

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148470.10 - phase: 0 /pseudo
(598 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
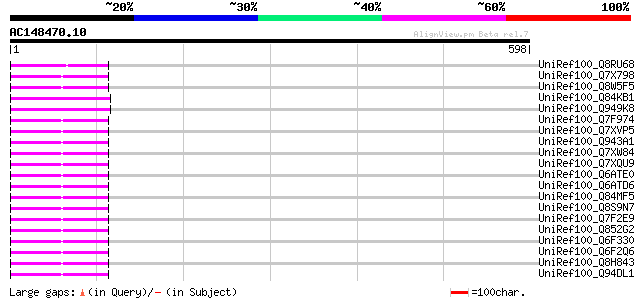
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8RU68 Putative 22 kDa kafirin cluster; Ty3-Gypsy type... 94 1e-17
UniRef100_Q7X798 OSJNBb0108J11.19 protein [Oryza sativa] 92 4e-17
UniRef100_Q8W5F5 Putative polyprotein [Oryza sativa] 92 6e-17
UniRef100_Q84KB1 Gag-protease polyprotein [Cucumis melo] 91 7e-17
UniRef100_Q949K8 Gag polyprotein [Cicer arietinum] 91 1e-16
UniRef100_Q7F974 OSJNBb0003A12.3 protein [Oryza sativa] 89 3e-16
UniRef100_Q7XVP5 OSJNBa0023J03.1 protein [Oryza sativa] 89 3e-16
UniRef100_Q943A1 Putative polyprotein [Oryza sativa] 89 4e-16
UniRef100_Q7XW84 OSJNBa0019J05.10 protein [Oryza sativa] 89 4e-16
UniRef100_Q7XQU9 OSJNBa0086B14.15 protein [Oryza sativa] 89 4e-16
UniRef100_Q6ATE0 Putative polyprotein [Oryza sativa] 89 4e-16
UniRef100_Q6ATD6 Putative polyprotein [Oryza sativa] 89 4e-16
UniRef100_Q84MF5 Putative retrotransposon gag protein [Oryza sat... 89 5e-16
UniRef100_Q8S9N7 Similar to polyprotein [Oryza sativa] 89 5e-16
UniRef100_Q7F2E9 Putative polyprotein [Oryza sativa] 88 6e-16
UniRef100_Q852G2 Putative polyprotein [Oryza sativa] 88 6e-16
UniRef100_Q6F330 Putative polyprotein [Oryza sativa] 88 6e-16
UniRef100_Q6F2Q6 Putative polyprotein [Oryza sativa] 88 6e-16
UniRef100_Q8H843 Putative polyprotein [Oryza sativa] 87 1e-15
UniRef100_Q94DL1 Putative polyprotein [Oryza sativa] 87 1e-15
>UniRef100_Q8RU68 Putative 22 kDa kafirin cluster; Ty3-Gypsy type [Oryza sativa]
Length = 1230
Score = 93.6 bits (231), Expect = 1e-17
Identities = 45/114 (39%), Positives = 67/114 (58%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
F + PPTF G +P A++W+ +E F M C++ +K+ + T+ML A +WW +
Sbjct: 74 FQKLKPPTFSGTANPLEAEEWIVAMEKSFEAMGCTDKEKIIYATYMLQSSAFEWWDAHKK 133
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ +TW +F+ F +Y PE V+ KE EFLELKQG+ SV EY EFS+
Sbjct: 134 SYSER-IFITWELFKEAFYKKYFPESVKRMKEKEFLELKQGNKSVAEYEIEFSR 186
>UniRef100_Q7X798 OSJNBb0108J11.19 protein [Oryza sativa]
Length = 1516
Score = 92.0 bits (227), Expect = 4e-17
Identities = 45/114 (39%), Positives = 65/114 (56%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL+ +E ++QC++ +KV F TH L A+ WW + +
Sbjct: 84 FLRIQPPTFSSTTNPMEANDWLRAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNYV- 142
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
D +TWA F + F +PE + Q++ EF L+QG MSVTEY EF++
Sbjct: 143 ATRLDATEVTWAEFCQSFRKAQIPEGIMAQQKREFRALQQGTMSVTEYLHEFNR 196
>UniRef100_Q8W5F5 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 91.7 bits (226), Expect = 6e-17
Identities = 46/114 (40%), Positives = 63/114 (54%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ A +TW FR F VPE + GQK+ EF L+QG +V EY EF++
Sbjct: 155 VTRPTGAEVTWTEFRHSFNKAQVPEGIVGQKKREFRSLQQGTKTVIEYLHEFNR 208
>UniRef100_Q84KB1 Gag-protease polyprotein [Cucumis melo]
Length = 429
Score = 91.3 bits (225), Expect = 7e-17
Identities = 47/117 (40%), Positives = 65/117 (55%), Gaps = 1/117 (0%)
Query: 1 FLRNHPPTFKGRY-DPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLL 59
F + +P TF G DP AQ WL +E+IFR M+C E QKV+ ML + WW +
Sbjct: 63 FRKYNPTTFDGSLEDPTRAQMWLSSLETIFRYMKCPEDQKVQCAVFMLTDRGTAWWETTE 122
Query: 60 PILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSKSS 116
+L D + +TW F+ F ++ +R K EFL L+QGDM+V +Y AEF S
Sbjct: 123 RMLGGDVSQITWQQFKESFYAKFFSASLRDAKRQEFLNLEQGDMTVEQYDAEFDMLS 179
>UniRef100_Q949K8 Gag polyprotein [Cicer arietinum]
Length = 277
Score = 90.5 bits (223), Expect = 1e-16
Identities = 44/116 (37%), Positives = 67/116 (56%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
F R +PP FKG + A +W++EVE IF ++ C KV + T+ML +A WW S
Sbjct: 59 FRRYNPPKFKGDEGSEKADQWIQEVEKIFDMINCQAGVKVSYATYMLLGDAEYWWRSARL 118
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSKSS 116
++ + W F+R+FL++Y P R + +FL+L QG ++V EYAA+F S
Sbjct: 119 LMGAAHEEVNWESFKRKFLDKYFPMSTRTKLGDDFLKLHQGSLTVGEYAAKFESLS 174
>UniRef100_Q7F974 OSJNBb0003A12.3 protein [Oryza sativa]
Length = 1471
Score = 89.4 bits (220), Expect = 3e-16
Identities = 45/114 (39%), Positives = 62/114 (53%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 70 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASVWWDNYM- 128
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF++
Sbjct: 129 VTRPTGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFNR 182
>UniRef100_Q7XVP5 OSJNBa0023J03.1 protein [Oryza sativa]
Length = 1827
Score = 89.4 bits (220), Expect = 3e-16
Identities = 46/114 (40%), Positives = 63/114 (54%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC++ +KV F TH L A+ WW + +
Sbjct: 401 FLRVRPPTFSSTTNPMEANDWLHAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNHM- 459
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+TWA F R F VP+ V QK+ EF L QG+M+VTEY EF++
Sbjct: 460 ATRPPGTEVTWAEFCRSFRKAQVPDGVVAQKKREFRALHQGNMTVTEYLHEFNR 513
>UniRef100_Q943A1 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 89.0 bits (219), Expect = 4e-16
Identities = 45/114 (39%), Positives = 62/114 (53%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF++
Sbjct: 155 VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFNR 208
>UniRef100_Q7XW84 OSJNBa0019J05.10 protein [Oryza sativa]
Length = 1525
Score = 89.0 bits (219), Expect = 4e-16
Identities = 45/114 (39%), Positives = 62/114 (53%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF++
Sbjct: 155 VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFNR 208
>UniRef100_Q7XQU9 OSJNBa0086B14.15 protein [Oryza sativa]
Length = 1516
Score = 89.0 bits (219), Expect = 4e-16
Identities = 42/114 (36%), Positives = 64/114 (55%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL+ +E ++QC++ +KV F TH L A+ WW + +
Sbjct: 84 FLRIQPPTFSSTTNPMEANDWLRAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNYV- 142
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ D +TW F + F +PE + Q++ EF L+QG +VTEY EF++
Sbjct: 143 VTRPDATEVTWVEFCQNFRKALIPEGIMAQQKREFRALQQGTRTVTEYLHEFNR 196
>UniRef100_Q6ATE0 Putative polyprotein [Oryza sativa]
Length = 1495
Score = 89.0 bits (219), Expect = 4e-16
Identities = 42/114 (36%), Positives = 64/114 (55%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL+ +E ++QC++ +KV F TH L A+ WW + +
Sbjct: 84 FLRIQPPTFSSTTNPMEANDWLRAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNYV- 142
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ D +TW F + F +PE + Q++ EF L+QG +VTEY EF++
Sbjct: 143 VTRPDATKVTWVEFCQNFRKAQIPEGIMAQQKREFRALQQGTRTVTEYLHEFNR 196
>UniRef100_Q6ATD6 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 89.0 bits (219), Expect = 4e-16
Identities = 45/114 (39%), Positives = 62/114 (53%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF++
Sbjct: 155 VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFNR 208
>UniRef100_Q84MF5 Putative retrotransposon gag protein [Oryza sativa]
Length = 486
Score = 88.6 bits (218), Expect = 5e-16
Identities = 42/114 (36%), Positives = 64/114 (55%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL+ +E ++QC++ +KV F TH L A+ WW + +
Sbjct: 84 FLRIQPPTFSSTTNPMEANDWLRAIEKKLNLLQCNDQEKVAFATHQLQGPASVWWDNYV- 142
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ D +TW F + F +PE + Q++ EF L+QG +VTEY EF++
Sbjct: 143 VTRPDATKVTWVEFCQNFRKAQIPEGIMAQQKREFRALQQGTRTVTEYLHEFNR 196
>UniRef100_Q8S9N7 Similar to polyprotein [Oryza sativa]
Length = 1524
Score = 88.6 bits (218), Expect = 5e-16
Identities = 45/114 (39%), Positives = 62/114 (53%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASVWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF++
Sbjct: 155 VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFHSLQQGTRTVIEYLHEFNR 208
>UniRef100_Q7F2E9 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 88.2 bits (217), Expect = 6e-16
Identities = 45/113 (39%), Positives = 61/113 (53%), Gaps = 1/113 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFS 113
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF+
Sbjct: 155 VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFN 207
>UniRef100_Q852G2 Putative polyprotein [Oryza sativa]
Length = 1079
Score = 88.2 bits (217), Expect = 6e-16
Identities = 45/113 (39%), Positives = 61/113 (53%), Gaps = 1/113 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFS 113
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF+
Sbjct: 155 VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFN 207
>UniRef100_Q6F330 Putative polyprotein [Oryza sativa]
Length = 1458
Score = 88.2 bits (217), Expect = 6e-16
Identities = 45/113 (39%), Positives = 61/113 (53%), Gaps = 1/113 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFS 113
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF+
Sbjct: 155 VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFN 207
>UniRef100_Q6F2Q6 Putative polyprotein [Oryza sativa]
Length = 1516
Score = 88.2 bits (217), Expect = 6e-16
Identities = 43/114 (37%), Positives = 64/114 (55%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL+ +E ++QC++ +KV F TH L A+ WW + +
Sbjct: 84 FLRIQPPTFSSTTNPMEANDWLRAIEKKLNLLQCNDREKVAFATHQLQGPASAWWDNYV- 142
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
D +TWA F + F +PE + Q++ EF L+QG +VTEY EF++
Sbjct: 143 ATRLDATEVTWAEFCQSFRKAQIPEGIMAQQKREFRALQQGTRTVTEYLHEFNR 196
>UniRef100_Q8H843 Putative polyprotein [Oryza sativa]
Length = 1796
Score = 87.4 bits (215), Expect = 1e-15
Identities = 45/114 (39%), Positives = 62/114 (53%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC++ +KV F TH L A+ WW + +
Sbjct: 370 FLRVRPPTFSSTTNPMEANDWLHAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNHM- 428
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+TWA F R F VP+ V QK+ EF L QG+ +VTEY EF++
Sbjct: 429 ATRPPGTEVTWAEFCRSFRKAQVPDGVVAQKKREFRALHQGNRTVTEYLQEFNR 482
>UniRef100_Q94DL1 Putative polyprotein [Oryza sativa]
Length = 1521
Score = 87.0 bits (214), Expect = 1e-15
Identities = 45/114 (39%), Positives = 62/114 (53%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC++ +KV F TH L A+ WW + +
Sbjct: 95 FLRVRPPTFSSTTNPMEANDWLHAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNHM- 153
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+TWA F R F VP+ V QK+ EF L QG+ +VTEY EF++
Sbjct: 154 ATRPPGTEVTWAEFCRSFRKAQVPDGVVAQKKREFRALHQGNRTVTEYLHEFNR 207
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.370 0.166 0.609
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 717,056,304
Number of Sequences: 2790947
Number of extensions: 22707122
Number of successful extensions: 126306
Number of sequences better than 10.0: 489
Number of HSP's better than 10.0 without gapping: 449
Number of HSP's successfully gapped in prelim test: 40
Number of HSP's that attempted gapping in prelim test: 124411
Number of HSP's gapped (non-prelim): 1177
length of query: 598
length of database: 848,049,833
effective HSP length: 133
effective length of query: 465
effective length of database: 476,853,882
effective search space: 221737055130
effective search space used: 221737055130
T: 11
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 78 (34.7 bits)
Medicago: description of AC148470.10


