

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
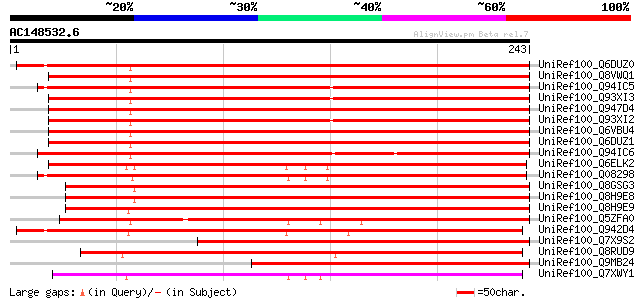
Query= AC148532.6 + phase: 0
(243 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6DUZ0 Dehydration-induced protein RD22-like protein 2... 269 4e-71
UniRef100_Q8VWQ1 Dehydration-induced protein RD22-like protein [... 269 4e-71
UniRef100_Q94IC5 BURP domain-containing protein [Bruguiera gymno... 267 2e-70
UniRef100_Q93XI3 BURP domain-containing protein [Bruguiera gymno... 266 3e-70
UniRef100_Q947D4 Dehydration-responsive protein RD22 [Prunus per... 265 8e-70
UniRef100_Q93XI2 BURP domain-containing protein [Bruguiera gymno... 264 2e-69
UniRef100_Q6VBU4 BURP domain-containing protein [Gossypium hirsu... 261 1e-68
UniRef100_Q6DUZ1 Dehydration-induced protein RD22-like protein 1... 256 5e-67
UniRef100_Q94IC6 BURP domain-containing protein [Bruguiera gymno... 248 1e-64
UniRef100_Q6ELK2 BURP domain-containing protein [Brassica napus] 245 6e-64
UniRef100_Q08298 Dehydration-responsive protein RD22 precursor [... 244 2e-63
UniRef100_Q8GSG3 Resistant specific protein-1(4) (Resistant spec... 240 2e-62
UniRef100_Q8H9E8 Resistant specific protein-3 [Vigna radiata] 240 2e-62
UniRef100_Q8H9E9 Resistant specific protein-2 [Vigna radiata] 238 1e-61
UniRef100_Q5ZFA0 BURP-domain containing protein [Plantago major] 236 5e-61
UniRef100_Q942D4 Putative dehydration-responsive protein RD22 [O... 235 7e-61
UniRef100_Q7X9S2 Putative dehydration-induced protein [Gossypium... 203 3e-51
UniRef100_Q8RUD9 Seed coat BURP domain protein 1 [Glycine max] 193 3e-48
UniRef100_Q9MB24 S1-3 protein [Vigna unguiculata] 187 2e-46
UniRef100_Q7XWY1 OSJNBa0036B17.11 protein [Oryza sativa] 186 3e-46
>UniRef100_Q6DUZ0 Dehydration-induced protein RD22-like protein 2 [Gossypium
arboreum]
Length = 376
Score = 269 bits (688), Expect = 4e-71
Identities = 128/242 (52%), Positives = 165/242 (67%), Gaps = 3/242 (1%)
Query: 4 GKARRMQPLLFIYWPYDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTG--ATFLPR 61
GK + F+Y Y AS TQ+HD N+ LFF EKDLH G +++ FT+ + FLP
Sbjct: 136 GKPVNVNVSPFLY-QYAASETQIHDDPNVALFFLEKDLHPGATMSLHFTENTEKSAFLPY 194
Query: 62 EVANSIPFSSNKVENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVD 121
+ A IPFSSN++ I N FS+K GS ++E +K I CE P IEGEEK C TSLESM+D
Sbjct: 195 QTAQKIPFSSNELPEIFNKFSVKPGSVKAEMMKNTIKECEQPAIEGEEKYCATSLESMID 254
Query: 122 FTTSKLGNNVEAVSTEANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRA 181
++ SKLG +AVSTE K++ +Y I GV+K++ +K VVCH +Y YAVFYCHK
Sbjct: 255 YSISKLGKVDQAVSTEVEKQTPTHKYTITAGVQKMTNDKAVVCHKQNYAYAVFYCHKSET 314
Query: 182 TKVYYLPMEGIDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWF 241
T+ Y +P+EG DGTK K A+CH DTS W+PKHLAF VLKV+PGT+PVCHFL H++W
Sbjct: 315 TRAYMVPLEGADGTKAKAVAVCHTDTSAWNPKHLAFQVLKVEPGTIPVCHFLPRDHIVWV 374
Query: 242 SK 243
K
Sbjct: 375 PK 376
>UniRef100_Q8VWQ1 Dehydration-induced protein RD22-like protein [Gossypium hirsutum]
Length = 335
Score = 269 bits (688), Expect = 4e-71
Identities = 124/227 (54%), Positives = 162/227 (70%), Gaps = 2/227 (0%)
Query: 19 YDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTG--ATFLPREVANSIPFSSNKVEN 76
Y AS TQ+H+ N+ LFF EKD+H G +++ FT+ + FLP + A IPFSS+K+
Sbjct: 109 YAASETQIHEDPNVALFFLEKDMHPGATMSLHFTENTEKSAFLPYQTAQKIPFSSDKLPE 168
Query: 77 ILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVST 136
I N FS+K GS ++E +K I CE P IEGEEK C TSLESM+D++ SKLG +AVST
Sbjct: 169 IFNKFSVKPGSLKAEMMKNTIKECEQPAIEGEEKYCATSLESMIDYSISKLGKVDQAVST 228
Query: 137 EANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTK 196
E K++ Q+Y IA GV+K++++K VVCH +Y YAVFYCHK T+ Y +P+EG DGTK
Sbjct: 229 EVEKQTPMQKYTIAAGVQKMTDDKAVVCHKQNYAYAVFYCHKSETTRAYMVPLEGADGTK 288
Query: 197 VKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
K A+CH DTS W+PKHLAF VLKV+PGT+PVCHFL H++W K
Sbjct: 289 AKAVAVCHTDTSAWNPKHLAFQVLKVEPGTIPVCHFLPRDHIVWVPK 335
>UniRef100_Q94IC5 BURP domain-containing protein [Bruguiera gymnorrhiza]
Length = 346
Score = 267 bits (683), Expect = 2e-70
Identities = 127/232 (54%), Positives = 170/232 (72%), Gaps = 4/232 (1%)
Query: 14 FIYWPYDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTG--ATFLPREVANSIPFSS 71
F+Y Y AS Q+HD N+ LFF EKD+ G +N+ F+++ ATFLPR VA+SIPFSS
Sbjct: 117 FVY-RYAASEDQIHDNPNVALFFLEKDMKPGKSMNLDFSESSNTATFLPRHVADSIPFSS 175
Query: 72 NKVENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNV 131
NK+++ FS+K GS E+E ++ + CE P IEGEEK C TSLESMVDF+TSKLG ++
Sbjct: 176 NKLQDAFREFSVKPGSMEAEIMENTVKECENPGIEGEEKYCATSLESMVDFSTSKLGKDI 235
Query: 132 EAVSTEANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEG 191
+A+STE K++ +QQY IA GVKK + +K VVCH +YPYAVFYCH ++T+ Y + ++G
Sbjct: 236 QAISTEVEKQTGRQQYTIA-GVKKTAGDKSVVCHKQNYPYAVFYCHSTQSTRAYLVQLKG 294
Query: 192 IDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
D T+++ AICH +TS W+PKHLAF VL V+PGTVP+CHFL HV+W K
Sbjct: 295 ADRTRLQAVAICHTNTSAWNPKHLAFQVLAVKPGTVPICHFLPQSHVVWVPK 346
>UniRef100_Q93XI3 BURP domain-containing protein [Bruguiera gymnorrhiza]
Length = 336
Score = 266 bits (681), Expect = 3e-70
Identities = 124/227 (54%), Positives = 168/227 (73%), Gaps = 3/227 (1%)
Query: 19 YDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTG--ATFLPREVANSIPFSSNKVEN 76
Y AS Q+HD NL LFF +KDL G +N+ F ++ ATFLPR VA+SIPFSSNK+++
Sbjct: 111 YAASEDQIHDNPNLALFFLQKDLKPGKSMNLDFPESSNTATFLPRHVADSIPFSSNKLQD 170
Query: 77 ILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVST 136
FS+K GS E+E ++ + CE P IEGEEK C TSLESMVDF+TSKLG +++A+ST
Sbjct: 171 AFREFSVKPGSLEAEIMENTVKECENPGIEGEEKYCATSLESMVDFSTSKLGKDIQAIST 230
Query: 137 EANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTK 196
E K++ +Q+Y IA GVKK++ +K VVCH +YPYAVFYCH +++T+ Y + ++G+D T+
Sbjct: 231 EVEKQTGRQKYTIA-GVKKIAGDKCVVCHKQNYPYAVFYCHSIQSTRAYLVQLKGVDRTR 289
Query: 197 VKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
++ AICH +TS W+PKH AF VL V+PGTVP+CHFL HV+W K
Sbjct: 290 LQAVAICHTNTSAWNPKHPAFQVLAVKPGTVPICHFLPQSHVVWVPK 336
>UniRef100_Q947D4 Dehydration-responsive protein RD22 [Prunus persica]
Length = 349
Score = 265 bits (677), Expect = 8e-70
Identities = 124/227 (54%), Positives = 161/227 (70%), Gaps = 2/227 (0%)
Query: 19 YDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTG--ATFLPREVANSIPFSSNKVEN 76
Y A+ TQLH +N +FF EKD+ GT + + F+ A F+PR+ A+SIPFSSNK+
Sbjct: 120 YAATETQLHGNRNAAIFFLEKDIRPGTSMTLTFSGNSNTAAFVPRKTADSIPFSSNKLPE 179
Query: 77 ILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVST 136
I + FS+K S E++ +K I CE+ I GEEK C TSLESMVDF+TSKLG N++A+ST
Sbjct: 180 IFSQFSVKPESVEADIIKGTIEECESSGIRGEEKYCATSLESMVDFSTSKLGGNIQAIST 239
Query: 137 EANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTK 196
EA K + Q+Y I GVKKL+ K VVCH +YPYAVFYCH + T+ Y +P++G DG +
Sbjct: 240 EAEKGATLQKYTITPGVKKLAAGKSVVCHKQTYPYAVFYCHATKTTRAYVVPLKGADGLE 299
Query: 197 VKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
K A+CH DTS+W+PKHLAF VLKV+PGTVPVCHFL H++W K
Sbjct: 300 AKAVAVCHTDTSEWNPKHLAFQVLKVKPGTVPVCHFLPKDHIVWVPK 346
>UniRef100_Q93XI2 BURP domain-containing protein [Bruguiera gymnorrhiza]
Length = 307
Score = 264 bits (674), Expect = 2e-69
Identities = 124/227 (54%), Positives = 166/227 (72%), Gaps = 3/227 (1%)
Query: 19 YDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTG--ATFLPREVANSIPFSSNKVEN 76
Y AS Q+HD N+ LFF +KDL G +N+ F + ATFLPR VA+SIPFSSNK+++
Sbjct: 82 YAASEDQIHDNPNVALFFLQKDLKPGKSMNLDFPASSNTATFLPRHVADSIPFSSNKLQD 141
Query: 77 ILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVST 136
FS+K GS E+E ++ + CE P IEGEEK C TSLESMVDF+TSKLG +V+A+ST
Sbjct: 142 AFREFSVKPGSLEAEIMENTVKECENPGIEGEEKYCATSLESMVDFSTSKLGKDVQAIST 201
Query: 137 EANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTK 196
E K++ +Q+Y IA GVKK++ +K VVCH +YPYAVFYCH +++T+ Y + ++G+D T+
Sbjct: 202 EVEKQTGRQKYTIA-GVKKIAGDKCVVCHKQNYPYAVFYCHSIQSTRAYLVQLKGVDRTR 260
Query: 197 VKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
++ AICH +TS W+PKH AF VL V+PGTVP CHFL HV+W K
Sbjct: 261 LQAVAICHTNTSAWNPKHPAFQVLAVKPGTVPTCHFLPQSHVVWVPK 307
>UniRef100_Q6VBU4 BURP domain-containing protein [Gossypium hirsutum]
Length = 335
Score = 261 bits (666), Expect = 1e-68
Identities = 120/227 (52%), Positives = 159/227 (69%), Gaps = 2/227 (0%)
Query: 19 YDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTG--ATFLPREVANSIPFSSNKVEN 76
Y AS TQ+H+ N+ LFF EKD+H G +++ F + + FLP + A IPFSS+K+
Sbjct: 109 YAASETQIHEDPNVALFFLEKDMHPGATMSLHFIENTEKSAFLPYQTAQKIPFSSDKLPE 168
Query: 77 ILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVST 136
I N FS+K GS ++E +K I CE P IEGEEK C TSLESM+D++ SKLG +AVST
Sbjct: 169 IFNKFSVKPGSVKAEMMKNTIKECEQPAIEGEEKYCATSLESMIDYSISKLGKVDQAVST 228
Query: 137 EANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTK 196
E K++ Q+Y IA GV+K++++K VVCH +Y YAVFYCHK T+ Y +P+EG GTK
Sbjct: 229 EVEKQTPMQKYTIAAGVQKMTDDKAVVCHKQNYAYAVFYCHKSETTRAYMVPLEGAGGTK 288
Query: 197 VKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
+ A+CH DTS W+PKHLAF LKV+PGT+PVCHFL H++W K
Sbjct: 289 AQALAVCHTDTSAWNPKHLAFQFLKVEPGTIPVCHFLPRDHIVWVPK 335
>UniRef100_Q6DUZ1 Dehydration-induced protein RD22-like protein 1 [Gossypium
arboreum]
Length = 335
Score = 256 bits (653), Expect = 5e-67
Identities = 119/227 (52%), Positives = 157/227 (68%), Gaps = 2/227 (0%)
Query: 19 YDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTG--ATFLPREVANSIPFSSNKVEN 76
Y AS TQ+H+ N+ LFF EKD+H G +++ F + + FLP + A FSS+K+
Sbjct: 109 YAASETQIHEDPNVALFFLEKDMHPGATMSLHFIENTEKSAFLPYQTAPKNTFSSDKLPE 168
Query: 77 ILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVST 136
I N FS+K GS ++E +K I CE P IEGEEK C TSLESM+D++ SKLG +AVST
Sbjct: 169 IFNKFSVKPGSVKAEMMKNTIKECEQPAIEGEEKYCATSLESMIDYSISKLGKVDQAVST 228
Query: 137 EANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTK 196
E K++ Q+Y IA GV+K++++K VVCH +Y YAVFYCHK T+ Y +P+EG GTK
Sbjct: 229 EVEKQTPMQKYTIAAGVQKMTDDKAVVCHKQNYAYAVFYCHKSETTRAYMVPLEGAGGTK 288
Query: 197 VKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
K A+CH DTS W+PKHLAF LKV+PGT+PVCHFL H++W K
Sbjct: 289 AKALAVCHTDTSAWNPKHLAFQFLKVEPGTIPVCHFLPRDHIVWVPK 335
>UniRef100_Q94IC6 BURP domain-containing protein [Bruguiera gymnorrhiza]
Length = 292
Score = 248 bits (632), Expect = 1e-64
Identities = 120/232 (51%), Positives = 165/232 (70%), Gaps = 4/232 (1%)
Query: 14 FIYWPYDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTG--ATFLPREVANSIPFSS 71
FI + AS Q+ D N+ +FF EKDL G +N+ F ++ ATFLPR VA+S+PFSS
Sbjct: 63 FINYYNAASEDQIRDNSNITIFFLEKDLKPGKSMNLYFFESSNTATFLPRHVADSMPFSS 122
Query: 72 NKVENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNV 131
NK++ FS+K GS E+E ++ I CE P IEGEEK CVTSLES+VDF++SKLG ++
Sbjct: 123 NKLQEAFREFSVKPGSLEAETMEDTIKECENPRIEGEEKYCVTSLESIVDFSSSKLGKDI 182
Query: 132 EAVSTEANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEG 191
+A+STE K++ +Q+Y IA VKK++ +K VVCH +YPYAVFYCH + ++ Y + ++G
Sbjct: 183 QAISTEVEKQTGRQKYTIAV-VKKMAGDKSVVCHKQNYPYAVFYCHTTQ-SRAYLVQLKG 240
Query: 192 IDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
D +++K A CH +TS W+PKH+AF VL V+PGTVPVCHFL HV+W K
Sbjct: 241 ADRSELKAVATCHTNTSAWNPKHMAFQVLAVKPGTVPVCHFLTQSHVVWAPK 292
>UniRef100_Q6ELK2 BURP domain-containing protein [Brassica napus]
Length = 387
Score = 245 bits (626), Expect = 6e-64
Identities = 129/232 (55%), Positives = 158/232 (67%), Gaps = 8/232 (3%)
Query: 19 YDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTK----TGAT-FLPREVANSIPFSSNK 73
Y AS TQLHD LFF EKD+ G +N++F G T FLPR A ++PF S K
Sbjct: 155 YAASETQLHDDPKAALFFLEKDMVPGKAMNLRFNAEDGYNGKTAFLPRGEAETVPFGSEK 214
Query: 74 VENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLG-NNVE 132
ILN FS+K GS E+E +K+ I CE + GEEK C TSLESMVDF+ SKLG ++V
Sbjct: 215 SSEILNTFSVKPGSGEAEMMKKTIEECEAKRVGGEEKYCATSLESMVDFSVSKLGKDHVR 274
Query: 133 AVSTE-ANKESDKQQY-IIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPME 190
AVSTE A K + Q+Y I A GVKKLS++K VVCH YP+AVFYCHK T VY +P+E
Sbjct: 275 AVSTEVAEKNAPMQKYRIAAAGVKKLSDDKSVVCHKQKYPFAVFYCHKAMMTSVYAVPLE 334
Query: 191 GIDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFS 242
G +G + K A+CH +TS W+P HLAF VLKV+PG+VPVCHFL HV+WFS
Sbjct: 335 GENGLRAKAVAVCHKNTSAWNPNHLAFKVLKVKPGSVPVCHFLPETHVVWFS 386
>UniRef100_Q08298 Dehydration-responsive protein RD22 precursor [Arabidopsis
thaliana]
Length = 392
Score = 244 bits (622), Expect = 2e-63
Identities = 128/237 (54%), Positives = 157/237 (66%), Gaps = 9/237 (3%)
Query: 14 FIYWPYDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTGA-----TFLPREVANSIP 68
F+Y Y A TQLHD N LFF EKDL G ++N++F FLPR A ++P
Sbjct: 156 FVY-NYAAKETQLHDDPNAALFFLEKDLVRGKEMNVRFNAEDGYGGKTAFLPRGEAETVP 214
Query: 69 FSSNKVENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLG 128
F S K L FS++ GS E+E +K+ I CE + GEEK C TSLESMVDF+ SKLG
Sbjct: 215 FGSEKFSETLKRFSVEAGSEEAEMMKKTIEECEARKVSGEEKYCATSLESMVDFSVSKLG 274
Query: 129 N-NVEAVSTE-ANKESDKQQY-IIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVY 185
+V AVSTE A K + Q+Y I A GVKKLS++K VVCH YP+AVFYCHK T VY
Sbjct: 275 KYHVRAVSTEVAKKNAPMQKYKIAAAGVKKLSDDKSVVCHKQKYPFAVFYCHKAMMTTVY 334
Query: 186 YLPMEGIDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFS 242
+P+EG +G + K A+CH +TS W+P HLAF VLKV+PGTVPVCHFL HV+WFS
Sbjct: 335 AVPLEGENGMRAKAVAVCHKNTSAWNPNHLAFKVLKVKPGTVPVCHFLPETHVVWFS 391
>UniRef100_Q8GSG3 Resistant specific protein-1(4) (Resistant specific protein- 1(8))
[Vigna radiata]
Length = 402
Score = 240 bits (613), Expect = 2e-62
Identities = 109/219 (49%), Positives = 148/219 (66%), Gaps = 2/219 (0%)
Query: 27 HDYQNLDLFFFEKDLHNGTKLNMQFTKTGAT--FLPREVANSIPFSSNKVENILNYFSIK 84
HD+ FF E+ L G KL++ F K + L RE+A +PFSS K+ IL ++K
Sbjct: 184 HDHLKPSNFFSEERLRRGAKLDVLFRKRNFSTPLLTREIAEHLPFSSEKINEILEILAVK 243
Query: 85 QGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDK 144
S +++N+++ + +CE P ++GEEK C TS+ESMVDF TSKLGNN STE S
Sbjct: 244 PDSKDAKNVEETLNHCEKPALKGEEKQCATSVESMVDFVTSKLGNNARVTSTELEIGSKF 303
Query: 145 QQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKTAAICH 204
Q++I+ GVK L+E KI+ CH +SYPY VFYCHK+ + ++LP+EG DGT+VK AICH
Sbjct: 304 QKFIVKDGVKILAEEKIIACHPMSYPYVVFYCHKMANSTAHFLPLEGEDGTRVKAVAICH 363
Query: 205 NDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
DTSQW P H+AF VLKV+PGT CHF GH++W++K
Sbjct: 364 KDTSQWDPHHVAFQVLKVKPGTSSACHFFPEGHLVWYAK 402
>UniRef100_Q8H9E8 Resistant specific protein-3 [Vigna radiata]
Length = 275
Score = 240 bits (613), Expect = 2e-62
Identities = 109/219 (49%), Positives = 148/219 (66%), Gaps = 2/219 (0%)
Query: 27 HDYQNLDLFFFEKDLHNGTKLNMQFTKTGAT--FLPREVANSIPFSSNKVENILNYFSIK 84
HD+ FF E+ L G KL++ F K + L RE+A +PFSS K+ IL ++K
Sbjct: 57 HDHLKPSNFFSEERLRRGAKLDVLFRKRNFSTPLLTREIAEHLPFSSEKINEILEILAVK 116
Query: 85 QGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDK 144
S +++N+++ + +CE P ++GEEK C TS+ESMVDF TSKLGNN STE S
Sbjct: 117 PDSKDAKNVEETLNHCEKPALKGEEKQCATSVESMVDFVTSKLGNNARVTSTELEIGSKF 176
Query: 145 QQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKTAAICH 204
Q++I+ GVK L+E KI+ CH +SYPY VFYCHK+ + ++LP+EG DGT+VK AICH
Sbjct: 177 QKFIVKDGVKILAEEKIIACHPMSYPYVVFYCHKMANSTAHFLPLEGEDGTRVKAVAICH 236
Query: 205 NDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
DTSQW P H+AF VLKV+PGT CHF GH++W++K
Sbjct: 237 KDTSQWDPHHVAFQVLKVKPGTSSACHFFPEGHLVWYAK 275
>UniRef100_Q8H9E9 Resistant specific protein-2 [Vigna radiata]
Length = 440
Score = 238 bits (606), Expect = 1e-61
Identities = 109/219 (49%), Positives = 146/219 (65%), Gaps = 2/219 (0%)
Query: 27 HDYQNLDLFFFEKDLHNGTKLNMQFTKT--GATFLPREVANSIPFSSNKVENILNYFSIK 84
HD+ +F E+ L G KL M F K L RE+A +PFSS K+ IL ++K
Sbjct: 222 HDHLKPSSYFSEEGLRRGAKLVMLFHKRKFSTPLLTREIAEHLPFSSEKINEILEILAVK 281
Query: 85 QGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDK 144
S ++N+++ + CE P ++GEEK C TS+ESMVDF TSKLGNN STE ES
Sbjct: 282 PDSKNAKNVEKTLNNCEEPALKGEEKHCATSVESMVDFVTSKLGNNARVTSTELEIESKF 341
Query: 145 QQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKTAAICH 204
Q++I+ GVK L+E +I+ CH +SYPY VFYCHK+ + + +P+EG DGT+VK ICH
Sbjct: 342 QKFIVKDGVKILAEEEIIACHPMSYPYVVFYCHKMSNSTAHVVPLEGEDGTRVKAIVICH 401
Query: 205 NDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
DTSQW P H+AF VLKV+PGT PVCHF +GH++W++K
Sbjct: 402 KDTSQWDPDHVAFQVLKVKPGTSPVCHFFPNGHLLWYAK 440
>UniRef100_Q5ZFA0 BURP-domain containing protein [Plantago major]
Length = 348
Score = 236 bits (601), Expect = 5e-61
Identities = 121/227 (53%), Positives = 153/227 (67%), Gaps = 9/227 (3%)
Query: 24 TQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTG--ATFLPREVANSIPFSSNKVENILNYF 81
TQLHD LFF EKD+H GTK+ +QF KT ATFLPR+VA+SIPFSS + IL F
Sbjct: 123 TQLHDDPTASLFFLEKDMHAGTKMKLQFVKTSNQATFLPRQVADSIPFSSKRFPEILRKF 182
Query: 82 SIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGN-NVEAVSTEANK 140
K S E++ +K I CE I+GEEK C TSLESM+DF TSKLG+ NVEA+ST A +
Sbjct: 183 --KPTSDEADIMKDTIKDCEDEGIKGEEKFCATSLESMIDFCTSKLGSRNVEAISTNAQE 240
Query: 141 ESDK---QQYIIAKGVKKLSENKIVV-CHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTK 196
++ ++Y + +K+ NK VV CH + Y YAVFYCHK+ T Y + + DG K
Sbjct: 241 NAENSSPKEYTLVGAPRKMPGNKAVVACHKMDYAYAVFYCHKIAKTVAYEVSLAAADGCK 300
Query: 197 VKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
V+ A+CH+DTSQW+PKH AF VLKVQPG+VPVCHFL ++W K
Sbjct: 301 VEAVAVCHHDTSQWNPKHFAFQVLKVQPGSVPVCHFLPQQQIVWVPK 347
>UniRef100_Q942D4 Putative dehydration-responsive protein RD22 [Oryza sativa]
Length = 429
Score = 235 bits (600), Expect = 7e-61
Identities = 123/244 (50%), Positives = 156/244 (63%), Gaps = 8/244 (3%)
Query: 4 GKARRMQPLLFIYWPYDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKT--GATFLPR 61
GK + FIY Y A+ TQLHD N+ LFF EKDLH G + + FT T G FLPR
Sbjct: 183 GKPVHVNVAPFIY-NYAATETQLHDDPNVALFFLEKDLHPGKTMAVHFTATTAGEKFLPR 241
Query: 62 EVANSIPFSSNKVENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVD 121
A+++PFSS KV IL+ FS+K GS E+ + Q + CE P +GE K+C TSLESMVD
Sbjct: 242 SEADAMPFSSEKVPEILSRFSVKPGSVEAAEMAQTLRDCEAPPAQGERKACATSLESMVD 301
Query: 122 FTTSKLG-NNVEAVSTEANKESDKQQYIIAKGVKKLS----ENKIVVCHLLSYPYAVFYC 176
F TS LG ++V A ST KE +Q VK+ + ++++V CH Y YAVF C
Sbjct: 302 FATSSLGTSHVRAASTVVGKEGSPEQEYTVTAVKRAAAGGDQDQLVACHAEPYAYAVFAC 361
Query: 177 HKLRATKVYYLPMEGIDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHG 236
H RAT+ Y + M G DGT V+ A+CH DT+ W+PKH+AF VLKV+PGTVPVCHFL
Sbjct: 362 HLTRATRAYAVSMAGRDGTGVEAVAVCHADTAGWNPKHVAFQVLKVKPGTVPVCHFLPQD 421
Query: 237 HVIW 240
HV+W
Sbjct: 422 HVVW 425
>UniRef100_Q7X9S2 Putative dehydration-induced protein [Gossypium barbadense]
Length = 156
Score = 203 bits (517), Expect = 3e-51
Identities = 90/155 (58%), Positives = 114/155 (73%)
Query: 89 ESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDKQQYI 148
++E +K I CE P IEGEEK C TSLESM+D++ SKLG +AVSTE K++ Q+Y
Sbjct: 2 KAEMMKNTIKECEQPAIEGEEKYCATSLESMIDYSISKLGKVDQAVSTEVEKQTPMQKYT 61
Query: 149 IAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKTAAICHNDTS 208
IA GV+K++++K VVCH +Y YAVFYCHK T+ Y +P+EG DGTK K A+CH DTS
Sbjct: 62 IAAGVQKMTDDKAVVCHKQNYAYAVFYCHKSETTRAYMVPLEGADGTKAKAVAVCHTDTS 121
Query: 209 QWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
W+PKHLAF VLK +PGTVPVCHFL H++W K
Sbjct: 122 AWNPKHLAFQVLKAEPGTVPVCHFLPRDHIVWVPK 156
>UniRef100_Q8RUD9 Seed coat BURP domain protein 1 [Glycine max]
Length = 305
Score = 193 bits (491), Expect = 3e-48
Identities = 90/211 (42%), Positives = 131/211 (61%), Gaps = 4/211 (1%)
Query: 34 LFFFEKDLHNGTKLNMQF---TKTGATFLPREVANSIPFSSNKVENILNYFSIKQGSAES 90
LFF E+DL G NM+F TK LPR+++ IPFS +K + +L ++ S+ +
Sbjct: 92 LFFLEEDLRAGKIFNMKFVNNTKATVPLLPRQISKQIPFSEDKKKQVLAMLGVEANSSNA 151
Query: 91 ENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDKQQYIIA 150
+ + + IG C+ P EGE K C TSLESMVDF S LG NV A STE +E++ ++++
Sbjct: 152 KIIAETIGLCQEPATEGERKHCATSLESMVDFVVSALGKNVGAFSTEKERETESGKFVVV 211
Query: 151 K-GVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKTAAICHNDTSQ 209
K GV+KL ++K++ CH +SYPY VF CH + + Y + ++G DG +VK CH DTS+
Sbjct: 212 KNGVRKLGDDKVIACHPMSYPYVVFGCHLVPRSSGYLVRLKGEDGVRVKAVVACHRDTSK 271
Query: 210 WSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
W H AF VL ++PG VCH G+++W
Sbjct: 272 WDHNHGAFKVLNLKPGNGTVCHVFTEGNLLW 302
>UniRef100_Q9MB24 S1-3 protein [Vigna unguiculata]
Length = 132
Score = 187 bits (475), Expect = 2e-46
Identities = 85/130 (65%), Positives = 101/130 (77%)
Query: 114 TSLESMVDFTTSKLGNNVEAVSTEANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAV 173
TSLESMVDF+TSKLG NV +STE ++E+ QQY IA GVKK+S + VVCH SYPYAV
Sbjct: 3 TSLESMVDFSTSKLGKNVAVLSTEVDQETGLQQYTIAPGVKKVSGDNAVVCHKQSYPYAV 62
Query: 174 FYCHKLRATKVYYLPMEGIDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFL 233
FYCHK T+ Y +P+EG +G +VK A+CH DTSQW+PKHLAF VLKV+PGTVPVCHFL
Sbjct: 63 FYCHKTETTRTYPVPLEGANGIRVKAVAVCHTDTSQWNPKHLAFEVLKVKPGTVPVCHFL 122
Query: 234 QHGHVIWFSK 243
HV+W K
Sbjct: 123 PEDHVVWVQK 132
>UniRef100_Q7XWY1 OSJNBa0036B17.11 protein [Oryza sativa]
Length = 285
Score = 186 bits (473), Expect = 3e-46
Identities = 94/226 (41%), Positives = 134/226 (58%), Gaps = 6/226 (2%)
Query: 21 ASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTK--TGATFLPREVANSIPFSSNKVENIL 78
A+ Q + LFF E +L + + + F G FLPR A+++PFSS ++ IL
Sbjct: 56 AASQQFFKDPTMGLFFLETNLQSSKSIKLHFANMMAGTKFLPRGEADAVPFSSKDLQEIL 115
Query: 79 NYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGN-NVEAVSTE 137
F ++ GS ++ +K + CE P +GE+K+C TSLESMVDF S LG +++A ST
Sbjct: 116 ARFGVRPGSVDASVVKNTLLECELPANKGEKKACATSLESMVDFVASSLGTRDIKAASTF 175
Query: 138 -ANKESDK--QQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDG 194
K+ D Q+Y + + +++ CH SYPYAVF CH AT+ Y + G DG
Sbjct: 176 LVGKDGDTPAQEYTVTGARRMAETGQLIACHPESYPYAVFMCHLTEATRAYKASLVGKDG 235
Query: 195 TKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
V+ A+CH DT++W+PKH AF VL V+PGTVPVCHF+Q V+W
Sbjct: 236 AAVEAVAVCHTDTAEWNPKHAAFQVLGVKPGTVPVCHFVQPDVVVW 281
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.133 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 402,254,265
Number of Sequences: 2790947
Number of extensions: 15482689
Number of successful extensions: 39942
Number of sequences better than 10.0: 102
Number of HSP's better than 10.0 without gapping: 72
Number of HSP's successfully gapped in prelim test: 30
Number of HSP's that attempted gapping in prelim test: 39729
Number of HSP's gapped (non-prelim): 106
length of query: 243
length of database: 848,049,833
effective HSP length: 124
effective length of query: 119
effective length of database: 501,972,405
effective search space: 59734716195
effective search space used: 59734716195
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 73 (32.7 bits)
Medicago: description of AC148532.6


