

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.15 + phase: 1 /partial
(178 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
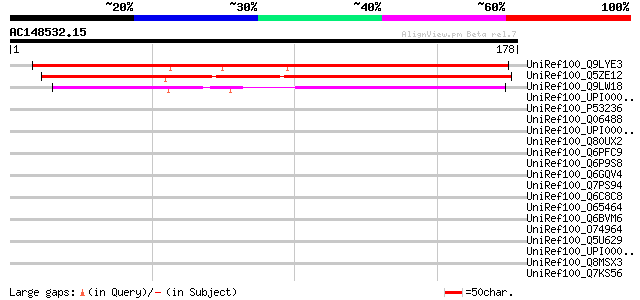
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LYE3 Hypothetical protein F15N18_60 [Arabidopsis tha... 173 2e-42
UniRef100_Q5ZE12 Hypothetical protein P0410E03.22 [Oryza sativa] 166 2e-40
UniRef100_Q9LW18 Emb|CAB87707.1 [Arabidopsis thaliana] 105 7e-22
UniRef100_UPI000043343E UPI000043343E UniRef100 entry 44 0.001
UniRef100_P53236 Chromatin structure remodeling complex protein ... 42 0.005
UniRef100_Q06488 Chromatin structure remodeling complex protein ... 40 0.021
UniRef100_UPI000021E173 UPI000021E173 UniRef100 entry 40 0.027
UniRef100_Q80UX2 BC060615 protein [Mus musculus] 40 0.027
UniRef100_Q6PFC9 BC060615 protein [Mus musculus] 40 0.027
UniRef100_Q6P9S8 BC060615 protein [Mus musculus] 40 0.027
UniRef100_Q6GQV4 Hypothetical protein BC060615 [Mus musculus] 40 0.027
UniRef100_Q7PS94 ENSANGP00000005615 [Anopheles gambiae str. PEST] 40 0.035
UniRef100_Q6C8C8 Similar to sp|Q06488 Saccharomyces cerevisiae Y... 40 0.035
UniRef100_O65464 Hypothetical protein F21P8.10 [Arabidopsis thal... 39 0.046
UniRef100_Q6BVM6 Similar to CA4169|CaRSC2 Candida albicans CaRSC... 39 0.046
UniRef100_O74964 Rsc1 protein [Schizosaccharomyces pombe] 39 0.046
UniRef100_Q5U629 KIAA1447 protein [Homo sapiens] 39 0.060
UniRef100_UPI000036632B UPI000036632B UniRef100 entry 39 0.078
UniRef100_Q8MSX3 LD15342p [Drosophila melanogaster] 38 0.10
UniRef100_Q7KS56 CG31151-PA, isoform A [Drosophila melanogaster] 38 0.10
>UniRef100_Q9LYE3 Hypothetical protein F15N18_60 [Arabidopsis thaliana]
Length = 691
Score = 173 bits (438), Expect = 2e-42
Identities = 90/174 (51%), Positives = 119/174 (67%), Gaps = 7/174 (4%)
Query: 9 SPESETLEFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKN----DFAEPRIGK 64
S SE LEFKWG+K+G GGKK+D+QFY+SFT G +Y L+D V V N D EP IG
Sbjct: 4 SVASEGLEFKWGKKKGVGGKKKDVQFYESFTYDGDEYRLYDCVLVGNASEPDSTEPFIGM 63
Query: 65 IIKIWETPT--LERKIKVQWFFRPIEVSKCLTWI-KIYFNELFFACGDGDGLATIHPLES 121
IIKIWE + +K+K+ WFF+P E++ L + + NE+F A G+G GLA + LE+
Sbjct: 64 IIKIWEHANKHIPKKVKLLWFFKPSEIAPYLEGVPNVLANEVFLASGEGLGLANTNQLEA 123
Query: 122 IAGKCNIVCISKDSRNPQLSDKVTWSAAFVFYRYFDVGQRKIVEEVDDKSIGIE 175
I GKC+++CISKD RNPQ SD+ SA FVF R FDVG K+V+ +DDK G++
Sbjct: 124 IGGKCSVLCISKDKRNPQPSDEKFNSADFVFCRAFDVGSCKVVDTIDDKIAGVD 177
>UniRef100_Q5ZE12 Hypothetical protein P0410E03.22 [Oryza sativa]
Length = 501
Score = 166 bits (421), Expect = 2e-40
Identities = 83/166 (50%), Positives = 113/166 (68%), Gaps = 3/166 (1%)
Query: 12 SETLEFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYV-KNDFAEPRIGKIIKIWE 70
+E ++F WG+KR KGG K D QFY SFT V YSL+DNVY+ K+ +EP IGKIIKIW+
Sbjct: 5 NENIQFSWGKKRAKGGIKMDTQFYDSFTFDNVKYSLYDNVYLFKSGESEPYIGKIIKIWQ 64
Query: 71 TPTLERKIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCNIVC 130
+K+K+ WFF P E+ K L+ + E+F ACG+G GLA I+PLE+I GKC ++C
Sbjct: 65 Q-NQAKKVKILWFFLPDEIRKHLSG-PVMEKEIFLACGEGVGLADINPLEAIGGKCTVLC 122
Query: 131 ISKDSRNPQLSDKVTWSAAFVFYRYFDVGQRKIVEEVDDKSIGIEG 176
ISKD RN Q S + A ++FYR+FDV + E++ +K G+EG
Sbjct: 123 ISKDERNRQPSPRELAMADYIFYRFFDVNSCTLSEQLPEKIAGVEG 168
>UniRef100_Q9LW18 Emb|CAB87707.1 [Arabidopsis thaliana]
Length = 604
Score = 105 bits (261), Expect = 7e-22
Identities = 63/163 (38%), Positives = 87/163 (52%), Gaps = 24/163 (14%)
Query: 16 EFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVK-NDFAEPRIGKIIKIWETPTL 74
+FKWG KRG G K ++FY+SFTL G++Y LFD Y + +E IGK++ T
Sbjct: 117 DFKWGAKRGVGRKDNKVRFYESFTLEGIEYRLFDCAYFYVHGQSETSIGKLVTF--TRHQ 174
Query: 75 ER---KIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCNIVCI 131
+R + + QW +ELF ACGD G++ I+ +E+I GKCN+VC
Sbjct: 175 QRFLGEYEPQW------------------DELFLACGDEKGVSNINDVETIMGKCNVVCT 216
Query: 132 SKDSRNPQLSDKVTWSAAFVFYRYFDVGQRKIVEEVDDKSIGI 174
S D RNP+ K A ++F R FD R I E+ D GI
Sbjct: 217 SDDRRNPRPGTKELRRAKYIFSRTFDTRLRIISEDFADAIAGI 259
>UniRef100_UPI000043343E UPI000043343E UniRef100 entry
Length = 953
Score = 44.3 bits (103), Expect = 0.001
Identities = 29/96 (30%), Positives = 41/96 (42%), Gaps = 6/96 (6%)
Query: 34 FYQSFTLRGVDYSLFDNVYVKNDFAEPRIGKIIKIWETPTLERKIKVQWFFRPIEVSKCL 93
+Y+ + D VYV D +I +I IW T + K W P EV
Sbjct: 790 YYEQYNTCAGSVKTGDFVYVATDGGRQQIAQIDAIWSTKDGKCYFKGPWLLMPAEVPHTP 849
Query: 94 TWIKIYFNELFFACGDGDGLATIHPLESIAGKCNIV 129
T + Y ELF + DG HP+ +I GKC ++
Sbjct: 850 TKL-FYKQELFLSTVDG-----THPIVAIVGKCAVL 879
>UniRef100_P53236 Chromatin structure remodeling complex protein RSC1 [Saccharomyces
cerevisiae]
Length = 928
Score = 42.4 bits (98), Expect = 0.005
Identities = 27/101 (26%), Positives = 46/101 (44%), Gaps = 7/101 (6%)
Query: 55 NDFAEPRIGKIIKIWETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLA 114
ND +P +G+I ++W T + + W+FRP + + + Y NE+ G
Sbjct: 382 NDINKPIVGQIFRLWSTTDGNKWLNACWYFRPEQTVHRVDRL-FYKNEVM-----KTGQY 435
Query: 115 TIHPLESIAGKCNIVCISKDSRNPQLSDKVTWSAAFVFYRY 155
HP++ I GKC ++ ++ R S KV +RY
Sbjct: 436 RDHPIQDIKGKCYVIHFTRFQRGDP-STKVNGPQFVCEFRY 475
>UniRef100_Q06488 Chromatin structure remodeling complex protein RSC2 [Saccharomyces
cerevisiae]
Length = 889
Score = 40.4 bits (93), Expect = 0.021
Identities = 25/98 (25%), Positives = 51/98 (51%), Gaps = 10/98 (10%)
Query: 44 DYSLFDNVYVKNDFAEPRIGKIIKIWETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNEL 103
D++L N +ND +P +G+I ++W+TP ++ + W++RP + + + Y NE+
Sbjct: 414 DWALLRN---QNDPQKPIVGQIFRLWKTPDGKQWLNACWYYRPEQTVHRVDRL-FYKNEV 469
Query: 104 FFACGDGDGLATIHPLESIAGKCNIVCISKDSR-NPQL 140
G H + ++ GKC ++ ++ R NP +
Sbjct: 470 M-----KTGQYRDHLVSNLVGKCYVIHFTRYQRGNPDM 502
>UniRef100_UPI000021E173 UPI000021E173 UniRef100 entry
Length = 258
Score = 40.0 bits (92), Expect = 0.027
Identities = 36/122 (29%), Positives = 61/122 (49%), Gaps = 19/122 (15%)
Query: 17 FKWG----QKRGKGGKKRDIQFYQSF-----TLRGVDYSLFDNVYVKNDFAEPRIGKIIK 67
+KW Q+RG GK R + FY++ TLR D ++F + N P IG+I
Sbjct: 103 WKWSGNPTQRRGMKGKARKL-FYKAIVRGKETLRIGDCAVFLSAGRPN---LPYIGRIES 158
Query: 68 IWETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCN 127
+WE+ +KV+WF+ P E +K N L+ +C + + + +++I+ KC
Sbjct: 159 LWESWGSNMVVKVKWFYHP-EETKLGKRQSDGKNALYQSCHEDE-----NDVQTISHKCQ 212
Query: 128 IV 129
+V
Sbjct: 213 VV 214
>UniRef100_Q80UX2 BC060615 protein [Mus musculus]
Length = 258
Score = 40.0 bits (92), Expect = 0.027
Identities = 36/122 (29%), Positives = 61/122 (49%), Gaps = 19/122 (15%)
Query: 17 FKWG----QKRGKGGKKRDIQFYQSF-----TLRGVDYSLFDNVYVKNDFAEPRIGKIIK 67
+KW Q+RG GK R + FY++ TLR D ++F + N P IG+I
Sbjct: 103 WKWSGNPTQRRGMKGKARKL-FYKAIVRGKETLRIGDCAVFLSAGRPN---LPYIGRIES 158
Query: 68 IWETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCN 127
+WE+ +KV+WF+ P E +K N L+ +C + + + +++I+ KC
Sbjct: 159 LWESWGSNMVVKVKWFYHP-EETKLGKRQSDGKNALYQSCHEDE-----NDVQTISHKCQ 212
Query: 128 IV 129
+V
Sbjct: 213 VV 214
>UniRef100_Q6PFC9 BC060615 protein [Mus musculus]
Length = 193
Score = 40.0 bits (92), Expect = 0.027
Identities = 36/122 (29%), Positives = 61/122 (49%), Gaps = 19/122 (15%)
Query: 17 FKWG----QKRGKGGKKRDIQFYQSF-----TLRGVDYSLFDNVYVKNDFAEPRIGKIIK 67
+KW Q+RG GK R + FY++ TLR D ++F + N P IG+I
Sbjct: 38 WKWSGNPTQRRGMKGKARKL-FYKAIVRGKETLRIGDCAVFLSAGRPN---LPYIGRIES 93
Query: 68 IWETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCN 127
+WE+ +KV+WF+ P E +K N L+ +C + + + +++I+ KC
Sbjct: 94 LWESWGSNMVVKVKWFYHP-EETKLGKRQSDGKNALYQSCHEDE-----NDVQTISHKCQ 147
Query: 128 IV 129
+V
Sbjct: 148 VV 149
>UniRef100_Q6P9S8 BC060615 protein [Mus musculus]
Length = 832
Score = 40.0 bits (92), Expect = 0.027
Identities = 36/122 (29%), Positives = 61/122 (49%), Gaps = 19/122 (15%)
Query: 17 FKWG----QKRGKGGKKRDIQFYQSF-----TLRGVDYSLFDNVYVKNDFAEPRIGKIIK 67
+KW Q+RG GK R + FY++ TLR D ++F + N P IG+I
Sbjct: 677 WKWSGNPTQRRGMKGKARKL-FYKAIVRGKETLRIGDCAVFLSAGRPN---LPYIGRIES 732
Query: 68 IWETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCN 127
+WE+ +KV+WF+ P E +K N L+ +C + + + +++I+ KC
Sbjct: 733 LWESWGSNMVVKVKWFYHP-EETKLGKRQSDGKNALYQSCHEDE-----NDVQTISHKCQ 786
Query: 128 IV 129
+V
Sbjct: 787 VV 788
>UniRef100_Q6GQV4 Hypothetical protein BC060615 [Mus musculus]
Length = 1191
Score = 40.0 bits (92), Expect = 0.027
Identities = 36/122 (29%), Positives = 61/122 (49%), Gaps = 19/122 (15%)
Query: 17 FKWG----QKRGKGGKKRDIQFYQSF-----TLRGVDYSLFDNVYVKNDFAEPRIGKIIK 67
+KW Q+RG GK R + FY++ TLR D ++F + N P IG+I
Sbjct: 1036 WKWSGNPTQRRGMKGKARKL-FYKAIVRGKETLRIGDCAVFLSAGRPN---LPYIGRIES 1091
Query: 68 IWETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCN 127
+WE+ +KV+WF+ P E +K N L+ +C + + + +++I+ KC
Sbjct: 1092 LWESWGSNMVVKVKWFYHP-EETKLGKRQSDGKNALYQSCHEDE-----NDVQTISHKCQ 1145
Query: 128 IV 129
+V
Sbjct: 1146 VV 1147
>UniRef100_Q7PS94 ENSANGP00000005615 [Anopheles gambiae str. PEST]
Length = 529
Score = 39.7 bits (91), Expect = 0.035
Identities = 25/75 (33%), Positives = 38/75 (50%), Gaps = 4/75 (5%)
Query: 20 GQKRGKGGKKRDIQFYQSFTLRGVDYSLFDN-VYVKNDFAE-PRIGKIIKIWETPTLERK 77
G +R G K+ QFY+S S+ D+ V++ + P IG I +WET T
Sbjct: 360 GYRRSTGRVKK--QFYKSIQRGKETISVGDSAVFLSTGRPDRPYIGHIESMWETSTNNMV 417
Query: 78 IKVQWFFRPIEVSKC 92
++V+WF+ P E C
Sbjct: 418 VRVKWFYHPEETQDC 432
>UniRef100_Q6C8C8 Similar to sp|Q06488 Saccharomyces cerevisiae YLR357w RSC2 member
of RSC complex [Yarrowia lipolytica]
Length = 759
Score = 39.7 bits (91), Expect = 0.035
Identities = 26/98 (26%), Positives = 49/98 (49%), Gaps = 8/98 (8%)
Query: 41 RGVDYSLFDNVYVKN--DFAEPRIGKIIKIWETPTLERKIKVQWFFRPIEVSKCLTWIKI 98
RG Y + D V++ N D ++P IG+I +IW+ P ++ + W++RP + + +
Sbjct: 354 RGDIYRVGDWVHIHNMNDPSKPTIGQIFRIWQAPDGQKWVNACWYYRPEQTVHRVDKV-F 412
Query: 99 YFNELFFACGDGDGLATIHPLESIAGKCNIVCISKDSR 136
Y NE+ G H ++ I KC ++ ++ R
Sbjct: 413 YENEVV-----KSGQYRDHLVDEILEKCFVMFFTRYQR 445
>UniRef100_O65464 Hypothetical protein F21P8.10 [Arabidopsis thaliana]
Length = 360
Score = 39.3 bits (90), Expect = 0.046
Identities = 41/164 (25%), Positives = 70/164 (42%), Gaps = 22/164 (13%)
Query: 24 GKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKNDFAEPRIGKIIKIWETPTLER--KIKVQ 81
GKG KK+ Y++F Y L D+V + + E IIK T E K++VQ
Sbjct: 40 GKGEKKKC--HYKTFQFHANKYGLEDSVLLVPEDGEKPYVAIIKDIYTQRKEGHVKLEVQ 97
Query: 82 WFFRPIEVSKCL--TWIKIYFNELFFACGDGDGLATIHPLESIAGKCNIVCISKDSRNP- 138
W +RP EV K W +LF++ + A ES+ C + + ++ + P
Sbjct: 98 WLYRPEEVEKKYVGNWKSKGSRDLFYSFHRDEVFA-----ESVKDDCIVHFVQENKQIPN 152
Query: 139 ----------QLSDKVTWSAAFVFYRYFDVGQRKIVEEVDDKSI 172
+ D V + + FD+ Q++ ++ +K+I
Sbjct: 153 RRKHPGFIVQHVYDNVKKKLRKLTFNGFDLQQKREIDHFVEKTI 196
>UniRef100_Q6BVM6 Similar to CA4169|CaRSC2 Candida albicans CaRSC2 [Debaryomyces
hansenii]
Length = 801
Score = 39.3 bits (90), Expect = 0.046
Identities = 26/100 (26%), Positives = 48/100 (48%), Gaps = 10/100 (10%)
Query: 37 SFTLRGVDYSLFDNVYVKN--DFAEPRIGKIIKIWETPTLERKIKVQWFFRPIEVSKCLT 94
S + G Y + D V + N D +P +G+I ++W T + V W++RP + C
Sbjct: 380 SIDINGYSYKIGDWVLIDNPSDPNKPTVGQIFRLWSTEDGTKYTNVCWYYRPEQT--CHR 437
Query: 95 WIKIYF-NELFFACGDGDGLATIHPLESIAGKCNIVCISK 133
+ +++F NE+ C G H I G C ++ +++
Sbjct: 438 YDRLFFMNEV---CKTGQ--YRDHLASEIVGPCYVIFLTR 472
>UniRef100_O74964 Rsc1 protein [Schizosaccharomyces pombe]
Length = 803
Score = 39.3 bits (90), Expect = 0.046
Identities = 30/125 (24%), Positives = 59/125 (47%), Gaps = 10/125 (8%)
Query: 11 ESETLEFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKN--DFAEPRIGKIIKI 68
+SE ++ + K DIQ + ++ G ++ D V ++N D ++P + +I +I
Sbjct: 321 DSEFFNYETDSRASPQLPKNDIQ--PAVSIDGTLLNVGDWVLIRNPADSSKPIVSQIYRI 378
Query: 69 WETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCNI 128
W++ + V W+ RP + + Y NE+F L HP+ I G+C +
Sbjct: 379 WKSDDDINYVTVCWYLRPEQTVHRADAV-FYENEVF-----KTSLYRDHPVSEIVGRCFV 432
Query: 129 VCISK 133
+ I++
Sbjct: 433 MYITR 437
>UniRef100_Q5U629 KIAA1447 protein [Homo sapiens]
Length = 189
Score = 38.9 bits (89), Expect = 0.060
Identities = 36/122 (29%), Positives = 61/122 (49%), Gaps = 19/122 (15%)
Query: 17 FKWG----QKRGKGGKKRDIQFYQSF-----TLRGVDYSLFDNVYVKNDFAEPRIGKIIK 67
+KW Q+RG GK R + FY++ TLR D ++F + N P IG+I
Sbjct: 34 WKWSGNPTQRRGMKGKARKL-FYKAIVRGEETLRVGDCAVFLSAGRPN---LPYIGRIES 89
Query: 68 IWETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCN 127
+WE+ +KV+WF+ P E +K N L+ +C + + + +++I+ KC
Sbjct: 90 MWESWGSNMVVKVKWFYHP-EETKLGKRQCDGKNALYQSCHEDE-----NDVQTISHKCQ 143
Query: 128 IV 129
+V
Sbjct: 144 VV 145
>UniRef100_UPI000036632B UPI000036632B UniRef100 entry
Length = 797
Score = 38.5 bits (88), Expect = 0.078
Identities = 33/114 (28%), Positives = 59/114 (50%), Gaps = 15/114 (13%)
Query: 21 QKRGKGGKKRDIQFYQSF-----TLRGVDYSLFDNVYVKNDFAEPRIGKIIKIWETPTLE 75
Q+RG GK R + FY++ T+R D ++F + N P IG+I WE+ T
Sbjct: 649 QRRGLKGKARKL-FYKAIVRGRDTVRVGDCAVFLSAGRPN---LPYIGRIENFWESWTSS 704
Query: 76 RKIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCNIV 129
+KV+WF+ P E +K + + L+ +C + + + +++I+ KC +V
Sbjct: 705 MVVKVKWFYHP-EETKLGKRHQDGKHALYQSCHEDE-----NDVQTISHKCQVV 752
>UniRef100_Q8MSX3 LD15342p [Drosophila melanogaster]
Length = 1322
Score = 38.1 bits (87), Expect = 0.10
Identities = 23/74 (31%), Positives = 40/74 (53%), Gaps = 3/74 (4%)
Query: 21 QKRGKGGKKRDIQFYQSFTLRGVDYSLFDN-VYVKNDFAE-PRIGKIIKIWETPTLERKI 78
+K G G+ R QFY++ ++ D+ V++ + P IG+I +WET T + +
Sbjct: 1153 RKAGVKGRARK-QFYKTIKRGKETITVGDSAVFLSTGRPDRPYIGRIESMWETTTGNKVV 1211
Query: 79 KVQWFFRPIEVSKC 92
+V WF+ P E + C
Sbjct: 1212 RVAWFYHPEETTGC 1225
>UniRef100_Q7KS56 CG31151-PA, isoform A [Drosophila melanogaster]
Length = 1543
Score = 38.1 bits (87), Expect = 0.10
Identities = 23/74 (31%), Positives = 40/74 (53%), Gaps = 3/74 (4%)
Query: 21 QKRGKGGKKRDIQFYQSFTLRGVDYSLFDN-VYVKNDFAE-PRIGKIIKIWETPTLERKI 78
+K G G+ R QFY++ ++ D+ V++ + P IG+I +WET T + +
Sbjct: 1374 RKAGVKGRARK-QFYKTIKRGKETITVGDSAVFLSTGRPDRPYIGRIESMWETTTGNKVV 1432
Query: 79 KVQWFFRPIEVSKC 92
+V WF+ P E + C
Sbjct: 1433 RVAWFYHPEETTGC 1446
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.140 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 337,558,358
Number of Sequences: 2790947
Number of extensions: 14052097
Number of successful extensions: 25889
Number of sequences better than 10.0: 83
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 67
Number of HSP's that attempted gapping in prelim test: 25843
Number of HSP's gapped (non-prelim): 101
length of query: 178
length of database: 848,049,833
effective HSP length: 119
effective length of query: 59
effective length of database: 515,927,140
effective search space: 30439701260
effective search space used: 30439701260
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 70 (31.6 bits)
Medicago: description of AC148532.15


