

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148525.11 + phase: 0 /pseudo
(88 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
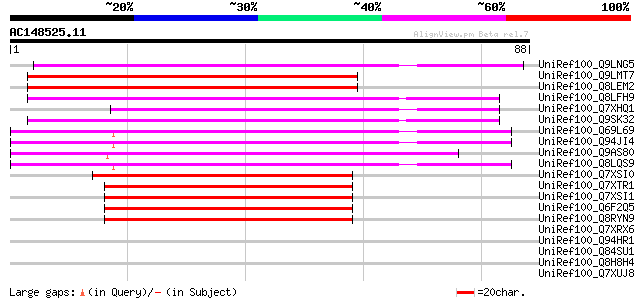
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LNG5 F21D18.16 [Arabidopsis thaliana] 52 5e-06
UniRef100_Q9LMT7 F2H15.15 protein [Arabidopsis thaliana] 52 5e-06
UniRef100_Q8LEM2 Hypothetical protein [Arabidopsis thaliana] 51 6e-06
UniRef100_Q8LFH9 Hypothetical protein [Arabidopsis thaliana] 50 1e-05
UniRef100_Q7XHQ1 Calcineurin-like phosphoesterase-like [Oryza sa... 50 2e-05
UniRef100_Q9SK32 Expressed protein [Arabidopsis thaliana] 50 2e-05
UniRef100_Q69L69 Calcineurin-like phosphoesterase-like protein [... 48 7e-05
UniRef100_Q94JI4 P0638D12.5 protein [Oryza sativa] 47 2e-04
UniRef100_Q9AS80 P0028E10.12 protein [Oryza sativa] 47 2e-04
UniRef100_Q8LQS9 P0702H08.15 protein [Oryza sativa] 47 2e-04
UniRef100_Q7XSI0 OSJNBa0073L13.8 protein [Oryza sativa] 45 6e-04
UniRef100_Q7XTR1 OSJNBb0085C12.10 protein [Oryza sativa] 44 8e-04
UniRef100_Q7XSI1 OSJNBa0073L13.6 protein [Oryza sativa] 44 8e-04
UniRef100_Q6F2Q5 Hypothetical protein OSJNBa0088M06.4 [Oryza sat... 44 8e-04
UniRef100_Q8RYN9 Putative mutator-like transposase [Oryza sativa] 44 8e-04
UniRef100_Q7XRX6 OSJNBb0054B09.15 protein [Oryza sativa] 44 0.001
UniRef100_Q94HR1 Hypothetical protein OSJNBa0065C16.8 [Oryza sat... 44 0.001
UniRef100_Q84SU1 Putative transposase [Oryza sativa] 44 0.001
UniRef100_Q8H8H4 Putative mutator-like transposase [Oryza sativa] 44 0.001
UniRef100_Q7XUJ8 OSJNBa0067K08.4 protein [Oryza sativa] 44 0.001
>UniRef100_Q9LNG5 F21D18.16 [Arabidopsis thaliana]
Length = 1340
Score = 51.6 bits (122), Expect = 5e-06
Identities = 28/83 (33%), Positives = 46/83 (54%), Gaps = 3/83 (3%)
Query: 5 GLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHMLY 64
GL + V+++ +D L++ + ERW T +FHLP E+ VTL D++ LL L ++ +
Sbjct: 68 GLYGVYKVAFIQLDYALITALVERWRPETHTFHLPAGEITVTLQDVNILLGLRVDGPAVT 127
Query: 65 HNIYLTRSEAVDLMVELLGSDSG 87
+ T+ DL +LLG G
Sbjct: 128 GS---TKYNWADLCEDLLGHRPG 147
>UniRef100_Q9LMT7 F2H15.15 protein [Arabidopsis thaliana]
Length = 478
Score = 51.6 bits (122), Expect = 5e-06
Identities = 20/56 (35%), Positives = 36/56 (63%)
Query: 4 SGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIE 59
+G W + V + ++ L+S + ERW T++FH P EM +TLD++S +L L+++
Sbjct: 49 AGFGWFRLVGSISLNNSLISALVERWRRETNTFHFPCGEMTITLDEVSLILGLAVD 104
>UniRef100_Q8LEM2 Hypothetical protein [Arabidopsis thaliana]
Length = 478
Score = 51.2 bits (121), Expect = 6e-06
Identities = 19/56 (33%), Positives = 36/56 (63%)
Query: 4 SGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIE 59
+G W + + + ++ L+S + ERW T++FH P EM +TLD++S +L L+++
Sbjct: 49 AGFGWFRLIGSISLNNSLISALVERWRRETNTFHFPCGEMTITLDEVSLILGLAVD 104
>UniRef100_Q8LFH9 Hypothetical protein [Arabidopsis thaliana]
Length = 509
Score = 50.1 bits (118), Expect = 1e-05
Identities = 24/80 (30%), Positives = 47/80 (58%), Gaps = 1/80 (1%)
Query: 4 SGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHML 63
+G + + + + ++ L+S + ERW T++FHLP+ EM +TLD+++ +L L I+ +
Sbjct: 58 AGFGYFRKIGPMSLNNSLISALVERWRRETNTFHLPLGEMTITLDEVALVLGLEIDGDPI 117
Query: 64 YHNIYLTRSEAVDLMVELLG 83
+ + A+D+ LLG
Sbjct: 118 VGS-KVDDEVAMDMCGRLLG 136
>UniRef100_Q7XHQ1 Calcineurin-like phosphoesterase-like [Oryza sativa]
Length = 327
Score = 49.7 bits (117), Expect = 2e-05
Identities = 26/66 (39%), Positives = 37/66 (55%), Gaps = 3/66 (4%)
Query: 18 DQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHMLYHNIYLTRSEAVDL 77
D+ L+S + ERW T++FHLP+ EM +TL D+SCL L I + VD+
Sbjct: 67 DKYLISALVERWRPKTNTFHLPVGEMTITLQDVSCLWGLPIHGRPITGQ---ADGSWVDM 123
Query: 78 MVELLG 83
+ LLG
Sbjct: 124 IERLLG 129
>UniRef100_Q9SK32 Expressed protein [Arabidopsis thaliana]
Length = 509
Score = 49.7 bits (117), Expect = 2e-05
Identities = 24/80 (30%), Positives = 47/80 (58%), Gaps = 1/80 (1%)
Query: 4 SGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHML 63
+G + + + + ++ L+S + ERW T++FHLP+ EM +TLD+++ +L L I+ +
Sbjct: 58 AGFGYFRKIGPMSLNNSLISALVERWRRETNTFHLPLGEMTITLDEVALVLGLEIDGDPI 117
Query: 64 YHNIYLTRSEAVDLMVELLG 83
+ + A+D+ LLG
Sbjct: 118 VGS-KVGDEVAMDMCGRLLG 136
>UniRef100_Q69L69 Calcineurin-like phosphoesterase-like protein [Oryza sativa]
Length = 766
Score = 47.8 bits (112), Expect = 7e-05
Identities = 29/87 (33%), Positives = 46/87 (52%), Gaps = 5/87 (5%)
Query: 1 MRVSGLQWLQHVSYLV--IDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 58
+R +GL L + + +D+ L++ + ERW T SFHL EM VTL D+S LL L I
Sbjct: 111 LRDAGLLGLSQICQKMPQLDKALITALVERWRPETHSFHLASGEMTVTLQDVSMLLALPI 170
Query: 59 ESHMLYHNIYLTRSEAVDLMVELLGSD 85
+ + T + ++++ LG D
Sbjct: 171 DGRPVCST---TDHDYAQMVIDCLGHD 194
>UniRef100_Q94JI4 P0638D12.5 protein [Oryza sativa]
Length = 761
Score = 46.6 bits (109), Expect = 2e-04
Identities = 28/87 (32%), Positives = 46/87 (52%), Gaps = 5/87 (5%)
Query: 1 MRVSGLQWLQHVSYLV--IDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 58
+R +GL L + + +D+ L++ + ERW T SFHL EM VTL D++ LL L I
Sbjct: 128 LRDAGLLGLSQICQKMPQLDKALITALVERWRPETHSFHLASGEMAVTLQDVAMLLALPI 187
Query: 59 ESHMLYHNIYLTRSEAVDLMVELLGSD 85
+ + T + ++++ LG D
Sbjct: 188 DGRPVCST---TDHDYAQMVIDCLGHD 211
>UniRef100_Q9AS80 P0028E10.12 protein [Oryza sativa]
Length = 792
Score = 46.6 bits (109), Expect = 2e-04
Identities = 25/79 (31%), Positives = 44/79 (55%), Gaps = 3/79 (3%)
Query: 1 MRVSGLQWLQHVSYL---VIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLS 57
+R +GL L ++ ID+ L+S + +RW T +FH+P E+ +TL D++ +L L
Sbjct: 238 LRPAGLMGLANICRAGLPSIDRALVSALVDRWRPETHTFHMPCGEITITLQDVAMILGLP 297
Query: 58 IESHMLYHNIYLTRSEAVD 76
I H + N ++E V+
Sbjct: 298 IAGHAVTVNPTEPQNELVE 316
>UniRef100_Q8LQS9 P0702H08.15 protein [Oryza sativa]
Length = 718
Score = 46.6 bits (109), Expect = 2e-04
Identities = 28/87 (32%), Positives = 46/87 (52%), Gaps = 5/87 (5%)
Query: 1 MRVSGLQWLQHVSYLV--IDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 58
+R +GL L + + +D+ L++ + ERW T SFHL EM VTL D++ LL L I
Sbjct: 142 LRDAGLLGLSQICQKMPQLDKALITALVERWRPETHSFHLASGEMAVTLQDVAMLLALPI 201
Query: 59 ESHMLYHNIYLTRSEAVDLMVELLGSD 85
+ + T + ++++ LG D
Sbjct: 202 DGRPVCST---TDHDYAQMVIDCLGHD 225
>UniRef100_Q7XSI0 OSJNBa0073L13.8 protein [Oryza sativa]
Length = 925
Score = 44.7 bits (104), Expect = 6e-04
Identities = 21/44 (47%), Positives = 28/44 (62%)
Query: 15 LVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 58
+ ID LLS + +RW T +FHL + EM+ TL D+S LL L I
Sbjct: 563 IYIDAALLSALVDRWRPETHTFHLTVGEMVPTLQDVSYLLGLPI 606
>UniRef100_Q7XTR1 OSJNBb0085C12.10 protein [Oryza sativa]
Length = 960
Score = 44.3 bits (103), Expect = 8e-04
Identities = 21/42 (50%), Positives = 27/42 (64%)
Query: 17 IDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 58
ID LLS + +RW T +FHL + EM+ TL D+S LL L I
Sbjct: 609 IDAALLSALVDRWRPETHTFHLTVGEMVPTLQDVSYLLGLPI 650
>UniRef100_Q7XSI1 OSJNBa0073L13.6 protein [Oryza sativa]
Length = 552
Score = 44.3 bits (103), Expect = 8e-04
Identities = 21/42 (50%), Positives = 27/42 (64%)
Query: 17 IDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 58
ID LLS + +RW T +FHL + EM+ TL D+S LL L I
Sbjct: 75 IDAALLSALVDRWRPETHTFHLTVGEMVPTLQDVSYLLGLPI 116
>UniRef100_Q6F2Q5 Hypothetical protein OSJNBa0088M06.4 [Oryza sativa]
Length = 1635
Score = 44.3 bits (103), Expect = 8e-04
Identities = 21/42 (50%), Positives = 27/42 (64%)
Query: 17 IDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 58
ID LLS + +RW T +FHL + EM+ TL D+S LL L I
Sbjct: 925 IDAALLSALVDRWRPETHTFHLTVGEMVPTLQDVSYLLGLPI 966
>UniRef100_Q8RYN9 Putative mutator-like transposase [Oryza sativa]
Length = 1701
Score = 44.3 bits (103), Expect = 8e-04
Identities = 21/42 (50%), Positives = 27/42 (64%)
Query: 17 IDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 58
ID LLS + +RW T +FHL + EM+ TL D+S LL L I
Sbjct: 1013 IDAALLSALVDRWRPETHTFHLTVGEMVPTLQDVSYLLGLPI 1054
>UniRef100_Q7XRX6 OSJNBb0054B09.15 protein [Oryza sativa]
Length = 1613
Score = 43.9 bits (102), Expect = 0.001
Identities = 21/42 (50%), Positives = 27/42 (64%)
Query: 17 IDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 58
ID LLS + +RW T +FHL + EM+ TL D+S LL L I
Sbjct: 913 IDAALLSALVDRWRPETHTFHLTVGEMVPTLQDVSYLLGLLI 954
>UniRef100_Q94HR1 Hypothetical protein OSJNBa0065C16.8 [Oryza sativa]
Length = 681
Score = 43.9 bits (102), Expect = 0.001
Identities = 20/56 (35%), Positives = 34/56 (60%), Gaps = 7/56 (12%)
Query: 15 LVIDQGL-------LSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHML 63
LV+ +G+ L+ + +RW T SFHLP EM +TL D++ +L L ++ H++
Sbjct: 158 LVVSRGMPVFNAPALTALVDRWRPETHSFHLPSGEMTITLQDVAMILALPLQGHVV 213
>UniRef100_Q84SU1 Putative transposase [Oryza sativa]
Length = 1511
Score = 43.5 bits (101), Expect = 0.001
Identities = 20/42 (47%), Positives = 27/42 (63%)
Query: 17 IDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 58
ID L+S + +RW T +FHL + EM+ TL D+S LL L I
Sbjct: 893 IDAALMSALVDRWRPETHTFHLTVGEMVPTLQDVSYLLGLPI 934
>UniRef100_Q8H8H4 Putative mutator-like transposase [Oryza sativa]
Length = 1436
Score = 43.5 bits (101), Expect = 0.001
Identities = 20/42 (47%), Positives = 27/42 (63%)
Query: 17 IDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 58
ID L+S + +RW T +FHL + EM+ TL D+S LL L I
Sbjct: 947 IDAALMSALVDRWRPETHTFHLTVGEMVPTLQDVSYLLGLPI 988
>UniRef100_Q7XUJ8 OSJNBa0067K08.4 protein [Oryza sativa]
Length = 1555
Score = 43.5 bits (101), Expect = 0.001
Identities = 20/42 (47%), Positives = 27/42 (63%)
Query: 17 IDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 58
ID L+S + +RW T +FHL + EM+ TL D+S LL L I
Sbjct: 904 IDAALMSALVDRWRPETHTFHLTVGEMVPTLQDVSYLLGLPI 945
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.323 0.137 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 141,989,762
Number of Sequences: 2790947
Number of extensions: 4558410
Number of successful extensions: 8704
Number of sequences better than 10.0: 110
Number of HSP's better than 10.0 without gapping: 107
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 8597
Number of HSP's gapped (non-prelim): 110
length of query: 88
length of database: 848,049,833
effective HSP length: 64
effective length of query: 24
effective length of database: 669,429,225
effective search space: 16066301400
effective search space used: 16066301400
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 68 (30.8 bits)
Medicago: description of AC148525.11


