDPSILHFLL 435
+ + + PRQRK LK++N+ L++LI K E++++ + +Y +D S+L FL+
Sbjct: 278 LARWLVPRQRKFHEDLKIINECLDSLIQGAKETRQEDDIEALQGRDYSKVKDASLLRFLV 337
Query: 436 -ASGDDVTSKQLRDDLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRF 494
G DV + QLRDDLMTMLIAGHET+AAVLTW +LL++ + K Q E+D++L R
Sbjct: 338 DMKGVDVDNGQLRDDLMTMLIAGHETTAAVLTWALFLLAQNTDKLVKAQAEIDTILDQRK 397
Query: 495 PTIEDMKKLKYTTRVINESLRLYPQPPVLIRRSIEDDVL--------GEYPIKRGEDIFI 546
PT ED+K+L+Y +I E+LRLYPQPP+LIRR++ D + Y I +G DIFI
Sbjct: 398 PTFEDIKRLQYIRLIIAEALRLYPQPPLLIRRALRQDTIPGGYRGDKDGYLIPKGTDIFI 457
Query: 547 SVWNLHRSPTLWN 559
SV+NLHRSP W+
Sbjct: 458 SVYNLHRSPYFWD 470
>UniRef100_UPI000028DB9A UPI000028DB9A UniRef100 entry
Length = 272
Score = 346 bits (887), Expect = 1e-93
Identities = 173/272 (63%), Positives = 215/272 (78%)
Query: 283 IVPALHLKFVAAMIGLFGQATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIGKAVFNYD 342
IVP+LH K+V +M+ +FG + +L A GE VEME+ +SR LD+IGKAVFNYD
Sbjct: 1 IVPSLHKKYVTSMVDMFGDCGLKGMSQLARAEKAGESVEMENFYSRYALDIIGKAVFNYD 60
Query: 343 FDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIWKDISPRQRKVTAALKLVNDTLNN 402
FDSL+ D +I+AVYTVLREAE RS++ IP W +P + + PRQRK AL++VNDTL+
Sbjct: 61 FDSLTTDDPVIKAVYTVLREAEYRSVTFIPYWKVPPLRWLVPRQRKCQEALEVVNDTLDE 120
Query: 403 LIAICKRMVDEEELQFHEEYMNEQDPSILHFLLASGDDVTSKQLRDDLMTMLIAGHETSA 462
LI+ CK++V+EE+ +F EEYMN DPSILHFL+ASGDDVTSKQLRDDLMT+LIAGHET+A
Sbjct: 121 LISRCKKVVEEEDEEFVEEYMNTDDPSILHFLIASGDDVTSKQLRDDLMTLLIAGHETTA 180
Query: 463 AVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMKKLKYTTRVINESLRLYPQPPV 522
AVLTWT +LL+K P V +K+ EEVD V+GDR PT+ DM++L YTTRVINES+RLYPQPPV
Sbjct: 181 AVLTWTTFLLAKHPEVKAKVFEEVDRVVGDRNPTVADMRELVYTTRVINESMRLYPQPPV 240
Query: 523 LIRRSIEDDVLGEYPIKRGEDIFISVWNLHRS 554
LIRR++E LG Y I G D FISVWNLHR+
Sbjct: 241 LIRRALEPVSLGGYNIDAGTDFFISVWNLHRN 272
>UniRef100_Q43078 Cytochrome P450 97B1 [Pisum sativum]
Length = 552
Score = 343 bits (881), Expect = 6e-93
Identities = 191/413 (46%), Positives = 268/413 (64%), Gaps = 23/413 (5%)
Query: 169 KKSLSNL--GVSNEILEFLFGL-YPKIPEAKGSISAIRSEAFFIPLYELYITYGGIFRLN 225
K+S N+ SN + L G +P A+G+++ + F LY+ ++ +G +++L
Sbjct: 62 KQSSRNVFDNASNLLTSLLSGANLGSMPIAEGAVTDLFDRPLFFSLYDWFLEHGSVYKLA 121
Query: 226 FGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILDFVMGKGLIPADGEIWRVRRRTIVP 285
FGPK+F++VSDP +A+HIL++N+ +Y KG+LA+IL+ +MGKGLIPAD E W+ RRR I P
Sbjct: 122 FGPKAFVVVSDPIVARHILRENAFSYDKGVLADILEPIMGKGLIPADLETWKQRRRVIAP 181
Query: 286 ALHLKFVAAMIGLFGQATDRLCQK----LDTAASDGE---DVEMESLFSRLTLDVIGKAV 338
H ++ AM+ LF ++R K L+ DG+ ++++E+ FS L L++IG V
Sbjct: 182 GFHTSYLEAMVQLFTSCSERTVLKVNELLEGEGRDGQKSVELDLEAEFSNLALEIIGLGV 241
Query: 339 FNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIWKDISPRQRKVTAALKLVND 398
FNYDF S++N++ +I+AVY L EAE RS IP W P+ + I PRQRK LK++N
Sbjct: 242 FNYDFGSVTNESPVIKAVYGTLFEAEHRSTFYIPYWKFPLARWIVPRQRKFQDDLKVINT 301
Query: 399 TLNNLIAICK---RMVDEEELQFHEEYMNEQDPSILHFLL-ASGDDVTSKQLRDDLMTML 454
L+ LI K + D E+LQ +Y N +D S+L FL+ G DV +QLRDDLMTML
Sbjct: 302 CLDGLIRNAKESRQETDVEKLQ-QRDYSNLKDASLLRFLVDMRGVDVDDRQLRDDLMTML 360
Query: 455 IAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMKKLKYTTRVINESL 514
IAGHET+AAVLTW +LL++ P M K Q EVD VLG PT E +KKL+Y ++ E+L
Sbjct: 361 IAGHETTAAVLTWAVFLLAQNPDKMKKAQAEVDLVLGMGKPTFELLKKLEYIRLIVVETL 420
Query: 515 RLYPQPPVLIRRSIEDDVL--------GEYPIKRGEDIFISVWNLHRSPTLWN 559
RLYPQPP+LIRRS++ DVL Y I G D+FISV+NLHRSP W+
Sbjct: 421 RLYPQPPLLIRRSLKPDVLPGGHKGDKDGYTIPAGTDVFISVYNLHRSPYFWD 473
>UniRef100_Q8W131 Cytochrome P450 [Skeletonema costatum]
Length = 659
Score = 335 bits (859), Expect = 2e-90
Identities = 185/383 (48%), Positives = 255/383 (66%), Gaps = 19/383 (4%)
Query: 197 GSISAIRSEAFFIPLYELYITYGGIFRLNFGPKSFLIVSDPAIAKHILKDNS-KAYSKGI 255
G + + F+ L + Y +G IF L+FGPKSFL++SDP +A+HIL+D+S + Y KG+
Sbjct: 129 GDLQTLAGGPLFLLLAKYYTDHGPIFNLSFGPKSFLVISDPVMARHILRDSSPEQYCKGM 188
Query: 256 LAEILDFVMGKGLIPADGEIWRVRRRTIVPALHLKFVAAMIGLFGQATDRLCQKLDT-AA 314
LAEIL+ +MG GLIPAD +IW+VRRR +VP H K++ +MIGLFG DRL L+ +
Sbjct: 189 LAEILEPIMGDGLIPADPKIWKVRRRAVVPGFHKKWLNSMIGLFGDCGDRLVDDLEKRST 248
Query: 315 SDGEDVEMESLFSRLTLDVIGKAVFNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVW 374
SD ++ME F +TLD+IGKAVFNYDF S++ ++ I++AVY VLREAE RS S IP W
Sbjct: 249 SDKPVIDMEERFCSVTLDIIGKAVFNYDFGSVTKESPIVKAVYRVLREAEHRSSSFIPYW 308
Query: 375 DLPIWKDISPRQRKVTAALKLVNDTLNNLIAICKRMVDEEELQFHEEYMNEQDPSILHFL 434
+LP + Q + + +++D L LI E ++ EE DPS+L FL
Sbjct: 309 NLPYAEKWMVGQVEFRKDMGMLDDILAKLINRAVETRQEATVEELEERETSDDPSLLRFL 368
Query: 435 L-ASGDDVTSKQLRDDLMTMLIAGHETSAAVLTWT-FYLLSKEPSVMSKLQEEVDSVLGD 492
+ G+D+TSK LRDDLMTMLIAGHET+AA+LTWT F L+S +P +M ++Q EV +V+G+
Sbjct: 369 VDMRGEDLTSKVLRDDLMTMLIAGHETTAAMLTWTMFGLVSNDPGMMKEIQAEVRTVMGN 428
Query: 493 R----FPTIEDMKKLKYTTRVINESLRLYPQPPVLIRRSIEDDVL--------GEYPIKR 540
+ + + MKKL+Y + E+LRLYP+PPVLIRR+ ++D L G + R
Sbjct: 429 KSRPDYDDVVAMKKLRY---ALIEALRLYPEPPVLIRRARQEDTLPPGGTGLSGGVKVLR 485
Query: 541 GEDIFISVWNLHRSPTLWNDADK 563
G DIFIS WNLHR+P W +ADK
Sbjct: 486 GTDIFISTWNLHRAPEYWENADK 508
>UniRef100_UPI000028DAE2 UPI000028DAE2 UniRef100 entry
Length = 311
Score = 269 bits (688), Expect = 1e-70
Identities = 142/308 (46%), Positives = 199/308 (64%), Gaps = 5/308 (1%)
Query: 202 IRSEAFFIPLYELYITYGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILD 261
+R+ F+PLY+ Y YGG++ L GPK F++VSDP + + KD + ++SKGIL +I++
Sbjct: 4 VRAAPLFVPLYDYYREYGGVYNLGAGPKWFVVVSDPVAVRTMFKDRADSFSKGILTDIME 63
Query: 262 FVMGKGLIPADGEIWRVRRRTIVPALHLKFVAAMIGLFGQATDRLCQKLDTAASDGEDVE 321
+MG GLIPA+ EIW RR I H ++ M LFG + RL KLD AA E VE
Sbjct: 64 PIMGDGLIPANKEIWAKRRPVIGAGFHNAWLKHMCDLFGASAMRLADKLDAAAETKETVE 123
Query: 322 MESLFSRLTLDVIGKAVFNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIWKD 381
+E + LDVIGKAVFNY+F +L +T II+AVY VLRE+E RS P+ W +P D
Sbjct: 124 LEGELYSMALDVIGKAVFNYEFGALREETPIIKAVYRVLRESEHRSTFPLQYWQIPGAMD 183
Query: 382 ISPRQRKVTAALKLVNDTLNNLI--AICKRMVDEEELQFHEEYMNEQDPSILHFLL-ASG 438
+ PRQ++ +K++N+ L+ LI AI R + E +Y +D S+L FL+ G
Sbjct: 184 VVPRQKQFKEDMKMINEELSVLINNAISSRTETDLEEMERRDYSKVEDASLLRFLVDIRG 243
Query: 439 DDVTSKQLRDDLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDR--FPT 496
D+ TS QLRDDLMTMLIAGHET+AAVLTWT YLL++ P + + EE+++ + D PT
Sbjct: 244 DEATSTQLRDDLMTMLIAGHETTAAVLTWTLYLLAQHPEIAEEAVEEINACVSDSNGVPT 303
Query: 497 IEDMKKLK 504
E+++KL+
Sbjct: 304 PEEVRKLQ 311
>UniRef100_UPI00002B0E7E UPI00002B0E7E UniRef100 entry
Length = 299
Score = 245 bits (626), Expect = 2e-63
Identities = 131/289 (45%), Positives = 182/289 (62%), Gaps = 8/289 (2%)
Query: 210 PLYELYITYGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILDFVMGKGLI 269
PLY+ Y YGG++ L GPK F++VSDP + +H+ KDN+ A+SKGIL +I++ +MG GLI
Sbjct: 1 PLYDYYRQYGGVYNLGAGPKWFVVVSDPVVVRHMFKDNADAFSKGILTDIMEPIMGDGLI 60
Query: 270 PADGEIWRVRRRTIVPALHLKFVAAMIGLFGQATDRLCQKLDTAASDGE-DVEMESLFSR 328
PA EIW RR T+ H ++ M LFG + L KL+ D + V +E
Sbjct: 61 PAPKEIWAKRRPTVGAGFHGAWLKHMTNLFGASATNLADKLEREWCDKDVAVNLEDELYA 120
Query: 329 LTLDVIGKAVFNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIWKDISPRQRK 388
+ LDVIGKAVFNY+F +L +T +I+AVY VLRE+E RS P+ W++P D+ PRQ++
Sbjct: 121 MALDVIGKAVFNYEFGALREETPLIKAVYRVLRESEHRSTFPLQYWNIPGAMDVVPRQKQ 180
Query: 389 VTAALKLVNDTLNNLIAICKRMVDEEELQFHE----EYMNEQDPSILHFLL-ASGDDVTS 443
+ +N L+ LIA + D E E +Y N +D S+L FL+ G+ V+S
Sbjct: 181 FKEDIAAINAELSKLIA--DALADRNETDLAEMESRDYANVEDASLLRFLVDVRGETVSS 238
Query: 444 KQLRDDLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGD 492
QLRDDLMTMLIAGHET+AAVLTWT YLL+ P + EV++++ D
Sbjct: 239 TQLRDDLMTMLIAGHETTAAVLTWTMYLLATHPEEAELARAEVEAIVAD 287
>UniRef100_Q84HA4 Cytochrome P-450 [Streptomyces carzinostaticus subsp.
neocarzinostaticus]
Length = 450
Score = 170 bits (431), Expect = 9e-41
Identities = 113/350 (32%), Positives = 179/350 (50%), Gaps = 28/350 (8%)
Query: 224 LNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILDFVMGKGLIPADGEIWRVRRRTI 283
++ GPK + + P AKH+L DN+ Y KGI ++G GL+ ++GE+WR +RRT+
Sbjct: 41 VSMGPKKLFVFNRPDYAKHVLADNAANYRKGIGLIESRKMLGDGLLTSEGELWREQRRTV 100
Query: 284 VPALHLKFVAAMIGLFGQATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIGKAVFNYDF 343
PA VAA + T L L +DG V++ + TL V+G+ V N D
Sbjct: 101 QPAFRPARVAAQADAVAEETMNLRDLLMRRGADGP-VDVLQEVTGFTLGVLGRTVLNTDL 159
Query: 344 DSLSNDTGIIEAVYTVLREAEDRSISPIPVWDL-PIWKDISPRQRKVTAALKLVNDTLNN 402
EAV +D+++ + ++ P W ++ ++R A +L+ T++
Sbjct: 160 GGYGGIAHAFEAV-------QDQAMFDMVTQNMVPTWAPLATQRRFRRARRELIR-TVDE 211
Query: 403 LIAI-CKRMVDEEELQFHEEYM-----NEQDPSILHFLLASGDDVTSKQLRDDLMTMLIA 456
L+A RM D EE M + DP +LRD+L+T+L+A
Sbjct: 212 LVADRSARMTDGEEADDAFSLMIAAARRQTDPR-----------TGQGRLRDELVTLLLA 260
Query: 457 GHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGD-RFPTIEDMKKLKYTTRVINESLR 515
GHET+A+ L WT LL++ P + ++EE VL D R P D++KL YTT+V+ E++R
Sbjct: 261 GHETTASTLAWTLLLLARHPHMRDLVREEARGVLADGRAPDAGDLRKLTYTTQVVQEAMR 320
Query: 516 LYPQPPVLIRRSIEDDVLGEYPIKRGEDIFISVWNLHRSPTLWNDADKLN 565
LYP +L R + + D +G Y + G D+ I + LHR P LW ++ +
Sbjct: 321 LYPPVWILPRVARQSDEVGPYSVSAGADVLICPYTLHRHPDLWERPEQFD 370
>UniRef100_Q70AR5 Putative cytochrome P450 [Streptomyces peucetius]
Length = 477
Score = 165 bits (417), Expect = 4e-39
Identities = 110/342 (32%), Positives = 172/342 (50%), Gaps = 20/342 (5%)
Query: 224 LNF-GPKSFLIVSDPAIAKHILKDNSKAYSKGILAE-ILDFVMGKGLIPADGEIWRVRRR 281
+NF GP +++ P +H+L+D K Y + + L ++G GL+ A+G W RR
Sbjct: 67 VNFRGPLRINLLAHPDYVQHVLRDQHKRYPRPRKVQGCLSTIVGDGLVAAEGTSWLRSRR 126
Query: 282 TIVPALHLKFVAAMIGLFGQATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIGKAVFNY 341
PA H + F +T L + A DGE ++++S L+L + +A+F
Sbjct: 127 LTQPAFHRDILRRFGDSFTASTAELLDGWERRARDGEPLDIKSEMMHLSLANLARALFRT 186
Query: 342 DF-DSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIWKDISPRQRKVTAALKLVNDTL 400
++ D +S I AV L R SP+ P S + + AL+ +N L
Sbjct: 187 EWTDEVSR---IEPAVQEALGFTHRRMTSPVDPLKFP-----SAARTRFRGALETINSLL 238
Query: 401 NNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLASGDDVTSKQLRDDLMTMLIAGHET 460
++A +R ++L ++ DP SG T +Q+RD++ +AGHET
Sbjct: 239 YPMVAERRRDGGGDDLV--SLLIDAVDPE-------SGGMFTDEQIRDEVSGFFVAGHET 289
Query: 461 SAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMKKLKYTTRVINESLRLYPQP 520
+ L+WT+YLLS P +LQ EVD VL R PT++D+ KL YTT V+ ES+RLYP
Sbjct: 290 VSTALSWTWYLLSLNPESRRRLQAEVDEVLAGRVPTVDDLPKLTYTTMVLQESMRLYPPI 349
Query: 521 PVLIRRSIEDDVLGEYPIKRGEDIFISVWNLHRSPTLWNDAD 562
V +R + EDDV+G Y I G + + + HR P W++ +
Sbjct: 350 FVYMRCAAEDDVIGGYHIPEGRWVVVCPYVTHRHPEFWDNPE 391
>UniRef100_Q9KFA6 Cytochrome P450 hydroxylase [Bacillus halodurans]
Length = 453
Score = 161 bits (408), Expect = 4e-38
Identities = 105/352 (29%), Positives = 181/352 (50%), Gaps = 27/352 (7%)
Query: 218 YGGIFRLNFG-PKSFLIVSDPAIAKHILKDNSKAYSKGILAE-ILDFVMGKGLIPADGEI 275
YG I RL F + +++DP +++ + +SKG + I+ V G GL+ ++G
Sbjct: 37 YGEIVRLRFERERDTFLLNDPKHIQYVFMNKGGEFSKGYQQDPIMGLVFGNGLLTSEGSF 96
Query: 276 WRVRRRTIVPALHLKFVAAMIGLFGQATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIG 335
W +RR PA H K +A + C+++ D + ++ +LT+ +
Sbjct: 97 WLRQRRLSQPAFHPKRIAD----YADTMVGYCERMLNTWMDNDTRDINDEMMQLTMAIAT 152
Query: 336 KAVFNYDFDS--LSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIWKDISPRQRKVTAAL 393
K +F+ D + ++ V T E + + + + K + P R++ A+
Sbjct: 153 KTLFDLDLHKGDTQEASRSLDTVMTAFNEQMTNVFRHV-LHLIGLGKLVPPVSRELREAV 211
Query: 394 KLVNDTLNNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLASGDD-----VTSKQLRD 448
+ ++ + ++I EE + H + +L L+++ D+ +T +QLRD
Sbjct: 212 ESLDKMIYSII---------EERRKHPGDRGD----LLSMLISTYDEDDGSYMTDRQLRD 258
Query: 449 DLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMKKLKYTTR 508
+++T+ +AGHET+A L+W FYLLS+ P V KL +EV VLG+R T+EDM KL Y
Sbjct: 259 EIITLFLAGHETTANTLSWAFYLLSQHPHVEEKLYQEVSQVLGNRPATLEDMPKLSYAEH 318
Query: 509 VINESLRLYPQPPVLIRRSIEDDVLGEYPIKRGEDIFISVWNLHRSPTLWND 560
VI E+LR+ P ++ RR+ +D LG+Y I G +I IS W +HR+P +ND
Sbjct: 319 VIKETLRVQPTVWLISRRAEKDVTLGDYHISAGSEIMISQWGMHRNPRYFND 370
>UniRef100_Q63HU3 Cytochrome P450 family protein [Burkholderia pseudomallei]
Length = 1373
Score = 159 bits (401), Expect = 3e-37
Identities = 104/353 (29%), Positives = 177/353 (49%), Gaps = 17/353 (4%)
Query: 211 LYELYITYGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILDFVMGKGLIP 270
L L+ YG + R GP ++ P +++L++N + Y +G + G GL+
Sbjct: 30 LLRLHQQYGDVARNRLGPFVTHALAHPDHIQYVLQENHRNYVRGRFYDNFKMFFGDGLLT 89
Query: 271 ADGEIWRVRRRTIVPALHLKFVAAMIGLFGQATDRLCQKLDTAASDGEDVEMESLFSRLT 330
DGE WR RR + P H K V A G A L + +A G+ +++ L+
Sbjct: 90 TDGEFWRRHRRVVQPLFHKKQVDAHTAAVGDAALALAHRW-SALPPGKALDVVEEMMHLS 148
Query: 331 LDVIGKAVFNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWD-LPIWKDISPRQRKV 389
L ++G VFN D S + EAV +R + + + D +P W + R++
Sbjct: 149 LRMLGLMVFNTDVSSHA------EAVGPAVRFGIEAMMPQGNLNDFIPRWAP-TRFNRRI 201
Query: 390 TAALKLVNDTLNNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLASGDDVTSKQLRDD 449
A + ++ + +IA R E +N +DP +G +T +++ D+
Sbjct: 202 AHARRAIDTIVAKIIAD-HREARCEPSDVISLLLNARDPD-------TGAPMTQQEVHDE 253
Query: 450 LMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMKKLKYTTRV 509
+MT+ +AGHET+ A L W Y L++ P+V+ +L++E+D+ LG R PT++D ++L Y ++V
Sbjct: 254 VMTVFLAGHETTGAGLAWALYALAQHPAVLRQLRDELDARLGGRAPTVQDFEQLPYLSQV 313
Query: 510 INESLRLYPQPPVLIRRSIEDDVLGEYPIKRGEDIFISVWNLHRSPTLWNDAD 562
++E LR+YP R +EDD +G Y I +F+S + HR P W + D
Sbjct: 314 VDEVLRVYPPIWGFTRDLVEDDEIGGYRIPARSSVFMSPYVTHRHPAFWRNPD 366
>UniRef100_Q8GME3 P450 hydroxylase [Streptomyces globisporus]
Length = 449
Score = 158 bits (400), Expect = 4e-37
Identities = 107/354 (30%), Positives = 178/354 (50%), Gaps = 26/354 (7%)
Query: 219 GGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILDFVMGKGLIPADGEIWRV 278
G R++ GPK I + P AKH+L DNS Y KGI V+G GL+ +DGE WR
Sbjct: 35 GDAVRVSMGPKKLYIFNRPDYAKHVLADNSDNYHKGIGLVQSRRVLGDGLLTSDGETWRE 94
Query: 279 RRRTIVPALHLKFVAAMIGLFGQATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIGKAV 338
+RR + PA + + +L L G V++ + LTL V+G+ +
Sbjct: 95 QRRIVQPAFKPGRINQQAAAVAEEAAKLVALL-RGHEGGGPVDVLQEVTGLTLGVLGRTL 153
Query: 339 FNYDFDSLSNDTGIIEAVYTVLREAEDRS-ISPIPVWDLPIWKDISPRQRKVTAALKLVN 397
L ++ E++ E +D++ + + +P W + P+ R A +L
Sbjct: 154 -------LDSNLTAHESLAHSFEEVQDQAMLEMVSQGTVPAWLPLPPQARFRRARRELYR 206
Query: 398 DTLNNLIAICK-RMVDEEELQFHEEYMNEQDPSILHFLLASG---DDVTS--KQLRDDLM 451
+ L+A + RM D D ++ ++A+ DD +LR++L+
Sbjct: 207 -VADLLVADRRSRMADG----------GPGDDALSRIIVAADRRRDDPARARNRLREELV 255
Query: 452 TMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMKKLKYTTRVIN 511
T+L+AGHET+A+ L WT +LL + P V +++ E + LGD P ED+ +L YTT V+
Sbjct: 256 TLLLAGHETTASTLGWTLHLLERHPEVRDRVRAEARAALGDGVPGPEDLHRLTYTTMVVQ 315
Query: 512 ESLRLYPQPPVLIRRSIEDDVLGEYPIKRGEDIFISVWNLHRSPTLWNDADKLN 565
E++RL+P +L R + + DV+G Y + G D+ + + +HR P LW D ++ +
Sbjct: 316 EAMRLFPPVWILPRVAQQRDVVGGYTVSAGSDVLVCPYIMHRHPGLWEDPERFD 369
>UniRef100_Q629N7 Cytochrome P450-related protein [Burkholderia mallei]
Length = 1373
Score = 157 bits (397), Expect = 8e-37
Identities = 102/354 (28%), Positives = 172/354 (47%), Gaps = 19/354 (5%)
Query: 211 LYELYITYGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILDFVMGKGLIP 270
L L+ YG + R GP ++ P +++L++N + Y +G + G GL+
Sbjct: 30 LLRLHQQYGDVARNRLGPFVTHALAHPDHIQYVLQENHRNYVRGRFYDNFKMFFGDGLLT 89
Query: 271 ADGEIWRVRRRTIVPALHLKFVAAMIGLFGQATDRLCQKLDTAASDGEDVEMESLFSRLT 330
DGE WR RR + P H K V A G A L + +A G+ +++ L+
Sbjct: 90 TDGEFWRRHRRVVQPLFHKKQVDAHTAAVGDAALALAYRW-SALPPGKALDVVEEMMHLS 148
Query: 331 LDVIGKAVFNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIWKDISPRQ--RK 388
L ++G VFN D S + G + IP W +P + R+
Sbjct: 149 LRMLGLMVFNTDVSSHAEAAGPAVRFGIEAMMPQGNLNDFIPRW--------APTRFNRR 200
Query: 389 VTAALKLVNDTLNNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLASGDDVTSKQLRD 448
+ A + ++ + +IA R E +N +DP +G +T +++ D
Sbjct: 201 IAHARRAIDTIVAKIIAD-HREARCEPSDVISLLLNARDPD-------TGAPMTQQEVHD 252
Query: 449 DLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMKKLKYTTR 508
++MT+ +AGHET+ A L W Y L++ P+V+ +L++E+D+ LG R PT++D ++L Y ++
Sbjct: 253 EVMTVFLAGHETTGAGLAWALYALAQHPAVLRQLRDELDARLGGRAPTVQDFEQLPYLSQ 312
Query: 509 VINESLRLYPQPPVLIRRSIEDDVLGEYPIKRGEDIFISVWNLHRSPTLWNDAD 562
V++E LR+YP R +EDD +G Y I +F+S + HR P W + D
Sbjct: 313 VVDEVLRVYPPIWGFTRDLVEDDEIGGYRIPARSSVFMSPYVTHRHPAFWRNPD 366
>UniRef100_Q5V5P9 Cytochrome P450 [Haloarcula marismortui]
Length = 445
Score = 153 bits (387), Expect = 1e-35
Identities = 100/350 (28%), Positives = 179/350 (50%), Gaps = 26/350 (7%)
Query: 218 YGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSK-GILAEILDFVMGKGLIPADGEIW 276
YG + R + GP +++ DP + +L + + K + L ++G GL+ ++GE W
Sbjct: 35 YGDVARFDMGPMDTVMLCDPTAIERVLVSEADRFRKPDFQGDALGDLLGDGLLLSEGETW 94
Query: 277 RVRRRTIVPALHLKFVAAMIG-LFGQATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIG 335
+R+ PA + ++ M + G A DR+ S G+ ++ E +R+TLDVI
Sbjct: 95 EQQRKLANPAFSMARLSGMADRITGHAEDRIADW-----SHGDVIDAEQSMTRVTLDVIL 149
Query: 336 KAVFNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPV-WDLPIWKDISPRQRKVTAALK 394
+ + T IE L + P P+ + +P W + P + A++
Sbjct: 150 DLMMGVELSEQRVQT--IEEQLLPLGQR----FEPDPIRFAMPQWMPM-PDDAEFNRAVR 202
Query: 395 LVNDTLNNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLASGDD--VTSKQLRDDLMT 452
+++ L+++I + + + +E + L LL + DD + +QLRD++MT
Sbjct: 203 TLDEVLDDIIEVREDSLGTDE---------DGPMDFLSVLLRARDDGNQSPEQLRDEMMT 253
Query: 453 MLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMKKLKYTTRVINE 512
ML+AGH+T+A LT+T++LLS+ P V ++ EE+D V+GD P +E +++L Y VI E
Sbjct: 254 MLLAGHDTTALTLTYTWFLLSEHPEVEQRVHEELDDVIGDDRPGMEHVRELDYLEWVIQE 313
Query: 513 SLRLYPQPPVLIRRSIEDDVLGEYPIKRGEDIFISVWNLHRSPTLWNDAD 562
++RLYP + R ED L Y ++ G + + W +HRS ++D +
Sbjct: 314 AMRLYPPVYTIFREPTEDVTLSGYEVEAGTTLMVPQWGVHRSERFYDDPE 363
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.136 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 947,727,355
Number of Sequences: 2790947
Number of extensions: 39968258
Number of successful extensions: 113532
Number of sequences better than 10.0: 4052
Number of HSP's better than 10.0 without gapping: 2647
Number of HSP's successfully gapped in prelim test: 1405
Number of HSP's that attempted gapping in prelim test: 104576
Number of HSP's gapped (non-prelim): 5342
length of query: 569
length of database: 848,049,833
effective HSP length: 133
effective length of query: 436
effective length of database: 476,853,882
effective search space: 207908292552
effective search space used: 207908292552
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 78 (34.7 bits)
Medicago: description of AC148994.8


