

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148819.3 + phase: 0 /pseudo/partial
(880 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
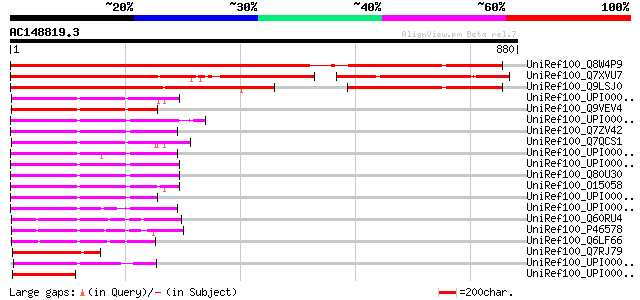
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8W4P9 Hypothetical protein [Arabidopsis thaliana] 937 0.0
UniRef100_Q7XVU7 OSJNBa0035B13.4 protein [Oryza sativa] 571 e-161
UniRef100_Q9LSJ0 Dbj|BAA20807.2 [Arabidopsis thaliana] 533 e-150
UniRef100_UPI0000430F66 UPI0000430F66 UniRef100 entry 196 2e-48
UniRef100_Q9VEV4 CG12753-PA [Drosophila melanogaster] 189 4e-46
UniRef100_UPI00003AAE5C UPI00003AAE5C UniRef100 entry 184 8e-45
UniRef100_Q7ZV42 Hypothetical protein zgc:56230 [Brachydanio rerio] 184 8e-45
UniRef100_Q7QCS1 ENSANGP00000022085 [Anopheles gambiae str. PEST] 184 1e-44
UniRef100_UPI000029BA9C UPI000029BA9C UniRef100 entry 183 2e-44
UniRef100_UPI00001D034F UPI00001D034F UniRef100 entry 182 4e-44
UniRef100_Q80U30 MKIAA0350 protein [Mus musculus] 177 9e-43
UniRef100_O15058 KIAA0350 protein [Homo sapiens] 177 1e-42
UniRef100_UPI0000429CF5 UPI0000429CF5 UniRef100 entry 177 2e-42
UniRef100_UPI00003651BC UPI00003651BC UniRef100 entry 166 2e-39
UniRef100_Q60RU4 Hypothetical protein CBG21183 [Caenorhabditis b... 153 2e-35
UniRef100_P46578 Hypothetical protein gop-1 [Caenorhabditis eleg... 151 9e-35
UniRef100_Q6LF66 Hypothetical protein [Plasmodium falciparum] 124 1e-26
UniRef100_Q7RJ79 GOP-1-related [Plasmodium yoelii yoelii] 105 8e-21
UniRef100_UPI00004991C2 UPI00004991C2 UniRef100 entry 91 1e-16
UniRef100_UPI000046CBB8 UPI000046CBB8 UniRef100 entry 86 4e-15
>UniRef100_Q8W4P9 Hypothetical protein [Arabidopsis thaliana]
Length = 837
Score = 937 bits (2421), Expect = 0.0
Identities = 495/856 (57%), Positives = 604/856 (69%), Gaps = 62/856 (7%)
Query: 1 DFVIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIM 60
D VIEALRSIAE++TYGDQHDP FFEFFMEKQV+G+FVRIL++S+T+++ +QLLQT+SIM
Sbjct: 36 DLVIEALRSIAEILTYGDQHDPLFFEFFMEKQVMGEFVRILRVSKTVTVSVQLLQTMSIM 95
Query: 61 IQNLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKT 120
IQNL+SE AIYY+FSNE++NYLITY+FDF++EELLSYYISFLRA+ GKLNK+TISLL+KT
Sbjct: 96 IQNLKSEQAIYYLFSNEYVNYLITYTFDFQHEELLSYYISFLRAVSGKLNKHTISLLLKT 155
Query: 121 RNDEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDDSVNRYITSEPHTDYFSNL 180
ND VVSFPLYVE I+FAFHEE+MIR AVRA+TLNVYHVGD+SVN Y+ S PHT+YFS L
Sbjct: 156 ENDVVVSFPLYVEGIQFAFHEENMIRTAVRALTLNVYHVGDESVNDYVVSPPHTEYFSKL 215
Query: 181 VSFFRKQCMDLNRLISETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVERLIT 240
+SFF+KQCMDL+ ++ TLK+P PDS + +AVD IED LYYFSD+ISAGIPD+ RLIT
Sbjct: 216 ISFFQKQCMDLSAMVLNTLKSPSPDSGGKLFSAVDGIEDTLYYFSDVISAGIPDIGRLIT 275
Query: 241 DSILMLLIFPVLLPSLRTVADNDMQSGVVTSLYLLCCILRIVKIKDLANTIVAALYYPLE 300
D IL L P+LLPSL + A ND+ VTSLYLL CILRIVKIKDLAN A L+ P++
Sbjct: 276 DHILQHLTLPLLLPSLCSEAVNDISVDPVTSLYLLSCILRIVKIKDLANMTAATLFCPVK 335
Query: 301 SFTKCSGGQVNGYIPDHGFTSKSDGIDNDNLAENSAKCLVVNVPCSSSSSDCHPQSITIL 360
+F S + N + G T + D E + +C S + T
Sbjct: 336 AFISSSLVKPNSSLAPEGLTYVNGHPDKGVTEEANQQCSSTAAGMSDDGNSHLCSEDTPK 395
Query: 361 NNGSSSNVALREVLLEYVTGGADVQVLGSLSVLATLLQTKELDESMLDGLGILPQRKQQK 420
+ ++S++ RE LL+Y++ G DVQ GSL VLATLLQTKEL+ESMLD GILPQRKQ K
Sbjct: 396 SIFNNSHMTFRETLLQYISEGDDVQAQGSLFVLATLLQTKELEESMLDAFGILPQRKQHK 455
Query: 421 KLLLQALVGEASGEEQLFSSESSLTRDGIACGLDVYLNKIKEQYGVSFQPSNVVSSPRVP 480
KLLLQ+LVGE +GEEQLFS + RDG++ LD YL +++EQ+GV PRV
Sbjct: 456 KLLLQSLVGEDTGEEQLFSPRNGSMRDGLSSELDWYLRRLEEQFGVCCSLPGAARCPRVH 515
Query: 481 RFQVLDALVSLFCRSNISAETLWDGGWLLRQLLPYSESEFNNHHLELLKSVVLPDI*LAK 540
R QV+D LV+L CR NISAETLWDGGWLLRQLLPYSE+EFN
Sbjct: 516 RHQVVDTLVTLLCRENISAETLWDGGWLLRQLLPYSEAEFN------------------- 556
Query: 541 TCFFLYEMIEIWCYGVPRSHIG*LNGFVIPP*QPAIKEFVSKF*KISYENSASALEKEVR 600
R H+ LN +SYE ++L +E++
Sbjct: 557 -----------------RKHLKMLN--------------------VSYEKCKNSLTREIK 579
Query: 601 GFWPDLLITVLCDEWRKCKRAMESSSPPKEPKCMLFPPRMFFSEEDIPEGSSFTAGERMH 660
G WPDLLI VL DEWRKCKR +E+ SP KEPK +L S ++ SSFTAGERM
Sbjct: 580 GIWPDLLIRVLLDEWRKCKRVIEAPSPQKEPKSVLLQLDRSSSNDNSVSESSFTAGERMC 639
Query: 661 ELVKVFVLLHQLQIFTHGRALPDQPLIYHPCDHRTNSRAHTSGL-AAVPKPGTEMNLVNA 719
E+VKVFVLLHQLQIF+ GR+LP+QP IY P D SRA +GL +VPKPGTE+ LV+A
Sbjct: 640 EVVKVFVLLHQLQIFSLGRSLPEQPPIYPPADRSETSRATRAGLDVSVPKPGTELKLVDA 699
Query: 720 VPCRIAFERGKERHFCFLAISVGTSGWLVLAEELPLKKPYGVVRVAAPLAGCNPRVDDKH 779
VPCRIAFERGKER F FLA+S G SGW+VLA+ G+VRV APLAGC PR+D+KH
Sbjct: 700 VPCRIAFERGKERDFSFLALSSGESGWIVLAD-----PDNGIVRVTAPLAGCKPRIDEKH 754
Query: 780 SKWLHLRIRPSALPFLDPVKYNRHGKLKTKAFVDGRWILAFRDEESCKNAFLMIREEINH 839
+WLHLRIRPS LP LDP K + KLK+K VDGRWILAFRD+ESC +A+ M+ EI+
Sbjct: 755 PRWLHLRIRPSTLPLLDPTKRGVYEKLKSKGLVDGRWILAFRDDESCHSAYSMVAGEIDL 814
Query: 840 LCDEVDRRIKPLLKLE 855
C EV+RR++PL LE
Sbjct: 815 QCSEVERRLRPLFDLE 830
>UniRef100_Q7XVU7 OSJNBa0035B13.4 protein [Oryza sativa]
Length = 859
Score = 571 bits (1471), Expect = e-161
Identities = 305/543 (56%), Positives = 394/543 (72%), Gaps = 34/543 (6%)
Query: 1 DFVIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIM 60
DFV+EALRSIAEL+ YGDQHDP++FEFFMEKQ++G+F RIL++S+ + LQLLQT+SIM
Sbjct: 35 DFVVEALRSIAELMIYGDQHDPSYFEFFMEKQIMGEFARILRISKLSRVSLQLLQTMSIM 94
Query: 61 IQNLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKT 120
IQNL++EH+IYY+FSNEH+N+LITY FDF+ +E+LSYYISFLRAI GKLNKNTISLLVKT
Sbjct: 95 IQNLRNEHSIYYIFSNEHINFLITYPFDFQIDEMLSYYISFLRAISGKLNKNTISLLVKT 154
Query: 121 RNDEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDDSVNRYITSEPHTDYFSNL 180
+NDEV+SFPLYVEA++FAFHE+SMIR A+R +TLNVYHVGD+SVNR+++ P +DYFS++
Sbjct: 155 KNDEVISFPLYVEALKFAFHEDSMIRVAIRTLTLNVYHVGDESVNRFVSRAPLSDYFSDM 214
Query: 181 VSFFRKQCMDLNRLISETLKN-PGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVERLI 239
V+ F+KQC+DL++L+ +++N ++V A+ +IED LYYFSD++S+GIPD+ R I
Sbjct: 215 VNHFQKQCIDLDKLVVRSVRNADSAVPTASVEDAIVQIEDTLYYFSDVMSSGIPDLGRFI 274
Query: 240 TDSILMLLIFPVLLPSLRTVADNDMQSGVVTSLYLLCCILRIVKIKDLANTIVAALYYPL 299
T++IL LL+F LLPSL+ D+ V TS+YL+CCIL I K KD+A+T+ AAL++
Sbjct: 275 TENILQLLVFRFLLPSLQR-QSTDLGISVTTSMYLICCILHIFKNKDMASTVAAALFHQP 333
Query: 300 ESFTKCSGGQVNGY-------IPDHGFTSKSDGIDNDN------LAENSAKCLVVNVPCS 346
+ + G NGY I D+ TS SD ID N L+ ++ CL P
Sbjct: 334 DCHDR-KQGTPNGYTSEHENGISDNQGTSTSD-IDQSNEDKSDILSSSNTHCL----PDD 387
Query: 347 SSSSDCHPQSITILNNGSSSNVALREVLLEYVTGGADVQVLGSLSVLATLLQTKELDESM 406
SSSDC LRE LL Y+ G D Q LGSL + +TLLQTKELDESM
Sbjct: 388 PSSSDC------------CQGNTLREHLLSYIISGDDFQALGSLCLFSTLLQTKELDESM 435
Query: 407 LDGLGILPQRKQQKKLLLQALVGEASGEEQLFSSESSLTRDGIACGLDVYLNKIKEQYGV 466
LD LGILPQRKQ KKLLLQALVGE E QLFSS S L D I D+Y+ K++++YG+
Sbjct: 436 LDALGILPQRKQHKKLLLQALVGEDLAERQLFSSSSGLADDSICSDFDMYVRKLQDKYGL 495
Query: 467 SFQPSNVVSSPRVPRFQVLDALVSLFCRSNISAETLWDGGWLLRQLLPYSESEFNNHHLE 526
++S + R+QVLDALV+LFCRSN+SA+ GGWL RQLLP+ E EF HL+
Sbjct: 496 KCHHPRQMTS-KFHRYQVLDALVALFCRSNVSADVRLVGGWLFRQLLPHGEEEFTAFHLK 554
Query: 527 LLK 529
LK
Sbjct: 555 WLK 557
Score = 271 bits (693), Expect = 6e-71
Identities = 149/302 (49%), Positives = 197/302 (64%), Gaps = 9/302 (2%)
Query: 568 VIPP*QPAIKEFVSKF*KISYENSASALEKEVRGFWPDLLITVLCDEWRKCKRAMESSSP 627
++P + F K+ K S+++ + L E G W DLL+ ++ + W+ CK+A+E+SSP
Sbjct: 540 LLPHGEEEFTAFHLKWLKDSHKDYSIKLLDESGGCWRDLLVPIVKEAWKNCKKAIETSSP 599
Query: 628 PKEPKCMLFPPRMFFSEEDIPEGSSFTAGERMHELVKVFVLLHQLQIFTHGRALPDQPLI 687
PK K + P SS ER++E+VK FVL Q+ +F G L DQP I
Sbjct: 600 PKGSKSAILP----LDSCSFGGDSSIAIAERIYEMVKGFVLQRQVILFCLGETLTDQPPI 655
Query: 688 YHPCDHRTNSRAHTSGL-AAVPKPGTEMNLVNAVPCRIAFERGKERHFCFLAISVGTSGW 746
+ P D N+RA +G +VPKPG E+NLV+AVPCRIAFERGKERHFCFLA+S GTSGW
Sbjct: 656 FSPTDLPVNNRATLAGFDGSVPKPGLEVNLVDAVPCRIAFERGKERHFCFLALSNGTSGW 715
Query: 747 LVLAEELPLKKPYGVVRVAAPLAGCNPRVDDKHSKWLHLRIRPSALPFLDPVKYNRHGKL 806
++L EELPLK+ G+VRV APLAG +PR+D+KH+KWLHLRIRPS +PFLD KY GK
Sbjct: 716 ILLLEELPLKEKRGIVRVTAPLAGSDPRIDEKHAKWLHLRIRPSTVPFLDTEKYK--GKT 773
Query: 807 KTKAFVDGRWILAFRDEESCKNAFLMIREEINHLCDEVDRRIKPLLKLETVLD-ISSSSA 865
+ K VDGRW LAFRDE+SCK A M+ EE+ V ++K LL+ + D I+ A
Sbjct: 774 R-KYLVDGRWTLAFRDEQSCKEAETMVIEEMKLQQGAVGEQLKLLLEFDMPEDGITGILA 832
Query: 866 PV 867
P+
Sbjct: 833 PL 834
>UniRef100_Q9LSJ0 Dbj|BAA20807.2 [Arabidopsis thaliana]
Length = 816
Score = 533 bits (1374), Expect = e-150
Identities = 284/475 (59%), Positives = 350/475 (72%), Gaps = 18/475 (3%)
Query: 1 DFVIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIM 60
D VIEALRSIAE++TYGDQHDP FFEFFMEKQV+G+FVRIL++S+T+++ +QLLQT+SIM
Sbjct: 11 DLVIEALRSIAEILTYGDQHDPLFFEFFMEKQVMGEFVRILRVSKTVTVSVQLLQTMSIM 70
Query: 61 IQNLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKT 120
IQNL+SE AIYY+FSNE++NYLITY+FDF++EELLSYYISFLRA+ GKLNK+TISLL+KT
Sbjct: 71 IQNLKSEQAIYYLFSNEYVNYLITYTFDFQHEELLSYYISFLRAVSGKLNKHTISLLLKT 130
Query: 121 RNDEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDDSVNRYITSEPHTDYFSNL 180
ND VVSFPLYVE I+FAFHEE+MIR AVRA+TLNVYHVGD+SVN Y+ S PHT+YFS L
Sbjct: 131 ENDVVVSFPLYVEGIQFAFHEENMIRTAVRALTLNVYHVGDESVNDYVVSPPHTEYFSKL 190
Query: 181 VSFFRKQCMDLNRLISETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVERLIT 240
+SFF+KQCMDL+ ++ TLK+P PDS + +AVD IED LYYFSD+ISAGIPD+ RLIT
Sbjct: 191 ISFFQKQCMDLSAMVLNTLKSPSPDSGGKLFSAVDGIEDTLYYFSDVISAGIPDIGRLIT 250
Query: 241 DSILMLLIFPVLLPSLRTVADNDMQSGVVTSLYLLCCILRIVKIKDLANTIVAALYYPLE 300
D IL L P+LLPSL + AD + VTSLYLL CILRIVKIKDLAN A L+ P++
Sbjct: 251 DHILQHLTLPLLLPSLCSEADISVDP--VTSLYLLSCILRIVKIKDLANMTAATLFCPVK 308
Query: 301 SFTKCSGGQVNGYIPDHGFTSKSDGIDNDNLAENSAKCLVVNVPCSSSSSDCHPQSITIL 360
+F S + N + G T + D E + +C S + T
Sbjct: 309 AFISSSLVKPNSSLAPEGLTYVNGHPDKGVTEEANQQCSSTAAGMSDDGNSHLCSEDTPK 368
Query: 361 NNGSSSNVALREVLLEYVTGGADVQVLGSLSVLATLLQTK----------------ELDE 404
+ ++S++ RE LL+Y++ G DVQ GSL VLATLLQTK EL+E
Sbjct: 369 SIFNNSHMTFRETLLQYISEGDDVQAQGSLFVLATLLQTKGIGSGIVIVQILLAFSELEE 428
Query: 405 SMLDGLGILPQRKQQKKLLLQALVGEASGEEQLFSSESSLTRDGIACGLDVYLNK 459
SMLD GILPQRKQ KKLLLQ+LVGE +GEEQLFS + RDG++ LD YL +
Sbjct: 429 SMLDAFGILPQRKQHKKLLLQSLVGEDTGEEQLFSPRNGSMRDGLSSELDWYLRR 483
Score = 330 bits (847), Expect = 9e-89
Identities = 165/271 (60%), Positives = 201/271 (73%), Gaps = 6/271 (2%)
Query: 586 ISYENSASALEKEVRGFWPDLLITVLCDEWRKCKRAMESSSPPKEPKCMLFPPRMFFSEE 645
+SYE ++L +E++G WPDLLI VL DEWRKCKR +E+ SP KEPK +L S +
Sbjct: 544 VSYEKCKNSLTREIKGIWPDLLIRVLLDEWRKCKRVIEAPSPQKEPKSVLLQLDRSSSND 603
Query: 646 DIPEGSSFTAGERMHELVKVFVLLHQLQIFTHGRALPDQPLIYHPCDHRTNSRAHTSGL- 704
+ SSFTAGERM E+VKVFVLLHQLQIF+ GR+LP+QP IY P D SRA +GL
Sbjct: 604 NSVSESSFTAGERMCEVVKVFVLLHQLQIFSLGRSLPEQPPIYPPADRSETSRATRAGLD 663
Query: 705 AAVPKPGTEMNLVNAVPCRIAFERGKERHFCFLAISVGTSGWLVLAEELPLKKPYGVVRV 764
+VPKPGTE+ LV+AVPCRIAFERGKER F FLA+S G SGW+VLA+ G+VRV
Sbjct: 664 VSVPKPGTELKLVDAVPCRIAFERGKERDFSFLALSSGESGWIVLAD-----PDNGIVRV 718
Query: 765 AAPLAGCNPRVDDKHSKWLHLRIRPSALPFLDPVKYNRHGKLKTKAFVDGRWILAFRDEE 824
APLAGC PR+D+KH +WLHLRIRPS LP LDP K + KLK+K VDGRWILAFRD+E
Sbjct: 719 TAPLAGCKPRIDEKHPRWLHLRIRPSTLPLLDPTKRGVYEKLKSKGLVDGRWILAFRDDE 778
Query: 825 SCKNAFLMIREEINHLCDEVDRRIKPLLKLE 855
SC +A+ M+ EI+ C EV+RR++PL LE
Sbjct: 779 SCHSAYSMVAGEIDLQCSEVERRLRPLFDLE 809
>UniRef100_UPI0000430F66 UPI0000430F66 UniRef100 entry
Length = 882
Score = 196 bits (499), Expect = 2e-48
Identities = 110/309 (35%), Positives = 182/309 (58%), Gaps = 22/309 (7%)
Query: 3 VIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIMIQ 62
++E LRSIAE++ +GDQ+D + F+FF+EK ++ F+RI+K + +QLLQT++I+ +
Sbjct: 21 LVETLRSIAEILIWGDQNDSSVFDFFLEKNMLSFFLRIMKQKCGSYVCVQLLQTLNILFE 80
Query: 63 NLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKTRN 122
N+++E ++YY+ SN H+N +I + FDF +EE+++YYISFL+ + KLN +TI N
Sbjct: 81 NIRNETSLYYLLSNNHVNSIIVHKFDFSDEEVMAYYISFLKTLSLKLNAHTIHFFY---N 137
Query: 123 DEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDDSVNRYITSEPHTDYFSNLVS 182
+ FPLY EAI+F H E M+R AVR +TLNVY V D S+ +I YFSNLV
Sbjct: 138 EHTNDFPLYTEAIKFFNHSEGMVRIAVRTLTLNVYRVEDASMLAFIRDRTAAPYFSNLVW 197
Query: 183 FFRKQCMDLNRLISETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVERLITDS 242
F ++L+ + + S + ++ V E D+L+Y +DI+ I D+ +++++
Sbjct: 198 FIGNHIIELDTCVRNDADH---QSQNRLSDLVAEHLDHLHYLNDILCLNISDLNKVLSEH 254
Query: 243 ILMLLIFPVLLPSL----RTVADNDMQS------------GVVTSLYLLCCILRIVKIKD 286
+L L+ P+ + SL + N + S +V +L+LL + IV
Sbjct: 255 LLHKLLVPLYVYSLMKHKNLLCQNQVCSQLDLKLFERKHVSIVVALFLLSQVFLIVSHGP 314
Query: 287 LANTIVAAL 295
L +T+ A +
Sbjct: 315 LVHTLAAVM 323
>UniRef100_Q9VEV4 CG12753-PA [Drosophila melanogaster]
Length = 1067
Score = 189 bits (479), Expect = 4e-46
Identities = 99/256 (38%), Positives = 162/256 (62%), Gaps = 8/256 (3%)
Query: 3 VIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRIL--KLSRTISIPLQLLQTVSIM 60
++E+LR IAE++ +GDQHD F+FF+EK ++ F+ I+ K + + +QLLQT++I+
Sbjct: 46 LVESLRCIAEILIWGDQHDSLVFDFFLEKNMLSYFLHIMRQKSGGSSFVCVQLLQTLNIL 105
Query: 61 IQNLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKT 120
+N+++E ++YY+ SN H+N ++ + FDF +E+++ YYI FL+ + KLN +TI
Sbjct: 106 FENIRNETSLYYLLSNNHVNSIMVHKFDFSDEDVMGYYILFLKTLSLKLNTHTIHFFY-- 163
Query: 121 RNDEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDDSVNRYITSEPHTDYFSNL 180
N+ FPLY EAI+F H ESM+R AVR ++LNVY V + S+ R+I + YFSNL
Sbjct: 164 -NEHTNDFPLYTEAIKFFNHPESMVRIAVRTISLNVYKVQNPSMLRFIRDKTAAPYFSNL 222
Query: 181 VSFFRKQCMDLNRLISETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVERLIT 240
V F K ++L+ + + + SN ++ V E D+L+Y SDI+ I D+ ++T
Sbjct: 223 VWFIGKHILELDTCVRTDIDH---QSNQKLSNLVAEHLDHLHYLSDILLLEIKDLNAVLT 279
Query: 241 DSILMLLIFPVLLPSL 256
+ +L L P+ + SL
Sbjct: 280 EHLLHKLFVPLYIFSL 295
>UniRef100_UPI00003AAE5C UPI00003AAE5C UniRef100 entry
Length = 404
Score = 184 bits (468), Expect = 8e-45
Identities = 112/344 (32%), Positives = 189/344 (54%), Gaps = 28/344 (8%)
Query: 1 DFVIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIM 60
+ ++E +RSI E++ +GDQ+D + F+FF+EK + F+ IL+ + +QLLQT++I+
Sbjct: 45 NLLVETIRSITEILIWGDQNDSSVFDFFLEKNMFVFFLNILRQKSGRYVCVQLLQTLNIL 104
Query: 61 IQNLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKT 120
+N+ E ++YY+ SN ++N +I + FDF +EE+++YYISFL+ + KLN +T+
Sbjct: 105 FENISHETSLYYLLSNNYVNSIIVHKFDFSDEEIMAYYISFLKTLSLKLNNHTVHFFY-- 162
Query: 121 RNDEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDDSVNRYITSEPHTDYFSNL 180
N+ F LY EAI+F H ESM+R AVR +TLNVY V + + YI + YFSNL
Sbjct: 163 -NEHTNDFALYTEAIKFFNHPESMVRIAVRTITLNVYKVDNQPMLHYIRDKTAVPYFSNL 221
Query: 181 VSFFRKQCMDLNRLI--SETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVERL 238
V F ++L+ + E +N G ++ V E D+L+Y +DI+ + +
Sbjct: 222 VWFIGSHVIELDNCVQTDEEHRNRG-----KLSDLVAEHLDHLHYLNDILIINCEFLNDV 276
Query: 239 ITDSILMLLIFPVLLPSL--RTVADNDMQSGVVTSLYLLCCILRIVKIKDLANTIVAALY 296
+TD +L L P+ + SL + + + SLYLL + I+ L N++ +
Sbjct: 277 LTDHLLNRLFLPLYVYSLVNQDKGGERPKISLQVSLYLLSQVFLIIHYAPLVNSLAEVI- 335
Query: 297 YPLESFTKCSGGQVNGYIPDHGFTSKSDGIDNDNLAENSAKCLV 340
+NG + F+SK++ N A++S +C +
Sbjct: 336 -------------LNGDL--SVFSSKAEQDVQKNSAKSSIRCFI 364
>UniRef100_Q7ZV42 Hypothetical protein zgc:56230 [Brachydanio rerio]
Length = 414
Score = 184 bits (468), Expect = 8e-45
Identities = 106/296 (35%), Positives = 173/296 (57%), Gaps = 13/296 (4%)
Query: 1 DFVIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIM 60
+ ++E +RSI E++ +GDQ+D + F+FF+EK + F+ I + + +QLLQT++I+
Sbjct: 43 NLLVETIRSITEILIWGDQNDSSVFDFFLEKNMFAFFLNIPRQKSGRYVCVQLLQTLNIL 102
Query: 61 IQNLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKT 120
+N+ E ++YY+ SN H+N +I + FDF +EE+++YYISFL+ + KLN +T+
Sbjct: 103 FENISHETSLYYLLSNSHVNSIIVHKFDFSDEEIMAYYISFLKTLSLKLNNHTVHFFY-- 160
Query: 121 RNDEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDDSVN--RYITSEPHTDYFS 178
N+ F LY EAI+F H ESM+R AVR +TLNVY V D+ + YI + YFS
Sbjct: 161 -NEHTNDFALYTEAIKFFNHPESMVRIAVRTITLNVYKVSLDNQHMLHYIRDKTAVPYFS 219
Query: 179 NLVSFFRKQCMDLNRLI--SETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVE 236
NLV F ++L++ + E KN G ++ V E D+L+Y +DI++ +
Sbjct: 220 NLVWFIGSHVIELDKCVQTDEEHKNRG-----KLSDLVAEHLDHLHYLNDILTINCEFLN 274
Query: 237 RLITDSILMLLIFPVLLPSLRTVAD-NDMQSGVVTSLYLLCCILRIVKIKDLANTI 291
++TD +L L P+ + SL T D + SLYLL + I+ + L +++
Sbjct: 275 DVLTDHLLNRLFLPLYVYSLVTQEQTEDRKINPQVSLYLLSQVFLIIHYQPLVHSL 330
>UniRef100_Q7QCS1 ENSANGP00000022085 [Anopheles gambiae str. PEST]
Length = 1001
Score = 184 bits (466), Expect = 1e-44
Identities = 113/340 (33%), Positives = 188/340 (55%), Gaps = 34/340 (10%)
Query: 3 VIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRIL--KLSRTISIPLQLLQTVSIM 60
++E+LRSIAE++ +GDQ+D + F+FF+EK ++ F+ I+ K + + +QLLQT++I+
Sbjct: 45 LVESLRSIAEILIWGDQNDSSVFDFFLEKNMLSYFLHIMRQKSGGSSFVCVQLLQTLNIL 104
Query: 61 IQNLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKT 120
+N+++E ++YY+ SN +N +I + FDF +E+++ YYI FL+ + KLN +TI
Sbjct: 105 FENIRNETSLYYLLSNNLVNSIIVHKFDFSDEDVMGYYILFLKTLSLKLNTHTIHFFY-- 162
Query: 121 RNDEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDDSVNRYITSEPHTDYFSNL 180
N+ FPLY EAI+F H ESM+R AVR +TLNVY V + ++ ++I + YFSNL
Sbjct: 163 -NEHTNDFPLYTEAIKFFNHPESMVRIAVRTITLNVYRVQNPNMLQFIRDKTAAPYFSNL 221
Query: 181 VSFFRKQCMDLNRLISETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVERLIT 240
V F K ++L+ + + + S + V E D+L+Y +DI+ I D+ ++T
Sbjct: 222 VWFIGKHILELDNCVRNDIDH---QSQHRLANLVAEHLDHLHYLNDILCLDIADLNAVLT 278
Query: 241 DSILMLLIFPV----LLPS---------------LRTVADNDMQS------GVVTSLYLL 275
+ +L L P+ L PS L V D D++ V +L+LL
Sbjct: 279 EHLLHKLFVPLYIFSLTPSPPMSMAEVTCNLAAVLNKVVDIDIKEMNNPRVASVVALFLL 338
Query: 276 CCILRIVKIKDLANTIVAALYYPLESFTKCSGGQV-NGYI 314
+ ++ + L N + + + K QV N YI
Sbjct: 339 SLVFLVITHRPLINALTWVILNADHTVFKEGAAQVLNAYI 378
>UniRef100_UPI000029BA9C UPI000029BA9C UniRef100 entry
Length = 1009
Score = 183 bits (465), Expect = 2e-44
Identities = 107/301 (35%), Positives = 170/301 (55%), Gaps = 18/301 (5%)
Query: 1 DFVIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIM 60
+ ++E +RSI E++ +GDQ+D + F+FF+EK + F+ IL+ + +QLLQT++I+
Sbjct: 43 NLLVETIRSITEILIWGDQNDSSVFDFFLEKNMFAFFLNILRQKSGRYVCVQLLQTLNIL 102
Query: 61 IQNLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKT 120
+N+ E ++YY+ SN H+N +I + FDF +EE+++YYISFL+ + KLN +T+
Sbjct: 103 FENISHETSLYYLLSNNHVNSIIVHKFDFSDEEIMAYYISFLKTLSLKLNNHTVHFFY-- 160
Query: 121 RNDEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYH-----VGDDSVNRYITSEPHTD 175
N+ F LY EAI+F H ESM+R AVR +TLNVY V + + YI +
Sbjct: 161 -NEHTNDFALYTEAIKFFNHPESMVRIAVRTITLNVYKALLLTVDNQHMLHYIRDKTAVP 219
Query: 176 YFSNLVSFFRKQCMDLNRLI--SETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIP 233
YFSNLV F ++L++ + E KN G ++ V E D+L+Y +DI+
Sbjct: 220 YFSNLVWFIGSHVIELDKCVQTDEEHKNRG-----KLSDLVAEHLDHLHYLNDILIINCE 274
Query: 234 DVERLITDSILMLLIFPVLLPSL---RTVADNDMQSGVVTSLYLLCCILRIVKIKDLANT 290
+ ++TD +L L P+ + SL TV SLYLL + I+ + L N
Sbjct: 275 FLNDVLTDHLLNRLFLPLYVYSLVSPETVRSPHWNINPQVSLYLLSQVFLIIHYQPLVNA 334
Query: 291 I 291
+
Sbjct: 335 L 335
>UniRef100_UPI00001D034F UPI00001D034F UniRef100 entry
Length = 793
Score = 182 bits (462), Expect = 4e-44
Identities = 103/299 (34%), Positives = 171/299 (56%), Gaps = 12/299 (4%)
Query: 1 DFVIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIM 60
+ ++E +RSI E++ +GDQ+D + F+FF+EK + F+ IL+ + +QLLQT++I+
Sbjct: 68 NLLVETIRSITEILIWGDQNDSSVFDFFLEKNMFVFFLNILRQKSGRYVCVQLLQTLNIL 127
Query: 61 IQNLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKT 120
+N+ E ++YY+ SN ++N +I + FDF +EE+++YYISFL+ + KLN +T+
Sbjct: 128 FENISHETSLYYLLSNNYVNSIIVHKFDFSDEEIMAYYISFLKTLSLKLNNHTVHFFY-- 185
Query: 121 RNDEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDDSVNRYITSEPHTDYFSNL 180
N+ F LY EAI+F H ESM+R AVR +TLNVY V + ++ YI + YFSNL
Sbjct: 186 -NEHTNDFALYTEAIKFFNHPESMVRIAVRTITLNVYKVDNQAMLHYIRDKTAVPYFSNL 244
Query: 181 VSFFRKQCMDLNRLI--SETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVERL 238
V F ++L+ + E +N G ++ V E D+L+Y +DI+ + +
Sbjct: 245 VWFIGSHVIELDNCVQTDEEHRNRG-----KLSDLVAEHLDHLHYLNDILIINCEFLNDV 299
Query: 239 ITDSILMLLIFPVLLPSLRT--VADNDMQSGVVTSLYLLCCILRIVKIKDLANTIVAAL 295
+TD +L L P+ + SL + + SLYLL + I+ L N++ +
Sbjct: 300 LTDHLLNRLFLPLYVHSLENPDKGGERPKISLPVSLYLLSQVFLIIHHAPLVNSLAEVI 358
>UniRef100_Q80U30 MKIAA0350 protein [Mus musculus]
Length = 1045
Score = 177 bits (450), Expect = 9e-43
Identities = 104/301 (34%), Positives = 171/301 (56%), Gaps = 14/301 (4%)
Query: 1 DFVIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIM 60
+ ++E +RSI E++ +GDQ+D + F+FF+EK + F+ IL+ + +QLLQT++I+
Sbjct: 54 NLLVETIRSITEILIWGDQNDSSVFDFFLEKNMFVFFLNILRQKSGRYVCVQLLQTLNIL 113
Query: 61 IQNLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKT 120
+N+ E ++YY+ SN ++N +I + FDF +EE+++YYISFL+ + KLN +T+
Sbjct: 114 FENISHETSLYYLLSNNYVNSIIVHKFDFSDEEIMAYYISFLKTLSLKLNNHTVHFFY-- 171
Query: 121 RNDEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDD--SVNRYITSEPHTDYFS 178
N+ F LY EAI+F H ESM+R AVR +TLNVY V D ++ YI + YFS
Sbjct: 172 -NEHTNDFALYTEAIKFFNHPESMVRIAVRTITLNVYKVSLDNQAMLHYIRDKTAVPYFS 230
Query: 179 NLVSFFRKQCMDLNRLI--SETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVE 236
NLV F ++L+ + E +N G ++ V E D+L+Y +DI+ +
Sbjct: 231 NLVWFIGSHVIELDNCVQTDEEHRNRG-----KLSDLVAEHLDHLHYLNDILIINCEFLN 285
Query: 237 RLITDSILMLLIFPVLLPSLRT--VADNDMQSGVVTSLYLLCCILRIVKIKDLANTIVAA 294
++TD +L L P+ + SL + + SLYLL + I+ L N++
Sbjct: 286 DVLTDHLLNRLFLPLYVYSLENPDKGGERPKISLPVSLYLLSQVFLIIHHAPLVNSLAEV 345
Query: 295 L 295
+
Sbjct: 346 I 346
>UniRef100_O15058 KIAA0350 protein [Homo sapiens]
Length = 1062
Score = 177 bits (449), Expect = 1e-42
Identities = 106/305 (34%), Positives = 174/305 (56%), Gaps = 22/305 (7%)
Query: 1 DFVIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIM 60
+ ++E +RSI E++ +GDQ+D + F+FF+EK + F+ IL+ + +QLLQT++I+
Sbjct: 54 NLLVETIRSITEILIWGDQNDSSVFDFFLEKNMFVFFLNILRQKSGRYVCVQLLQTLNIL 113
Query: 61 IQNLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKT 120
+N+ E ++YY+ SN ++N +I + FDF +EE+++YYISFL+ + KLN +T+
Sbjct: 114 FENISHETSLYYLLSNNYVNSIIVHKFDFSDEEIMAYYISFLKTLSLKLNNHTVHFFY-- 171
Query: 121 RNDEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDD--SVNRYITSEPHTDYFS 178
N+ F LY EAI+F H ESM+R AVR +TLNVY V D ++ YI + YFS
Sbjct: 172 -NEHTNDFALYTEAIKFFNHPESMVRIAVRTITLNVYKVSLDNQAMLHYIRDKTAVPYFS 230
Query: 179 NLVSFFRKQCMDLNRLI--SETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVE 236
NLV F ++L+ + E +N G ++ V E D+L+Y +DI+ +
Sbjct: 231 NLVWFIGSHVIELDDCVQTDEEHRNRG-----KLSDLVAEHLDHLHYLNDILIINCEFLN 285
Query: 237 RLITDSILMLLIFPVLLPSLRTVADNDMQSG------VVTSLYLLCCILRIVKIKDLANT 290
++TD +L L P+ + SL +N + G + SLYLL + I+ L N+
Sbjct: 286 DVLTDHLLNRLFLPLYVYSL----ENQDKGGERPKISLPVSLYLLSQVFLIIHHAPLVNS 341
Query: 291 IVAAL 295
+ +
Sbjct: 342 LAEVI 346
>UniRef100_UPI0000429CF5 UPI0000429CF5 UniRef100 entry
Length = 1522
Score = 177 bits (448), Expect = 2e-42
Identities = 95/258 (36%), Positives = 156/258 (59%), Gaps = 10/258 (3%)
Query: 1 DFVIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIM 60
+ ++E +RSI E++ +GDQ+D + F+FF+EK + F+ IL+ + +QLLQT++I+
Sbjct: 92 NLLVETIRSITEILIWGDQNDSSVFDFFLEKNMFVFFLNILRQKSGRYVCVQLLQTLNIL 151
Query: 61 IQNLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKT 120
+N+ E ++YY+ SN ++N +I + FDF +EE+++YYISFL+ + KLN +T+
Sbjct: 152 FENISHETSLYYLLSNNYVNSIIVHKFDFSDEEIMAYYISFLKTLSLKLNNHTVHFFY-- 209
Query: 121 RNDEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDDSVNRYITSEPHTDYFSNL 180
N+ F LY EAI+F H ESM+R AVR +TLNVY V + ++ YI + YFSNL
Sbjct: 210 -NEHTNDFALYTEAIKFFNHPESMVRIAVRTITLNVYKVDNQAMLHYIRDKTAVPYFSNL 268
Query: 181 VSFFRKQCMDLNRLI--SETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVERL 238
V F ++L+ + E +N G ++ V E D+L+Y +DI+ + +
Sbjct: 269 VWFIGSHVIELDNCVQTDEEHRNRG-----KLSDLVAEHLDHLHYLNDILIINCEFLNDV 323
Query: 239 ITDSILMLLIFPVLLPSL 256
+TD +L L P+ + SL
Sbjct: 324 LTDHLLNRLFLPLYVYSL 341
>UniRef100_UPI00003651BC UPI00003651BC UniRef100 entry
Length = 384
Score = 166 bits (421), Expect = 2e-39
Identities = 100/292 (34%), Positives = 164/292 (55%), Gaps = 24/292 (8%)
Query: 1 DFVIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIM 60
+ ++E +RSI E++ +GDQ+D + F+FF+EK + F+ IL+ + +QLLQT++I+
Sbjct: 43 NLLVETIRSITEILIWGDQNDSSVFDFFLEKNMFAFFLNILRQKSGRYVCVQLLQTLNIL 102
Query: 61 IQNLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKT 120
+N+ E ++YY+ SN H+N +I + FDF +EE+++YYISFL+ + KLN +T+
Sbjct: 103 FENISHETSLYYLLSNNHVNSIIVHKFDFSDEEIMAYYISFLKTLSLKLNNHTVHFFY-- 160
Query: 121 RNDEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDDSVNRYITSEPHTDYFSNL 180
N+ F LY EAI+F H ESM+R AVR +TLNVY V V+ + + YFS L
Sbjct: 161 -NEHTNDFALYTEAIKFFNHPESMVRIAVRTITLNVYKV---IVDHFHYLNETSIYFSAL 216
Query: 181 VSFFRKQCMDLNRLISETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVERLIT 240
F + KN G ++ V E D+L+Y +DI+ + ++T
Sbjct: 217 SVFVSRH------------KNRG-----KLSDLVAEHLDHLHYLNDILIINCEFLNDVLT 259
Query: 241 DSILMLLIFPVLLPSLRTVADNDMQS-GVVTSLYLLCCILRIVKIKDLANTI 291
D +L L P+ + SL + ++ ++ SLYLL + I+ + L N +
Sbjct: 260 DHLLNRLFLPLYVYSLVSPETSEERNINPQVSLYLLSQVFLIIHYQPLVNAL 311
>UniRef100_Q60RU4 Hypothetical protein CBG21183 [Caenorhabditis briggsae]
Length = 906
Score = 153 bits (387), Expect = 2e-35
Identities = 90/299 (30%), Positives = 171/299 (57%), Gaps = 16/299 (5%)
Query: 3 VIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIMIQ 62
++EALR+IAE++ +GDQ+D + F+FF+E+Q++ F++I++ T + +QLLQT++I+ +
Sbjct: 44 LVEALRAIAEILIWGDQNDASVFDFFLERQMLTHFLKIMEQGIT-QLNVQLLQTLNILFE 102
Query: 63 NLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKTRN 122
N++ E ++Y++ SN H+N +I++ FD +N+E+++YYISFL+ + KLN TI N
Sbjct: 103 NIRHETSLYFLLSNNHVNSIISHKFDLQNDEIMAYYISFLKTLSFKLNPATIHFFF---N 159
Query: 123 DEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDDSVNRYITSEPHTDYFSNLVS 182
+ FPL VE ++ ESM+R AVR + LN+ V DDS+ ++ +Y S L+
Sbjct: 160 ETTEEFPLLVEVLKLYNWNESMVRIAVRNILLNIVRVDDDSMTIFVIRH-IKEYLSELID 218
Query: 183 FFRKQCMDLN---RLISETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVERLI 239
++++ R L N + VD++ D ++Y +++ + V +
Sbjct: 219 SLVALSVEMDTFVRSAENVLAN-----RERLRGKVDDLIDLIHYIGELLE--VEAVAETL 271
Query: 240 TDSILMLLIFPVLLPSLRTVADN-DMQSGVVTSLYLLCCILRIVKIKDLANTIVAALYY 297
+ + + P+LL SL DN + V++L+ L +V+ + +T +++ +
Sbjct: 272 SVFVTTRYLSPLLLSSLSPRRDNHSLLLTPVSALFFFSQFLLVVRHHETIHTFLSSFLF 330
>UniRef100_P46578 Hypothetical protein gop-1 [Caenorhabditis elegans]
Length = 892
Score = 151 bits (381), Expect = 9e-35
Identities = 96/308 (31%), Positives = 179/308 (57%), Gaps = 26/308 (8%)
Query: 3 VIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIMIQ 62
++EALR+IAE++ +GDQ+D + F+FF+E+Q++ F++I++ T + +QLLQT++I+ +
Sbjct: 44 LVEALRAIAEILIWGDQNDASVFDFFLERQMLLYFLKIMEQGNT-PLNVQLLQTLNILFE 102
Query: 63 NLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKTRN 122
N++ E ++Y++ SN H+N +I++ FD +N+E+++YYISFL+ + KLN TI N
Sbjct: 103 NIRHETSLYFLLSNNHVNSIISHKFDLQNDEIMAYYISFLKTLSFKLNPATIHFFF---N 159
Query: 123 DEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDDSVNRYITSEPHT-DYFSNLV 181
+ FPL VE ++ ESM+R AVR + LN+ V DDS+ I + HT +Y S L+
Sbjct: 160 ETTEEFPLLVEVLKLYNWNESMVRIAVRNILLNIVRVQDDSM--IIFAIKHTKEYLSELI 217
Query: 182 SFFRKQCMDLN---RLISETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVERL 238
++++ R L N + VD++ D ++Y +++ DVE
Sbjct: 218 DSLVGLSLEMDTFVRSAENVLAN-----RERLRGKVDDLIDLIHYIGELL-----DVE-A 266
Query: 239 ITDSILMLL----IFPVLLPSLRTVADN-DMQSGVVTSLYLLCCILRIVKIKDLANTIVA 293
+ +S+ +L+ + P+LL S+ DN + +++L+ L IV+ + T ++
Sbjct: 267 VAESLSILVTTRYLSPLLLSSISPRRDNHSLLLTPISALFFFSEFLLIVRHHETIYTFLS 326
Query: 294 ALYYPLES 301
+ + ++
Sbjct: 327 SFLFDTQN 334
>UniRef100_Q6LF66 Hypothetical protein [Plasmodium falciparum]
Length = 1869
Score = 124 bits (311), Expect = 1e-26
Identities = 65/251 (25%), Positives = 144/251 (56%), Gaps = 11/251 (4%)
Query: 4 IEALRSIAELVTYGDQHDPTFFEFFM--EKQVVGDFVRILKLSRTISIPLQLLQTVSIMI 61
IE +R I ++ +GD++D F++ + +K+ + + K+++ I I Q+ Q+++++I
Sbjct: 63 IELIRKITQITIWGDKYDDQIFQYVIRIKKKFISLKCSVQKINKNIRI--QIYQSLTMLI 120
Query: 62 QNLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKTR 121
QN+Q + ++YY+FSN +N LI +F ++++E++ YYIS +++I LN +T
Sbjct: 121 QNIQKDISLYYIFSNNKINNLIYTTFIYQDDEIMPYYISMIKSISFFLNYHTSKFFF--- 177
Query: 122 NDEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYHVGDDSVNRYITSEPHTDYFSNLV 181
N++ + FPLY E++R + + R V+ + LN+ + D+ + I + +F+ LV
Sbjct: 178 NEKSMKFPLYTESLRLYKFNDIITRTYVKNIILNILKIDDEGIENLILL--RSSFFTILV 235
Query: 182 SFFRKQCMDLNRLISETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVERLITD 241
+ R + +N++ ++T ++ N ++EI+D LY+F D+++ + ++
Sbjct: 236 CYLRNNLIKMNKIYTKTKES--KKLNKHFNNYLEEIDDVLYFFEDMLNCEKKRINDMLIV 293
Query: 242 SILMLLIFPVL 252
+++ FP++
Sbjct: 294 KLIIYFYFPII 304
>UniRef100_Q7RJ79 GOP-1-related [Plasmodium yoelii yoelii]
Length = 179
Score = 105 bits (261), Expect = 8e-21
Identities = 50/153 (32%), Positives = 94/153 (60%), Gaps = 3/153 (1%)
Query: 5 EALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIMIQNL 64
E LR I +++ +GD+HD F++F E + F+ +L+ +I +Q+ Q+++++IQNL
Sbjct: 21 ELLRKITQIIIWGDKHDDQIFQYFCEDNIFTHFIYLLRQDINKTIRIQVYQSLTLLIQNL 80
Query: 65 QSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKTRNDE 124
Q + ++YY+FSN +N LI +F ++E+++ YYIS +++I LN +T +N +
Sbjct: 81 QKDISLYYIFSNNKINNLIYTTFINQDEDIIPYYISMIKSISFFLNYDTSKFFFNEKNKK 140
Query: 125 VVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVY 157
FPLY E++R + + R V+ + LN++
Sbjct: 141 ---FPLYTESLRLYKFNDIITRTYVKNIILNIF 170
>UniRef100_UPI00004991C2 UPI00004991C2 UniRef100 entry
Length = 600
Score = 91.3 bits (225), Expect = 1e-16
Identities = 59/250 (23%), Positives = 123/250 (48%), Gaps = 25/250 (10%)
Query: 7 LRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIMIQNLQS 66
+R + E++ +GD+H+ + F+E + I+K + ++ LQ LQ +I+++N++S
Sbjct: 1 MRDVVEMLVWGDKHNTGIIDSFLEDNSFSTIIEIVKKGKKPAL-LQFLQCFAILVENIKS 59
Query: 67 EHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKTRNDEVV 126
+ +Y++ S + ++++ D +E+L+S YIS L++I +L +T+ N +
Sbjct: 60 TNFLYFLLSQNTIETILSFPIDRNDEDLISPYISMLKSISLRLTDDTLQFFY---NHQKN 116
Query: 127 SFPLYVEAIRFAFHEESMIRAAVRAVTLN-VYHVGDDSVNRYITSEPHTDYFSNLVSFFR 185
+ P+ EA+ H++SMI + R + L D + N Y S ++ +L S
Sbjct: 117 TCPILSEAVSLLNHKDSMIVSHSRNIILQFAAFTNDKAFNDYYMSYSQKHFYPHLASLIL 176
Query: 186 KQCMDLNRLISETLKNPGPDSNSTVTTAVDEIEDNLYYFSDIISAGIPDVERLITDSILM 245
L + ++ + D++Y+ SD+++ P ++T S+
Sbjct: 177 PHIHSLPQ-------------------SLQLLIDDIYFISDLLTLPHP-YPTMLTSSLFD 216
Query: 246 LLIFPVLLPS 255
LI+P +LPS
Sbjct: 217 NLIYPHILPS 226
>UniRef100_UPI000046CBB8 UPI000046CBB8 UniRef100 entry
Length = 130
Score = 86.3 bits (212), Expect = 4e-15
Identities = 39/109 (35%), Positives = 72/109 (65%)
Query: 5 EALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQLLQTVSIMIQNL 64
E LR I ++ +GD+HD F++F E + F+ +L+ +I +Q+ Q+++++IQNL
Sbjct: 21 ELLRKITQITIWGDKHDDQIFQYFCEDNIFTHFIYLLRQDINKTIRIQVYQSLTLLIQNL 80
Query: 65 QSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNT 113
Q + ++YY+FSN +N LI +F ++E+++ YYIS +++I LN +T
Sbjct: 81 QKDISLYYIFSNNKINNLIYTTFINQDEDIIPYYISMIKSISFFLNYDT 129
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.323 0.139 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,426,911,511
Number of Sequences: 2790947
Number of extensions: 59255783
Number of successful extensions: 132439
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 25
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 132343
Number of HSP's gapped (non-prelim): 46
length of query: 880
length of database: 848,049,833
effective HSP length: 136
effective length of query: 744
effective length of database: 468,481,041
effective search space: 348549894504
effective search space used: 348549894504
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 79 (35.0 bits)
Medicago: description of AC148819.3


