

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148816.5 + phase: 0 /pseudo
(211 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
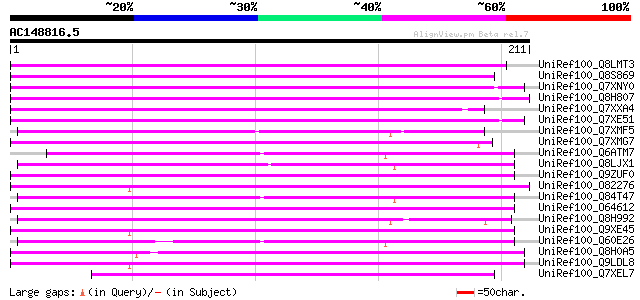
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8LMT3 Putative retroelement [Oryza sativa] 148 8e-35
UniRef100_Q8S869 Putative retroelement [Oryza sativa] 144 1e-33
UniRef100_Q7XNY0 OSJNBb0015N08.11 protein [Oryza sativa] 142 6e-33
UniRef100_Q8H807 Putative non-LTR retroelement reverse transcrip... 133 3e-30
UniRef100_Q7XXA4 OSJNBa0019G23.12 protein [Oryza sativa] 132 5e-30
UniRef100_Q7XE51 Putative non-LTR retroelement reverse transcrip... 128 9e-29
UniRef100_Q7XMF5 OSJNBa0061G20.4 protein [Oryza sativa] 127 2e-28
UniRef100_Q7XMG7 OSJNBa0028I23.15 protein [Oryza sativa] 126 3e-28
UniRef100_Q6ATM7 Hypothetical protein OSJNBb0021K20.15 [Oryza sa... 124 2e-27
UniRef100_Q8LJX1 Putative reverse transcriptase [Sorghum bicolor] 123 3e-27
UniRef100_Q9ZUF0 Putative non-LTR retroelement reverse transcrip... 122 6e-27
UniRef100_O82276 Putative non-LTR retroelement reverse transcrip... 122 6e-27
UniRef100_Q84T47 Putative reverse transcriptase [Oryza sativa] 120 2e-26
UniRef100_O64612 Putative reverse transcriptase [Arabidopsis tha... 120 2e-26
UniRef100_Q8H992 Oryza sativa (japonica cultivar-group) retrotra... 120 3e-26
UniRef100_Q9XE45 Putative non-LTR retrolelement reverse transcri... 119 4e-26
UniRef100_Q60E26 Hypothetical protein OSJNBb0012L23.11 [Oryza sa... 117 2e-25
UniRef100_Q8H0A5 Putative reverse transcriptase [Oryza sativa] 117 3e-25
UniRef100_Q9LDL8 Hypothetical protein AT4g08830 [Arabidopsis tha... 117 3e-25
UniRef100_Q7XEL7 Putative non-LTR retroelement reverse transcrip... 117 3e-25
>UniRef100_Q8LMT3 Putative retroelement [Oryza sativa]
Length = 764
Score = 148 bits (374), Expect = 8e-35
Identities = 74/202 (36%), Positives = 119/202 (58%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
++HLQYADDT+ + +N+ T+K +L ++ +SGLKIN+ KS ++ I E
Sbjct: 214 LTHLQYADDTILFMTNTEENIVTVKFLLYCYEAMSGLKINYQKSEIMVIGGDEMETQRVA 273
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
D NC AG + F YLG+P+ N + + + I RL TW Y+S+GG+ +L+NS
Sbjct: 274 DLFNCQAGKMPFTYLGIPISMNKLTNADLDIPPNKIEKRLATWKCGYLSYGGKAILINSC 333
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
L+S+P++ + + GV ++ I+ F W G+ +K + W+++C+ KE GG+
Sbjct: 334 LSSIPLYMMGVYLLPEGVHNKMDSIRARFFWEGLEKKRKYHMIKWEALCRPKEFGGLGFI 393
Query: 181 DIRVMNISLLAKWRWRLIDGRE 202
D R MNI+LL KW +RL G+E
Sbjct: 394 DTRKMNIALLCKWIYRLESGKE 415
>UniRef100_Q8S869 Putative retroelement [Oryza sativa]
Length = 779
Score = 144 bits (364), Expect = 1e-33
Identities = 71/197 (36%), Positives = 118/197 (59%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
+SHLQYA+DT+ KA+ +N+ +K +L F+ +SG+KIN+ KS + + V ++ +
Sbjct: 73 LSHLQYANDTILFAKATKENVLALKFLLFCFEEMSGMKINYQKSEVYVLGVPKEEEELYV 132
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
D LNC GS+ F YLGLP+G P+++ I RL +W + ++S+GGR + +N+
Sbjct: 133 DMLNCKVGSLPFTYLGLPMGVGKVGKRDLLPIMQKIEKRLQSWHSGHLSYGGREIRINTF 192
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
L+S+P++ + F K+ K + I+ F W G G KK + + +C+ K+ GG+
Sbjct: 193 LSSVPMYAMGFFKLPEEFHKNLETIRGRFYWQGNGKKKKYHLIKLQGLCRPKDFGGLGFL 252
Query: 181 DIRVMNISLLAKWRWRL 197
D R+MN LL+KW +L
Sbjct: 253 DTRIMNFCLLSKWIMKL 269
>UniRef100_Q7XNY0 OSJNBb0015N08.11 protein [Oryza sativa]
Length = 1026
Score = 142 bits (358), Expect = 6e-33
Identities = 73/209 (34%), Positives = 118/209 (55%), Gaps = 1/209 (0%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
++ LQYADDT+ + + + +K IL F+ +SGLKINF+KS + +++ +
Sbjct: 483 IAILQYADDTIFLINDKLDHAKNLKYILCLFEQLSGLKINFNKSEVFCFGEAKEKQDLYS 542
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
+ C GS+ KYLG+P+ W+ + +L W R S GGR++LLNS
Sbjct: 543 NIFTCKVGSLPLKYLGIPIDQKRILNKDWKLAENKMEHKLGCWQGRLQSIGGRLILLNST 602
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
L+S+P++ +SF ++ GV +RI ++ FLW G +K VNW VC ++ GG+ V
Sbjct: 603 LSSVPMYMISFYRLPKGVQERIDYFRKRFLWQEDQGIRKYHLVNWPLVCSPRDQGGLGVL 662
Query: 181 DIRVMNISLLAKWRWRLIDGREALWK*VL 209
D+ MN ++L KW WRL + E W+ ++
Sbjct: 663 DLEAMNKAMLGKWIWRL-ENEEGWWQEII 690
>UniRef100_Q8H807 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 603
Score = 133 bits (335), Expect = 3e-30
Identities = 70/211 (33%), Positives = 120/211 (56%), Gaps = 1/211 (0%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
+S LQYADDT+ + + + +K +L F+ ++GLKINF KS L S+
Sbjct: 130 LSILQYADDTILFMEHDLDEVKNLKLVLSTFEKLAGLKINFHKSELFCYGKSKFAENEYV 189
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
C +G+ F+YLG+P+ S W+ + E +L++W + +S GGR+VL+NSV
Sbjct: 190 MLFGCRSGTYPFRYLGIPMHHKKLSNKDWQIIEERFQRKLSSWKGKCLSVGGRLVLINSV 249
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
L+S+ +F LSF ++ G+ K++ + F W G +K W +C K+ GG+ ++
Sbjct: 250 LSSLAMFMLSFFEVPKGILKKLDYYRSRFFWQSDGHKRKYRLARWSVICTPKDCGGLGIQ 309
Query: 181 DIRVMNISLLAKWRWRLIDGREALWK*VLVK 211
++ V N LL+KW ++LI+ + +W+ +L K
Sbjct: 310 NLNVQNKCLLSKWLYKLIN-EDGVWQKLLRK 339
>UniRef100_Q7XXA4 OSJNBa0019G23.12 protein [Oryza sativa]
Length = 1140
Score = 132 bits (333), Expect = 5e-30
Identities = 64/193 (33%), Positives = 108/193 (55%), Gaps = 2/193 (1%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
+S LQY DD + ++ MK++L F+ +SGLKINF KS L + ++
Sbjct: 909 LSILQYVDDIVLFMNHDLEKAQNMKSVLLAFEQLSGLKINFHKSELYCFGEALEYRDQYA 968
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
C G+ F+YLG+P+ + W+ +VE +L++W + +S GGR+ L+NSV
Sbjct: 969 QLFGCQVGNFPFRYLGIPIHCRKLRNAEWKEVVERFEKKLSSWKGKLLSLGGRLTLINSV 1028
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
L+S+P++ +SFL + GV K++ ++ F W G G KK W +C+ K+ G + +
Sbjct: 1029 LSSLPMYMMSFLAIPSGVLKKLDYLRSRFYWQGDGHKKKYQLAKWDIICRPKDQGRLGIH 1088
Query: 181 DIRVMNISLLAKW 193
D+ V I+L+ +W
Sbjct: 1089 DLEV--ITLVTRW 1099
>UniRef100_Q7XE51 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 1652
Score = 128 bits (322), Expect = 9e-29
Identities = 69/209 (33%), Positives = 115/209 (55%), Gaps = 1/209 (0%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
+S LQYADDT+ + +K +L F+ +S LKINF KS L ++D
Sbjct: 1392 LSILQYADDTILFMDHDLDEARDLKLVLSTFEKLSSLKINFYKSELFCYGKAKDVEHEYV 1451
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
C FKYLG+ + + W+ + E I +L++W +++S GGR+VL+NSV
Sbjct: 1452 KLFGCDTEDYPFKYLGIRMHHKRINNKDWQGVEERIQKKLSSWKGKFLSVGGRLVLINSV 1511
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
L+++ IF LSF ++ G+ K++ + F W KK W +C+ KE GG+ ++
Sbjct: 1512 LSNLAIFMLSFFEIPKGILKKLDYYRSRFFWQCDEHKKKYRLARWSVLCKPKECGGLGIQ 1571
Query: 181 DIRVMNISLLAKWRWRLIDGREALWK*VL 209
++ + N LL+KW ++LI+ E +W+ +L
Sbjct: 1572 NLEIQNKCLLSKWLYKLIN-EEGVWQDIL 1599
>UniRef100_Q7XMF5 OSJNBa0061G20.4 protein [Oryza sativa]
Length = 632
Score = 127 bits (319), Expect = 2e-28
Identities = 77/192 (40%), Positives = 112/192 (58%), Gaps = 4/192 (2%)
Query: 4 LQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMACDFL 63
+QYADDTL KAS + L+T+KA+L+ FQ+ + LK+NF+KS L+ INV E
Sbjct: 145 VQYADDTLPYLKASGKELFTLKALLQTFQLGTSLKVNFNKSCLIPINVDEGKAQNLAAVF 204
Query: 64 NCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSVLNS 123
C GS+ F YLGLP+G + + PLV+ RL T + ++S+G R+ L+NSV +S
Sbjct: 205 GCQLGSLPFTYLGLPLGTIKPKVIDFAPLVDRTERRL-TSNAYFLSYGVRLTLVNSVFSS 263
Query: 124 MPIFYLSFLKMLVGVWKRIVRIQR*FLWGG--VGGGKKISWVNWKSVCQQKENGGVRVKD 181
+P +Y+ L + V I R +R LW G V KK S W VC+ K GG+ V +
Sbjct: 264 LPTYYMCTLMLPKTVIDSIDRARRHCLWRGSEVNSNKK-SLAAWHKVCKPKRKGGLGVLN 322
Query: 182 IRVMNISLLAKW 193
+ + N +LL K+
Sbjct: 323 LSIQNQALLIKF 334
>UniRef100_Q7XMG7 OSJNBa0028I23.15 protein [Oryza sativa]
Length = 593
Score = 126 bits (317), Expect = 3e-28
Identities = 70/199 (35%), Positives = 109/199 (54%), Gaps = 3/199 (1%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
+S LQYADDT+ + +++ +K +L F+ +SGLKINF KS L+ + +
Sbjct: 127 LSILQYADDTILFMEHDLEDAKNLKLVLSAFERLSGLKINFHKSELLCFGKAIEVEREYA 186
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
C GS T KYLGLP+ S W+ + E RL +W + +S GGR+VL+NSV
Sbjct: 187 LLFGCKTGSYTLKYLGLPMHYRKLSNKDWKEVEERFQKRLGSWKGKLLSVGGRLVLINSV 246
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
L+S+ ++ LSF ++ G+ K++ + F W KK W +C+ KE GG+ ++
Sbjct: 247 LSSLAMYMLSFFEVPKGIIKKLDYYRSRFFWQSDEHKKKYRLARWSVLCKPKECGGLGIQ 306
Query: 181 DIRVMNISL---LAKWRWR 196
++ V N L LAK W+
Sbjct: 307 NLEVQNKCLLRGLAKLVWQ 325
>UniRef100_Q6ATM7 Hypothetical protein OSJNBb0021K20.15 [Oryza sativa]
Length = 352
Score = 124 bits (310), Expect = 2e-27
Identities = 72/191 (37%), Positives = 110/191 (56%), Gaps = 2/191 (1%)
Query: 16 ASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMACDFLNCSAGSITFKYL 75
AS + L+ +KA+L+ F +GLK+NF KS L+ INVSED M GS+ F YL
Sbjct: 9 ASGKELFVLKALLQSFAQGTGLKVNFHKSCLIPINVSEDKAEMLAGVFGYRTGSMPFTYL 68
Query: 76 GLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSVLNSMPIFYLSFLKML 135
GLP+G + + PLV+ I RL+ ++ ++S G R+ L+NSV +S+P +Y+ L++
Sbjct: 69 GLPLGTTKPRVIDFAPLVDRIERRLSA-NSAFLSNGDRLTLMNSVFSSLPTYYMCILQLP 127
Query: 136 VGVWKRIVRIQR*FLW-GGVGGGKKISWVNWKSVCQQKENGGVRVKDIRVMNISLLAKWR 194
V + I R +R LW GG K S V W VC+ K GG+ V ++ + N +LL K+
Sbjct: 128 KSVIENIDRARRHCLWRGGDINSNKKSLVAWDKVCKPKVKGGLGVINLSIQNQALLLKFL 187
Query: 195 WRLIDGREALW 205
+ + R+ W
Sbjct: 188 DKFYNRRQVPW 198
>UniRef100_Q8LJX1 Putative reverse transcriptase [Sorghum bicolor]
Length = 1998
Score = 123 bits (309), Expect = 3e-27
Identities = 69/203 (33%), Positives = 110/203 (53%), Gaps = 2/203 (0%)
Query: 4 LQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMACDFL 63
+QYADDTL I + + L +K+I+ F +GLK+N KS ++ IN+ E+ +
Sbjct: 1490 IQYADDTLIIAEGDTRQLLILKSIINTFSEATGLKVNLQKSMMLPINMDEERLDTLARTF 1549
Query: 64 NCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSVLNS 123
CS GS F YLGLP+G ++ ++PL+ RL + +T ++S GR+ L N+V +
Sbjct: 1550 GCSKGSFPFTYLGLPLGITKPTIQDYQPLINKCEARLGSVAT-FLSEAGRLELTNAVFTA 1608
Query: 124 MPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVG-GGKKISWVNWKSVCQQKENGGVRVKDI 182
+P FY+ L + V +I + ++ LW G G +K WK VC+ K GG+ V DI
Sbjct: 1609 LPTFYMCTLAIPKSVIHKIDKFRKHCLWRGNGINARKPPKAAWKLVCKPKNEGGLGVIDI 1668
Query: 183 RVMNISLLAKWRWRLIDGREALW 205
N +LL K + + ++ W
Sbjct: 1669 EKQNEALLMKNLDKFFNKKDTPW 1691
>UniRef100_Q9ZUF0 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1352
Score = 122 bits (306), Expect = 6e-27
Identities = 65/205 (31%), Positives = 110/205 (52%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
++HL +ADD + + +++ AI F +S LKI+ KS++ +S +
Sbjct: 845 LTHLCFADDIMVFSDGTSKSIQGTLAIFEKFAAMSWLKISLEKSTIFMAGISPNAKTSIL 904
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
G++ KYLGLP+ + S + PLVE I R+ +W+ R++SF GR+ L+ SV
Sbjct: 905 QQFPFELGTLPVKYLGLPLLTKRMTQSDYLPLVEKIRARITSWTNRFLSFAGRLQLIKSV 964
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
L+S+ F+LS ++ + I ++ FLW G K + + W VC+ KE GG+ +K
Sbjct: 965 LSSITNFWLSVFRLPKACLQEIEKMFSAFLWSGPDLNTKKAKIAWSEVCKLKEEGGLGLK 1024
Query: 181 DIRVMNISLLAKWRWRLIDGREALW 205
++ N L K WR++ R++LW
Sbjct: 1025 PLKEANEVSLLKLIWRILSARDSLW 1049
>UniRef100_O82276 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1231
Score = 122 bits (306), Expect = 6e-27
Identities = 67/212 (31%), Positives = 112/212 (52%), Gaps = 1/212 (0%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLV-GINVSEDFMAMA 59
+SH+ +ADD + +ASV + ++ +L F SG K++ KS + NVS + +
Sbjct: 539 LSHVCFADDLILFAEASVAQIRIIRRVLERFCEASGQKVSLEKSKIFFSHNVSREMEQLI 598
Query: 60 CDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNS 119
+ KYLG+P+ + T+ ++E + RL W R +S GRI L +
Sbjct: 599 SEESGIGCTKELGKYLGMPILQKRMNKETFGEVLERVSARLAGWKGRSLSLAGRITLTKA 658
Query: 120 VLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRV 179
VL+S+P+ +S + + V + R R FLWG KK ++W+ +C+ K GG+ +
Sbjct: 659 VLSSIPVHVMSAILLPVSTLDTLDRYSRTFLWGSTMEKKKQHLLSWRKICKPKAEGGIGL 718
Query: 180 KDIRVMNISLLAKWRWRLIDGREALWK*VLVK 211
+ R MN +L+AK WRL+ +E+LW V+ K
Sbjct: 719 RSARDMNKALVAKVGWRLLQDKESLWARVVRK 750
>UniRef100_Q84T47 Putative reverse transcriptase [Oryza sativa]
Length = 746
Score = 120 bits (302), Expect = 2e-26
Identities = 69/204 (33%), Positives = 112/204 (54%), Gaps = 3/204 (1%)
Query: 4 LQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMACDFL 63
+QYADDTL I KA + ++ +K IL+ + +GLKIN+ KS ++ INV E+ +
Sbjct: 434 IQYADDTLIIMKACQKEIFNLKGILQSYSESTGLKINYQKSCMIHINVLEEDKDILSKTF 493
Query: 64 NCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSVLNS 123
C +G++ F YLGLPVG + + PL++ + RL T +++ G R+ L+N VL+S
Sbjct: 494 GCQSGNLPFTYLGLPVGTTKPKIIDFAPLIDRVERRLPA-VTMFLNHGQRLTLVNFVLSS 552
Query: 124 MPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVG--GGKKISWVNWKSVCQQKENGGVRVKD 181
+P +Y+ LK+ V I R +R LW S W VC+ K GG+ + +
Sbjct: 553 LPTYYMCSLKIPKKVIDHIDRARRHCLWRKSPNLSANSHSLAAWDLVCRPKSKGGLGIIN 612
Query: 182 IRVMNISLLAKWRWRLIDGREALW 205
+ + NI+LL K + + ++ W
Sbjct: 613 LELQNIALLMKHLDKFYNRKDVPW 636
>UniRef100_O64612 Putative reverse transcriptase [Arabidopsis thaliana]
Length = 1412
Score = 120 bits (301), Expect = 2e-26
Identities = 64/205 (31%), Positives = 109/205 (52%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
++HL +ADD + S +L + AI + F SGL I+ KS+L ++S + A
Sbjct: 953 LTHLCFADDIMVFSAGSAHSLEGVLAIFKDFAAFSGLNISLEKSTLFMASISSETCASIL 1012
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
+GS+ +YLGLP+ +++ PL+E I R+++W R++S+ GR+ LLNSV
Sbjct: 1013 ARFPFDSGSLPVRYLGLPLMTKRMTLADCLPLLEKIRSRISSWKNRFLSYAGRLQLLNSV 1072
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
++S+ F++S ++ + I +I FLW G + V W VC+ K GG+ ++
Sbjct: 1073 ISSLTKFWISAFRLPRACIREIEQISAAFLWSGTDLNPHKAKVAWHDVCKPKSEGGLGLR 1132
Query: 181 DIRVMNISLLAKWRWRLIDGREALW 205
+ N K WRL+ + +LW
Sbjct: 1133 SLVDANKICCFKLIWRLVSAKHSLW 1157
>UniRef100_Q8H992 Oryza sativa (japonica cultivar-group) retrotransposon Karma DNA,
complete sequence [Oryza sativa]
Length = 1197
Score = 120 bits (300), Expect = 3e-26
Identities = 67/218 (30%), Positives = 114/218 (51%), Gaps = 19/218 (8%)
Query: 4 LQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMACDFL 63
+QYADDTL I +A ++ +++ L F +GL+INF+KS+++ +++ + + L
Sbjct: 684 IQYADDTLVICRAEEDDVLALRSTLLQFSKATGLQINFAKSTMISLHIDRSKESSLSELL 743
Query: 64 NCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSVLNS 123
C S+ YLGLP+ + + + +P+V + L W +S R++L+N+VL+S
Sbjct: 744 QCKLESLPMSYLGLPLSLHKLTNNDLQPIVVKVDSFLTGWEASLLSQAERLILVNAVLSS 803
Query: 124 MPIFYLSFLKMLVGVWKRIVRIQR*FLWGG---VGGGKKISWVNWKSVCQQKENGGVRVK 180
+P++ +S K+ V + I + +R F W G G K + V W VC KE GG+ +K
Sbjct: 804 VPVYAMSAFKLPPKVIEAIDKRRRAFFWTGDDTCSGAKCL--VAWDEVCTAKEKGGLGIK 861
Query: 181 DIRVMNISLLAK--------------WRWRLIDGREAL 204
++ N +LL K W W+ DGR L
Sbjct: 862 SLKTQNEALLLKRLFNLFSDNSSWTNWIWKEFDGRSLL 899
>UniRef100_Q9XE45 Putative non-LTR retrolelement reverse transcriptase [Arabidopsis
thaliana]
Length = 732
Score = 119 bits (299), Expect = 4e-26
Identities = 65/206 (31%), Positives = 111/206 (53%), Gaps = 1/206 (0%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLV-GINVSEDFMAMA 59
+SH+ +ADD + +ASV + ++ +L F + SG K++ KS + NVS D +
Sbjct: 201 LSHICFADDLILFAEASVAQIRVIRRVLERFCVASGQKVSLEKSKIFFSENVSRDLGKLI 260
Query: 60 CDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNS 119
D S+ KYLG+PV + T+ ++E + RL W R++S GR+ L +
Sbjct: 261 SDESGISSTRELGKYLGMPVLQRRINKDTFGDILEKLTTRLAGWKGRFLSLAGRVTLTKA 320
Query: 120 VLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRV 179
VL+S+P+ +S + + + ++ R FLWG +K ++WK VC+ + GG+ +
Sbjct: 321 VLSSIPVHTMSTIALPKSTLDGLDKVSRSFLWGSSVTQRKQHLISWKRVCKPRSEGGLGI 380
Query: 180 KDIRVMNISLLAKWRWRLIDGREALW 205
+ + MN +LL+K WRLI +LW
Sbjct: 381 RKAQDMNKALLSKVGWRLIQDYHSLW 406
>UniRef100_Q60E26 Hypothetical protein OSJNBb0012L23.11 [Oryza sativa]
Length = 720
Score = 117 bits (294), Expect = 2e-25
Identities = 69/204 (33%), Positives = 115/204 (55%), Gaps = 10/204 (4%)
Query: 4 LQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMACDFL 63
+QYADDTL I KA + ++ +KA+L + M +GLKINF KS ++ IN M +
Sbjct: 386 IQYADDTLIIMKADQKEIFCLKALLNTYAMCTGLKINFHKSFMIPINTDSTKMEVL---- 441
Query: 64 NCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSVLNS 123
AG++ F YLGLP+ +S + PL++ + RL + +T ++++G R+ ++NSVL+S
Sbjct: 442 ---AGTLPFTYLGLPLVTTKPKVSDFAPLIDRVERRLPS-TTIFLNYGQRLTMVNSVLSS 497
Query: 124 MPIFYLSFLKMLVGVWKRIVRIQR*FLW--GGVGGGKKISWVNWKSVCQQKENGGVRVKD 181
+P +Y+ LK+ V I R +R LW + S W VC+ K+ GG+ + +
Sbjct: 498 LPTYYMCTLKLPKKVILHIDRARRHCLWRKNNELETRTHSLAAWDIVCKPKKKGGLGIIN 557
Query: 182 IRVMNISLLAKWRWRLIDGREALW 205
+ + N +LL K + + R+ W
Sbjct: 558 LEIQNTALLMKHLHKFFNNRDLPW 581
>UniRef100_Q8H0A5 Putative reverse transcriptase [Oryza sativa]
Length = 1557
Score = 117 bits (292), Expect = 3e-25
Identities = 69/213 (32%), Positives = 110/213 (51%), Gaps = 7/213 (3%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGIN----VSEDFM 56
+SHL +ADDTL KA + +K + + +G IN +K S++ N S D +
Sbjct: 1045 ISHLLFADDTLLFFKADLSQSQAIKEVFGSYATSTGQLINPTKCSILFGNSLPIASRDAI 1104
Query: 57 AMACDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVL 116
+ L ++ KYLGLP ++ L E I ++ W Y+S GG+ VL
Sbjct: 1105 T---NCLQIASTEFEDKYLGLPTPGGRMHKGRFQSLRERIWKKILQWGENYLSSGGKEVL 1161
Query: 117 LNSVLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGG 176
+ +V+ ++P++ + K+ V + + + R F WG G +K W WKS+ + K GG
Sbjct: 1162 IKAVIQAIPVYVMGIFKLPESVCEDLTSLTRNFWWGAEKGKRKTHWKAWKSLTKSKSLGG 1221
Query: 177 VRVKDIRVMNISLLAKWRWRLIDGREALWK*VL 209
+ KDIR+ N +LLA+ WRLID ++L VL
Sbjct: 1222 LGFKDIRLFNQALLARQAWRLIDNPDSLCARVL 1254
>UniRef100_Q9LDL8 Hypothetical protein AT4g08830 [Arabidopsis thaliana]
Length = 947
Score = 117 bits (292), Expect = 3e-25
Identities = 67/210 (31%), Positives = 110/210 (51%), Gaps = 1/210 (0%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLV-GINVSEDFMAMA 59
+SH+ +ADD + +ASV + ++ IL F + SG K++ KS + NVS D +
Sbjct: 353 ISHICFADDLILFAEASVSQIRVIRRILETFCIASGQKVSLDKSKIFFSKNVSRDLEKLI 412
Query: 60 CDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNS 119
+ KYLG+P+ + T+ ++E + RL W R +SF GR+ L S
Sbjct: 413 SKESGIKSTRELGKYLGMPILQRRINKDTFGEVLERVSSRLAGWKGRSLSFAGRLTLTKS 472
Query: 120 VLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRV 179
VL+ +PI +S + + + + ++ R FL G KK+ V W VC K GG+ +
Sbjct: 473 VLSLIPIHTMSTISLPQSTLEGLDKLARVFLLGSSAEKKKLHLVAWDRVCLPKSEGGLGI 532
Query: 180 KDIRVMNISLLAKWRWRLIDGREALWK*VL 209
+ + MN +L++K WRLI+ R +LW +L
Sbjct: 533 RTSKCMNKALVSKVGWRLINDRYSLWARIL 562
>UniRef100_Q7XEL7 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 339
Score = 117 bits (292), Expect = 3e-25
Identities = 57/164 (34%), Positives = 95/164 (57%)
Query: 34 VSGLKINFSKSSLVGINVSEDFMAMACDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLV 93
+SG+KIN+ KS + + V ++ + D LNC GS+ F YLGLP+G P++
Sbjct: 1 MSGMKINYQKSEVYVLGVPKEEEELYVDMLNCKVGSLPFTYLGLPMGVGKVGKRDLLPIM 60
Query: 94 ETIGGRLNTWSTRYISFGGRIVLLNSVLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGG 153
+ I RL +W + ++S+GGR + +N+ L+S+P++ + F K+ K + I+ F W G
Sbjct: 61 QKIEKRLQSWHSGHLSYGGREIRINTFLSSVPMYAMGFFKLPEEFHKNLETIRGRFYWQG 120
Query: 154 VGGGKKISWVNWKSVCQQKENGGVRVKDIRVMNISLLAKWRWRL 197
G KK + + +C+ K+ GG+ D R+MN LL+KW +L
Sbjct: 121 NGKKKKYHLIKLQGLCRPKDFGGLGFLDTRIMNFCLLSKWIMKL 164
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.330 0.142 0.465
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 351,749,176
Number of Sequences: 2790947
Number of extensions: 13537175
Number of successful extensions: 36012
Number of sequences better than 10.0: 363
Number of HSP's better than 10.0 without gapping: 229
Number of HSP's successfully gapped in prelim test: 134
Number of HSP's that attempted gapping in prelim test: 35489
Number of HSP's gapped (non-prelim): 435
length of query: 211
length of database: 848,049,833
effective HSP length: 122
effective length of query: 89
effective length of database: 507,554,299
effective search space: 45172332611
effective search space used: 45172332611
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 72 (32.3 bits)
Medicago: description of AC148816.5


