

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148526.1 + phase: 0
(598 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
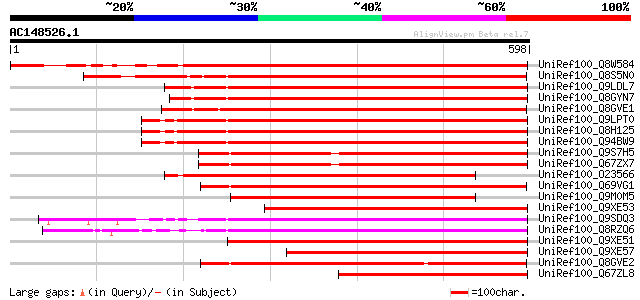
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8W584 AT4g17230/dl4650c [Arabidopsis thaliana] 486 e-135
UniRef100_Q8S5N0 Putative SCARECROW gene regulator-like [Oryza s... 483 e-135
UniRef100_Q9LDL7 SCARECROW gene regulator-like [Arabidopsis thal... 467 e-130
UniRef100_Q8GYN7 Putative SCARECROW gene regulator [Arabidopsis ... 464 e-129
UniRef100_Q8GVE1 Chitin-inducible gibberellin-responsive protein... 454 e-126
UniRef100_Q9LPT0 Putative transcription factor [Arabidopsis thal... 439 e-122
UniRef100_Q8H125 Putative scarecrow protein [Arabidopsis thaliana] 439 e-122
UniRef100_Q94BW9 F17J6.12/F17J6.12 [Arabidopsis thaliana] 439 e-121
UniRef100_Q9S7H5 Putative SCARECROW gene regulator [Arabidopsis ... 435 e-120
UniRef100_Q67ZX7 Putative SCARECROW gene regulator [Arabidopsis ... 434 e-120
UniRef100_O23566 SCARECROW like protein [Arabidopsis thaliana] 429 e-118
UniRef100_Q69VG1 Putative chitin-inducible gibberellin-responsiv... 404 e-111
UniRef100_Q9M0M5 Scarecrow-like 13 [Arabidopsis thaliana] 391 e-107
UniRef100_Q9XE53 Scarecrow-like 5 [Arabidopsis thaliana] 367 e-100
UniRef100_Q9SDQ3 Scarecrow-like 1 [Arabidopsis thaliana] 355 2e-96
UniRef100_Q8RZQ6 Scarecrow-like protein [Oryza sativa] 351 3e-95
UniRef100_Q9XE51 Scarecrow-like 1 [Arabidopsis thaliana] 332 1e-89
UniRef100_Q9XE57 Scarecrow-like 13 [Arabidopsis thaliana] 330 6e-89
UniRef100_Q8GVE2 Chitin-inducible gibberellin-responsive protein... 298 2e-79
UniRef100_Q67ZL8 Putative SCARECROW gene regulator [Arabidopsis ... 281 3e-74
>UniRef100_Q8W584 AT4g17230/dl4650c [Arabidopsis thaliana]
Length = 529
Score = 486 bits (1250), Expect = e-135
Identities = 278/600 (46%), Positives = 368/600 (61%), Gaps = 79/600 (13%)
Query: 1 MYTSRKKPNSAGIQLYYQQQSLR-QIDSNKNR-FKTLQTNSNNGGRRQRTNVSLETSNWE 58
M TS+K ++AG+ + Y Q Q + N+ F + + N
Sbjct: 1 MQTSQKHHSAAGLHMLYPQVYCSPQFQAKDNKGFSDIPSKEN------------------ 42
Query: 59 SNWEKQFCTLESSQVANDVIAFDSPSSA--SGDPFNMRSTFSPQISRYCSSDNTTNVSPL 116
F TLESS + + ++DSPS + SG +S FSPQ S+ C SD
Sbjct: 43 ------FFTLESSTASGSLPSYDSPSVSITSG-----QSPFSPQGSQSCISD-------- 83
Query: 117 GVYSSADDDSFEFKMKEIELSLVGTDIEVFGSCLSNSNGSLHQGTSQDNEFHIDEMISEQ 176
++ S D+ V+GS LS + + DE +
Sbjct: 84 -LHHSPDN--------------------VYGSPLSGVSSLAY-----------DEAGVKS 111
Query: 177 GFKESEISLLGPSSDIVDSCQSNLNGSLHQGTSQYDWSQFEEIIPKLDLKEELIRCAQFV 236
+E E+SLL + + + +G ++W + + P+LDLKE L+ A+ V
Sbjct: 112 KIRELEVSLLSGDTKVEE-----FSGFSPAAGKSWNWDELLALTPQLDLKEVLVEAARAV 166
Query: 237 FDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGSAIYKALKCEEPTS 296
DGDF A GF++ VL +MVSV+GSPIQRLG YM EGLRAR+E SGS IYK+LKC EPT
Sbjct: 167 ADGDFATAYGFLD-VLEQMVSVSGSPIQRLGTYMAEGLRARLEGSGSNIYKSLKCNEPTG 225
Query: 297 IELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKH 356
ELMS M +LY+ICPY++FAY ++N I E + E+R+HIIDFQIAQGSQ+M L+ L
Sbjct: 226 RELMSYMSVLYEICPYWKFAYTTANVEILEAIAGETRVHIIDFQIAQGSQYMFLIQELAK 285
Query: 357 KPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLED 416
+PGGPP +RVTG+DDSQS +ARGG L +VG++L A++C VPFEF+ M GC+VQ E
Sbjct: 286 RPGGPPLLRVTGVDDSQSTYARGGGLSLVGERLATLAQSCGVPFEFHDAIMSGCKVQREH 345
Query: 417 FEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPF 476
++ +VVNFP+ LHH+PDESVS+ENHRDRLL L+K LSPK+V VEQESNTNTSPF
Sbjct: 346 LGLEPGFAVVVNFPYVLHHMPDESVSVENHRDRLLHLIKSLSPKLVTLVEQESNTNTSPF 405
Query: 477 LPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFG 536
L RF ETL+YYTAMFESID A PRDDK+RI+AEQHCVARDIVN+IACE ER ERHE+ G
Sbjct: 406 LSRFVETLDYYTAMFESIDAARPRDDKQRISAEQHCVARDIVNMIACEESERVERHEVLG 465
Query: 537 KWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKDYRIEQTDVAINLAWKSKVMCTSSAWR 596
KW+ R MAGF +S S + +LK ++K+Y++ + A+ L WK + M T S W+
Sbjct: 466 KWRVRMMMAGFTGWPVSTSAAFAASEMLKAYDKNYKLGGHEGALYLFWKRRPMATCSVWK 525
>UniRef100_Q8S5N0 Putative SCARECROW gene regulator-like [Oryza sativa]
Length = 524
Score = 483 bits (1243), Expect = e-135
Identities = 266/515 (51%), Positives = 350/515 (67%), Gaps = 24/515 (4%)
Query: 86 ASGDPFNMRSTFSPQI--SRYCSSDNTTNVSPLGVYSSADDDSFEFKMKEIELSLVGTDI 143
AS D + + SPQ + YC+ ++++ +SSA S
Sbjct: 28 ASSDDGSQKIGSSPQAFEAPYCTLESSSANGAHPAHSSASSHSIS--------------- 72
Query: 144 EVFGSCLSNSNG-SLHQGTSQDNEFHIDEMISEQ-GFKESEISLLGPSSDIV-DSCQSNL 200
+ GS LS+ + S H S + + E+ Q +E E ++LGP DI DS +S L
Sbjct: 73 PISGSPLSHHDSHSDHTYNSPPSASCVTEITDLQIKLRELENAILGPELDIAYDSPESAL 132
Query: 201 NGSLHQGTSQYDWSQFEEIIPKLDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAG 260
++ + +W Q I DLK+ +I C + V + D + +++ LG+MVSV+G
Sbjct: 133 QPNIM--ATPENWRQLLGINTG-DLKQVIIACGKAVAENDVRLTELLISE-LGQMVSVSG 188
Query: 261 SPIQRLGAYMLEGLRARVESSGSAIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISS 320
P+QRLGAYMLEGL AR+ SSGS IYK+LKC+EPTS ELMS MH+LY+ICP+F+F Y+S+
Sbjct: 189 DPLQRLGAYMLEGLVARLSSSGSKIYKSLKCKEPTSSELMSYMHLLYEICPFFKFGYMSA 248
Query: 321 NAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGG 380
N I E ++ E+ +HIIDFQIAQGSQWM L+ AL +PGGPPF+R+TGIDDS S +ARGG
Sbjct: 249 NGAIAEAIKGENFVHIIDFQIAQGSQWMTLIQALAARPGGPPFLRITGIDDSNSAYARGG 308
Query: 381 KLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDES 440
LDIVG +L A++ +PFEFN+V EV LE +++ EV+VVNF + LHH PDES
Sbjct: 309 GLDIVGMRLYKVAQSFGLPFEFNAVPAASHEVYLEHLDIRVGEVIVVNFAYQLHHTPDES 368
Query: 441 VSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPR 500
VS ENHRDR+LR+VK LSP++V VEQESNTNT PF PR+ ETL+YYTAMFESIDVALPR
Sbjct: 369 VSTENHRDRILRMVKSLSPRLVTLVEQESNTNTRPFFPRYLETLDYYTAMFESIDVALPR 428
Query: 501 DDKKRINAEQHCVARDIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSV 560
DDK+R++AEQHCVARDIVN+IACEG ER ERHE+FGKWKAR +MAGF P LS V ++
Sbjct: 429 DDKRRMSAEQHCVARDIVNLIACEGAERVERHEVFGKWKARLTMAGFRPYPLSSVVNSTI 488
Query: 561 RTLLKDFNKDYRIEQTDVAINLAWKSKVMCTSSAW 595
+TLL +N YR+E+ D + L WK++V+ SSAW
Sbjct: 489 KTLLHTYNSFYRLEERDGVLYLGWKNRVLVVSSAW 523
>UniRef100_Q9LDL7 SCARECROW gene regulator-like [Arabidopsis thaliana]
Length = 490
Score = 467 bits (1202), Expect = e-130
Identities = 235/420 (55%), Positives = 302/420 (70%), Gaps = 5/420 (1%)
Query: 179 KESEISLLGPSS-DIVDSCQSNLNGSLHQGTSQYDWSQFEEIIPKLDLKEELIRCAQFVF 237
+E E ++GP S D++ C + + + Q + W E I + DL+ +L+ CA+ +
Sbjct: 74 REIETVMMGPDSLDLLVDCTDSFDSTASQEIN--GWRSTLEAISRRDLRADLVSCAKAMS 131
Query: 238 DGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGSAIYKAL-KCEEPTS 296
+ D A M K L +MVSV+G PIQRLGAY+LEGL A++ SSGS+IYKAL +C EP S
Sbjct: 132 ENDLMMAHSMMEK-LRQMVSVSGEPIQRLGAYLLEGLVAQLASSGSSIYKALNRCPEPAS 190
Query: 297 IELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKH 356
EL+S MHILY++CPYF+F Y+S+N I E M+ E+R+HIIDFQI QGSQW+ L+ A
Sbjct: 191 TELLSYMHILYEVCPYFKFGYMSANGAIAEAMKEENRVHIIDFQIGQGSQWVTLIQAFAA 250
Query: 357 KPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLED 416
+PGGPP IR+TGIDD S +ARGG L IVG +L AK VPFEFNSV + EV+ ++
Sbjct: 251 RPGGPPRIRITGIDDMTSAYARGGGLSIVGNRLAKLAKQFNVPFEFNSVSVSVSEVKPKN 310
Query: 417 FEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPF 476
V+ E L VNF F LHH+PDESVS ENHRDRLLR+VK LSPKVV VEQESNTNT+ F
Sbjct: 311 LGVRPGEALAVNFAFVLHHMPDESVSTENHRDRLLRMVKSLSPKVVTLVEQESNTNTAAF 370
Query: 477 LPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFG 536
PRF ET+NYY AMFESIDV LPRD K+RIN EQHC+ARD+VNIIACEG +R ERHEL G
Sbjct: 371 FPRFMETMNYYAAMFESIDVTLPRDHKQRINVEQHCLARDVVNIIACEGADRVERHELLG 430
Query: 537 KWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKDYRIEQTDVAINLAWKSKVMCTSSAWR 596
KW++RF MAGF P LSP V ++++LL++++ YR+E+ D A+ L W + + S AW+
Sbjct: 431 KWRSRFGMAGFTPYPLSPLVNSTIKSLLRNYSDKYRLEERDGALYLGWMHRDLVASCAWK 490
>UniRef100_Q8GYN7 Putative SCARECROW gene regulator [Arabidopsis thaliana]
Length = 411
Score = 464 bits (1195), Expect = e-129
Identities = 233/414 (56%), Positives = 299/414 (71%), Gaps = 5/414 (1%)
Query: 185 LLGPSS-DIVDSCQSNLNGSLHQGTSQYDWSQFEEIIPKLDLKEELIRCAQFVFDGDFQK 243
++GP S D++ C + + + Q + W E I + DL+ +L+ CA+ + + D
Sbjct: 1 MMGPDSLDLLVDCTDSFDSTASQEIN--GWRSTLEAISRRDLRADLVSCAKAMSENDLMM 58
Query: 244 AIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGSAIYKAL-KCEEPTSIELMSA 302
A M K L +MVSV+G PIQRLGAY+LEGL A++ SSGS+IYKAL +C EP S EL+S
Sbjct: 59 AHSMMEK-LRQMVSVSGEPIQRLGAYLLEGLVAQLASSGSSIYKALNRCPEPASTELLSY 117
Query: 303 MHILYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKHKPGGPP 362
MHILY++CPYF+F Y+S+N I E M+ E+R+HIIDFQI QGSQW+ L+ A +PGGPP
Sbjct: 118 MHILYEVCPYFKFGYMSANGAIAEAMKEENRVHIIDFQIGQGSQWVTLIQAFAARPGGPP 177
Query: 363 FIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHD 422
IR+TGIDD S +ARGG L IVG +L AK VPFEFNSV + EV+ ++ V+
Sbjct: 178 RIRITGIDDMTSAYARGGGLSIVGNRLAKLAKQFNVPFEFNSVSVSVSEVKPKNLGVRPG 237
Query: 423 EVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAE 482
E L VNF F LHH+PDESVS ENHRDRLLR+VK LSPKVV VEQESNTNT+ F PRF E
Sbjct: 238 EALAVNFAFVLHHMPDESVSTENHRDRLLRMVKSLSPKVVTLVEQESNTNTAAFFPRFME 297
Query: 483 TLNYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFGKWKARF 542
T+NYY AMFESIDV LPRD K+RIN EQHC+ARD+VNIIACEG +R ERHEL GKW++RF
Sbjct: 298 TMNYYAAMFESIDVTLPRDHKQRINVEQHCLARDVVNIIACEGADRVERHELLGKWRSRF 357
Query: 543 SMAGFVPLLLSPSVIDSVRTLLKDFNKDYRIEQTDVAINLAWKSKVMCTSSAWR 596
MAGF P LSP V ++++LL++++ YR+E+ D A+ L W + + S AW+
Sbjct: 358 GMAGFTPYPLSPLVNSTIKSLLRNYSDKYRLEERDGALYLGWMHRDLVASCAWK 411
>UniRef100_Q8GVE1 Chitin-inducible gibberellin-responsive protein [Oryza sativa]
Length = 544
Score = 454 bits (1169), Expect = e-126
Identities = 233/421 (55%), Positives = 307/421 (72%), Gaps = 4/421 (0%)
Query: 175 EQGFKESEISLLGPSSDIVDSCQSNLNGSLHQGTSQYDWSQFEEIIPKLDLKEELIRCAQ 234
+Q K+ E +LGP S+IV+S ++++ L + W + I P+ +LKE LI CA+
Sbjct: 127 KQKLKDLEAVMLGPDSEIVNSLENSVANQLSLEPEK--WVRMMGI-PRGNLKELLIACAR 183
Query: 235 FVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGSAIYKALKCEEP 294
V + + AI M L K+VSV+G P++RLGAYM+EGL AR+ SSG +IYKALKC+EP
Sbjct: 184 AVEEKN-SFAIDMMIPELRKIVSVSGEPLERLGAYMVEGLVARLASSGISIYKALKCKEP 242
Query: 295 TSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHAL 354
S +L+S MH LY+ CPYF+F Y+S+N I E ++ E RIHIIDF I+QG+QW+ LL AL
Sbjct: 243 KSSDLLSYMHFLYEACPYFKFGYMSANGAIAEAVKGEDRIHIIDFHISQGAQWISLLQAL 302
Query: 355 KHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQL 414
+PGGPP +R+TGIDDS S +ARGG L++VG++L A CKVPFEF+ + + G +V+
Sbjct: 303 AARPGGPPTVRITGIDDSVSAYARGGGLELVGRRLSHIASLCKVPFEFHPLAISGSKVEA 362
Query: 415 EDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTS 474
V E L VNF LHHIPDESVS NHRDRLLR+VK LSPKV+ VE ESNTNT+
Sbjct: 363 AHLGVIPGEALAVNFTLELHHIPDESVSTANHRDRLLRMVKSLSPKVLTLVEMESNTNTA 422
Query: 475 PFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHEL 534
PF RFAETL+YYTA+FESID+ LPRDD++RIN EQHC+AR+IVN+IACEG+ER ER+E
Sbjct: 423 PFPQRFAETLDYYTAIFESIDLTLPRDDRERINMEQHCLAREIVNLIACEGEERAERYEP 482
Query: 535 FGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKDYRIEQTDVAINLAWKSKVMCTSSA 594
FGKWKAR +MAGF P LS V ++RTLL+ ++ +Y++ + D A+ L WKS+ + SSA
Sbjct: 483 FGKWKARLTMAGFRPSPLSSLVNATIRTLLQSYSDNYKLAERDGALYLGWKSRPLVVSSA 542
Query: 595 W 595
W
Sbjct: 543 W 543
>UniRef100_Q9LPT0 Putative transcription factor [Arabidopsis thaliana]
Length = 526
Score = 439 bits (1130), Expect = e-122
Identities = 230/447 (51%), Positives = 313/447 (69%), Gaps = 11/447 (2%)
Query: 152 NSNGSLHQGTSQD-NEFHIDEMISEQGFKESEISLLGPSSDIVDSCQSNLNGSLHQ-GTS 209
N+N L ++ + NE + M+ K+ E +++ P VD+ +N G Q G
Sbjct: 89 NNNSPLSGSSATNTNETELSLML-----KDLETAMMEPD---VDNSYNNQGGFGQQHGVV 140
Query: 210 QYDWSQFEEIIPKLDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAY 269
+ E+I + DLK L CA+ V + D + +++ L +MVSV+G P+QRLGAY
Sbjct: 141 SSAMYRSMEMISRGDLKGVLYECAKAVENYDLEMTDWLISQ-LQQMVSVSGEPVQRLGAY 199
Query: 270 MLEGLRARVESSGSAIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQ 329
MLEGL AR+ SSGS+IYKAL+C++PT EL++ MHILY+ CPYF+F Y S+N I E ++
Sbjct: 200 MLEGLVARLASSGSSIYKALRCKDPTGPELLTYMHILYEACPYFKFGYESANGAIAEAVK 259
Query: 330 NESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKL 389
NES +HIIDFQI+QG QW+ L+ AL +PGGPP +R+TGIDD +S AR G L++VG++L
Sbjct: 260 NESFVHIIDFQISQGGQWVSLIRALGARPGGPPNVRITGIDDPRSSFARQGGLELVGQRL 319
Query: 390 EDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDR 449
A+ C VPFEF+ + EV++E V++ E L VNFP LHH+PDESV++ENHRDR
Sbjct: 320 GKLAEMCGVPFEFHGAALCCTEVEIEKLGVRNGEALAVNFPLVLHHMPDESVTVENHRDR 379
Query: 450 LLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAE 509
LLRLVK LSP VV VEQE+NTNT+PFLPRF ET+N+Y A+FESIDV L RD K+RIN E
Sbjct: 380 LLRLVKHLSPNVVTLVEQEANTNTAPFLPRFVETMNHYLAVFESIDVKLARDHKERINVE 439
Query: 510 QHCVARDIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNK 569
QHC+AR++VN+IACEG ER ERHE GKW++RF MAGF P LS V +++ LL+ +++
Sbjct: 440 QHCLAREVVNLIACEGVEREERHEPLGKWRSRFHMAGFKPYPLSSYVNATIKGLLESYSE 499
Query: 570 DYRIEQTDVAINLAWKSKVMCTSSAWR 596
Y +E+ D A+ L WK++ + TS AWR
Sbjct: 500 KYTLEERDGALYLGWKNQPLITSCAWR 526
>UniRef100_Q8H125 Putative scarecrow protein [Arabidopsis thaliana]
Length = 597
Score = 439 bits (1130), Expect = e-122
Identities = 230/447 (51%), Positives = 313/447 (69%), Gaps = 11/447 (2%)
Query: 152 NSNGSLHQGTSQD-NEFHIDEMISEQGFKESEISLLGPSSDIVDSCQSNLNGSLHQ-GTS 209
N+N L ++ + NE + M+ K+ E +++ P VD+ +N G Q G
Sbjct: 160 NNNSPLSGSSATNTNETELSLML-----KDLETAMMEPD---VDNSYNNQGGFGQQHGVV 211
Query: 210 QYDWSQFEEIIPKLDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAY 269
+ E+I + DLK L CA+ V + D + +++ L +MVSV+G P+QRLGAY
Sbjct: 212 SSAMYRSMEMISRGDLKGVLYECAKAVENYDLEMTDWLISQ-LQQMVSVSGEPVQRLGAY 270
Query: 270 MLEGLRARVESSGSAIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQ 329
MLEGL AR+ SSGS+IYKAL+C++PT EL++ MHILY+ CPYF+F Y S+N I E ++
Sbjct: 271 MLEGLVARLASSGSSIYKALRCKDPTGPELLTYMHILYEACPYFKFGYESANGAIAEAVK 330
Query: 330 NESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKL 389
NES +HIIDFQI+QG QW+ L+ AL +PGGPP +R+TGIDD +S AR G L++VG++L
Sbjct: 331 NESFVHIIDFQISQGGQWVSLIRALGARPGGPPNVRITGIDDPRSSFARQGGLELVGQRL 390
Query: 390 EDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDR 449
A+ C VPFEF+ + EV++E V++ E L VNFP LHH+PDESV++ENHRDR
Sbjct: 391 GKLAEMCGVPFEFHGAALCCTEVEIEKLGVRNGEALAVNFPLVLHHMPDESVTVENHRDR 450
Query: 450 LLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAE 509
LLRLVK LSP VV VEQE+NTNT+PFLPRF ET+N+Y A+FESIDV L RD K+RIN E
Sbjct: 451 LLRLVKHLSPNVVTLVEQEANTNTAPFLPRFVETMNHYLAVFESIDVKLARDHKERINVE 510
Query: 510 QHCVARDIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNK 569
QHC+AR++VN+IACEG ER ERHE GKW++RF MAGF P LS V +++ LL+ +++
Sbjct: 511 QHCLAREVVNLIACEGVEREERHEPLGKWRSRFHMAGFKPYPLSSYVNATIKGLLESYSE 570
Query: 570 DYRIEQTDVAINLAWKSKVMCTSSAWR 596
Y +E+ D A+ L WK++ + TS AWR
Sbjct: 571 KYTLEERDGALYLGWKNQPLITSCAWR 597
>UniRef100_Q94BW9 F17J6.12/F17J6.12 [Arabidopsis thaliana]
Length = 526
Score = 439 bits (1128), Expect = e-121
Identities = 230/447 (51%), Positives = 312/447 (69%), Gaps = 11/447 (2%)
Query: 152 NSNGSLHQGTSQD-NEFHIDEMISEQGFKESEISLLGPSSDIVDSCQSNLNGSLHQ-GTS 209
N+N L ++ + NE + M+ K+ E +++ P VD+ +N G Q G
Sbjct: 89 NNNSPLSGSSATNTNETELSLML-----KDLETAMMEPD---VDNSYNNQGGFGQQHGVV 140
Query: 210 QYDWSQFEEIIPKLDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAY 269
+ E+I + DLK L CA+ V + D + +++ L +MVSV+G P+QRLGAY
Sbjct: 141 SSAMYRSMEMISRGDLKGVLYECAKAVENYDLEMTDWLISQ-LQQMVSVSGEPVQRLGAY 199
Query: 270 MLEGLRARVESSGSAIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQ 329
MLEGL AR+ SSGS+IYKAL+C++PT EL++ MHILY+ CPYF+F Y S+N I E ++
Sbjct: 200 MLEGLVARLASSGSSIYKALRCKDPTGPELLTYMHILYEACPYFKFGYESANGAIAEAVK 259
Query: 330 NESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKL 389
NES +HIIDFQI+QG QW+ L+ AL +PGGPP +R+TGIDD +S AR G L++VG++L
Sbjct: 260 NESFVHIIDFQISQGGQWVSLIRALGARPGGPPNVRITGIDDPRSSFARQGGLELVGQRL 319
Query: 390 EDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDR 449
A+ C VPFEF+ + EV++E V++ E L VNFP LHH+PDESV++ENHRDR
Sbjct: 320 GKLAEMCGVPFEFHGAALCCTEVEIEKLGVRNGEALAVNFPLVLHHMPDESVTVENHRDR 379
Query: 450 LLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAE 509
LLRLVK LSP VV VEQE+NTNT+PFLPRF ET+N+Y A+FESIDV L RD K+RIN E
Sbjct: 380 LLRLVKHLSPNVVTLVEQEANTNTAPFLPRFVETMNHYLAVFESIDVKLARDHKERINVE 439
Query: 510 QHCVARDIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNK 569
QHC+AR++VN+IACEG ER ERHE GKW++RF MAGF P LS V ++ LL+ +++
Sbjct: 440 QHCLAREVVNLIACEGVEREERHEPLGKWRSRFHMAGFKPYPLSSYVNATIEGLLESYSE 499
Query: 570 DYRIEQTDVAINLAWKSKVMCTSSAWR 596
Y +E+ D A+ L WK++ + TS AWR
Sbjct: 500 KYTLEERDGALYLGWKNQPLITSCAWR 526
>UniRef100_Q9S7H5 Putative SCARECROW gene regulator [Arabidopsis thaliana]
Length = 413
Score = 435 bits (1119), Expect = e-120
Identities = 218/379 (57%), Positives = 278/379 (72%), Gaps = 8/379 (2%)
Query: 218 EIIPKLDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRAR 277
E I + DLK L+ CA+ V + + A M ++ G MVS++G PIQRLGAYMLEGL AR
Sbjct: 43 EAISRGDLKLVLVACAKAVSENNLLMARWCMGELRG-MVSISGEPIQRLGAYMLEGLVAR 101
Query: 278 VESSGSAIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHII 337
+ +SGS+IYK+L+ EP S E +S +++L+++CPYF+F Y+S+N I E M++E RIHII
Sbjct: 102 LAASGSSIYKSLQSREPESYEFLSYVYVLHEVCPYFKFGYMSANGAIAEAMKDEERIHII 161
Query: 338 DFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCK 397
DFQI QGSQW+ L+ A +PGG P IR+TG+ D G L V K+LE AK
Sbjct: 162 DFQIGQGSQWIALIQAFAARPGGAPNIRITGVGD-------GSVLVTVKKRLEKLAKKFD 214
Query: 398 VPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKIL 457
VPF FN+V CEV++E+ +V+ E L VNF + LHH+PDESVSMENHRDRLLR+VK L
Sbjct: 215 VPFRFNAVSRPSCEVEVENLDVRDGEALGVNFAYMLHHLPDESVSMENHRDRLLRMVKSL 274
Query: 458 SPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDI 517
SPKVV VEQE NTNTSPFLPRF ETL+YYTAMFESIDV LPR+ K+RIN EQHC+ARD+
Sbjct: 275 SPKVVTLVEQECNTNTSPFLPRFLETLSYYTAMFESIDVMLPRNHKERINIEQHCMARDV 334
Query: 518 VNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKDYRIEQTD 577
VNIIACEG ER ERHEL GKWK+RFSMAGF P LS + ++R LL+D++ Y IE+ D
Sbjct: 335 VNIIACEGAERIERHELLGKWKSRFSMAGFEPYPLSSIISATIRALLRDYSNGYAIEERD 394
Query: 578 VAINLAWKSKVMCTSSAWR 596
A+ L W +++ +S AW+
Sbjct: 395 GALYLGWMDRILVSSCAWK 413
>UniRef100_Q67ZX7 Putative SCARECROW gene regulator [Arabidopsis thaliana]
Length = 413
Score = 434 bits (1116), Expect = e-120
Identities = 217/379 (57%), Positives = 278/379 (73%), Gaps = 8/379 (2%)
Query: 218 EIIPKLDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRAR 277
E I + DLK L+ CA+ V + + A M ++ G MVS++G PIQRLGAYMLEGL AR
Sbjct: 43 EAISRGDLKLVLVACAKAVSENNLLMARWCMGELRG-MVSISGEPIQRLGAYMLEGLVAR 101
Query: 278 VESSGSAIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHII 337
+ +SGS+IYK+L+ EP S E +S +++L+++CPYF+F Y+S+N I E M++E RIHII
Sbjct: 102 LAASGSSIYKSLQSREPESYEFLSYVYVLHEVCPYFKFGYMSANGAIAEAMKDEERIHII 161
Query: 338 DFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCK 397
DFQI QGSQW+ L+ A +PGG P IR+TG+ D G L V K+LE AK
Sbjct: 162 DFQIGQGSQWIALIQAFAARPGGAPNIRITGVGD-------GSVLVTVKKRLEKLAKKFD 214
Query: 398 VPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKIL 457
VPF FN+V CEV++E+ +V+ E L VNF + LHH+PDESVSMENHRDRLLR+VK L
Sbjct: 215 VPFRFNAVSRPSCEVEVENLDVRDGEALGVNFAYMLHHLPDESVSMENHRDRLLRMVKSL 274
Query: 458 SPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDI 517
SPKVV VEQE NTNTSPFLPRF ETL+YYTAMFESIDV LPR+ K+RIN EQHC+ARD+
Sbjct: 275 SPKVVTLVEQECNTNTSPFLPRFLETLSYYTAMFESIDVMLPRNHKERINIEQHCMARDV 334
Query: 518 VNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKDYRIEQTD 577
VNI+ACEG ER ERHEL GKWK+RFSMAGF P LS + ++R LL+D++ Y IE+ D
Sbjct: 335 VNIMACEGAERIERHELLGKWKSRFSMAGFEPYPLSSIISATIRALLRDYSNGYAIEERD 394
Query: 578 VAINLAWKSKVMCTSSAWR 596
A+ L W +++ +S AW+
Sbjct: 395 GALYLGWMDRILVSSCAWK 413
>UniRef100_O23566 SCARECROW like protein [Arabidopsis thaliana]
Length = 375
Score = 429 bits (1102), Expect = e-118
Identities = 215/358 (60%), Positives = 268/358 (74%), Gaps = 6/358 (1%)
Query: 179 KESEISLLGPSSDIVDSCQSNLNGSLHQGTSQYDWSQFEEIIPKLDLKEELIRCAQFVFD 238
+E E+SLL + + + +G ++W + + P+LDLKE L+ A+ V D
Sbjct: 19 RELEVSLLSGDTKVEE-----FSGFSPAAGKSWNWDELLALTPQLDLKEVLVEAARAVAD 73
Query: 239 GDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGSAIYKALKCEEPTSIE 298
GDF A GF++ VL +MVSV+GSPIQRLG YM EGLRAR+E SGS IYK+LKC EPT E
Sbjct: 74 GDFATAYGFLD-VLEQMVSVSGSPIQRLGTYMAEGLRARLEGSGSNIYKSLKCNEPTGRE 132
Query: 299 LMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKHKP 358
LMS M +LY+ICPY++FAY ++N I E + E+R+HIIDFQIAQGSQ+M L+ L P
Sbjct: 133 LMSYMSVLYEICPYWKFAYTTANVEILEAIAGETRVHIIDFQIAQGSQYMFLIQELAKHP 192
Query: 359 GGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLEDFE 418
GGPP +RVTG+DDSQS +ARGG L +VG++L A++C VPFEF+ M GC+VQ E
Sbjct: 193 GGPPLLRVTGVDDSQSTYARGGGLSLVGERLATLAQSCGVPFEFHDAIMSGCKVQREHLG 252
Query: 419 VQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLP 478
++ +VVNFP+ LHH+PDESVS+ENHRDRLL L+K LSPK+V VEQESNTNTSPFL
Sbjct: 253 LEPGFAVVVNFPYVLHHMPDESVSVENHRDRLLHLIKSLSPKLVTLVEQESNTNTSPFLS 312
Query: 479 RFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFG 536
RF ETL+YYTAMFESID A PRDDK+RI+AEQHCVARDIVN+IACE ER ERHE+ G
Sbjct: 313 RFVETLDYYTAMFESIDAARPRDDKQRISAEQHCVARDIVNMIACEESERVERHEVLG 370
>UniRef100_Q69VG1 Putative chitin-inducible gibberellin-responsive protein [Oryza
sativa]
Length = 571
Score = 404 bits (1039), Expect = e-111
Identities = 202/375 (53%), Positives = 269/375 (70%), Gaps = 1/375 (0%)
Query: 221 PKLDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVES 280
P++ +K+ L RCA+ + + ++ + + G +VS+ G PIQRLGAY+LEGL AR +
Sbjct: 197 PQIIVKQLLTRCAEALSEDRTEEFHKLVQEARG-VVSINGEPIQRLGAYLLEGLVARHGN 255
Query: 281 SGSAIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQ 340
SG+ IY+ALKC EP S EL+S M ILY ICPYF+F Y+++N I E ++ E+ IHIIDFQ
Sbjct: 256 SGTNIYRALKCREPESKELLSYMRILYNICPYFKFGYMAANGAIAEALRTENNIHIIDFQ 315
Query: 341 IAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPF 400
IAQG+QW+ L+ AL +PGGPP +R+TGIDD S +ARG LDIVGK L+ ++ K+P
Sbjct: 316 IAQGTQWITLIQALAARPGGPPRVRITGIDDPVSEYARGEGLDIVGKMLKSMSEEFKIPL 375
Query: 401 EFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPK 460
EF + +Y +V E E++ E L VNF LHH PDESV + N RD LLR+VK LSPK
Sbjct: 376 EFTPLSVYATQVTKEMLEIRPGEALSVNFTLQLHHTPDESVDVNNPRDGLLRMVKGLSPK 435
Query: 461 VVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNI 520
V VEQES+TNT+PFL RF ET+ YY+AMFESID LPRD+K+RI+ EQHC+A+DIVNI
Sbjct: 436 VTTLVEQESHTNTTPFLMRFGETMEYYSAMFESIDANLPRDNKERISVEQHCLAKDIVNI 495
Query: 521 IACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKDYRIEQTDVAI 580
IACEG +R ERHEL GKWK+R +MAGF P LS V +R LL ++ Y +++ D A+
Sbjct: 496 IACEGKDRVERHELLGKWKSRLTMAGFRPYPLSSYVNSVIRKLLACYSDKYTLDEKDGAM 555
Query: 581 NLAWKSKVMCTSSAW 595
L W+S+ + ++SAW
Sbjct: 556 LLGWRSRKLISASAW 570
>UniRef100_Q9M0M5 Scarecrow-like 13 [Arabidopsis thaliana]
Length = 287
Score = 391 bits (1004), Expect = e-107
Identities = 190/282 (67%), Positives = 227/282 (80%)
Query: 255 MVSVAGSPIQRLGAYMLEGLRARVESSGSAIYKALKCEEPTSIELMSAMHILYQICPYFQ 314
MVSV+GSPIQRLG YM EGLRAR+E SGS IYK+LKC EPT ELMS M +LY+ICPY++
Sbjct: 1 MVSVSGSPIQRLGTYMAEGLRARLEGSGSNIYKSLKCNEPTGRELMSYMSVLYEICPYWK 60
Query: 315 FAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQS 374
FAY ++N I E + E+R+HIIDFQIAQGSQ+M L+ L PGGPP +RVTG+DDSQS
Sbjct: 61 FAYTTANVEILEAIAGETRVHIIDFQIAQGSQYMFLIQELAKHPGGPPLLRVTGVDDSQS 120
Query: 375 FHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALH 434
+ARGG L +VG++L A++C VPFEF+ M GC+VQ E ++ +VVNFP+ LH
Sbjct: 121 TYARGGGLSLVGERLATLAQSCGVPFEFHDAIMSGCKVQREHLGLEPGFAVVVNFPYVLH 180
Query: 435 HIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESI 494
H+PDESVS+ENHRDRLL L+K LSPK+V VEQESNTNTSPFL RF ETL+YYTAMFESI
Sbjct: 181 HMPDESVSVENHRDRLLHLIKSLSPKLVTLVEQESNTNTSPFLSRFVETLDYYTAMFESI 240
Query: 495 DVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFG 536
D A PRDDK+RI+AEQHCVARDIVN+IACE ER ERHE+ G
Sbjct: 241 DAARPRDDKQRISAEQHCVARDIVNMIACEESERVERHEVLG 282
>UniRef100_Q9XE53 Scarecrow-like 5 [Arabidopsis thaliana]
Length = 306
Score = 367 bits (942), Expect = e-100
Identities = 175/303 (57%), Positives = 229/303 (74%)
Query: 294 PTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHA 353
PT EL++ MHILY+ CPYF+F Y S+N I E ++NES +HIIDFQI+QG QW+ L+ A
Sbjct: 4 PTGPELLTYMHILYEACPYFKFGYESANGAIAEAVKNESFVHIIDFQISQGGQWVSLIRA 63
Query: 354 LKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQ 413
L +PGGPP +R+TGIDD +S AR G L++VG++L A+ C VPFEF+ + EV+
Sbjct: 64 LGARPGGPPNVRITGIDDPRSSFARQGGLELVGQRLGKLAEMCGVPFEFHGAALCCTEVE 123
Query: 414 LEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNT 473
+E V++ E L VNFP LHH+PDESV++ENHRDRLLRLVK LSP VV VEQE+NTNT
Sbjct: 124 IEKLGVRNGEALAVNFPLVLHHMPDESVTVENHRDRLLRLVKHLSPNVVTLVEQEANTNT 183
Query: 474 SPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHE 533
+PFLPRF ET+N+Y A+FESIDV L RD K+RIN EQHC+AR++VN+IACEG ER ERHE
Sbjct: 184 APFLPRFVETMNHYLAVFESIDVKLARDHKERINVEQHCLAREVVNLIACEGVEREERHE 243
Query: 534 LFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKDYRIEQTDVAINLAWKSKVMCTSS 593
GKW++RF MAGF P LS V +++ LL+ +++ Y +E+ D A+ L WK++ + TS
Sbjct: 244 PLGKWRSRFHMAGFKPYPLSSYVNATIKGLLESYSEKYTLEERDGALYLGWKNQPLITSC 303
Query: 594 AWR 596
AWR
Sbjct: 304 AWR 306
>UniRef100_Q9SDQ3 Scarecrow-like 1 [Arabidopsis thaliana]
Length = 593
Score = 355 bits (911), Expect = 2e-96
Identities = 212/591 (35%), Positives = 323/591 (53%), Gaps = 59/591 (9%)
Query: 34 TLQTNSNNGG---------RRQRTNVSLETSNWESNWEKQFCTLESSQVANDVIAFDSPS 84
TL N NN G R + + S ++EK F + + I +
Sbjct: 34 TLNENGNNNGVSSAQIFDPDRSKNPCLTDDSYPSQSYEKYFLDSPTDEFVQHPIGSGASV 93
Query: 85 SASGD----PFNMRSTF--SPQISRYCSSDNTTNVSPLGVYSSA------------DDDS 126
S+ G P+ R S + S +T++ LG Y + DD+
Sbjct: 94 SSFGSLDSFPYQSRPVLGCSMEFQLPLDSTSTSSTRLLGDYQAVSYSPSMDVVEEFDDEQ 153
Query: 127 FEFKMKEIELSLVGTDIEVFGSCLSNSNGSLHQGTSQDNEFHIDEMISEQGFKESEISLL 186
K++E+E +L+G + + DN ID S Q ESE
Sbjct: 154 MRSKIQELERALLGDEDDKM--------------VGIDNLMEIDSEWSYQN--ESEQHQD 197
Query: 187 GPSSDIVDSCQSNLNGSLHQGTSQYDWSQFEEIIPKLDLKEELIRCAQFVFDGDFQKAIG 246
P S SN + S +E++ + K+ LI CA+ + +G ++A+
Sbjct: 198 SPKES--SSADSNSHVSS------------KEVVSQATPKQILISCARALSEGKLEEALS 243
Query: 247 FMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGSAIYKALKCEEPTSIELMSAMHIL 306
+N+ L ++VS+ G P QR+ AYM+EGL AR+ +SG IY+ALKC+EP S E ++AM +L
Sbjct: 244 MVNE-LRQIVSIQGDPSQRIAAYMVEGLAARMAASGKFIYRALKCKEPPSDERLAAMQVL 302
Query: 307 YQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRV 366
+++CP F+F ++++N I E ++ E +HIIDF I QG+Q+M L+ ++ PG P +R+
Sbjct: 303 FEVCPCFKFGFLAANGAILEAIKGEEEVHIIDFDINQGNQYMTLIRSIAELPGKRPRLRL 362
Query: 367 TGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLV 426
TGIDD +S G L I+G +LE A+ V F+F ++ V + E L+
Sbjct: 363 TGIDDPESVQRSIGGLRIIGLRLEQLAEDNGVSFKFKAMPSKTSIVSPSTLGCKPGETLI 422
Query: 427 VNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNY 486
VNF F LHH+PDESV+ N RD LL +VK L+PK+V VEQ+ NTNTSPF PRF E Y
Sbjct: 423 VNFAFQLHHMPDESVTTVNQRDELLHMVKSLNPKLVTVVEQDVNTNTSPFFPRFIEAYEY 482
Query: 487 YTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFGKWKARFSMAG 546
Y+A+FES+D+ LPR+ ++R+N E+ C+ARDIVNI+ACEG+ER ER+E GKW+AR MAG
Sbjct: 483 YSAVFESLDMTLPRESQERMNVERQCLARDIVNIVACEGEERIERYEAAGKWRARMMMAG 542
Query: 547 FVPLLLSPSVIDSVRTLLK-DFNKDYRIEQTDVAINLAWKSKVMCTSSAWR 596
F P +S V ++++ L+K + Y++++ ++ W+ K + +SAWR
Sbjct: 543 FNPKPMSAKVTNNIQNLIKQQYCNKYKLKEEMGELHFCWEEKSLIVASAWR 593
>UniRef100_Q8RZQ6 Scarecrow-like protein [Oryza sativa]
Length = 553
Score = 351 bits (901), Expect = 3e-95
Identities = 212/564 (37%), Positives = 317/564 (55%), Gaps = 41/564 (7%)
Query: 38 NSNNGGRRQRTNVSLETSNWES-NWEKQFCTLESSQVANDVIAFD-SPSSASGDPFNMRS 95
NSN RR ++ L E N E F ++S + V A SPSSA ++
Sbjct: 26 NSNFAARRYASDTQLFRYGPEPYNPENSFYNQQASPMPYMVTADGHSPSSADNSCSDVAK 85
Query: 96 TFSPQISRYCSSDNTTNVSP---LGVYSSADDDSFEFKMKEIELSLVGTDIEVFGSCLSN 152
SP +S S N+ ++S + D+D K++E+E +L+ ++ +
Sbjct: 86 D-SPLVSNV-SQQNSQSISDNQSSELEVEFDEDDIRMKLQELEHALLDDSDDILYEI--S 141
Query: 153 SNGSLHQGTSQDNEFHIDEMISEQGFKESEISLLGPSSDIVDSCQSNLNGSLHQGTSQYD 212
GS++ + + +I KESE S+ SC + NG
Sbjct: 142 QAGSINDEWADP----MKNVILPNSPKESESSI---------SCAGSNNGE--------- 179
Query: 213 WSQFEEIIPKLDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLE 272
P+ K+ L CA + D + +A + L +MVS+ G P QR+ AY++E
Sbjct: 180 --------PRTP-KQLLFDCAMALSDYNVDEAQAIITD-LRQMVSIQGDPSQRIAAYLVE 229
Query: 273 GLRARVESSGSAIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNES 332
GL AR+ +SG IYKAL C+EP ++ +SAM IL++ICP F+F ++++N I E + E
Sbjct: 230 GLAARIVASGKGIYKALSCKEPPTLYQLSAMQILFEICPCFRFGFMAANYAILEACKGED 289
Query: 333 RIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDC 392
R+HIIDF I QGSQ++ L+ LK+ P +R+TG+DD ++ G L ++G++LE
Sbjct: 290 RVHIIDFDINQGSQYITLIQFLKNNANKPRHLRITGVDDPETVQRTVGGLKVIGQRLEKL 349
Query: 393 AKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLR 452
A+ C + FEF +V +V + E LVVNF F LHH+PDESVS+ N RD+LLR
Sbjct: 350 AEDCGISFEFRAVGANIGDVTPAMLDCCPGEALVVNFAFQLHHLPDESVSIMNERDQLLR 409
Query: 453 LVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHC 512
+VK L PK+V VEQ++NTNT+PF RF E +YY A+F+S+D LPR+ R+N E+ C
Sbjct: 410 MVKGLQPKLVTLVEQDANTNTAPFQTRFREVYDYYAALFDSLDATLPRESPDRMNVERQC 469
Query: 513 VARDIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKDYR 572
+AR+IVNI+ACEG +R ER+E+ GKW+AR +MAGF P S +VI +R+LLK + Y+
Sbjct: 470 LAREIVNILACEGPDRVERYEVAGKWRARMTMAGFTPCPFSSNVISGIRSLLKSYCDRYK 529
Query: 573 IEQTDVAINLAWKSKVMCTSSAWR 596
E+ ++ W K + SSAW+
Sbjct: 530 FEEDHGGLHFGWGEKTLIVSSAWQ 553
>UniRef100_Q9XE51 Scarecrow-like 1 [Arabidopsis thaliana]
Length = 352
Score = 332 bits (852), Expect = 1e-89
Identities = 160/346 (46%), Positives = 235/346 (67%), Gaps = 1/346 (0%)
Query: 252 LGKMVSVAGSPIQRLGAYMLEGLRARVESSGSAIYKALKCEEPTSIELMSAMHILYQICP 311
L ++VS+ G P QR+ AYM+EGL AR+ +SG IY+ALKC+EP S E ++AM +L+++CP
Sbjct: 7 LRQIVSIQGDPSQRIAAYMVEGLAARMAASGKFIYRALKCKEPPSDERLAAMQVLFEVCP 66
Query: 312 YFQFAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDD 371
F+F ++++N I E ++ E +HIIDF I QG+Q+M L+ ++ PG P +R+TGIDD
Sbjct: 67 CFKFGFLAANGAILEAIKGEEEVHIIDFDINQGNQYMTLIRSIAELPGKRPRLRLTGIDD 126
Query: 372 SQSFHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPF 431
+S G L I+G +LE A+ V F+F ++ V + E L+VNF F
Sbjct: 127 PESVQRSIGGLRIIGLRLEQLAEDNGVSFKFKAMPSKTSIVSPSTLGCKPGETLIVNFAF 186
Query: 432 ALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMF 491
LHH+PDESV+ N RD LL +VK L+PK+V VEQ+ NTNTSPF PRF E YY+A+F
Sbjct: 187 QLHHMPDESVTTVNQRDELLHMVKSLNPKLVTVVEQDVNTNTSPFFPRFIEAYEYYSAVF 246
Query: 492 ESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFGKWKARFSMAGFVPLL 551
ES+D+ LPR+ ++R+N E+ C+ARDIVNI+ACEG+ER ER+E GKW+AR MAGF P
Sbjct: 247 ESLDMTLPRESQERMNVERQCLARDIVNIVACEGEERIERYEAAGKWRARMMMAGFNPKP 306
Query: 552 LSPSVIDSVRTLLK-DFNKDYRIEQTDVAINLAWKSKVMCTSSAWR 596
+S V ++++ L+K + Y++++ ++ W+ K + +SAWR
Sbjct: 307 MSAKVTNNIQNLIKQQYCNKYKLKEEMGELHFCWEEKSLIVASAWR 352
>UniRef100_Q9XE57 Scarecrow-like 13 [Arabidopsis thaliana]
Length = 284
Score = 330 bits (847), Expect = 6e-89
Identities = 160/277 (57%), Positives = 205/277 (73%)
Query: 320 SNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARG 379
+N I E + E+R+HIIDFQIAQGSQ+M L+ L +PGGPP +RVTG+DDSQS +ARG
Sbjct: 1 ANVEILEAIAGETRVHIIDFQIAQGSQYMFLIQELAKRPGGPPLLRVTGVDDSQSTYARG 60
Query: 380 GKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDE 439
G L +VG++L A++C VPFEF+ M GC+VQ E ++ +VVNFP+ LHH+PDE
Sbjct: 61 GGLSLVGERLATLAQSCGVPFEFHDAIMSGCKVQREHLGLEPGFAVVVNFPYVLHHMPDE 120
Query: 440 SVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALP 499
SVS+E +RDRLL L+K LSPK+V VEQESNTNTSP + RF ETL+YYTAMFESID A P
Sbjct: 121 SVSVEKYRDRLLHLIKSLSPKLVTLVEQESNTNTSPLVSRFVETLDYYTAMFESIDAARP 180
Query: 500 RDDKKRINAEQHCVARDIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDS 559
RDDK+RI+AEQHCVARDIVN+IACE ER ERHE+ GKW+ R MAGF +S S +
Sbjct: 181 RDDKQRISAEQHCVARDIVNMIACEESERVERHEVLGKWRVRMMMAGFTGWPVSTSAAFA 240
Query: 560 VRTLLKDFNKDYRIEQTDVAINLAWKSKVMCTSSAWR 596
+LK ++K+Y++ + A+ L WK + M T S W+
Sbjct: 241 ASEMLKAYDKNYKLGGHEGALYLFWKRRPMATCSVWK 277
>UniRef100_Q8GVE2 Chitin-inducible gibberellin-responsive protein [Oryza sativa]
Length = 572
Score = 298 bits (764), Expect = 2e-79
Identities = 169/379 (44%), Positives = 235/379 (61%), Gaps = 8/379 (2%)
Query: 221 PKLDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVES 280
P++ +K+ L RCA+ + + ++ + + G +VS+ G PIQRLGAY+LEGL AR +
Sbjct: 197 PQIIVKQLLTRCAEALSEDRTEEFHKLVQEARG-VVSINGEPIQRLGAYLLEGLVARHGN 255
Query: 281 SGSAIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQ 340
SG+ IY+ALKC EP S EL+S M ILY ICPYF+F Y+++N I E ++ E+ IHIIDFQ
Sbjct: 256 SGTNIYRALKCREPESKELLSYMRILYNICPYFKFGYMAANGAIAEALRTENNIHIIDFQ 315
Query: 341 IAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPF 400
IAQG+QW+ L+ AL +PGGPP +R+TGIDD S +ARG LDIVGK L+ ++ K+P
Sbjct: 316 IAQGTQWITLIQALAARPGGPPRVRITGIDDPVSEYARGEGLDIVGKMLKSMSEEFKIPL 375
Query: 401 EFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPK 460
EF + +Y +V E E++ E L VNF LHH PDESV + N RD LL + P+
Sbjct: 376 EFTPLSVYATQVTKEMLEIRPGEALSVNFTLQLHHTPDESVDVNNPRDGLLPDGERAVPE 435
Query: 461 VVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDV-ALP--RDDKKRINAEQHCVA-RD 516
F + FL E + + + V LP R +R +A + R
Sbjct: 436 GDYFGRAGVTHQHNAFLD---EVWGDHGVLLRHVRVDRLPTCRGTTRRGSAWSSTASPRH 492
Query: 517 IVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKDYRIEQT 576
IVNIIACEG +R ERHEL GKWK+R +MAGF P LS V +R LL ++ Y +++
Sbjct: 493 IVNIIACEGKDRVERHELLGKWKSRLTMAGFRPYPLSSYVNSVIRKLLACYSDKYTLDEK 552
Query: 577 DVAINLAWKSKVMCTSSAW 595
D A+ L W+S+ + ++SAW
Sbjct: 553 DGAMLLGWRSRKLISASAW 571
>UniRef100_Q67ZL8 Putative SCARECROW gene regulator [Arabidopsis thaliana]
Length = 220
Score = 281 bits (720), Expect = 3e-74
Identities = 137/218 (62%), Positives = 166/218 (75%)
Query: 379 GGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPD 438
G L V K+LE AK VPF FN+V CEV++E+ +V+ E L VNF + LHH+PD
Sbjct: 3 GSVLVTVKKRLEKLAKKFDVPFRFNAVSRPSCEVEVENLDVRDGEALGVNFAYMLHHLPD 62
Query: 439 ESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVAL 498
ESVSMENHRDRLLR+VK LSPKVV VEQE NTNTSPFLPRF ETL+YYTAMFESIDV L
Sbjct: 63 ESVSMENHRDRLLRMVKSLSPKVVTLVEQECNTNTSPFLPRFLETLSYYTAMFESIDVML 122
Query: 499 PRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVID 558
PR+ K+RIN EQHC+ARD+VNIIACEG ER ERHEL GKWK+RFSMAGF P LS +
Sbjct: 123 PRNHKERINIEQHCMARDVVNIIACEGAERIERHELLGKWKSRFSMAGFEPYPLSSIISA 182
Query: 559 SVRTLLKDFNKDYRIEQTDVAINLAWKSKVMCTSSAWR 596
++R LL+D++ Y IE+ D A+ L W +++ +S AW+
Sbjct: 183 TIRALLRDYSNGYAIEERDGALYLGWMDRILVSSCAWK 220
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.134 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,014,274,826
Number of Sequences: 2790947
Number of extensions: 43073012
Number of successful extensions: 105057
Number of sequences better than 10.0: 216
Number of HSP's better than 10.0 without gapping: 169
Number of HSP's successfully gapped in prelim test: 48
Number of HSP's that attempted gapping in prelim test: 104235
Number of HSP's gapped (non-prelim): 371
length of query: 598
length of database: 848,049,833
effective HSP length: 133
effective length of query: 465
effective length of database: 476,853,882
effective search space: 221737055130
effective search space used: 221737055130
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 78 (34.7 bits)
Medicago: description of AC148526.1


