

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148652.15 + phase: 0 /pseudo
(121 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
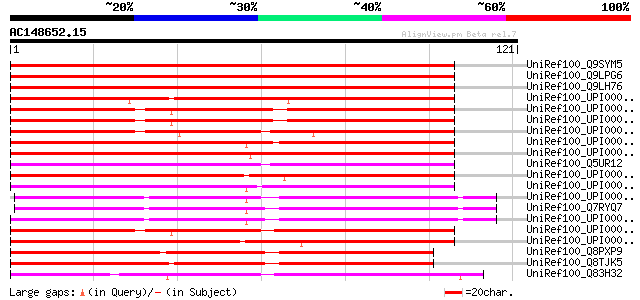
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SYM5 T30F21.10 protein [Arabidopsis thaliana] 147 4e-35
UniRef100_Q9LPG6 T3F20.18 protein [Arabidopsis thaliana] 147 5e-35
UniRef100_Q9LH76 Arabidopsis thaliana genomic DNA, chromosome 3,... 146 9e-35
UniRef100_UPI0000289B35 UPI0000289B35 UniRef100 entry 106 1e-22
UniRef100_UPI000049976E UPI000049976E UniRef100 entry 99 2e-20
UniRef100_UPI0000499547 UPI0000499547 UniRef100 entry 98 5e-20
UniRef100_UPI00002F6816 UPI00002F6816 UniRef100 entry 93 2e-18
UniRef100_UPI00002ED239 UPI00002ED239 UniRef100 entry 91 6e-18
UniRef100_UPI00002EAC57 UPI00002EAC57 UniRef100 entry 90 1e-17
UniRef100_Q5UR12 GDP mannose 4,6-dehydratase [Mimivirus] 87 9e-17
UniRef100_UPI00002BDFEA UPI00002BDFEA UniRef100 entry 87 1e-16
UniRef100_UPI0000270452 UPI0000270452 UniRef100 entry 86 1e-16
UniRef100_UPI000021B5CA UPI000021B5CA UniRef100 entry 85 3e-16
UniRef100_Q7RYQ7 Hypothetical protein [Neurospora crassa] 85 3e-16
UniRef100_UPI000023F568 UPI000023F568 UniRef100 entry 82 2e-15
UniRef100_UPI00002F4D3D UPI00002F4D3D UniRef100 entry 81 6e-15
UniRef100_UPI000025E9FF UPI000025E9FF UniRef100 entry 80 1e-14
UniRef100_Q8PXP9 DTDP-glucose 4,6-dehydratase [Methanosarcina ma... 80 1e-14
UniRef100_Q8TJK5 DTDP-glucose 4,6-dehydratase [Methanosarcina ac... 77 1e-13
UniRef100_Q83H32 DTDP-glucose 4,6-dehydratase [Tropheryma whipplei] 75 3e-13
>UniRef100_Q9SYM5 T30F21.10 protein [Arabidopsis thaliana]
Length = 669
Score = 147 bits (372), Expect = 4e-35
Identities = 73/106 (68%), Positives = 83/106 (77%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MHFAAQTHVDNS+ NSFEFT+ + Q++ FIHVSTD+ YGETDE+A+
Sbjct: 85 MHFAAQTHVDNSFGNSFEFTKNNIYGTHVLLEACKVTGQIRRFIHVSTDEVYGETDEDAL 144
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
VGNHEASQLL TNPYSA K GAEMLVM+YGRSYGLPVITTRG+ VY
Sbjct: 145 VGNHEASQLLPTNPYSATKAGAEMLVMAYGRSYGLPVITTRGNNVY 190
>UniRef100_Q9LPG6 T3F20.18 protein [Arabidopsis thaliana]
Length = 667
Score = 147 bits (371), Expect = 5e-35
Identities = 73/106 (68%), Positives = 82/106 (76%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MHFAAQTHVDNS+ NSFEFT+ + Q++ FIHVSTD+ YGETDE+A
Sbjct: 87 MHFAAQTHVDNSFGNSFEFTKNNIYGTHVLLEACKVTGQIRRFIHVSTDEVYGETDEDAA 146
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
VGNHEASQLL TNPYSA K GAEMLVM+YGRSYGLPVITTRG+ VY
Sbjct: 147 VGNHEASQLLPTNPYSATKAGAEMLVMAYGRSYGLPVITTRGNNVY 192
>UniRef100_Q9LH76 Arabidopsis thaliana genomic DNA, chromosome 3, BAC clone: T21E2
[Arabidopsis thaliana]
Length = 664
Score = 146 bits (369), Expect = 9e-35
Identities = 73/106 (68%), Positives = 82/106 (76%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MHFAAQTHVDNS+ NSFEFT+ + Q++ FIHVSTD+ YGETDE+A
Sbjct: 85 MHFAAQTHVDNSFGNSFEFTKNNIYGTHVLLEACKVTGQIRRFIHVSTDEVYGETDEDAS 144
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
VGNHEASQLL TNPYSA K GAEMLVM+YGRSYGLPVITTRG+ VY
Sbjct: 145 VGNHEASQLLPTNPYSATKAGAEMLVMAYGRSYGLPVITTRGNNVY 190
>UniRef100_UPI0000289B35 UPI0000289B35 UniRef100 entry
Length = 248
Score = 106 bits (265), Expect = 1e-22
Identities = 58/108 (53%), Positives = 77/108 (70%), Gaps = 3/108 (2%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMEL-MSFRSLQVSKNQVKGFIHVSTDKFYGETDENA 59
+HFAAQ+HVDNS+ NSFEFT+ + + S+++S++ V+ F+HVSTD+ YGE+
Sbjct: 2 LHFAAQSHVDNSFGNSFEFTKNNIEGTHVLLESVRLSRS-VRRFVHVSTDEVYGESSFEL 60
Query: 60 VVGNHE-ASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
N E AS L TNPYSA K GAEMLVM+YGRS+ +P I TRG+ VY
Sbjct: 61 DASNTEHASLLAPTNPYSATKAGAEMLVMAYGRSHDVPFIITRGNNVY 108
>UniRef100_UPI000049976E UPI000049976E UniRef100 entry
Length = 341
Score = 99.4 bits (246), Expect = 2e-20
Identities = 55/107 (51%), Positives = 71/107 (65%), Gaps = 6/107 (5%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSK-NQVKGFIHVSTDKFYGETDENA 59
+HFAAQTHVDNS+ NSF+FT + L+VSK N +K FIHVSTD+ YG+ NA
Sbjct: 86 LHFAAQTHVDNSFGNSFQFTHNNIYGTHVL--LEVSKANHIKRFIHVSTDEVYGQVIGNA 143
Query: 60 VVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
N S L TNPY+A K GAE + S+ +S+GLP+I TRG+ V+
Sbjct: 144 ATEN---SLLNPTNPYAATKAGAEFIARSFYQSFGLPLIITRGNNVF 187
>UniRef100_UPI0000499547 UPI0000499547 UniRef100 entry
Length = 342
Score = 97.8 bits (242), Expect = 5e-20
Identities = 53/107 (49%), Positives = 71/107 (65%), Gaps = 6/107 (5%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSK-NQVKGFIHVSTDKFYGETDENA 59
+HFAAQTHVDNS+ NSF+FT + L+++K N +K FIHVSTD+ YG+ NA
Sbjct: 84 LHFAAQTHVDNSFGNSFQFTHNNIYGTHVL--LEIAKANHIKRFIHVSTDEVYGQVIGNA 141
Query: 60 VVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
N S L TNPYSA K GAE + ++ +S+GLP+I TRG+ V+
Sbjct: 142 ATEN---SLLNPTNPYSATKAGAEFIARAFYQSFGLPLIITRGNNVF 185
>UniRef100_UPI00002F6816 UPI00002F6816 UniRef100 entry
Length = 227
Score = 92.8 bits (229), Expect = 2e-18
Identities = 55/108 (50%), Positives = 70/108 (63%), Gaps = 6/108 (5%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQ-VKGFIHVSTDKFYGETDENA 59
+H AAQTHVDNS+ NSF FTQ + M ++ +K +K FIHVSTD+ YG + EN
Sbjct: 101 IHAAAQTHVDNSFGNSFTFTQNNVMGTHVL--IETAKTHGIKRFIHVSTDEVYGSSYEND 158
Query: 60 VVGNHEASQLLE-TNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
H S +LE TNPY+A K AE +V SY RS+ LPVI TRG+ V+
Sbjct: 159 P--RHIESDVLEPTNPYAATKAAAENIVKSYYRSFNLPVIITRGNNVF 204
>UniRef100_UPI00002ED239 UPI00002ED239 UniRef100 entry
Length = 265
Score = 90.9 bits (224), Expect = 6e-18
Identities = 47/108 (43%), Positives = 67/108 (61%), Gaps = 3/108 (2%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGET--DEN 58
+HFAAQ+HV NS+E+S +T + + + + + FIHVSTD+ YGE+ D N
Sbjct: 55 IHFAAQSHVQNSFEDSLNYTNDNIVGTHTLLEVNRKNKHLIKFIHVSTDEVYGESMLDVN 114
Query: 59 AVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
N E S L TNPY+A K GAE++ SY S+G+P++ TRG+ VY
Sbjct: 115 EKHKN-EHSILCPTNPYAATKAGAELIAQSYNHSFGMPIVITRGNNVY 161
>UniRef100_UPI00002EAC57 UPI00002EAC57 UniRef100 entry
Length = 229
Score = 89.7 bits (221), Expect = 1e-17
Identities = 47/107 (43%), Positives = 67/107 (61%), Gaps = 1/107 (0%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETD-ENA 59
+HFAAQ+HV NS+ +S ++TQ + M + ++K FIHVSTD+ YG++ E
Sbjct: 105 IHFAAQSHVCNSFTDSLQYTQDNIMGTHNLLECCRLYGRIKRFIHVSTDEVYGQSLLEKN 164
Query: 60 VVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
E S L TNPY+A K GAE++ SYG S+ +P+I TRG+ VY
Sbjct: 165 EDQKTEQSVLCPTNPYAATKAGAELIAQSYGHSFKMPIIITRGNNVY 211
>UniRef100_Q5UR12 GDP mannose 4,6-dehydratase [Mimivirus]
Length = 323
Score = 87.0 bits (214), Expect = 9e-17
Identities = 47/106 (44%), Positives = 62/106 (58%), Gaps = 2/106 (1%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
+HFAA +HVDNS++NS FT+T+ ++K F H+STD+ YGE D
Sbjct: 77 VHFAAHSHVDNSFKNSLAFTETNVFGTHVLLECSRMYGKLKLFFHMSTDEVYGEIDTTDT 136
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
+ E S L TNPY+A K GAE +V SY SY LP+I R + VY
Sbjct: 137 --SREVSLLCPTNPYAATKAGAEHIVKSYFLSYKLPIIIARCNNVY 180
>UniRef100_UPI00002BDFEA UPI00002BDFEA UniRef100 entry
Length = 299
Score = 86.7 bits (213), Expect = 1e-16
Identities = 46/108 (42%), Positives = 67/108 (61%), Gaps = 3/108 (2%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
+HFAAQ+HV +S+ +S +T + + + L +++ FIHVSTD+ YGE+ N V
Sbjct: 121 IHFAAQSHVQDSFTHSLRYTNDNILGTHNLLELSRLYGKIERFIHVSTDEVYGESI-NDV 179
Query: 61 VGNH--EASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
H E S L TNPY+A K GAE++ SY S+ +P+I TRG+ VY
Sbjct: 180 NEKHKTEHSILCPTNPYAATKAGAELIAQSYAHSFKMPIIITRGNNVY 227
>UniRef100_UPI0000270452 UPI0000270452 UniRef100 entry
Length = 261
Score = 86.3 bits (212), Expect = 1e-16
Identities = 45/108 (41%), Positives = 65/108 (59%), Gaps = 3/108 (2%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGET--DEN 58
+HFAAQ+HV NS+E+S +T + + + + + FIHVSTD+ YGE+ D N
Sbjct: 59 IHFAAQSHVQNSFEDSLNYTNDNIVGTHTLLEVNRKNKHLIKFIHVSTDEVYGESMLDVN 118
Query: 59 AVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
E S L TNPY+A K GAE++ SY S+ +P++ TRG+ VY
Sbjct: 119 E-KHKTEHSVLCPTNPYAATKAGAELIAQSYNHSFKMPIVITRGNNVY 165
>UniRef100_UPI000021B5CA UPI000021B5CA UniRef100 entry
Length = 343
Score = 85.1 bits (209), Expect = 3e-16
Identities = 53/118 (44%), Positives = 69/118 (57%), Gaps = 9/118 (7%)
Query: 2 HFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGET---DEN 58
HFAAQ+HVD S+ NS+ FT T+ K ++ FIH+STD+ YGE DE+
Sbjct: 44 HFAAQSHVDLSFGNSYSFTHTNVYGTHVLLE-SAKKVGIRRFIHISTDEVYGEVKDDDED 102
Query: 59 AVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVYPFSHERKESI 116
+ E+S L TNPY+A K AEMLV SY +S+ LPVI R + VY H+ E I
Sbjct: 103 LL----ESSILAPTNPYAASKAAAEMLVHSYQKSFKLPVIIVRSNNVYG-PHQYPEKI 155
>UniRef100_Q7RYQ7 Hypothetical protein [Neurospora crassa]
Length = 428
Score = 85.1 bits (209), Expect = 3e-16
Identities = 54/117 (46%), Positives = 68/117 (57%), Gaps = 7/117 (5%)
Query: 2 HFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGET--DENA 59
HFAAQ+HVD S+ N F FT T+ K +K FIHVSTD+ YGE DE+
Sbjct: 121 HFAAQSHVDLSFGNPFGFTHTNVYGTHVLLE-SARKAGIKRFIHVSTDEVYGEVKDDEDD 179
Query: 60 VVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVYPFSHERKESI 116
++ E S L TNPY+A K AEMLV SY +S+ LP+I R + VY H+ E I
Sbjct: 180 LL---ETSILAPTNPYAASKAAAEMLVNSYKKSFKLPIIIVRSNNVYG-PHQYPEKI 232
>UniRef100_UPI000023F568 UPI000023F568 UniRef100 entry
Length = 449
Score = 82.4 bits (202), Expect = 2e-15
Identities = 52/118 (44%), Positives = 68/118 (57%), Gaps = 7/118 (5%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGET--DEN 58
+HFAAQ+HVD S+ NS+ FT T+ K + FIHVSTD+ YGE D++
Sbjct: 128 LHFAAQSHVDLSFGNSYGFTHTNVYGTHVLLE-SAKKVGIGRFIHVSTDEVYGEVKEDDD 186
Query: 59 AVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVYPFSHERKESI 116
++ E S L TNPY+A K AEMLV SY +S+ LP I R + VY H+ E I
Sbjct: 187 DLL---ETSILAPTNPYAASKAAAEMLVQSYQKSFKLPAIIVRSNNVYG-PHQYPEKI 240
>UniRef100_UPI00002F4D3D UPI00002F4D3D UniRef100 entry
Length = 196
Score = 80.9 bits (198), Expect = 6e-15
Identities = 45/107 (42%), Positives = 69/107 (64%), Gaps = 6/107 (5%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSK-NQVKGFIHVSTDKFYGETDENA 59
+H AA++HVD S+ + +F QT+ M + ++ SK N++K IH+STD+ YGE + +
Sbjct: 50 IHAAAESHVDRSFVLNKKFIQTNVMGTRNV--MEASKINKIKKIIHISTDEVYGEIYKGS 107
Query: 60 VVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
+E+S L +NPYS+ K AEM+V Y RSY LPVI RG+ ++
Sbjct: 108 F---NESSNLKPSNPYSSSKAAAEMIVNGYIRSYNLPVIIIRGNNIF 151
>UniRef100_UPI000025E9FF UPI000025E9FF UniRef100 entry
Length = 257
Score = 80.1 bits (196), Expect = 1e-14
Identities = 41/108 (37%), Positives = 67/108 (61%), Gaps = 3/108 (2%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
+HFAAQ+HV NS+ ++ ++++ + + + +++ FIHVSTD+ YGE+ V
Sbjct: 28 IHFAAQSHVQNSFSDALQYSKDNIIGTHTLLESVRVYGKIQKFIHVSTDEVYGES-MLTV 86
Query: 61 VGNHEASQ--LLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
H+ Q L +NPY+A K GAE++ SY S+ +P+I TRG+ VY
Sbjct: 87 NEKHKTEQSVLCPSNPYAATKAGAELIAQSYIHSFNMPIIITRGNNVY 134
>UniRef100_Q8PXP9 DTDP-glucose 4,6-dehydratase [Methanosarcina mazei]
Length = 321
Score = 79.7 bits (195), Expect = 1e-14
Identities = 42/101 (41%), Positives = 64/101 (62%), Gaps = 4/101 (3%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
+HFAA++HVD S E+ F +T+ + + ++ N++K FIHVSTD+ YG T E +
Sbjct: 80 VHFAAESHVDRSIEDGSVFVRTNVLGTNTLLQSALA-NKIKKFIHVSTDEVYGSTMEGSF 138
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTR 101
E +L ++PYS+ K G+++L MSY +YGLPV TR
Sbjct: 139 T---ETDKLNPSSPYSSSKAGSDLLAMSYHTTYGLPVCITR 176
>UniRef100_Q8TJK5 DTDP-glucose 4,6-dehydratase [Methanosarcina acetivorans]
Length = 318
Score = 76.6 bits (187), Expect = 1e-13
Identities = 39/101 (38%), Positives = 62/101 (60%), Gaps = 4/101 (3%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
+HFAA++HVD S E+ F +T+ + + ++ N +K F+H+STD+ YG E +
Sbjct: 77 VHFAAESHVDRSIEDGSVFVRTNVLGTNTLLQSALANN-IKKFVHISTDEVYGSIKEGSF 135
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTR 101
E +L ++PYS+ K G+++L MSY +YGLPV TR
Sbjct: 136 T---ETDKLNPSSPYSSSKAGSDLLAMSYYTTYGLPVCITR 173
>UniRef100_Q83H32 DTDP-glucose 4,6-dehydratase [Tropheryma whipplei]
Length = 327
Score = 75.5 bits (184), Expect = 3e-13
Identities = 45/115 (39%), Positives = 67/115 (58%), Gaps = 7/115 (6%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVS-KNQVKGFIHVSTDKFYGETDENA 59
+HFAA+THVD S +N+ F +T+ L + R L+ + +N + FIH+STD+ YG + +
Sbjct: 82 VHFAAETHVDRSIKNASAFVETNV--LGTGRLLEAAARNNIDRFIHISTDEVYGSIESGS 139
Query: 60 VVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY-PFSHERK 113
E LL +PYSA K +++LV SY ++Y LPV TR Y P+ K
Sbjct: 140 W---DENQPLLPNSPYSASKAASDLLVRSYHKTYDLPVCITRCSNNYGPYQFPEK 191
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.130 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 183,510,110
Number of Sequences: 2790947
Number of extensions: 5939590
Number of successful extensions: 17918
Number of sequences better than 10.0: 1355
Number of HSP's better than 10.0 without gapping: 590
Number of HSP's successfully gapped in prelim test: 765
Number of HSP's that attempted gapping in prelim test: 16241
Number of HSP's gapped (non-prelim): 1364
length of query: 121
length of database: 848,049,833
effective HSP length: 97
effective length of query: 24
effective length of database: 577,327,974
effective search space: 13855871376
effective search space used: 13855871376
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC148652.15


