

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148607.14 - phase: 0
(359 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
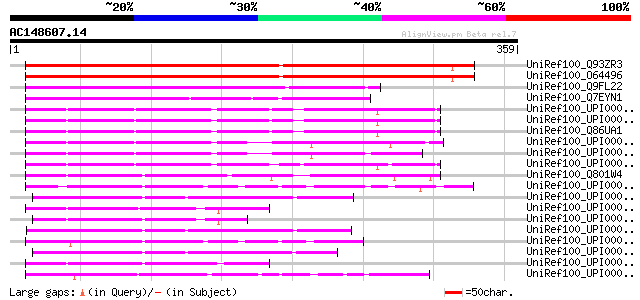
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q93ZR3 Hypothetical protein At1g04080 [Arabidopsis tha... 424 e-117
UniRef100_O64496 F20D22.14 protein [Arabidopsis thaliana] 424 e-117
UniRef100_Q9FL22 Arabidopsis thaliana genomic DNA, chromosome 5,... 175 2e-42
UniRef100_Q7EYN1 Hypothetical protein OSJNBb0011E04.121 [Oryza s... 165 2e-39
UniRef100_UPI00001BBB26 UPI00001BBB26 UniRef100 entry 97 7e-19
UniRef100_UPI0000369C87 UPI0000369C87 UniRef100 entry 97 7e-19
UniRef100_Q86UA1 PRPF39 protein [Homo sapiens] 97 7e-19
UniRef100_UPI00003AE104 UPI00003AE104 UniRef100 entry 94 4e-18
UniRef100_UPI00003AE105 UPI00003AE105 UniRef100 entry 94 8e-18
UniRef100_UPI00001CF9F1 UPI00001CF9F1 UniRef100 entry 92 3e-17
UniRef100_Q801W4 Novel protein similar to pre-mRNA processing pr... 90 8e-17
UniRef100_UPI0000431839 UPI0000431839 UniRef100 entry 88 3e-16
UniRef100_UPI000021A861 UPI000021A861 UniRef100 entry 88 4e-16
UniRef100_UPI000036473B UPI000036473B UniRef100 entry 86 1e-15
UniRef100_UPI00002BB17A UPI00002BB17A UniRef100 entry 85 3e-15
UniRef100_UPI000023E0E1 UPI000023E0E1 UniRef100 entry 84 6e-15
UniRef100_UPI0000431838 UPI0000431838 UniRef100 entry 82 3e-14
UniRef100_UPI0000234C5F UPI0000234C5F UniRef100 entry 80 7e-14
UniRef100_UPI000035FA82 UPI000035FA82 UniRef100 entry 78 3e-13
UniRef100_UPI0000007649 UPI0000007649 UniRef100 entry 75 2e-12
>UniRef100_Q93ZR3 Hypothetical protein At1g04080 [Arabidopsis thaliana]
Length = 768
Score = 424 bits (1091), Expect = e-117
Identities = 212/321 (66%), Positives = 254/321 (79%), Gaps = 5/321 (1%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
+++VKLYERCV+ CANYPEYWIRYV MEAS S DLA N LARA+QVFVK+QPEIHLF A
Sbjct: 378 NKVVKLYERCVVTCANYPEYWIRYVTNMEASGSADLAENALARATQVFVKKQPEIHLFAA 437
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKG 131
R KEQ GDI GARAAYQLVH+EISPGLLEA+I+HANME+RLG L+DAFSLYEQ IA+EKG
Sbjct: 438 RLKEQNGDIAGARAAYQLVHSEISPGLLEAVIKHANMEYRLGNLDDAFSLYEQVIAVEKG 497
Query: 132 KEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPK 191
KEHS LP+L+AQYSRF YL S ++EKAR I+V L++ SKPL+EAL+HFEAIQP P+
Sbjct: 498 KEHSTILPLLYAQYSRFSYLVSRDAEKARRIIVEALDHVQPSKPLMEALIHFEAIQPPPR 557
Query: 192 RVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFGDVQSIKRAEDRHAKL 251
+ID+LE LV K I P+ + +AS+TEREELS I++EFL +FGDV+SIK+AED+H KL
Sbjct: 558 --EIDYLEPLVEKVIKPDADAQNIASSTEREELSLIYIEFLGIFGDVKSIKKAEDQHVKL 615
Query: 252 FLPNRGLSELKKRHAEDFLASDKTKVSRAYSAQSPAQSVAGAYPNGPNQWP-NYGVQPQT 310
F P+R SELKKR A+DFLASD+TK+++ Y+ PAQ V+ AYPN QW Y QPQT
Sbjct: 616 FYPHRSTSELKKRSADDFLASDRTKMAKTYNGTPPAQPVSNAYPNAQAQWSGGYAAQPQT 675
Query: 311 WP--ATTQAQGQQWPAGYTQQ 329
WP AQ QQW Y QQ
Sbjct: 676 WPPAQAAPAQPQQWNPAYGQQ 696
>UniRef100_O64496 F20D22.14 protein [Arabidopsis thaliana]
Length = 1345
Score = 424 bits (1091), Expect = e-117
Identities = 212/321 (66%), Positives = 254/321 (79%), Gaps = 5/321 (1%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
+++VKLYERCV+ CANYPEYWIRYV MEAS S DLA N LARA+QVFVK+QPEIHLF A
Sbjct: 378 NKVVKLYERCVVTCANYPEYWIRYVTNMEASGSADLAENALARATQVFVKKQPEIHLFAA 437
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKG 131
R KEQ GDI GARAAYQLVH+EISPGLLEA+I+HANME+RLG L+DAFSLYEQ IA+EKG
Sbjct: 438 RLKEQNGDIAGARAAYQLVHSEISPGLLEAVIKHANMEYRLGNLDDAFSLYEQVIAVEKG 497
Query: 132 KEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPK 191
KEHS LP+L+AQYSRF YL S ++EKAR I+V L++ SKPL+EAL+HFEAIQP P+
Sbjct: 498 KEHSTILPLLYAQYSRFSYLVSRDAEKARRIIVEALDHVQPSKPLMEALIHFEAIQPPPR 557
Query: 192 RVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFGDVQSIKRAEDRHAKL 251
+ID+LE LV K I P+ + +AS+TEREELS I++EFL +FGDV+SIK+AED+H KL
Sbjct: 558 --EIDYLEPLVEKVIKPDADAQNIASSTEREELSLIYIEFLGIFGDVKSIKKAEDQHVKL 615
Query: 252 FLPNRGLSELKKRHAEDFLASDKTKVSRAYSAQSPAQSVAGAYPNGPNQWP-NYGVQPQT 310
F P+R SELKKR A+DFLASD+TK+++ Y+ PAQ V+ AYPN QW Y QPQT
Sbjct: 616 FYPHRSTSELKKRSADDFLASDRTKMAKTYNGTPPAQPVSNAYPNAQAQWSGGYAAQPQT 675
Query: 311 WP--ATTQAQGQQWPAGYTQQ 329
WP AQ QQW Y QQ
Sbjct: 676 WPPAQAAPAQPQQWNPAYGQQ 696
>UniRef100_Q9FL22 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MPL12
[Arabidopsis thaliana]
Length = 1022
Score = 175 bits (443), Expect = 2e-42
Identities = 98/252 (38%), Positives = 150/252 (58%), Gaps = 4/252 (1%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
D + LYERC+I CANY E+W RYV +E+ +LAN LARASQ FVK IHLF A
Sbjct: 315 DWAINLYERCLIPCANYTEFWFRYVDFVESKGGRELANFALARASQTFVKSASVIHLFNA 374
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAI-AIEK 130
RFKE GD A A E+ G +E + + ANME RLG E A + Y +A+
Sbjct: 375 RFKEHVGDASAASVALSRCGEELGFGFVENVTKKANMEKRLGNFEAAVTTYREALNKTLI 434
Query: 131 GKEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQP 190
GKE+ +T L+ Q+SR Y+ + +++ A +IL+ G EN K LLE L+ +
Sbjct: 435 GKENLETTARLYVQFSRLKYVITNSADDAAQILLEGNENVPHCKLLLEELMRLLMMHGGS 494
Query: 191 KRVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFGDVQSIKRAEDRHAK 250
++VD+ L+ ++ K ++ ++ SA ++EE+SN+++EF++L G + +++A RH K
Sbjct: 495 RQVDL--LDPIIDKELSHQADSSDGLSAEDKEEISNLYMEFIDLSGTIHDVRKALGRHIK 552
Query: 251 LFLPNRGLSELK 262
LF P+ ++L+
Sbjct: 553 LF-PHSARAKLR 563
>UniRef100_Q7EYN1 Hypothetical protein OSJNBb0011E04.121 [Oryza sativa]
Length = 1161
Score = 165 bits (418), Expect = 2e-39
Identities = 91/244 (37%), Positives = 144/244 (58%), Gaps = 4/244 (1%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
D VKLYERC+I CANY E+WIRY ++A ++A+ L RAS FVK P H++ A
Sbjct: 278 DWAVKLYERCLIPCANYSEFWIRYAEFVDAKGGREIASYALGRASSYFVKGVPTFHMYYA 337
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKG 131
FKEQ GD GAR+ + ++ I R ANME R+G + A +YE AI +
Sbjct: 338 MFKEQIGDAQGARSLFIEGSNNLTSNFCANINRLANMEKRMGNTKAASEIYETAIQ-DAM 396
Query: 132 KEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPK 191
+++ + LP L+ +++F Y + N +A+E+ V G++ A K L++ + F + P
Sbjct: 397 QKNVKILPDLYTNFAQFKYAVNHNISEAKEVFVEGIKQAP-CKALIKGFMQFMSTHGGP- 454
Query: 192 RVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFGDVQSIKRAEDRHAKL 251
+I L+S++ + P + V S +RE++S +FLEF++L+GDV+ +++A RH+KL
Sbjct: 455 -TEIPILDSVISNAVVPGSDISTVLSREDREDISLLFLEFVDLYGDVRDLRKAWARHSKL 513
Query: 252 FLPN 255
F N
Sbjct: 514 FPHN 517
>UniRef100_UPI00001BBB26 UPI00001BBB26 UniRef100 entry
Length = 548
Score = 97.1 bits (240), Expect = 7e-19
Identities = 88/306 (28%), Positives = 133/306 (42%), Gaps = 26/306 (8%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
+++V L+ERCVI+CA Y E+WI+Y ME + S++ +V +RA + + ++P +H+ A
Sbjct: 249 ERVVVLFERCVISCALYEEFWIKYAKYME-NHSIEGVRHVFSRACTIHLPKKPMVHMLWA 307
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKG 131
F+EQ G+I AR + E GL +R ++E R G LE+A L + AI K
Sbjct: 308 AFEEQQGNINEARNILK-TFEECVLGLAMVRLRRVSLERRHGNLEEAEHLLQDAIKNAKS 366
Query: 132 KEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPK 191
S + A R ++ N K+R++L+ +E + L LL E K
Sbjct: 367 NNESSFYAVKLA---RHLFKIQKNLPKSRKVLLEAIERDKENTKLYLNLLEME-YSGDLK 422
Query: 192 RVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFG-DVQSIKRAEDRHAK 250
+ + + L + G R S +EFL FG DV + A D H
Sbjct: 423 QNEENILNCF-------DKAVHGSLPIKMRITFSQRKVEFLEDFGSDVNKLLNAYDEHQT 475
Query: 251 LFLPNRGLS----------ELKKRHAEDFLASDKTKVSRAYSAQSPAQSVAGAYP-NGPN 299
L L E KK H ED +S + A + + Y N N
Sbjct: 476 LLKEQDSLKRKAENGSEEPEEKKAHTEDTTSSSTQMIDGDLQANQAVYNYSAWYQYNYQN 535
Query: 300 QWPNYG 305
W NYG
Sbjct: 536 PW-NYG 540
>UniRef100_UPI0000369C87 UPI0000369C87 UniRef100 entry
Length = 548
Score = 97.1 bits (240), Expect = 7e-19
Identities = 88/306 (28%), Positives = 133/306 (42%), Gaps = 26/306 (8%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
+++V L+ERCVI+CA Y E+WI+Y ME + S++ +V +RA + + ++P +H+ A
Sbjct: 249 ERVVVLFERCVISCALYEEFWIKYAKYME-NHSIEGVRHVFSRACTIHLPKKPMVHMLWA 307
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKG 131
F+EQ G+I AR + E GL +R ++E R G LE+A L + AI K
Sbjct: 308 AFEEQQGNINEARNILR-TFEECVLGLAMVRLRRVSLERRHGNLEEAEHLLQDAIKNAKS 366
Query: 132 KEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPK 191
S + A R ++ N K+R++L+ +E + L LL E K
Sbjct: 367 NNESSFYAVKLA---RHLFKIQKNLPKSRKVLLEAIERDKENTKLYLNLLEME-YSGDLK 422
Query: 192 RVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFG-DVQSIKRAEDRHAK 250
+ + + L + G R S +EFL FG DV + A D H
Sbjct: 423 QNEENILNCF-------DKAVHGSLPIKMRITFSQRKVEFLEDFGSDVNKLLNAYDEHQT 475
Query: 251 LFLPNRGLS----------ELKKRHAEDFLASDKTKVSRAYSAQSPAQSVAGAYP-NGPN 299
L L E KK H ED +S + A + + Y N N
Sbjct: 476 LLKEQDSLKRKAENGSEEPEEKKAHTEDTTSSSTQMIDGDLQANQAVYNYSAWYQYNYQN 535
Query: 300 QWPNYG 305
W NYG
Sbjct: 536 PW-NYG 540
>UniRef100_Q86UA1 PRPF39 protein [Homo sapiens]
Length = 479
Score = 97.1 bits (240), Expect = 7e-19
Identities = 88/306 (28%), Positives = 133/306 (42%), Gaps = 26/306 (8%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
+++V L+ERCVI+CA Y E+WI+Y ME + S++ +V +RA + + ++P +H+ A
Sbjct: 180 ERVVVLFERCVISCALYEEFWIKYAKYME-NHSIEGVRHVFSRACTIHLPKKPMVHMLWA 238
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKG 131
F+EQ G+I AR + E GL +R ++E R G LE+A L + AI K
Sbjct: 239 AFEEQQGNINEARNILK-TFEECVLGLAMVRLRRVSLERRHGNLEEAEHLLQDAIKNAKS 297
Query: 132 KEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPK 191
S + A R ++ N K+R++L+ +E + L LL E K
Sbjct: 298 NNESSFYAVKLA---RHLFKIQKNLPKSRKVLLEAIERDKENTKLYLNLLEME-YSGDLK 353
Query: 192 RVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFG-DVQSIKRAEDRHAK 250
+ + + L + G R S +EFL FG DV + A D H
Sbjct: 354 QNEENILNCF-------DKAVHGSLPIKMRITFSQRKVEFLEDFGSDVNKLLNAYDEHQT 406
Query: 251 LFLPNRGLS----------ELKKRHAEDFLASDKTKVSRAYSAQSPAQSVAGAYP-NGPN 299
L L E KK H ED +S + A + + Y N N
Sbjct: 407 LLKEQDSLKRKAENGSEEPEEKKAHTEDTTSSSTQMIDGDLQANQAVYNYSAWYQYNYQN 466
Query: 300 QWPNYG 305
W NYG
Sbjct: 467 PW-NYG 471
>UniRef100_UPI00003AE104 UPI00003AE104 UniRef100 entry
Length = 475
Score = 94.4 bits (233), Expect = 4e-18
Identities = 84/310 (27%), Positives = 135/310 (43%), Gaps = 38/310 (12%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
+++V L+ERCVI+CA Y ++WI+Y ME + S++ +V +RA + + ++P +H+ A
Sbjct: 179 ERVVVLFERCVISCALYEDFWIKYAKYME-NHSIEGVRHVYSRACTIHLPKKPMVHMLWA 237
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKG 131
F+EQ G+I AR + E GL +R ++E R G +E+A L E+A+ K
Sbjct: 238 AFEEQQGNIDEARRILK-TFEECILGLAMVRLRRVSLERRHGNMEEAERLLEEAVRNAKS 296
Query: 132 KEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPK 191
S + A R ++ N KAR++L +E I +
Sbjct: 297 VSESSFYAIKLA---RHLFKVQKNLPKARKVLSDAIE-----------------IDKENT 336
Query: 192 RVDIDFLESLVVKFITPNPEN---------PGVASATEREELSNIFLEFLNLFG-DVQSI 241
++ ++ LE +T N EN G S R S +EFL FG DV +
Sbjct: 337 KLYLNLLEMEYCGDLTQNEENILSCFDKAVNGSLSIKMRVTFSQRKVEFLEDFGSDVNKL 396
Query: 242 KRAEDRHAKLFLPNRGLSELKKRHAED----FLASDKTKVSRAYSAQSPAQSVAGAYPNG 297
A D H L L + +E+ + +D+ ++ A Q AY
Sbjct: 397 LDAYDEHQALLKEQDTLKRRAENGSEEPDEKKMLTDEQTMASAQMMDGDMQVNQAAY--N 454
Query: 298 PNQWPNYGVQ 307
N W Y Q
Sbjct: 455 YNAWYQYNYQ 464
>UniRef100_UPI00003AE105 UPI00003AE105 UniRef100 entry
Length = 578
Score = 93.6 bits (231), Expect = 8e-18
Identities = 81/291 (27%), Positives = 130/291 (43%), Gaps = 35/291 (12%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
+++V L+ERCVI+CA Y ++WI+Y ME + S++ +V +RA + + ++P +H+ A
Sbjct: 297 ERVVVLFERCVISCALYEDFWIKYAKYME-NHSIEGVRHVYSRACTIHLPKKPMVHMLWA 355
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKG 131
F+EQ G+I AR + E GL +R ++E R G +E+A L E+A+ K
Sbjct: 356 AFEEQQGNIDEARRILK-TFEECILGLAMVRLRRVSLERRHGNMEEAERLLEEAVRNAKS 414
Query: 132 KEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPK 191
S + A R ++ N KAR++L +E I +
Sbjct: 415 VSESSFYAIKLA---RHLFKVQKNLPKARKVLSDAIE-----------------IDKENT 454
Query: 192 RVDIDFLESLVVKFITPNPEN---------PGVASATEREELSNIFLEFLNLFG-DVQSI 241
++ ++ LE +T N EN G S R S +EFL FG DV +
Sbjct: 455 KLYLNLLEMEYCGDLTQNEENILSCFDKAVNGSLSIKMRVTFSQRKVEFLEDFGSDVNKL 514
Query: 242 KRAEDRHAKLFLPNRGLSELKKRHAEDFLASDKTKVSRAYSAQSPAQSVAG 292
A D H L + LK+R D+ K+ + AQ + G
Sbjct: 515 LDAYDEHQALL---KEQDTLKRRAENGSEEPDEKKMLTDEQTMASAQMMDG 562
>UniRef100_UPI00001CF9F1 UPI00001CF9F1 UniRef100 entry
Length = 480
Score = 91.7 bits (226), Expect = 3e-17
Identities = 89/306 (29%), Positives = 132/306 (43%), Gaps = 28/306 (9%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
+++V L+ERCVI+CA Y E+WI+Y ME + S++ +V +RA V + ++P H+ A
Sbjct: 183 ERVVVLFERCVISCALYEEFWIKYAKYME-NHSIEGVRHVFSRACTVHLPKKPMAHMLWA 241
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKG 131
F+EQ G+I AR + E GL +R ++E R G +E+A L + AI K
Sbjct: 242 AFEEQQGNINEARIILR-TFEECVLGLAMVRLRRVSLERRHGNMEEAEHLLQDAIRNAKS 300
Query: 132 KEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPK 191
S + A R ++ N K+R++L+ +E + L LL E
Sbjct: 301 NNESSFYAIKLA---RHLFKIQKNLPKSRKVLLEAIEKDKENTKLYLNLLEME------Y 351
Query: 192 RVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFG-DVQSIKRAEDRHAK 250
D+ E ++ + G R S +EFL FG DV + A D H
Sbjct: 352 SCDLKQNEENILNCF--DKAIHGSLPIKMRITFSQRKVEFLEDFGSDVNKLLNAYDEHQT 409
Query: 251 LFLPNRGLS----------ELKKRHAEDFLASDKTKVSRAYSAQSPAQSVAGAYP-NGPN 299
L L E KK H ED S + A A + + Y N N
Sbjct: 410 LLKEQDTLKRKAENGSEEPEEKKAHTED--VSSAQIIDGDLQANQAAYNYSAWYQYNYQN 467
Query: 300 QWPNYG 305
W NYG
Sbjct: 468 PW-NYG 472
>UniRef100_Q801W4 Novel protein similar to pre-mRNA processing proteins [Brachydanio
rerio]
Length = 752
Score = 90.1 bits (222), Expect = 8e-17
Identities = 78/310 (25%), Positives = 133/310 (42%), Gaps = 32/310 (10%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
+++V L+ERC+IACA Y E+WI+Y +E S S + ++ +A V + ++P +HL A
Sbjct: 445 ERVVVLFERCLIACALYEEFWIKYAKYLE-SYSTEAVRHIYKKACTVHLPKKPNVHLLWA 503
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKG 131
F+EQ G I AR+ + V + PGL +R ++E R G +E+A +L + AI +
Sbjct: 504 AFEEQQGSIDEARSILKAVEVSV-PGLAMVRLRRVSLERRHGNMEEAEALLQDAITNGRN 562
Query: 132 KEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFE---AIQP 188
S + A+ V + G +A+++L+ +E + L LL E +Q
Sbjct: 563 SSESSFYSVKLARQLVKVQKSIG---RAKKVLLEAVEKDETNPKLYLNLLELEYSGDVQQ 619
Query: 189 QPKRVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFG-DVQSIKRAEDR 247
+ F +L + R S ++FL FG D+ ++ A ++
Sbjct: 620 NEAEIIACFDRAL-----------SSSMALESRITFSQRKVDFLEDFGSDINTLMAAYEQ 668
Query: 248 HAKLFLPNRGLSELKKRHAEDFLA----SDKTKVSRAYSAQSPAQSVAGAYPN------- 296
H +L + +E+ A +D V+ A Y N
Sbjct: 669 HQRLLAEQESFKRKAENGSEEPDAKRQRTDDQSVASGQMMDMQANHAGYNYNNWYQYNSW 728
Query: 297 -GPNQWPNYG 305
N W YG
Sbjct: 729 GSQNSWGQYG 738
>UniRef100_UPI0000431839 UPI0000431839 UniRef100 entry
Length = 628
Score = 88.2 bits (217), Expect = 3e-16
Identities = 89/322 (27%), Positives = 141/322 (43%), Gaps = 38/322 (11%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
++I+ L+ERC+IACA Y E+W+R S++++ +V RA V ++P +HL A
Sbjct: 326 NRIIILFERCLIACALYDEFWMR------VSDNVEKIRDVYTRACTVHHPKKPNLHLQWA 379
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKG 131
F+E G+ A + + I P +L+ R N+E R G L+ A +LYE I+ K
Sbjct: 380 TFEEGQGNFEKAANILENIDNVI-PNMLQVAYRRINLERRRGDLDKACTLYENYISNSKN 438
Query: 132 KEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPK 191
+ + + +Y+RF+ + +KA ++ LE P L L +Q P
Sbjct: 439 RTIANN---IVVKYARFLCKVKNDVDKAIKV----LEEKDKDNPRLYLQLIDLGMQRTP- 490
Query: 192 RVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFG-DVQSIKRAEDRHAK 250
VD + + FI A +R + +EFL F D++ I +A ++ K
Sbjct: 491 -VDTQEIVGYMDMFIEREH-----ADLEQRVLFAQRKVEFLEDFSPDIRQILKAHEQFQK 544
Query: 251 LFLPNRGLSELKKRHAEDFLASDKTKVSRAYSAQSPAQS----VAGAYPNGPNQWPNYGV 306
+ E KK +D KT S Y +Q QS G Y G Q+P
Sbjct: 545 CI---KQAKERKKTKNDD----TKTGPSGPYQSQQQFQSGQYGGQGGYQQGYQQYP---- 593
Query: 307 QPQTWPATTQAQGQQWPAGYTQ 328
P + P Q Q+ G Q
Sbjct: 594 -PPSDPNYANYQNWQYGQGGPQ 614
>UniRef100_UPI000021A861 UPI000021A861 UniRef100 entry
Length = 480
Score = 87.8 bits (216), Expect = 4e-16
Identities = 62/229 (27%), Positives = 107/229 (46%), Gaps = 7/229 (3%)
Query: 17 LYERCVIACANYPEYWIRYVLCMEAS-ESMDLANNVLARASQVFVK-RQPEIHLFCARFK 74
LYERC++ CA Y E+W RY M A + + N+ RA+ +FV +P I L A F+
Sbjct: 193 LYERCLVTCAFYDEFWFRYARWMSAQPDKTEEVRNIYLRAATIFVPISRPGIRLQFAYFE 252
Query: 75 EQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKGKEH 134
E G + AR + + + PG +E II AN+E R ++ A + +Q IE +
Sbjct: 253 ESCGRVAMAREVHNAILLRL-PGCIEVIISLANLERRHNDIDTAIEVLKQ--QIESPEVD 309
Query: 135 SQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPKRVD 194
T +L +++ ++ G +E+AR + + S+ + FE QP ++
Sbjct: 310 IWTKAVLVTEWASLLWTVKGTAEEARAVFQKNAQWYGGSRHFWMQWIQFELEQPTSAELE 369
Query: 195 IDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFGDVQSIKR 243
E L + I S+ ++EL ++L +L G ++K+
Sbjct: 370 AQHSERL--REIIDKIRTESNMSSAAKKELCGVYLAYLQHRGGKDAMKQ 416
>UniRef100_UPI000036473B UPI000036473B UniRef100 entry
Length = 542
Score = 86.3 bits (212), Expect = 1e-15
Identities = 62/177 (35%), Positives = 93/177 (52%), Gaps = 12/177 (6%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
+QI L+ERC+IACA Y E+W RY +E S +++ A V RA ++ + R+P I + A
Sbjct: 300 NQIRVLFERCLIACALYEEFWTRYTRYLE-SHNVEEARAVFKRACEIHLTRRPNICMQWA 358
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKG 131
F+E+ G++ AR + + E PGL +R +E R G+LE + +L ++A+A K
Sbjct: 359 TFEERHGNLSEARRVLETIE-ETVPGLAVVRLRRVALERRAGQLERSEALLQEAVAGSKE 417
Query: 132 KEHSQTLPMLFAQYS----RFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFE 184
K P L A YS R + N KAR +L LE + + L LL E
Sbjct: 418 K------PTLHAFYSIKLARLLLKLGRNPSKARTVLQEALEISPNNDKLHINLLEVE 468
>UniRef100_UPI00002BB17A UPI00002BB17A UniRef100 entry
Length = 509
Score = 85.1 bits (209), Expect = 3e-15
Identities = 54/156 (34%), Positives = 85/156 (53%), Gaps = 12/156 (7%)
Query: 17 LYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCARFKEQ 76
L+ERC+IACA Y E+W RY +E S +++ A V RA ++ + R+P I + A F+E+
Sbjct: 242 LFERCLIACALYEEFWTRYARYLE-SHNVEEARAVFKRACEIHLTRRPNICMQWATFEER 300
Query: 77 AGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKGKEHSQ 136
++ AR + + + PGL +R +E R G+L+ A +L ++A+A K K
Sbjct: 301 HNNLAEARRVLAAIESRV-PGLAVVRLRRVALERRAGQLDQAVALLQEAVAESKEK---- 355
Query: 137 TLPMLFAQYS----RFVYLASGNSEKAREILVGGLE 168
P L A YS R + + N KAR++L LE
Sbjct: 356 --PTLHAFYSIKLARLLLKLARNPSKARKVLQEALE 389
>UniRef100_UPI000023E0E1 UPI000023E0E1 UniRef100 entry
Length = 587
Score = 84.0 bits (206), Expect = 6e-15
Identities = 63/232 (27%), Positives = 108/232 (46%), Gaps = 7/232 (3%)
Query: 13 QIVKLYERCVIACANYPEYWIRYVLCMEASE-SMDLANNVLARASQVFVK-RQPEIHLFC 70
+IV LYERC++ CA Y + W RY M E + N+ RAS +FV +P I L
Sbjct: 295 RIVALYERCLVTCAFYDDLWFRYARWMSGQEGKAEEVRNIYVRASTMFVPISRPGIRLQW 354
Query: 71 ARFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEK 130
A F+E G + A ++ + + P +E I+ AN+E R ++ A +Y+ I+
Sbjct: 355 AYFEESTGRVDVALDIHEAILLRL-PDSVEVIVSWANVERRQNGIDAAIQVYKN--QIDA 411
Query: 131 GKEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQP 190
T L A+++ ++ G++E+ARE+ + S+ + FE QP
Sbjct: 412 PTVDIYTKAALVAEWALLLWKVKGSTEEAREVFTKNVTWYGDSRLFWDRWFQFELDQPSS 471
Query: 191 KRVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFGDVQSIK 242
+ E + K + SA +++L+ I+L +L GD ++K
Sbjct: 472 AETEAQHGECM--KKVFDELRERSQLSAPVKKDLAQIYLNYLVERGDKDAMK 521
>UniRef100_UPI0000431838 UPI0000431838 UniRef100 entry
Length = 637
Score = 81.6 bits (200), Expect = 3e-14
Identities = 67/242 (27%), Positives = 116/242 (47%), Gaps = 18/242 (7%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEA--SESMDLANNVLARASQVFVKRQPEIHLF 69
++I+ L+ERC+IACA Y E+W+R + +E+ ++++ +V RA V ++P +HL
Sbjct: 401 NRIIILFERCLIACALYDEFWMRVIRYLESLKGDNVEKIRDVYTRACTVHHPKKPNLHLQ 460
Query: 70 CARFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIE 129
A F+E G+ A + + I P +L+ R N+E R G L+ A +LYE I+
Sbjct: 461 WATFEEGQGNFEKAANILENIDNVI-PNMLQVAYRRINLERRRGDLDKACTLYENYISNS 519
Query: 130 KGKEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQ 189
K + + + +Y+RF+ + +KA ++ LE P L L +Q
Sbjct: 520 KNRTIANN---IVVKYARFLCKVKNDVDKAIKV----LEEKDKDNPRLYLQLIDLGMQRT 572
Query: 190 PKRVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFG-DVQSIKRAEDRH 248
P VD + + FI A +R + +EFL F D++ I +A ++
Sbjct: 573 P--VDTQEIVGYMDMFIEREH-----ADLEQRVLFAQRKVEFLEDFSPDIRQILKAHEQF 625
Query: 249 AK 250
K
Sbjct: 626 QK 627
>UniRef100_UPI0000234C5F UPI0000234C5F UniRef100 entry
Length = 588
Score = 80.5 bits (197), Expect = 7e-14
Identities = 59/218 (27%), Positives = 103/218 (47%), Gaps = 7/218 (3%)
Query: 17 LYERCVIACANYPEYWIRYVLCMEASESMDL-ANNVLARASQVFVK-RQPEIHLFCARFK 74
LYERC++ CA+Y E+W RY M A + N+ RAS ++V P L A F+
Sbjct: 299 LYERCLVTCAHYDEFWQRYARWMSAQPGKEEDVRNIYQRASYLYVPIANPATRLQYAYFE 358
Query: 75 EQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKGKEH 134
E G + A+ ++ + I P +E I+ ANM R G LE A +Y+ ++ +
Sbjct: 359 EMCGRVSVAKEIHEAILINI-PNHVETIVSLANMCRRHGGLEAAIEVYKS--QLDSPQCE 415
Query: 135 SQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPKRVD 194
T L A+++R ++ G++E+AR++ + S+ + L FE QP +
Sbjct: 416 MSTKAALVAEWARLLWKIKGSTEEARQVFQKNQQYYLDSQAFWHSYLTFELDQPTSAATE 475
Query: 195 IDFLESLVVKFITPNPENPGVASATEREELSNIFLEFL 232
E +K + + + S+ +L I++ +L
Sbjct: 476 SAQYER--IKQVVEDIRSKSALSSNVARDLVQIYMVYL 511
>UniRef100_UPI000035FA82 UPI000035FA82 UniRef100 entry
Length = 622
Score = 78.2 bits (191), Expect = 3e-13
Identities = 54/174 (31%), Positives = 88/174 (50%), Gaps = 7/174 (4%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
+++V L+ERC+IACA Y E+WI+Y +E S D ++ +A + ++P IHL A
Sbjct: 314 ERVVVLFERCLIACALYEEFWIKYAKYLE-GYSTDGMRHIYKKACITHLPKKPAIHLLWA 372
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAI-AIEK 130
F+EQ GD AR + + + PGL +R ++E R G L +A +L +++ +
Sbjct: 373 AFEEQQGDAEEARRILKSLEATV-PGLAMVRLRRVSLERRHGNLTEAEALLRESMESAST 431
Query: 131 GKEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFE 184
E S L Q+ + N KA+ +L+ LE+ + L LL E
Sbjct: 432 IAERSFYAVKLARQHMK----VQRNLSKAKAVLLDALESDHTNPKLYLNLLELE 481
>UniRef100_UPI0000007649 UPI0000007649 UniRef100 entry
Length = 969
Score = 75.5 bits (184), Expect = 2e-12
Identities = 73/290 (25%), Positives = 130/290 (44%), Gaps = 21/290 (7%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASES----MDLANNVLARASQVFVKRQPEIH 67
++++ L+ERC+IACA Y E+W++ + +E+ E +DL +V RA ++ +P +H
Sbjct: 625 ERVLVLFERCLIACALYDEFWLKMLRYLESLEDQSGVVDLVRDVYRRACRIHHPDKPSLH 684
Query: 68 LFCARFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIA 127
L A F+E + A Q + + P LL+ R N+E R G L+ LY+ I
Sbjct: 685 LMWAAFEECQMNFDDAAEILQRI-DQRCPNLLQLSYRRINVERRRGALDKCRELYKHYIE 743
Query: 128 IEKGKEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQ 187
K K + +L + +Y+RF+ + + L LE + + AL +
Sbjct: 744 STKNKGIAGSLAI---KYARFLNKICHDLDAGLAALQQALERDPANTRV--ALQMIDLCL 798
Query: 188 PQPKRVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFGDVQSIKRAEDR 247
+PK VD + ++ KF+ P ++ + +EFL FG + + +D
Sbjct: 799 QRPK-VDEQEVVEIMDKFMARADIEP-----DQKVLFAQRKVEFLEDFG--STARGLQDA 850
Query: 248 HAKLFLPNRGLSELKKRHAEDFLASDKTKVSRAYSAQSPAQSVAGAYPNG 297
L + L++ K+ + + + S + P S A AY NG
Sbjct: 851 QRAL---QQALTKAKEAQKKSDGSPSRKNSSSSKEGPVPTGSAAAAYNNG 897
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 573,058,675
Number of Sequences: 2790947
Number of extensions: 22965783
Number of successful extensions: 56627
Number of sequences better than 10.0: 120
Number of HSP's better than 10.0 without gapping: 28
Number of HSP's successfully gapped in prelim test: 92
Number of HSP's that attempted gapping in prelim test: 56473
Number of HSP's gapped (non-prelim): 185
length of query: 359
length of database: 848,049,833
effective HSP length: 128
effective length of query: 231
effective length of database: 490,808,617
effective search space: 113376790527
effective search space used: 113376790527
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 75 (33.5 bits)
Medicago: description of AC148607.14


