

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148652.13 - phase: 0 /pseudo
(395 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
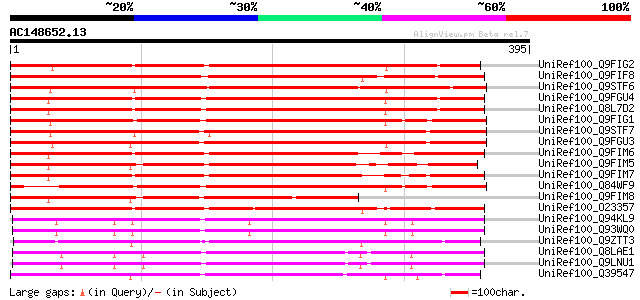
UniRef100_Q9FIG2 Serine protease-like protein [Arabidopsis thali... 361 2e-98
UniRef100_Q9FIF8 Serine protease-like protein [Arabidopsis thali... 358 2e-97
UniRef100_Q9STF6 Subtilisin-like proteinase homolog [Arabidopsis... 350 4e-95
UniRef100_Q9FGU4 Subtilisin-like protease [Arabidopsis thaliana] 348 2e-94
UniRef100_Q8L7D2 Subtilisin-like serine protease [Arabidopsis th... 348 2e-94
UniRef100_Q9FIG1 Serine protease-like protein [Arabidopsis thali... 347 4e-94
UniRef100_Q9STF7 Subtilisin-like proteinase homolog [Arabidopsis... 337 3e-91
UniRef100_Q9FGU3 Serine protease-like protein [Arabidopsis thali... 337 5e-91
UniRef100_Q9FIM6 Subtilisin-like serine protease [Arabidopsis th... 331 2e-89
UniRef100_Q9FIM5 Subtilisin-like serine protease [Arabidopsis th... 330 4e-89
UniRef100_Q9FIM7 Subtilisin-like serine protease [Arabidopsis th... 320 6e-86
UniRef100_Q84WF9 Putative subtilisin-like serine protease [Arabi... 310 3e-83
UniRef100_Q9FIM8 Subtilisin-like serine protease [Arabidopsis th... 301 3e-80
UniRef100_O23357 CUCUMISIN [Arabidopsis thaliana] 299 8e-80
UniRef100_Q94KL9 Subtilisin-like protein [Glycine max] 295 2e-78
UniRef100_Q93WQ0 Subtilisin-type protease [Glycine max] 295 2e-78
UniRef100_Q9ZTT3 Subtilisin-like protease C1 [Glycine max] 285 2e-75
UniRef100_Q8LAE1 Subtilisin-like serine protease [Arabidopsis th... 281 2e-74
UniRef100_Q9LNU1 T20H2.6 protein [Arabidopsis thaliana] 281 2e-74
UniRef100_Q39547 Pre-pro-cucumisin precursor [Cucumis melo] 278 2e-73
>UniRef100_Q9FIG2 Serine protease-like protein [Arabidopsis thaliana]
Length = 732
Score = 361 bits (927), Expect = 2e-98
Identities = 184/362 (50%), Positives = 247/362 (67%), Gaps = 11/362 (3%)
Query: 1 MKGVISVFPSEEFHIQTTRSWDFLGLRQSIK--RDQIIETDLVIGVIDIRIWPESESFND 58
M GV+SVFP+++ +QTT SWDF+GL++ IK R+ +E+D +IGVID I PES+SF+D
Sbjct: 94 MVGVVSVFPNKKLQLQTTTSWDFMGLKEGIKTKRNPTVESDTIIGVIDSGITPESQSFSD 153
Query: 59 KGLGPIPKKWRGVCAGGGNFSCNNKIIGARFYGDGDVSARYSFGHRTHTASTAGGREVED 118
KG GP P+KW+GVC+GG NF+CNNK+IGAR Y R GH THTASTA G V D
Sbjct: 154 KGFGPPPQKWKGVCSGGKNFTCNNKLIGARDYTSE--GTRDMDGHGTHTASTAAGNAVVD 211
Query: 119 VSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGP 178
SF+G GT RGGVP+SR+AAYK+C CS +A+L+AFDDAI+DGVD+IT+S+G
Sbjct: 212 ASFFGIGNGTVRGGVPASRVAAYKVCTPTG---CSSEALLSAFDDAIADGVDLITISIGD 268
Query: 179 EHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFID 238
+ AS F NDPIAIG+FHAM KG+LT +AGN GP P SV APW+++VAA++ +R F+
Sbjct: 269 KTASMFQNDPIAIGAFHAMAKGVLTVNSAGNSGPKPISVSGVAPWILTVAASTTNRGFVT 328
Query: 239 KVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLAGGNASPEMCD--CIDENMVKGKL 296
KV+LGNGKT VGKS+N GK P+ + A A S +C+ C+D++ VKGK+
Sbjct: 329 KVVLGNGKTLVGKSVNAYEMKGKDYPLVYGKSAASSACDAESAGLCELSCVDKSRVKGKI 388
Query: 297 VLCGSRNGKDLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTNSSKYP 356
++CG G + + GA+G I+ K + +F+ P P+ L T DF + SY S+ P
Sbjct: 389 LVCGGPGGLKIVESVGAVGLIYRTPKPDV--AFIHPLPAAGLLTEDFESLVSYLESTDSP 446
Query: 357 VA 358
A
Sbjct: 447 QA 448
>UniRef100_Q9FIF8 Serine protease-like protein [Arabidopsis thaliana]
Length = 729
Score = 358 bits (918), Expect = 2e-97
Identities = 189/366 (51%), Positives = 245/366 (66%), Gaps = 14/366 (3%)
Query: 1 MKGVISVFPSEEFHIQTTRSWDFLGLRQSIKRDQIIETDLVIGVIDIRIWPESESFNDKG 60
MK V+SVFPS+ + TTRSWDF+G + +R+ + E+D+++GVID IWPESESF+D+G
Sbjct: 94 MKEVVSVFPSKSHELTTTRSWDFVGFGEKARRESVKESDVIVGVIDSGIWPESESFDDEG 153
Query: 61 LGPIPKKWRGVCAGGGNFSCNNKIIGARFYGDGDVSARYSFGHRTHTASTAGGREVEDVS 120
GP PKKW+G C GG F+CNNK+IGARFY SAR GH THTASTA G V+ S
Sbjct: 154 FGPPPKKWKGSCKGGLKFACNNKLIGARFYNKFADSARDEEGHGTHTASTAAGNAVQAAS 213
Query: 121 FYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEH 180
FYG A+GTARGGVPS+RIAAYK+C + RC+ ILAAFDDAI+DGVDVI++S+ ++
Sbjct: 214 FYGLAQGTARGGVPSARIAAYKVCFN----RCNDVDILAAFDDAIADGVDVISISISADY 269
Query: 181 ASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKV 240
S+ LN +AIGSFHAM +GI+T +AGN GP SV + +PW+++VAA+ DRQFID+V
Sbjct: 270 VSNLLNASVAIGSFHAMMRGIITAGSAGNNGPDQGSVANVSPWMITVAASGTDRQFIDRV 329
Query: 241 ILGNGKTFVGKSINITPSNGKKIPIAV-----RNAQACLAGGNASPEMCDCIDENMVKGK 295
+LGNGK G S+N NG K PI RN AG +S C+D +VKGK
Sbjct: 330 VLGNGKALTGISVNTFNLNGTKFPIVYGQNVSRNCSQAQAGYCSS----GCVDSELVKGK 385
Query: 296 LVLCGSRNGKDLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTNSSKY 355
+VLC G AY GAIG I T ++FV P P+ +L D+ I+SY S++
Sbjct: 386 IVLCDDFLGYREAYLAGAIGVIVQNTLLP-DSAFVVPFPASSLGFEDYKSIKSYIESAEP 444
Query: 356 PVADFL 361
P A+ L
Sbjct: 445 PQAEIL 450
>UniRef100_Q9STF6 Subtilisin-like proteinase homolog [Arabidopsis thaliana]
Length = 739
Score = 350 bits (898), Expect = 4e-95
Identities = 193/371 (52%), Positives = 250/371 (67%), Gaps = 12/371 (3%)
Query: 1 MKGVISVFPSEEFHIQTTRSWDFLGLRQS--IKRDQIIETDLVIGVIDIRIWPESESFND 58
M V+SVFP+++ +QTT SW+F+GL++S KR+ IIE+D +IGVID I+PES+SF+
Sbjct: 97 MDEVVSVFPNKKLKLQTTTSWNFMGLKESKRTKRNTIIESDTIIGVIDSGIYPESDSFSG 156
Query: 59 KGLGPIPKKWRGVCAGGGNFSCNNKIIGARFYGDG----DVSARYSFGHRTHTASTAGGR 114
KG GP PKKW+GVC GG NF+ NNK+IGAR+Y SAR GH +HTASTA G
Sbjct: 157 KGFGPPPKKWKGVCKGGKNFTWNNKLIGARYYTPKLEGFPESARDYMGHGSHTASTAAGN 216
Query: 115 EVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITV 174
V+ VSFYG GTARGGVP++RIA YK+C +G C+ D ILAAFDDAI+D VD+IT+
Sbjct: 217 AVKHVSFYGLGNGTARGGVPAARIAVYKVCDPGVDG-CTTDGILAAFDDAIADKVDIITI 275
Query: 175 SLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDR 234
S+G +++S F DPIAIG+FHAM KGIL +AGN GP PS+V S APW+ +VAA++ +R
Sbjct: 276 SIGGDNSSPFEEDPIAIGAFHAMAKGILIVNSAGNSGPEPSTVASIAPWMFTVAASNTNR 335
Query: 235 QFIDKVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLAGGNASPEMCD--CIDENMV 292
F+ KV+LGNGKT VG+S+N NGKK P+ V A + G AS C C+D V
Sbjct: 336 AFVTKVVLGNGKTVVGRSVNSFDLNGKKYPL-VYGKSASSSCGAASAGFCSPGCLDSKRV 394
Query: 293 KGKLVLCGSRNGKDLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTNS 352
KGK+VLC S D A A GAI SI ++ + + F P L +D+ + SY NS
Sbjct: 395 KGKIVLCDSPQNPDEAQAMGAIASIVRSHRTDVASIFSFPVSV--LLEDDYNTVLSYMNS 452
Query: 353 SKYPVADFLFS 363
+K P A L S
Sbjct: 453 TKNPKAAVLKS 463
>UniRef100_Q9FGU4 Subtilisin-like protease [Arabidopsis thaliana]
Length = 707
Score = 348 bits (893), Expect = 2e-94
Identities = 180/365 (49%), Positives = 246/365 (67%), Gaps = 11/365 (3%)
Query: 1 MKGVISVFPSEEFHIQTTRSWDFLGLRQ--SIKRDQIIETDLVIGVIDIRIWPESESFND 58
++GV+SVFP++ + TT SWDF+G+++ + KR+ IE+D +IGVID IWPES+SF+D
Sbjct: 66 IEGVVSVFPNKILQLHTTTSWDFMGVKEGKNTKRNLAIESDTIIGVIDTGIWPESKSFSD 125
Query: 59 KGLGPIPKKWRGVCAGGGNFSCNNKIIGARFYGDGDVSARYSFGHRTHTASTAGGREVED 118
KG GP PKKW+GVC+GG NF+CNNK+IGAR Y R + GH THTASTA G V+D
Sbjct: 126 KGFGPPPKKWKGVCSGGKNFTCNNKLIGARDYTSE--GTRDTSGHGTHTASTAAGNAVKD 183
Query: 119 VSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGP 178
SF+G GT RGGVP+SRIAAYK+C D CS +A+L++FDDAI+DGVD+IT+S+G
Sbjct: 184 TSFFGIGNGTVRGGVPASRIAAYKVCTD---SGCSSEALLSSFDDAIADGVDLITISIGF 240
Query: 179 EHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFID 238
+ S F +DPIAIG+FHAM KGILT +AGN GP P++V APW+ +VAA++ +R FI
Sbjct: 241 QFPSIFEDDPIAIGAFHAMAKGILTVSSAGNSGPKPTTVSHVAPWIFTVAASTTNRGFIT 300
Query: 239 KVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLAGGNASPEMC--DCIDENMVKGKL 296
KV+LGNGKT G+S+N GKK P+ + A A + +C C++++ VKGK+
Sbjct: 301 KVVLGNGKTLAGRSVNAFDMKGKKYPLVYGKSAASSACDAKTAALCAPACLNKSRVKGKI 360
Query: 297 VLCGSRNGKDLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTNSSKYP 356
++CG +G +A + GAI I + + +F P+ LK DF + SY S P
Sbjct: 361 LVCGGPSGYKIAKSVGAIAIIDKSPRPDV--AFTHHLPASGLKAKDFKSLVSYIESQDSP 418
Query: 357 VADFL 361
A L
Sbjct: 419 QAAVL 423
>UniRef100_Q8L7D2 Subtilisin-like serine protease [Arabidopsis thaliana]
Length = 736
Score = 348 bits (893), Expect = 2e-94
Identities = 180/365 (49%), Positives = 246/365 (67%), Gaps = 11/365 (3%)
Query: 1 MKGVISVFPSEEFHIQTTRSWDFLGLRQ--SIKRDQIIETDLVIGVIDIRIWPESESFND 58
++GV+SVFP++ + TT SWDF+G+++ + KR+ IE+D +IGVID IWPES+SF+D
Sbjct: 95 IEGVVSVFPNKILQLHTTTSWDFMGVKEGKNTKRNLAIESDTIIGVIDTGIWPESKSFSD 154
Query: 59 KGLGPIPKKWRGVCAGGGNFSCNNKIIGARFYGDGDVSARYSFGHRTHTASTAGGREVED 118
KG GP PKKW+GVC+GG NF+CNNK+IGAR Y R + GH THTASTA G V+D
Sbjct: 155 KGFGPPPKKWKGVCSGGKNFTCNNKLIGARDYTSE--GTRDTSGHGTHTASTAAGNAVKD 212
Query: 119 VSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGP 178
SF+G GT RGGVP+SRIAAYK+C D CS +A+L++FDDAI+DGVD+IT+S+G
Sbjct: 213 TSFFGIGNGTVRGGVPASRIAAYKVCTD---SGCSSEALLSSFDDAIADGVDLITISIGF 269
Query: 179 EHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFID 238
+ S F +DPIAIG+FHAM KGILT +AGN GP P++V APW+ +VAA++ +R FI
Sbjct: 270 QFPSIFEDDPIAIGAFHAMAKGILTVSSAGNSGPKPTTVSHVAPWIFTVAASTTNRGFIT 329
Query: 239 KVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLAGGNASPEMC--DCIDENMVKGKL 296
KV+LGNGKT G+S+N GKK P+ + A A + +C C++++ VKGK+
Sbjct: 330 KVVLGNGKTLAGRSVNAFDMKGKKYPLVYGKSAASSACDAKTAALCAPACLNKSRVKGKI 389
Query: 297 VLCGSRNGKDLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTNSSKYP 356
++CG +G +A + GAI I + + +F P+ LK DF + SY S P
Sbjct: 390 LVCGGPSGYKIAKSVGAIAIIDKSPRPDV--AFTHHLPASGLKAKDFKSLVSYIESQDSP 447
Query: 357 VADFL 361
A L
Sbjct: 448 QAAVL 452
>UniRef100_Q9FIG1 Serine protease-like protein [Arabidopsis thaliana]
Length = 697
Score = 347 bits (889), Expect = 4e-94
Identities = 183/367 (49%), Positives = 244/367 (65%), Gaps = 14/367 (3%)
Query: 1 MKGVISVFPSEEFHIQTTRSWDFLGLRQS--IKRDQIIETDLVIGVIDIRIWPESESFND 58
M+GV+SVFP+++ +QT+ SWDF+GL++ KR+ +E+D +IGV D IWPESESF+D
Sbjct: 59 MEGVVSVFPNKKLKLQTSASWDFMGLKEGKGTKRNPSVESDTIIGVFDGGIWPESESFSD 118
Query: 59 KGLGPIPKKWRGVCAGGGNFSCNNKIIGARFYGDGDVSARYSFGHRTHTASTAGGREVED 118
KG GP PKKW+G+CAGG NF+CNNK+IGAR Y GD AR S GH THTAS A G V +
Sbjct: 119 KGFGPPPKKWKGICAGGKNFTCNNKLIGARHYSPGD--ARDSTGHGTHTASIAAGNAVAN 176
Query: 119 VSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGP 178
SF+G GT RG VP+SRIA Y++C G C DAIL+AFDDAISDGVD+IT+S+G
Sbjct: 177 TSFFGIGNGTVRGAVPASRIAVYRVCA----GECRDDAILSAFDDAISDGVDIITISIGD 232
Query: 179 EHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFID 238
+ F DPIAIG+FHAM KGILT AAGN GP +S+ S APWL++VAA++ +R+F+
Sbjct: 233 INVYPFEKDPIAIGAFHAMSKGILTVNAAGNTGPDTASITSLAPWLLTVAASTANREFVS 292
Query: 239 KVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLAGGNASPEMC--DCIDENMVKGKL 296
KV+LG+GKT VGKS+N GKK P+ + A E C +C+D ++VKGK+
Sbjct: 293 KVVLGDGKTLVGKSVNGFDLKGKKFPLVYGKSAALSLSQAKCAEDCTPECLDASLVKGKI 352
Query: 297 VLCGSRNGKDLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTNSSKYP 356
++C +R +AY A+ +I + + + P L+ +DF + SY S K P
Sbjct: 353 LVC-NRFLPYVAYTKRAVAAIF---EDGSDWAQINGLPVSGLQKDDFESVLSYFKSEKSP 408
Query: 357 VADFLFS 363
A L S
Sbjct: 409 EAAVLKS 415
>UniRef100_Q9STF7 Subtilisin-like proteinase homolog [Arabidopsis thaliana]
Length = 736
Score = 337 bits (864), Expect = 3e-91
Identities = 187/372 (50%), Positives = 243/372 (65%), Gaps = 14/372 (3%)
Query: 1 MKGVISVFPSEEFHIQTTRSWDFLGLRQS--IKRDQIIETDLVIGVIDIRIWPESESFND 58
M V+SVFPS+ ++QTT SW+F+GL++ KR+ +IE+D +IGVID I+PES+SF+
Sbjct: 96 MDEVVSVFPSKNLNLQTTTSWNFMGLKEGKRTKRNPLIESDTIIGVIDSGIYPESDSFSG 155
Query: 59 KGLGPIPKKWRGVCAGGGNFSCNNKIIGARFYGDG----DVSARYSFGHRTHTASTAGGR 114
KG GP PKKW+GVC GG NF+CNNK+IGAR+Y SAR + GH +HTAS A G
Sbjct: 156 KGFGPPPKKWKGVCKGGTNFTCNNKLIGARYYTPKLEGFPESARDNTGHGSHTASIAAGN 215
Query: 115 EVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNG--RCSGDAILAAFDDAISDGVDVI 172
V+ VSFYG GT RGGVP++RIA YK+C D G RC+ D ILAAFDDAI+D VD+I
Sbjct: 216 AVKHVSFYGLGNGTVRGGVPAARIAVYKVC---DPGVIRCTSDGILAAFDDAIADKVDII 272
Query: 173 TVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSI 232
TVSLG + F D +AIG+FHAM KGILT AGN GP ++ S APWL +VAA+++
Sbjct: 273 TVSLGADAVGTFEEDTLAIGAFHAMAKGILTVNGAGNNGPERRTIVSMAPWLFTVAASNM 332
Query: 233 DRQFIDKVILGNGKTFVGKSINITPSNGKKIPIAV-RNAQACLAGGNASPEMCDCIDENM 291
+R FI KV+LGNGKT VG+S+N NGKK P+ ++A + +A C+D
Sbjct: 333 NRAFITKVVLGNGKTIVGRSVNSFDLNGKKYPLVYGKSASSRCDASSAGFCSPGCLDSKR 392
Query: 292 VKGKLVLCGSRNGKDLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTN 351
VKGK+VLC ++ A A GA+ SI V A+ V P L +D+ + SY N
Sbjct: 393 VKGKIVLCDTQRNPGEAQAMGAVASI--VRNPYEDAASVFSFPVSVLSEDDYNIVLSYVN 450
Query: 352 SSKYPVADFLFS 363
S+K P A L S
Sbjct: 451 STKNPKAAVLKS 462
>UniRef100_Q9FGU3 Serine protease-like protein [Arabidopsis thaliana]
Length = 741
Score = 337 bits (863), Expect = 5e-91
Identities = 180/370 (48%), Positives = 242/370 (64%), Gaps = 12/370 (3%)
Query: 1 MKGVISVFPSEEFHIQTTRSWDFLGLRQSIK--RDQIIETDLVIGVIDIRIWPESESFND 58
M+ V+SVFPS + +QTT SW+F+GL++ IK R + IE+D +IGVID I+PES+SF+D
Sbjct: 97 MERVVSVFPSRKLKLQTTSSWNFMGLKEGIKTKRTRSIESDTIIGVIDSGIYPESDSFSD 156
Query: 59 KGLGPIPKKWRGVCAGGGNFSCNNKIIGARFY---GDGDVSARYSFGHRTHTASTAGGRE 115
+G GP PKKW+G CAGG NF+CNNK+IGAR Y + +AR GH THTAS A G
Sbjct: 157 QGFGPPPKKWKGTCAGGKNFTCNNKVIGARDYTAKSKANQTARDYSGHGTHTASIAAGNA 216
Query: 116 VEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVS 175
V + +FYG GTARGGVP++RIA YK+C DN C G+A+++AFDDAI+DGVDVI++S
Sbjct: 217 VANSNFYGLGNGTARGGVPAARIAVYKVC---DNEGCDGEAMMSAFDDAIADGVDVISIS 273
Query: 176 LGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQ 235
+ ++ F DPIAIG+FHAM G+LT AAGN GP S+V S APW+ SVAA+ +R
Sbjct: 274 IVLDNIPPFEEDPIAIGAFHAMAVGVLTVNAAGNNGPKISTVTSTAPWVFSVAASVTNRA 333
Query: 236 FIDKVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLAGGNASPEMCD--CIDENMVK 293
F+ KV+LG+GK +G+S+N NG P+ + A +C+ C+D +VK
Sbjct: 334 FMAKVVLGDGKILIGRSVNTYDMNGTNYPLVYGKSAALSTCSVDKARLCEPKCLDGKLVK 393
Query: 294 GKLVLCGSRNGKDLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTNSS 353
GK+VLC S G A GA+GSI V + +F+ P L +D+ + SY NS+
Sbjct: 394 GKIVLCDSTKGLIEAQKLGAVGSI--VKNPEPDRAFIRSFPVSFLSNDDYKSLVSYMNST 451
Query: 354 KYPVADFLFS 363
K P A L S
Sbjct: 452 KNPKATVLKS 461
>UniRef100_Q9FIM6 Subtilisin-like serine protease [Arabidopsis thaliana]
Length = 710
Score = 331 bits (849), Expect = 2e-89
Identities = 175/363 (48%), Positives = 238/363 (65%), Gaps = 31/363 (8%)
Query: 1 MKGVISVFPSEEFHIQTTRSWDFLGLRQ--SIKRDQIIETDLVIGVIDIRIWPESESFND 58
M+GV+SVF S+ + +QTT SWDF+G+++ + KR+ +E+D +IG ID IWPESESF+D
Sbjct: 96 MEGVVSVFRSKNYKLQTTASWDFMGMKEGKNTKRNFAVESDTIIGFIDSGIWPESESFSD 155
Query: 59 KGLGPIPKKWRGVCAGGGNFSCNNKIIGARFYGDGDVSARYSFGHRTHTASTAGGREVED 118
KG GP PKKW+GVC GG NF+CNNK+IGAR Y R GH THT STA G V D
Sbjct: 156 KGFGPPPKKWKGVCKGGKNFTCNNKLIGARDYTSE--GTRDLQGHGTHTTSTAAGNAVAD 213
Query: 119 VSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGP 178
SF+G GTARGGVP+SR+AAYK+C CS D +L+AFDDAI+DGVD+I+VSLG
Sbjct: 214 TSFFGIGNGTARGGVPASRVAAYKVCTITG---CSDDNVLSAFDDAIADGVDLISVSLGG 270
Query: 179 EHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFID 238
++ S + D IAIG+FHAM KGILT +AGN GP P++V S APW+++VAAT+ +R+F+
Sbjct: 271 DYPSLYAEDTIAIGAFHAMAKGILTVHSAGNAGPNPTTVVSVAPWMLTVAATTTNRRFLT 330
Query: 239 KVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLAGGNASPEMCDCIDENMVKGKLVL 298
KV+LGNGKT VGKS+N GKK P+ E D ++E++VKGK+++
Sbjct: 331 KVVLGNGKTLVGKSVNAFDLKGKKYPL----------------EYGDYLNESLVKGKILV 374
Query: 299 CGSRNGKDLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTNSSKYPVA 358
Y +G+ ++ +T + ++ RP L +DF + SY NS++ P
Sbjct: 375 S--------RYLSGSEVAVSFITTDNKDYASISSRPLSVLSQDDFDSLVSYINSTRSPQG 426
Query: 359 DFL 361
L
Sbjct: 427 SVL 429
>UniRef100_Q9FIM5 Subtilisin-like serine protease [Arabidopsis thaliana]
Length = 713
Score = 330 bits (846), Expect = 4e-89
Identities = 177/361 (49%), Positives = 238/361 (65%), Gaps = 38/361 (10%)
Query: 1 MKGVISVFPSEEFHIQTTRSWDFLGLRQ--SIKRDQIIETDLVIGVIDIRIWPESESFND 58
M+GV+SVFP + +QTT SWDFLGL++ + KR+ IE+D +IG ID IWPESESF+D
Sbjct: 98 MEGVVSVFPDINYKLQTTASWDFLGLKEGKNTKRNLAIESDTIIGFIDSGIWPESESFSD 157
Query: 59 KGLGPIPKKWRGVCAGGGNFSCNNKIIGARFY---GDGDVSARYSFGHRTHTASTAGGRE 115
KG GP PKKW+GVC+ G NF+CNNK+IGAR Y G D+ GH THTASTA G
Sbjct: 158 KGFGPPPKKWKGVCSAGKNFTCNNKLIGARDYTNEGTRDIE-----GHGTHTASTAAGNA 212
Query: 116 VEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVS 175
V++ SFYG GTARGGVP+SRIAAYK C ++ C+ +++L+AFDDAI+DGVD+I++S
Sbjct: 213 VKNTSFYGIGNGTARGGVPASRIAAYKACSEMG---CTTESVLSAFDDAIADGVDLISIS 269
Query: 176 LGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQ 235
LG + DPIAIG+FHAM KGILT Q+AGN GP P SV S APW+++VAA++ +R
Sbjct: 270 LGANLVRTYETDPIAIGAFHAMVKGILTVQSAGNGGPNPGSVMSVAPWILTVAASNTNRG 329
Query: 236 FIDKVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLAGGNASPEMCDCIDENMVKGK 295
F+ KV+LGNGKTFVGKS+N GK P L GG+ D +++GK
Sbjct: 330 FVTKVVLGNGKTFVGKSLNAFDLKGKNYP---------LYGGST--------DGPLLRGK 372
Query: 296 LVLCGSRNGKDLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTNSSKY 355
+++ + ++ A N+ ++ ++V+ PS L +DF + SY NS+K
Sbjct: 373 ILVSEDKVSSEIVVA--------NINENYHDYAYVSILPSSALSKDDFDSVISYVNSTKS 424
Query: 356 P 356
P
Sbjct: 425 P 425
>UniRef100_Q9FIM7 Subtilisin-like serine protease [Arabidopsis thaliana]
Length = 677
Score = 320 bits (819), Expect = 6e-86
Identities = 174/363 (47%), Positives = 233/363 (63%), Gaps = 30/363 (8%)
Query: 1 MKGVISVFPSEEFHIQTTRSWDFLGLRQ--SIKRDQIIETDLVIGVIDIRIWPESESFND 58
M+GV+SVFP+ + +QTT SWDFLGL++ + KR+ IE+D +IG ID IWPESESF+D
Sbjct: 66 MEGVVSVFPNINYKLQTTASWDFLGLKEGKNTKRNLAIESDTIIGFIDSGIWPESESFSD 125
Query: 59 KGLGPIPKKWRGVCAGGGNFSCNNKIIGARFYGDGDVSARYSFGHRTHTASTAGGREVED 118
KG GP PKKW+GVC+GG NF+CNNK+IGAR Y R GH THTASTA G V D
Sbjct: 126 KGFGPPPKKWKGVCSGGKNFTCNNKLIGARDYTSE--GTRDLQGHGTHTASTAAGNAVAD 183
Query: 119 VSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGP 178
SF+G GTARGGVP+SRIAAYK+C + D C+ ++L+AFDDAI+DGVD+I++SL
Sbjct: 184 ASFFGIGNGTARGGVPASRIAAYKVCSEKD---CTAASLLSAFDDAIADGVDLISISLAS 240
Query: 179 EHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFID 238
E + D IAIG+FHA KGILT +AGN G PS+ S APW++SVAA++ +R F
Sbjct: 241 EFPQKYYKDAIAIGAFHANVKGILTVNSAGNSGSFPSTTASVAPWILSVAASNTNRGFFT 300
Query: 239 KVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLAGGNASPEMCDCIDENMVKGKLVL 298
KV+LGNGKT VG+S+N GKK P+ D +E++V+GK+++
Sbjct: 301 KVVLGNGKTLVGRSVNSFDLKGKKYPLVYG----------------DNFNESLVQGKILV 344
Query: 299 CGSRNGKDLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTNSSKYPVA 358
K + A+GSI + + ++ +P L +DF + SY NS++ P
Sbjct: 345 -----SKFPTSSKVAVGSI--LIDDYQHYALLSSKPFSLLPPDDFDSLVSYINSTRSPQG 397
Query: 359 DFL 361
FL
Sbjct: 398 TFL 400
>UniRef100_Q84WF9 Putative subtilisin-like serine protease [Arabidopsis thaliana]
Length = 708
Score = 310 bits (795), Expect = 3e-83
Identities = 172/365 (47%), Positives = 225/365 (61%), Gaps = 38/365 (10%)
Query: 1 MKGVISVFPSEEFHIQTTRSWDFLGLRQSIKRDQIIETDLVIGVIDIRIWPESESFNDKG 60
M+GV+SVFP++ +D +IGV D IWPESESF+DKG
Sbjct: 98 MEGVVSVFPNK--------------------------SDTIIGVFDGGIWPESESFSDKG 131
Query: 61 LGPIPKKWRGVCAGGGNFSCNNKIIGARFYGDGDVSARYSFGHRTHTASTAGGREVEDVS 120
GP PKKW+G+CAGG NF+CNNK+IGAR Y GD AR S GH THTAS A G V + S
Sbjct: 132 FGPPPKKWKGICAGGKNFTCNNKLIGARHYSPGD--ARDSTGHGTHTASIAAGNAVANTS 189
Query: 121 FYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEH 180
F+G GT RG VP+SRIA Y++C G C DAIL+AFDDAISDGVD+IT+S+G +
Sbjct: 190 FFGIGNGTVRGAVPASRIAVYRVCA----GECRDDAILSAFDDAISDGVDIITISIGDIN 245
Query: 181 ASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKV 240
F DPIAIG+FHAM KGILT AAGN GP +S+ S APWL++VAA++ +R+F+ KV
Sbjct: 246 VYPFEKDPIAIGAFHAMSKGILTVNAAGNTGPDTASITSLAPWLLTVAASTANREFVSKV 305
Query: 241 ILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLAGGNASPEMC--DCIDENMVKGKLVL 298
+LG+GKT VGKS+N GKK P+ + A E C +C+D ++VKGK+++
Sbjct: 306 VLGDGKTLVGKSVNGFDLKGKKFPLVYGKSAALSLSQAKCAEDCTPECLDASLVKGKILV 365
Query: 299 CGSRNGKDLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTNSSKYPVA 358
C +R +AY A+ +I + + + P L+ +DF + SY S K P A
Sbjct: 366 C-NRFLPYVAYTKRAVAAIF---EDGSDWAQINGLPVSGLQKDDFESVLSYFKSEKSPEA 421
Query: 359 DFLFS 363
L S
Sbjct: 422 AVLKS 426
>UniRef100_Q9FIM8 Subtilisin-like serine protease [Arabidopsis thaliana]
Length = 693
Score = 301 bits (770), Expect = 3e-80
Identities = 153/270 (56%), Positives = 197/270 (72%), Gaps = 15/270 (5%)
Query: 1 MKGVISVFPSEEFHIQTTRSWDFLGLRQ--SIKRDQIIETDLVIGVIDIRIWPESESFND 58
M+GV+SVFPS+++ + TT SWDF+GL++ + KR+ +E+D ++GV D I PESESF+
Sbjct: 98 MEGVVSVFPSKKYKLHTTASWDFMGLKEGKNTKRNLAVESDTIVGVFDTGISPESESFSG 157
Query: 59 KGLGPIPKKWRGVCAGGGNFSCNNKIIGARFY---GDGDVSARYSFGHRTHTASTAGGRE 115
KG GP PKKW+GVC GG NF+CNNK+IGAR Y G D+ GH THTASTA G
Sbjct: 158 KGFGPPPKKWKGVCKGGKNFTCNNKLIGARDYTNEGTRDIE-----GHGTHTASTAAGNV 212
Query: 116 VEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVS 175
VE+ SFYG GTARGGVP SRIAAYK+C CS + IL+AFDDAI+DGVDVI+ S
Sbjct: 213 VENTSFYGIGNGTARGGVPDSRIAAYKVC---SGAGCSSEYILSAFDDAIADGVDVISAS 269
Query: 176 LGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQ 235
LG + A + DPIAIG+FHAM KGILT Q+AGN GP P+ S APW+++VAA++ +R+
Sbjct: 270 LGGDTAYMYEKDPIAIGAFHAMAKGILTVQSAGNNGPNPT--VSVAPWILTVAASTTNRR 327
Query: 236 FIDKVILGNGKTFVGKSINITPSNGKKIPI 265
+ KV+LGNGKT VG+S+N GK+ P+
Sbjct: 328 IVTKVVLGNGKTLVGQSVNAFDLKGKQYPL 357
>UniRef100_O23357 CUCUMISIN [Arabidopsis thaliana]
Length = 687
Score = 299 bits (766), Expect = 8e-80
Identities = 175/366 (47%), Positives = 230/366 (62%), Gaps = 19/366 (5%)
Query: 1 MKGVISVFPSEEFHIQTTRSWDFLGLRQSIKRDQIIETDLVIGVIDIRIWPESESFNDKG 60
M+GV+SVFPS + + TTRS++F+GL +E+++++GVID IWPES+SF+D+G
Sbjct: 59 MEGVVSVFPSTVYKLFTTRSYEFMGLGDKSNNVPEVESNVIVGVIDGGIWPESKSFSDEG 118
Query: 61 LGPIPKKWRGVCAGGGNFSCNNKIIGARFYGDGDVSARYSFGHRTHTASTAGGREVEDVS 120
+GPIPKKW+G CAGG NF+CN K+IGAR Y SAR S H +HTASTA G +V+ VS
Sbjct: 119 IGPIPKKWKGTCAGGTNFTCNRKVIGARHYVHD--SARDSDAHGSHTASTAAGNKVKGVS 176
Query: 121 FYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEH 180
G A+GTARGGVP RIA YK+C + C+G+ ILAAFDDAI+DGVDV+T+SLG
Sbjct: 177 VNGVAEGTARGGVPLGRIAVYKVCEPLG---CNGERILAAFDDAIADGVDVLTISLGGGV 233
Query: 181 ASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKV 240
+ DPIAIGSFHAM KGI+TT A GN G + + APWL+SVAA S DR+F+ V
Sbjct: 234 TKVDI-DPIAIGSFHAMTKGIVTTVAVGNAGTALAKADNLAPWLISVAAGSTDRKFVTNV 292
Query: 241 ILGNGKTFVGKSINITPSNGKKIPIAV-----RNAQACLAGGNASPEMCDCIDENMVKGK 295
+ G+ K G+SIN GKK P+A N LA G AS C+ N V+GK
Sbjct: 293 VNGDDKMLPGRSINDFDLEGKKYPLAYGKTASNNCTEELARGCAS----GCL--NTVEGK 346
Query: 296 LVLCGSRNGKDLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTNSSKY 355
+V+C N A GA+G+I +VT + + P L ++ ++SY SS
Sbjct: 347 IVVCDVPNNVMEQKAAGAVGTILHVT--DVDTPGLGPIAVATLDDTNYEELRSYVLSSPN 404
Query: 356 PVADFL 361
P L
Sbjct: 405 PQGTIL 410
>UniRef100_Q94KL9 Subtilisin-like protein [Glycine max]
Length = 766
Score = 295 bits (754), Expect = 2e-78
Identities = 162/382 (42%), Positives = 229/382 (59%), Gaps = 26/382 (6%)
Query: 3 GVISVFPSEEFHIQTTRSWDFLGLRQSIKRDQ----IIETDLVIGVIDIRIWPESESFND 58
GV+SVFP + TTRSWDFL + +K D + ++ VIG++D IWPE+ SF+D
Sbjct: 102 GVVSVFPGPVLKLHTTRSWDFLKYQTQVKIDTKPNAVSKSSSVIGILDTGIWPEAASFSD 161
Query: 59 KGLGPIPKKWRGVCAGGGNF---SCNNKIIGARFYGD----GDVSARYSFGHRTHTASTA 111
KG+GP+P +W+G C +F +CN K+IGAR+Y D GD +AR S GH TH A TA
Sbjct: 162 KGMGPVPSRWKGTCMKSQDFYSSNCNRKLIGARYYADPNDSGDNTARDSNGHGTHVAGTA 221
Query: 112 GGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDV 171
G V + S+YG A G A+GG P SR+A Y++C N C G +ILAAFDDAI+DGVD+
Sbjct: 222 AGVMVTNASYYGVATGCAKGGSPESRLAVYRVCS---NFGCRGSSILAAFDDAIADGVDL 278
Query: 172 ITVSLGPEHA--SDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAA 229
++VSLG D +DPI++G+FHAME GIL +AGN GP ++ + APW+++VAA
Sbjct: 279 LSVSLGASTGFRPDLTSDPISLGAFHAMEHGILVVCSAGNDGPSSYTLVNDAPWILTVAA 338
Query: 230 TSIDRQFIDKVILGNGKTFVGKSINITP-SNGKKIPIAVRNAQACLAGGNASPEMC--DC 286
++IDR F+ ++LG+ K GK+IN++P SN K P+ + + C +
Sbjct: 339 STIDRNFLSNIVLGDNKIIKGKAINLSPLSNSPKYPLIYGESAKANSTSLVEARQCHPNS 398
Query: 287 IDENMVKGKLVLCGSRNGK-------DLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLK 339
+D N VKGK+V+C +N K A G IG +H +++ AS P+ +
Sbjct: 399 LDGNKVKGKIVVCDDKNDKYSTRKKVATVKAVGGIGLVHITDQNEAIASNYGDFPATVIS 458
Query: 340 TNDFVHIQSYTNSSKYPVADFL 361
+ D V I Y NS+ PVA L
Sbjct: 459 SKDGVTILQYINSTSNPVATIL 480
>UniRef100_Q93WQ0 Subtilisin-type protease [Glycine max]
Length = 766
Score = 295 bits (754), Expect = 2e-78
Identities = 162/382 (42%), Positives = 229/382 (59%), Gaps = 26/382 (6%)
Query: 3 GVISVFPSEEFHIQTTRSWDFLGLRQSIKRDQ----IIETDLVIGVIDIRIWPESESFND 58
GV+SVFP + TTRSWDFL + +K D + ++ VIG++D IWPE+ SF+D
Sbjct: 102 GVVSVFPGPVLKLHTTRSWDFLKYQTQVKIDTKPNAVSKSSSVIGILDTGIWPEAASFSD 161
Query: 59 KGLGPIPKKWRGVCAGGGNF---SCNNKIIGARFYGD----GDVSARYSFGHRTHTASTA 111
KG+GP+P +W+G C +F +CN K+IGAR+Y D GD +AR S GH TH A TA
Sbjct: 162 KGMGPVPSRWKGTCMKSQDFYSSNCNRKLIGARYYADPNDSGDNTARDSNGHGTHVAGTA 221
Query: 112 GGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDV 171
G V + S+YG A G A+GG P SR+A Y++C N C G +ILAAFDDAI+DGVD+
Sbjct: 222 AGVMVTNASYYGVATGCAKGGSPESRLAVYRVCS---NFGCRGSSILAAFDDAIADGVDL 278
Query: 172 ITVSLGPEHA--SDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAA 229
++VSLG D +DPI++G+FHAME GIL +AGN GP ++ + APW+++VAA
Sbjct: 279 LSVSLGASTGFRPDLTSDPISLGAFHAMEHGILVVCSAGNDGPSSYTLVNDAPWILTVAA 338
Query: 230 TSIDRQFIDKVILGNGKTFVGKSINITP-SNGKKIPIAVRNAQACLAGGNASPEMC--DC 286
++IDR F+ ++LG+ K GK+IN++P SN K P+ + + C +
Sbjct: 339 STIDRNFLSNIVLGDNKIIKGKAINLSPLSNSPKYPLIYGESAKANSTSLVEARQCRPNS 398
Query: 287 IDENMVKGKLVLCGSRNGK-------DLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLK 339
+D N VKGK+V+C +N K A G IG +H +++ AS P+ +
Sbjct: 399 LDGNKVKGKIVVCDDKNDKYSTRKKVATVKAVGGIGLVHITDQNEAIASNYGDFPATVIS 458
Query: 340 TNDFVHIQSYTNSSKYPVADFL 361
+ D V I Y NS+ PVA L
Sbjct: 459 SKDGVTILQYINSTSNPVATIL 480
>UniRef100_Q9ZTT3 Subtilisin-like protease C1 [Glycine max]
Length = 738
Score = 285 bits (728), Expect = 2e-75
Identities = 165/367 (44%), Positives = 216/367 (57%), Gaps = 18/367 (4%)
Query: 4 VISVFPSEEFHIQTTRSWDFLGLRQSIKRDQIIETDLVIGVIDIRIWPESESFNDKGLGP 63
V++VFP+++ + TTRSWDF+G R E+D++I V D IWPESESFNDKG GP
Sbjct: 98 VVAVFPNKKKQLHTTRSWDFIGFPLQANRAPA-ESDVIIAVFDSGIWPESESFNDKGFGP 156
Query: 64 IPKKWRGVCAGGGNFSCNNKIIGARFYG-------DGDVSARYSFGHRTHTASTAGGREV 116
P KW+G C NF+CNNKIIGA+ Y D S R GH TH ASTA G V
Sbjct: 157 PPSKWKGTCQTSKNFTCNNKIIGAKIYKVDGFFSKDDPKSVRDIDGHGTHVASTAAGNPV 216
Query: 117 EDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSL 176
S G +GT+RGGV +RIA YK+C D C+ ILAAFDDAI+DGVD+ITVSL
Sbjct: 217 STASMLGLGQGTSRGGVTKARIAVYKVCW-FDG--CTDADILAAFDDAIADGVDIITVSL 273
Query: 177 GPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQF 236
G ++ D IAIG+FHA+ G+LT +AGN GP PSS+ + +PW +SVAA++IDR+F
Sbjct: 274 GGFSDENYFRDGIAIGAFHAVRNGVLTVTSAGNSGPRPSSLSNFSPWSISVAASTIDRKF 333
Query: 237 IDKVILGNGKTFVGKSINITPSNGKKIPIA----VRNAQACLAGGNASPEMCDCIDENMV 292
+ KV LGN T+ G SIN G+ PI N + G ++ +D+ +V
Sbjct: 334 VTKVELGNKITYEGTSINTFDLKGELYPIIYGGDAPNKGEGIDGSSSRYCSSGSLDKKLV 393
Query: 293 KGKLVLCGSRNGKDLAYANGAIGS-IHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTN 351
KGK+VLC SR+ + GA+G+ I L S P P L D + Y N
Sbjct: 394 KGKIVLCESRSKALGPFDAGAVGALIQGQGFRDLPPSL--PLPGSYLALQDGASVYDYIN 451
Query: 352 SSKYPVA 358
S++ P+A
Sbjct: 452 STRTPIA 458
>UniRef100_Q8LAE1 Subtilisin-like serine protease [Arabidopsis thaliana]
Length = 769
Score = 281 bits (720), Expect = 2e-74
Identities = 166/390 (42%), Positives = 232/390 (58%), Gaps = 38/390 (9%)
Query: 3 GVISVFPSEEFHIQTTRSWDFLGLRQSIKRDQIIET-------DLVIGVIDIRIWPESES 55
GV+SVFP F + TT SWDFL + S+K D + D ++G++D IWPESES
Sbjct: 95 GVVSVFPDPHFQLHTTHSWDFLKYQTSVKVDSGPPSSASDGXYDSIVGILDTGIWPESES 154
Query: 56 FNDKGLGPIPKKWRGVCAGGGNF---SCNNKIIGARFYGDGDVSARYS-----FGHRTHT 107
FNDK +GPIP +W+G C +F +CN KIIGAR+Y + D + Y GH +H
Sbjct: 155 FNDKDMGPIPSRWKGTCMEAKDFKSSNCNRKIIGARYYKNPDDDSEYYTTRDVIGHGSHV 214
Query: 108 ASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISD 167
+ST G VE+ S+YG A GTA+GG ++RIA YK+C G C+G +ILAAFDDAI+D
Sbjct: 215 SSTIAGSAVENASYYGVASGTAKGGSQNARIAMYKVCNP---GGCTGSSILAAFDDAIAD 271
Query: 168 GVDVITVSLG-PEHASDFLN-DPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLV 225
GVDV+++SLG P +A LN DPIAIG+FHA+E+GIL +AGN GP +V + APW++
Sbjct: 272 GVDVLSLSLGAPAYARIDLNTDPIAIGAFHAVEQGILVICSAGNDGPDGGTVTNTAPWIM 331
Query: 226 SVAATSIDRQFIDKVILGNGKTFVGKSINITPSNGKKIPI-------AVRNAQACLAGGN 278
+VAA +IDR F V+LG K G+ I+ SN K P+ + ++A A + G+
Sbjct: 332 TVAANTIDRDFESDVVLGGNKVIKGEGIHF--SNVSKSPVYPLIHGKSAKSADA--SEGS 387
Query: 279 ASPEMCDCIDENMVKGKLVLCGSRNG-------KDLAYANGAIGSIHNVTKSQLGASFVT 331
A D +D+ VKGK+VLC + G +D + G G + +++ AS
Sbjct: 388 ARACDSDSLDQEKVKGKIVLCENVGGSYYASSARDKVKSKGGTGCVFVDDRTRAVASAYG 447
Query: 332 PRPSLNLKTNDFVHIQSYTNSSKYPVADFL 361
P+ + + + I SY NS+K PVA L
Sbjct: 448 SFPTTVIDSKEAAEIFSYLNSTKDPVATIL 477
>UniRef100_Q9LNU1 T20H2.6 protein [Arabidopsis thaliana]
Length = 769
Score = 281 bits (719), Expect = 2e-74
Identities = 166/390 (42%), Positives = 232/390 (58%), Gaps = 38/390 (9%)
Query: 3 GVISVFPSEEFHIQTTRSWDFLGLRQSIKRDQIIET-------DLVIGVIDIRIWPESES 55
GV+SVFP F + TT SWDFL + S+K D + D ++G++D IWPESES
Sbjct: 95 GVVSVFPDPHFQLHTTHSWDFLKYQTSVKVDSGPPSSASDGSYDSIVGILDTGIWPESES 154
Query: 56 FNDKGLGPIPKKWRGVCAGGGNF---SCNNKIIGARFYGDGDVSARYS-----FGHRTHT 107
FNDK +GPIP +W+G C +F +CN KIIGAR+Y + D + Y GH +H
Sbjct: 155 FNDKDMGPIPSRWKGTCMEAKDFKSSNCNRKIIGARYYKNPDDDSEYYTTRDVIGHGSHV 214
Query: 108 ASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISD 167
+ST G VE+ S+YG A GTA+GG ++RIA YK+C G C+G +ILAAFDDAI+D
Sbjct: 215 SSTIAGSAVENASYYGVASGTAKGGSQNARIAMYKVCNP---GGCTGSSILAAFDDAIAD 271
Query: 168 GVDVITVSLG-PEHASDFLN-DPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLV 225
GVDV+++SLG P +A LN DPIAIG+FHA+E+GIL +AGN GP +V + APW++
Sbjct: 272 GVDVLSLSLGAPAYARIDLNTDPIAIGAFHAVEQGILVICSAGNDGPDGGTVTNTAPWIM 331
Query: 226 SVAATSIDRQFIDKVILGNGKTFVGKSINITPSNGKKIPI-------AVRNAQACLAGGN 278
+VAA +IDR F V+LG K G+ I+ SN K P+ + ++A A + G+
Sbjct: 332 TVAANTIDRDFESDVVLGGNKVIKGEGIHF--SNVSKSPVYPLIHGKSAKSADA--SEGS 387
Query: 279 ASPEMCDCIDENMVKGKLVLCGSRNG-------KDLAYANGAIGSIHNVTKSQLGASFVT 331
A D +D+ VKGK+VLC + G +D + G G + +++ AS
Sbjct: 388 ARACDSDSLDQEKVKGKIVLCENVGGSYYASSARDEVKSKGGTGCVFVDDRTRAVASAYG 447
Query: 332 PRPSLNLKTNDFVHIQSYTNSSKYPVADFL 361
P+ + + + I SY NS+K PVA L
Sbjct: 448 SFPTTVIDSKEAAEIFSYLNSTKDPVATIL 477
>UniRef100_Q39547 Pre-pro-cucumisin precursor [Cucumis melo]
Length = 731
Score = 278 bits (710), Expect = 2e-73
Identities = 160/369 (43%), Positives = 222/369 (59%), Gaps = 18/369 (4%)
Query: 1 MKGVISVFPSEEFHIQTTRSWDFLGLRQSIKRDQIIETDLVIGVIDIRIWPESESFNDKG 60
M+GV+SVF +E + TTRSWDFLG ++ R +E+++V+GV+D IWPES SF+D+G
Sbjct: 95 MEGVVSVFLNEMNELHTTRSWDFLGFPLTVPRRSQVESNIVVGVLDTGIWPESPSFDDEG 154
Query: 61 LGPIPKKWRGVCAGGGNFSCNNKIIGARFY------GDGDVSA-RYSFGHRTHTASTAGG 113
P P KW+G C NF CN KIIGAR Y GDV+ R + GH THTASTA G
Sbjct: 155 FSPPPPKWKGTCETSNNFRCNRKIIGARSYHIGRPISPGDVNGPRDTNGHGTHTASTAAG 214
Query: 114 REVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVIT 173
V + YG GTARGGVP +RIAAYK+C N CS ILAA+DDAI+DGVD+I+
Sbjct: 215 GLVSQANLYGLGLGTARGGVPLARIAAYKVCW---NDGCSDTDILAAYDDAIADGVDIIS 271
Query: 174 VSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSID 233
+S+G + + D IAIGSFHA+E+GILT+ +AGN GP + S +PWL+SVAA+++D
Sbjct: 272 LSVGGANPRHYFVDAIAIGSFHAVERGILTSNSAGNGGPNFFTTASLSPWLLSVAASTMD 331
Query: 234 RQFIDKVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLAGGNASPEMC--DCIDENM 291
R+F+ +V +GNG++F G SIN + + P+ ++ C ++ N+
Sbjct: 332 RKFVTQVQIGNGQSFQGVSIN--TFDNQYYPLVSGRDIPNTGFDKSTSRFCTDKSVNPNL 389
Query: 292 VKGKLVLCGSRNGKDLAY--ANGAIGSIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSY 349
+KGK+V+C + G + +GA G + S+ P PS L ND + Y
Sbjct: 390 LKGKIVVCEASFGPHEFFKSLDGAAGVLMTSNTRDYADSY--PLPSSVLDPNDLLATLRY 447
Query: 350 TNSSKYPVA 358
S + P A
Sbjct: 448 IYSIRSPGA 456
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.137 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 704,573,127
Number of Sequences: 2790947
Number of extensions: 31780927
Number of successful extensions: 61245
Number of sequences better than 10.0: 449
Number of HSP's better than 10.0 without gapping: 193
Number of HSP's successfully gapped in prelim test: 256
Number of HSP's that attempted gapping in prelim test: 60122
Number of HSP's gapped (non-prelim): 489
length of query: 395
length of database: 848,049,833
effective HSP length: 129
effective length of query: 266
effective length of database: 488,017,670
effective search space: 129812700220
effective search space used: 129812700220
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 76 (33.9 bits)
Medicago: description of AC148652.13


