

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148469.7 - phase: 0
(540 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
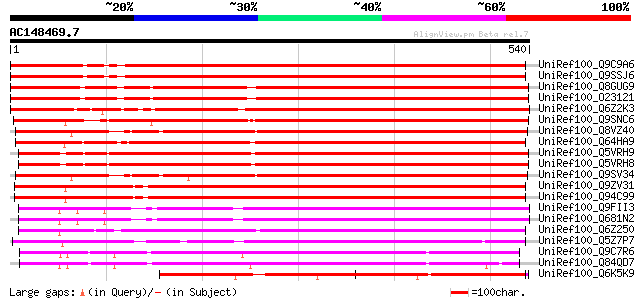
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9C9A6 Hypothetical protein F23N20.1 [Arabidopsis thal... 634 e-180
UniRef100_Q9SSJ6 F15H11.22 protein [Arabidopsis thaliana] 634 e-180
UniRef100_Q8GUG9 Hypothetical protein At1g23030 [Arabidopsis tha... 633 e-180
UniRef100_O23121 F10G19.3 protein [Arabidopsis thaliana] 633 e-180
UniRef100_Q6Z2K3 Putative Avr9/Cf-9 rapidly elicited protein 276... 566 e-160
UniRef100_Q9SNC6 Arm repeat containing protein homolog [Arabidop... 452 e-125
UniRef100_Q8VZ40 Hypothetical protein At3g54850 [Arabidopsis tha... 440 e-122
UniRef100_Q64HA9 Cell death-related protein SPL11 [Oryza sativa] 437 e-121
UniRef100_Q5VRH9 Putative cell death-related protein SPL11 [Oryz... 434 e-120
UniRef100_Q5VRH8 Putative cell death-related protein SPL11 [Oryz... 434 e-120
UniRef100_Q9SV34 Hypothetical protein F28P10.170 [Arabidopsis th... 433 e-120
UniRef100_Q9ZV31 Expressed protein [Arabidopsis thaliana] 416 e-114
UniRef100_Q94C99 Hypothetical protein At2g28830 [Arabidopsis tha... 414 e-114
UniRef100_Q9FII3 Arm repeat containing protein [Arabidopsis thal... 365 2e-99
UniRef100_Q681N2 Arm repeat containing protein [Arabidopsis thal... 362 2e-98
UniRef100_Q6Z250 Putative Avr9/Cf-9 rapidly elicited protein [Or... 360 6e-98
UniRef100_Q5Z7P7 Putative cell death-related protein SPL11 [Oryz... 342 2e-92
UniRef100_Q9C7R6 Arm repeat-containing protein, putative; 6839-9... 279 1e-73
UniRef100_Q84QD7 Avr9/Cf-9 rapidly elicited protein 276 [Nicotia... 276 8e-73
UniRef100_Q6K5K9 Avr9/Cf-9 rapidly elicited protein-like [Oryza ... 273 9e-72
>UniRef100_Q9C9A6 Hypothetical protein F23N20.1 [Arabidopsis thaliana]
Length = 628
Score = 634 bits (1636), Expect = e-180
Identities = 354/538 (65%), Positives = 411/538 (75%), Gaps = 18/538 (3%)
Query: 1 DGAAKKIIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGFMISK 60
DGAAK+I FQFQ VTWKLEK L L YD DIS+EV+EQV+L R QLRRA +YG + SK
Sbjct: 99 DGAAKRISFQFQCVTWKLEKALGDLTYDRYDISDEVREQVELARLQLRRAMQRYGSLNSK 158
Query: 61 MPSFDSSQPLAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPFGAGSRLE 120
S S+P+ ++ S + + K S PE + L S+E K +P
Sbjct: 159 KFSSGLSEPMEKDASS----NRKVIEKLESIPETVHSL-----SDEKKFESP-------P 202
Query: 121 RTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELMRDPVIV 180
+S S S E E + + N + +K D + IPEDFLCPISLELM+DP IV
Sbjct: 203 PWKSSSVSLAFFLSKDGDDERLEKAVTENSDDSQKSDNLTIPEDFLCPISLELMKDPAIV 262
Query: 181 ATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLT 240
+TGQTYERS+IQRWIDCGN +CPKTQQKL++ TLTPNYVLRSL+SQWC +HNIEQP G
Sbjct: 263 STGQTYERSFIQRWIDCGNLSCPKTQQKLENFTLTPNYVLRSLISQWCTKHNIEQPGGYM 322
Query: 241 NGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEA 300
NG+ K SDGSFRD++GD++AI LV KLS +S+E+ R AV+EIRSLSKRSTDNRILIAEA
Sbjct: 323 NGRTKNSDGSFRDLSGDMSAIRALVCKLSSQSIEDRRTAVSEIRSLSKRSTDNRILIAEA 382
Query: 301 GAIPVLVSLLTSE-DVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEAR 359
GAIPVLV LLTS+ D TQENAVT ILNLSIYE+NK LIMLAGA+ SIV VLRAG+MEAR
Sbjct: 383 GAIPVLVKLLTSDGDTETQENAVTCILNLSIYEHNKELIMLAGAVTSIVLVLRAGSMEAR 442
Query: 360 ENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGR 419
ENAAATLFSLSLADENKIIIGASGAI ALVDLLQ GS RGKKDAATALFNLCIYQGNKGR
Sbjct: 443 ENAAATLFSLSLADENKIIIGASGAIMALVDLLQYGSVRGKKDAATALFNLCIYQGNKGR 502
Query: 420 AIRAGIITALLNMLTD-SSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGL 478
A+RAGI+ L+ MLTD SS+ M DEALTI+SVLAS+Q AK +I++A+ IP LID L+
Sbjct: 503 AVRAGIVKPLVKMLTDSSSERMADEALTILSVLASNQVAKTAILRANAIPPLIDCLQKDQ 562
Query: 479 PRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRK 536
PRN+ENAAAILL LCKRDT+ L I RLGAV+PL EL+R GTERAKRKA SLLE LRK
Sbjct: 563 PRNRENAAAILLCLCKRDTEKLISIGRLGAVVPLMELSRDGTERAKRKANSLLELLRK 620
>UniRef100_Q9SSJ6 F15H11.22 protein [Arabidopsis thaliana]
Length = 530
Score = 634 bits (1636), Expect = e-180
Identities = 354/538 (65%), Positives = 411/538 (75%), Gaps = 18/538 (3%)
Query: 1 DGAAKKIIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGFMISK 60
DGAAK+I FQFQ VTWKLEK L L YD DIS+EV+EQV+L R QLRRA +YG + SK
Sbjct: 1 DGAAKRISFQFQCVTWKLEKALGDLTYDRYDISDEVREQVELARLQLRRAMQRYGSLNSK 60
Query: 61 MPSFDSSQPLAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPFGAGSRLE 120
S S+P+ ++ S + + K S PE + L S+E K +P
Sbjct: 61 KFSSGLSEPMEKDASS----NRKVIEKLESIPETVHSL-----SDEKKFESP-------P 104
Query: 121 RTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELMRDPVIV 180
+S S S E E + + N + +K D + IPEDFLCPISLELM+DP IV
Sbjct: 105 PWKSSSVSLAFFLSKDGDDERLEKAVTENSDDSQKSDNLTIPEDFLCPISLELMKDPAIV 164
Query: 181 ATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLT 240
+TGQTYERS+IQRWIDCGN +CPKTQQKL++ TLTPNYVLRSL+SQWC +HNIEQP G
Sbjct: 165 STGQTYERSFIQRWIDCGNLSCPKTQQKLENFTLTPNYVLRSLISQWCTKHNIEQPGGYM 224
Query: 241 NGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEA 300
NG+ K SDGSFRD++GD++AI LV KLS +S+E+ R AV+EIRSLSKRSTDNRILIAEA
Sbjct: 225 NGRTKNSDGSFRDLSGDMSAIRALVCKLSSQSIEDRRTAVSEIRSLSKRSTDNRILIAEA 284
Query: 301 GAIPVLVSLLTSE-DVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEAR 359
GAIPVLV LLTS+ D TQENAVT ILNLSIYE+NK LIMLAGA+ SIV VLRAG+MEAR
Sbjct: 285 GAIPVLVKLLTSDGDTETQENAVTCILNLSIYEHNKELIMLAGAVTSIVLVLRAGSMEAR 344
Query: 360 ENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGR 419
ENAAATLFSLSLADENKIIIGASGAI ALVDLLQ GS RGKKDAATALFNLCIYQGNKGR
Sbjct: 345 ENAAATLFSLSLADENKIIIGASGAIMALVDLLQYGSVRGKKDAATALFNLCIYQGNKGR 404
Query: 420 AIRAGIITALLNMLTD-SSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGL 478
A+RAGI+ L+ MLTD SS+ M DEALTI+SVLAS+Q AK +I++A+ IP LID L+
Sbjct: 405 AVRAGIVKPLVKMLTDSSSERMADEALTILSVLASNQVAKTAILRANAIPPLIDCLQKDQ 464
Query: 479 PRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRK 536
PRN+ENAAAILL LCKRDT+ L I RLGAV+PL EL+R GTERAKRKA SLLE LRK
Sbjct: 465 PRNRENAAAILLCLCKRDTEKLISIGRLGAVVPLMELSRDGTERAKRKANSLLELLRK 522
>UniRef100_Q8GUG9 Hypothetical protein At1g23030 [Arabidopsis thaliana]
Length = 612
Score = 633 bits (1633), Expect = e-180
Identities = 350/540 (64%), Positives = 414/540 (75%), Gaps = 22/540 (4%)
Query: 1 DGAAKKIIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGFMISK 60
DGAAK+I FQFQ VTWKLEK LS+LPYD DIS+EV EQV+L R+QLRRA +YG + S
Sbjct: 94 DGAAKRISFQFQCVTWKLEKALSNLPYDLYDISDEVGEQVELARSQLRRAMQRYGSLNSN 153
Query: 61 MPSFDSSQPLAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPFGAGSRLE 120
S S+P+ ++ G S K E++SE + E +S P L
Sbjct: 154 KFSSALSEPMERD-----GFSNVIKIKAEEKLESVSETLHFGEEEEKQSSPP------LR 202
Query: 121 RTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELMRDPVIV 180
R+ SI + +S T + ++ N E KK D + IP DFLCP+SLELM+DPVIV
Sbjct: 203 RSSSISLAYYLSKDADTDRLDKMVNK--NTDESKKSDKLTIPVDFLCPVSLELMKDPVIV 260
Query: 181 ATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLT 240
ATGQTYER+YIQRWIDCGN TCPKTQQKL++ TLTPNYVLRSL+S+WC EHNIEQP G
Sbjct: 261 ATGQTYERAYIQRWIDCGNLTCPKTQQKLENFTLTPNYVLRSLISRWCAEHNIEQPAGYI 320
Query: 241 NGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEA 300
NG+ K S GD++ I LV++LS RS E+ R AV+EIRSLSKRSTDNRILIAEA
Sbjct: 321 NGRTKNS--------GDMSVIRALVQRLSSRSTEDRRNAVSEIRSLSKRSTDNRILIAEA 372
Query: 301 GAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARE 360
GAIPVLV+LLTSEDV TQENA+T +LNLSIYENNK LIM AGA+ SIVQVLRAGTMEARE
Sbjct: 373 GAIPVLVNLLTSEDVATQENAITCVLNLSIYENNKELIMFAGAVTSIVQVLRAGTMEARE 432
Query: 361 NAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRA 420
NAAATLFSLSLADENKIIIG SGAI ALVDLL+NG+PRGKKDAATALFNLCIY GNKGRA
Sbjct: 433 NAAATLFSLSLADENKIIIGGSGAIPALVDLLENGTPRGKKDAATALFNLCIYHGNKGRA 492
Query: 421 IRAGIITALLNMLTDSSK-SMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLP 479
+RAGI+TAL+ ML+DS++ MVDEALTI+SVLA++Q+AK +IVKA+T+P LI +L+T
Sbjct: 493 VRAGIVTALVKMLSDSTRHRMVDEALTILSVLANNQDAKSAIVKANTLPALIGILQTDQT 552
Query: 480 RNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRKLQQ 539
RN+ENAAAILL+LCKRDT+ L I RLGAV+PL +L++ GTER KRKA SLLE LRK Q
Sbjct: 553 RNRENAAAILLSLCKRDTEKLITIGRLGAVVPLMDLSKNGTERGKRKAISLLELLRKACQ 612
>UniRef100_O23121 F10G19.3 protein [Arabidopsis thaliana]
Length = 618
Score = 633 bits (1633), Expect = e-180
Identities = 350/540 (64%), Positives = 414/540 (75%), Gaps = 22/540 (4%)
Query: 1 DGAAKKIIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGFMISK 60
DGAAK+I FQFQ VTWKLEK LS+LPYD DIS+EV EQV+L R+QLRRA +YG + S
Sbjct: 100 DGAAKRISFQFQCVTWKLEKALSNLPYDLYDISDEVGEQVELARSQLRRAMQRYGSLNSN 159
Query: 61 MPSFDSSQPLAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPFGAGSRLE 120
S S+P+ ++ G S K E++SE + E +S P L
Sbjct: 160 KFSSALSEPMERD-----GFSNVIKIKAEEKLESVSETLHFGEEEEKQSSPP------LR 208
Query: 121 RTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELMRDPVIV 180
R+ SI + +S T + ++ N E KK D + IP DFLCP+SLELM+DPVIV
Sbjct: 209 RSSSISLAYYLSKDADTDRLDKMVNK--NTDESKKSDKLTIPVDFLCPVSLELMKDPVIV 266
Query: 181 ATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLT 240
ATGQTYER+YIQRWIDCGN TCPKTQQKL++ TLTPNYVLRSL+S+WC EHNIEQP G
Sbjct: 267 ATGQTYERAYIQRWIDCGNLTCPKTQQKLENFTLTPNYVLRSLISRWCAEHNIEQPAGYI 326
Query: 241 NGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEA 300
NG+ K S GD++ I LV++LS RS E+ R AV+EIRSLSKRSTDNRILIAEA
Sbjct: 327 NGRTKNS--------GDMSVIRALVQRLSSRSTEDRRNAVSEIRSLSKRSTDNRILIAEA 378
Query: 301 GAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARE 360
GAIPVLV+LLTSEDV TQENA+T +LNLSIYENNK LIM AGA+ SIVQVLRAGTMEARE
Sbjct: 379 GAIPVLVNLLTSEDVATQENAITCVLNLSIYENNKELIMFAGAVTSIVQVLRAGTMEARE 438
Query: 361 NAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRA 420
NAAATLFSLSLADENKIIIG SGAI ALVDLL+NG+PRGKKDAATALFNLCIY GNKGRA
Sbjct: 439 NAAATLFSLSLADENKIIIGGSGAIPALVDLLENGTPRGKKDAATALFNLCIYHGNKGRA 498
Query: 421 IRAGIITALLNMLTDSSK-SMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLP 479
+RAGI+TAL+ ML+DS++ MVDEALTI+SVLA++Q+AK +IVKA+T+P LI +L+T
Sbjct: 499 VRAGIVTALVKMLSDSTRHRMVDEALTILSVLANNQDAKSAIVKANTLPALIGILQTDQT 558
Query: 480 RNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRKLQQ 539
RN+ENAAAILL+LCKRDT+ L I RLGAV+PL +L++ GTER KRKA SLLE LRK Q
Sbjct: 559 RNRENAAAILLSLCKRDTEKLITIGRLGAVVPLMDLSKNGTERGKRKAISLLELLRKACQ 618
>UniRef100_Q6Z2K3 Putative Avr9/Cf-9 rapidly elicited protein 276 [Oryza sativa]
Length = 637
Score = 566 bits (1459), Expect = e-160
Identities = 326/560 (58%), Positives = 400/560 (71%), Gaps = 38/560 (6%)
Query: 1 DGAAKKIIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGFM-IS 59
D + QF+ VTW+L+ +L+ LP IS+EV+E+VDLVR QLRR +K G + ++
Sbjct: 96 DAVCNNVAVQFKFVTWQLQTVLARLPQSCFQISDEVQEEVDLVRAQLRREMEKKGDIDVN 155
Query: 60 KMPSFDSSQPLAQEISQVLGQSVSGLHKQHSCP--ENL--------------SELGSIPK 103
F LA +S V QS H Q P ENL SE+ +PK
Sbjct: 156 IFSKFHDI--LALHVSTVGSQSEQS-HGQPDTPQMENLCNGHLELQNIIMLVSEISGVPK 212
Query: 104 SNEGKSCNPFGAGSRLERTRSIHASSEVSFSIKTAPESQEISGSGNLPEV-KKPDAIVIP 162
S+ + + G LE R + VS S S E S PE KK DA+ IP
Sbjct: 213 SDAERITSQLIEG--LENMRVTDSKKPVSVS----QSSDETKAS---PETHKKSDAVAIP 263
Query: 163 EDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRS 222
EDF CPISLELMRDPVIV+TGQTYER++IQRWIDCGN TCPKTQ KLQ++TLTPNYVLRS
Sbjct: 264 EDFRCPISLELMRDPVIVSTGQTYERAFIQRWIDCGNRTCPKTQLKLQNITLTPNYVLRS 323
Query: 223 LVSQWCIEHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAE 282
L+ QWC E IE PT K+DG++ +V G+ AIETLVR LS S++E ++A AE
Sbjct: 324 LILQWCEEKGIEPPTR------SKNDGAYLEVGGERVAIETLVRNLSSSSLDERKSAAAE 377
Query: 283 IRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAG 342
IRSL+K+STDNRIL+AE+GAI LV LL+S+D+ TQE+AVT++LNLSIY+ NK LI++AG
Sbjct: 378 IRSLAKKSTDNRILLAESGAISALVKLLSSKDLKTQEHAVTALLNLSIYDQNKELIVVAG 437
Query: 343 AIPSIVQVLRAGTMEARENAAATLFSLSLADENKIIIGAS-GAISALVDLLQNGSPRGKK 401
AI I+QVLR G MEARENAAA +FSLSL D+NKI IG++ GAI ALV+LLQ+GSPRG+K
Sbjct: 438 AIVPIIQVLRKGGMEARENAAAAIFSLSLIDDNKITIGSTPGAIEALVELLQSGSPRGRK 497
Query: 402 DAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKS-MVDEALTIMSVLASHQEAKVS 460
DAATALFNLCIYQ NK RA+RAGI+ L+ ML DSS++ +DEALTI+SVL SH E K++
Sbjct: 498 DAATALFNLCIYQANKVRAVRAGILAPLIQMLQDSSRNGAIDEALTILSVLVSHHECKIA 557
Query: 461 IVKASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGT 520
I KA IP LIDLLR+ RNKENAAAILLALCK+D +NL+CI RLGA IPL+EL++TGT
Sbjct: 558 IAKAHAIPFLIDLLRSSQARNKENAAAILLALCKKDAENLACIGRLGAQIPLTELSKTGT 617
Query: 521 ERAKRKATSLLEHLRKLQQL 540
+RAKRKATSLLEHL KLQ L
Sbjct: 618 DRAKRKATSLLEHLSKLQVL 637
>UniRef100_Q9SNC6 Arm repeat containing protein homolog [Arabidopsis thaliana]
Length = 660
Score = 452 bits (1162), Expect = e-125
Identities = 258/557 (46%), Positives = 360/557 (64%), Gaps = 41/557 (7%)
Query: 5 KKIIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGF-------- 56
+++ + V+ KLE+ LS +PY++LDIS+EV+EQV+LV +Q RRA +
Sbjct: 95 EQVTSKLMEVSVKLEQSLSQIPYEELDISDEVREQVELVLSQFRRAKGRVDVSDDELYED 154
Query: 57 ---MISKMPSFDSSQPLAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPF 113
+ +K D+ QP+ + +++ L +L E+ + + E + +
Sbjct: 155 LQSLCNKSSDVDAYQPVLERVAKKL---------------HLMEIPDL--AQESVALHEM 197
Query: 114 GAGSRLERTRSIHASSEVSFSIKTAPESQEISG-----------SGNLPEVKKPDAIVIP 162
A S + +I + V IK ++++ +G +G VIP
Sbjct: 198 VASSGGDVGENIEEMAMVLKMIKDFVQTEDDNGEEQKVGVNSRSNGQTSTAASQKIPVIP 257
Query: 163 EDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRS 222
+DF CPISLE+MRDPVIV++GQTYER+ I++WI+ G++TCPKTQQ L TLTPNYVLRS
Sbjct: 258 DDFRCPISLEMMRDPVIVSSGQTYERTCIEKWIEGGHSTCPKTQQALTSTTLTPNYVLRS 317
Query: 223 LVSQWCIEHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAE 282
L++QWC ++IE P ++ + +K SF + IE L+ +L+ + E+ R+A E
Sbjct: 318 LIAQWCEANDIEPPKPPSSLRPRKVS-SFSS-PAEANKIEDLMWRLAYGNPEDQRSAAGE 375
Query: 283 IRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAG 342
IR L+KR+ DNR+ IAEAGAIP+LV LL++ D QE++VT++LNLSI ENNKG I+ AG
Sbjct: 376 IRLLAKRNADNRVAIAEAGAIPLLVGLLSTPDSRIQEHSVTALLNLSICENNKGAIVSAG 435
Query: 343 AIPSIVQVLRAGTMEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKD 402
AIP IVQVL+ G+MEARENAAATLFSLS+ DENK+ IGA GAI LV LL G+ RGKKD
Sbjct: 436 AIPGIVQVLKKGSMEARENAAATLFSLSVIDENKVTIGALGAIPPLVVLLNEGTQRGKKD 495
Query: 403 AATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIV 462
AATALFNLCIYQGNKG+AIRAG+I L +LT+ MVDEAL I+++L+SH E K I
Sbjct: 496 AATALFNLCIYQGNKGKAIRAGVIPTLTRLLTEPGSGMVDEALAILAILSSHPEGKAIIG 555
Query: 463 KASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTER 522
+ +P L++ +RTG PRN+ENAAA+L+ LC D +L +LG + PL +LA GT+R
Sbjct: 556 SSDAVPSLVEFIRTGSPRNRENAAAVLVHLCSGDPQHLVEAQKLGLMGPLIDLAGNGTDR 615
Query: 523 AKRKATSLLEHLRKLQQ 539
KRKA LLE + +L +
Sbjct: 616 GKRKAAQLLERISRLAE 632
>UniRef100_Q8VZ40 Hypothetical protein At3g54850 [Arabidopsis thaliana]
Length = 632
Score = 440 bits (1132), Expect = e-122
Identities = 251/542 (46%), Positives = 354/542 (65%), Gaps = 28/542 (5%)
Query: 7 IIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGFMISKMPS--- 63
++ +F+ +T ++E LS +PY+ +++SEEV+EQV L+ Q +RA +++ ++
Sbjct: 101 LVEKFRDMTVEIEAALSQIPYEKIEVSEEVREQVQLLHFQFKRAKERWEESDLQLSHDLA 160
Query: 64 -----FDSSQPLAQEISQVLG-QSVSGLHKQ-HSCPENLSELGSIPKSNEGKSCNPFGAG 116
D + + +SQ L ++ L K+ H+ E P C
Sbjct: 161 MAENVMDPDPIILKRLSQELQLTTIDELKKESHAIHEYFLSYDGDPDD-----C------ 209
Query: 117 SRLERTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELMRD 176
ER S+ + V F + + +GS + + P VIPE F CPISLELM+D
Sbjct: 210 --FERMSSL-LKNLVDFVTMESSDPDPSTGSRIVSRHRSP---VIPEYFRCPISLELMKD 263
Query: 177 PVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQP 236
PVIV+TGQTYERS IQ+W+D G+ TCPK+Q+ L H LTPNYVL+SL++ WC + IE P
Sbjct: 264 PVIVSTGQTYERSSIQKWLDAGHKTCPKSQETLLHAGLTPNYVLKSLIALWCESNGIELP 323
Query: 237 TGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRIL 296
+ + K GS D + +L+ KL+ + E+ RAA E+R L+KR+ DNR+
Sbjct: 324 QNQGSCRTTKIGGSSSSDC-DRTFVLSLLEKLANGTTEQQRAAAGELRLLAKRNVDNRVC 382
Query: 297 IAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTM 356
IAEAGAIP+LV LL+S D TQE++VT++LNLSI E NKG I+ AGAI IV+VL+ G+M
Sbjct: 383 IAEAGAIPLLVELLSSPDPRTQEHSVTALLNLSINEGNKGAIVDAGAITDIVEVLKNGSM 442
Query: 357 EARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGN 416
EARENAAATLFSLS+ DENK+ IGA+GAI AL+ LL+ G+ RGKKDAATA+FNLCIYQGN
Sbjct: 443 EARENAAATLFSLSVIDENKVAIGAAGAIQALISLLEEGTRRGKKDAATAIFNLCIYQGN 502
Query: 417 KGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRT 476
K RA++ GI+ L +L D+ MVDEAL I+++L+++QE K +I +A +IPVL++++RT
Sbjct: 503 KSRAVKGGIVDPLTRLLKDAGGGMVDEALAILAILSTNQEGKTAIAEAESIPVLVEIIRT 562
Query: 477 GLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRK 536
G PRN+ENAAAIL LC + + L+ +GA + L EL GT+RAKRKA SLLE +++
Sbjct: 563 GSPRNRENAAAILWYLCIGNIERLNVAREVGADVALKELTENGTDRAKRKAASLLELIQQ 622
Query: 537 LQ 538
+
Sbjct: 623 TE 624
>UniRef100_Q64HA9 Cell death-related protein SPL11 [Oryza sativa]
Length = 694
Score = 437 bits (1125), Expect = e-121
Identities = 251/537 (46%), Positives = 347/537 (63%), Gaps = 16/537 (2%)
Query: 7 IIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYG-----FMISKM 61
++ +FQ V +LE+ L +PY++LDIS+EV+EQV+LV QL+RA ++ F +
Sbjct: 119 VMKKFQGVILQLEQALCDIPYNELDISDEVREQVELVHAQLKRAKERIDMPDDEFYNDLL 178
Query: 62 PSFDSSQPLAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPFGAGSRLER 121
+D + + E++ +LG+ LH L G +P G +ER
Sbjct: 179 SVYDKNYDPSAELA-ILGRLSEKLHLMTITDLTQESLALHEMVASGGGQDP---GEHIER 234
Query: 122 TRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELMRDPVIVA 181
S+ F P+ S L I IP++F CPISLELM+DPVIV+
Sbjct: 235 M-SMLLKKIKDFVQTQNPDMGPPMASRVLDSNGDSRPITIPDEFRCPISLELMKDPVIVS 293
Query: 182 TGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLTN 241
TGQTYER+ I++WI G+ TCP TQQK+ LTPNYVLRSL+SQWC + +E P T
Sbjct: 294 TGQTYERACIEKWIASGHHTCPTTQQKMSTSALTPNYVLRSLISQWCETNGMEPPKRSTQ 353
Query: 242 GKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAG 301
S + + A I+ L+ KL EE R+A AE+R L+KR+ +NRI IAEAG
Sbjct: 354 PNKPTPACS----SSERANIDALLSKLCSPDTEEQRSAAAELRLLAKRNANNRICIAEAG 409
Query: 302 AIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEAREN 361
AIP+L+SLL+S D+ TQE+AVT++LNLSI+E+NK I+ +GA+PSIV VL+ G+MEAREN
Sbjct: 410 AIPLLLSLLSSSDLRTQEHAVTALLNLSIHEDNKASIISSGAVPSIVHVLKNGSMEAREN 469
Query: 362 AAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAI 421
AAATLFSLS+ DE K+ IG GAI ALV LL GS RGKKDAA ALFNLCIYQGNKGRAI
Sbjct: 470 AAATLFSLSVIDEYKVTIGGMGAIPALVVLLGEGSQRGKKDAAAALFNLCIYQGNKGRAI 529
Query: 422 RAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRN 481
RAG++ ++ ++T+ + +++DEA+ I+S+L+SH E K +I A +PVL++++ +G PRN
Sbjct: 530 RAGLVPLIMGLVTNPTGALMDEAMAILSILSSHPEGKAAIGAAEPVPVLVEMIGSGTPRN 589
Query: 482 KENAAAILLALCKRDTD--NLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRK 536
+ENAAA++L LC + +L+ G ++PL ELA GT+R KRKA LLE + +
Sbjct: 590 RENAAAVMLHLCSGEHHLVHLARAQECGIMVPLRELALNGTDRGKRKAVQLLERMSR 646
Score = 39.7 bits (91), Expect = 0.23
Identities = 29/118 (24%), Positives = 54/118 (45%), Gaps = 1/118 (0%)
Query: 422 RAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRN 481
RA I L + + ++ A + + + ++ I +A IP+L+ LL + R
Sbjct: 366 RANIDALLSKLCSPDTEEQRSAAAELRLLAKRNANNRICIAEAGAIPLLLSLLSSSDLRT 425
Query: 482 KENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRKLQQ 539
+E+A LL L + DN + I GAV + + + G+ A+ A + L L + +
Sbjct: 426 QEHAVTALLNLSIHE-DNKASIISSGAVPSIVHVLKNGSMEARENAAATLFSLSVIDE 482
>UniRef100_Q5VRH9 Putative cell death-related protein SPL11 [Oryza sativa]
Length = 611
Score = 434 bits (1116), Expect = e-120
Identities = 245/532 (46%), Positives = 346/532 (64%), Gaps = 14/532 (2%)
Query: 10 QFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGFMISKMPSFDSSQP 69
+F V ++ L +LPY+ + +EV+EQV LV +Q +RA+ + + P S
Sbjct: 84 EFAGVNRQIHLALDALPYNTFHMPQEVQEQVALVHSQFQRASTR-----TDPPDTQLSMD 138
Query: 70 LAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPFGAGSRLERTRSIHASS 129
LA ++ S L + S L + + NE + + + E + S
Sbjct: 139 LAWALTD--NPSDPALLTRISHKLQLHTMADM--KNESIALHNMVISTAGEPDGCVDQMS 194
Query: 130 EVSFSIKTAPESQEISGSGNLPEVK--KPDAIVIPEDFLCPISLELMRDPVIVATGQTYE 187
+ +K +++ + K + +IP++F CPISLELM+DPVIV++GQTYE
Sbjct: 195 SLLKKLKDCVVTEDHANDALTTRSASIKHRSPIIPDEFRCPISLELMQDPVIVSSGQTYE 254
Query: 188 RSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLTNGKIKKS 247
RS IQ+W+D G+ TCPKTQQ L H +LTPN+VL+SL+SQWC + IE P N + KK+
Sbjct: 255 RSCIQKWLDSGHKTCPKTQQPLSHTSLTPNFVLKSLISQWCEANGIELPKNKQNSRDKKA 314
Query: 248 DGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLV 307
S D A + +L+ +L + +E RAA EIR L+KR+ +NRI IAEAGAIP+LV
Sbjct: 315 AKSS---DYDHAGLVSLMNRLRSGNQDEQRAAAGEIRLLAKRNVNNRICIAEAGAIPLLV 371
Query: 308 SLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLF 367
+LL+S D TQE+AVT++LNLSI+ENNK I+ + AIP IV+VL+ G+ME RENAAATLF
Sbjct: 372 NLLSSSDPRTQEHAVTALLNLSIHENNKASIVDSHAIPKIVEVLKTGSMETRENAAATLF 431
Query: 368 SLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIIT 427
SLS+ DENK+ IGA+GAI L++LL +GSPRGKKDAATA+FNLCIYQGNK RA++AGI+
Sbjct: 432 SLSVVDENKVTIGAAGAIPPLINLLCDGSPRGKKDAATAIFNLCIYQGNKVRAVKAGIVI 491
Query: 428 ALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAA 487
L+N L D + M+DEAL+++S+LA + E K+ I ++ IP L+++++TG PRN+ENAAA
Sbjct: 492 HLMNFLVDPTGGMIDEALSLLSILAGNPEGKIVIARSEPIPPLVEVIKTGSPRNRENAAA 551
Query: 488 ILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRKLQQ 539
IL LC DT+ G L EL+ TGT+RAKRKA+S+LE + + +
Sbjct: 552 ILWLLCSADTEQTLAAKAAGVEDALKELSETGTDRAKRKASSILELMHQANE 603
>UniRef100_Q5VRH8 Putative cell death-related protein SPL11 [Oryza sativa]
Length = 604
Score = 434 bits (1116), Expect = e-120
Identities = 245/532 (46%), Positives = 346/532 (64%), Gaps = 14/532 (2%)
Query: 10 QFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGFMISKMPSFDSSQP 69
+F V ++ L +LPY+ + +EV+EQV LV +Q +RA+ + + P S
Sbjct: 84 EFAGVNRQIHLALDALPYNTFHMPQEVQEQVALVHSQFQRASTR-----TDPPDTQLSMD 138
Query: 70 LAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPFGAGSRLERTRSIHASS 129
LA ++ S L + S L + + NE + + + E + S
Sbjct: 139 LAWALTD--NPSDPALLTRISHKLQLHTMADM--KNESIALHNMVISTAGEPDGCVDQMS 194
Query: 130 EVSFSIKTAPESQEISGSGNLPEVK--KPDAIVIPEDFLCPISLELMRDPVIVATGQTYE 187
+ +K +++ + K + +IP++F CPISLELM+DPVIV++GQTYE
Sbjct: 195 SLLKKLKDCVVTEDHANDALTTRSASIKHRSPIIPDEFRCPISLELMQDPVIVSSGQTYE 254
Query: 188 RSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLTNGKIKKS 247
RS IQ+W+D G+ TCPKTQQ L H +LTPN+VL+SL+SQWC + IE P N + KK+
Sbjct: 255 RSCIQKWLDSGHKTCPKTQQPLSHTSLTPNFVLKSLISQWCEANGIELPKNKQNSRDKKA 314
Query: 248 DGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLV 307
S D A + +L+ +L + +E RAA EIR L+KR+ +NRI IAEAGAIP+LV
Sbjct: 315 AKSS---DYDHAGLVSLMNRLRSGNQDEQRAAAGEIRLLAKRNVNNRICIAEAGAIPLLV 371
Query: 308 SLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLF 367
+LL+S D TQE+AVT++LNLSI+ENNK I+ + AIP IV+VL+ G+ME RENAAATLF
Sbjct: 372 NLLSSSDPRTQEHAVTALLNLSIHENNKASIVDSHAIPKIVEVLKTGSMETRENAAATLF 431
Query: 368 SLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIIT 427
SLS+ DENK+ IGA+GAI L++LL +GSPRGKKDAATA+FNLCIYQGNK RA++AGI+
Sbjct: 432 SLSVVDENKVTIGAAGAIPPLINLLCDGSPRGKKDAATAIFNLCIYQGNKVRAVKAGIVI 491
Query: 428 ALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAA 487
L+N L D + M+DEAL+++S+LA + E K+ I ++ IP L+++++TG PRN+ENAAA
Sbjct: 492 HLMNFLVDPTGGMIDEALSLLSILAGNPEGKIVIARSEPIPPLVEVIKTGSPRNRENAAA 551
Query: 488 ILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRKLQQ 539
IL LC DT+ G L EL+ TGT+RAKRKA+S+LE + + +
Sbjct: 552 ILWLLCSADTEQTLAAKAAGVEDALKELSETGTDRAKRKASSILELMHQANE 603
>UniRef100_Q9SV34 Hypothetical protein F28P10.170 [Arabidopsis thaliana]
Length = 639
Score = 433 bits (1114), Expect = e-120
Identities = 251/549 (45%), Positives = 354/549 (63%), Gaps = 35/549 (6%)
Query: 7 IIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGFMISKMPS--- 63
++ +F+ +T ++E LS +PY+ +++SEEV+EQV L+ Q +RA +++ ++
Sbjct: 101 LVEKFRDMTVEIEAALSQIPYEKIEVSEEVREQVQLLHFQFKRAKERWEESDLQLSHDLA 160
Query: 64 -----FDSSQPLAQEISQVLG-QSVSGLHKQ-HSCPENLSELGSIPKSNEGKSCNPFGAG 116
D + + +SQ L ++ L K+ H+ E P C
Sbjct: 161 MAENVMDPDPIILKRLSQELQLTTIDELKKESHAIHEYFLSYDGDPDD-----C------ 209
Query: 117 SRLERTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELMRD 176
ER S+ + V F + + +GS + + P VIPE F CPISLELM+D
Sbjct: 210 --FERMSSL-LKNLVDFVTMESSDPDPSTGSRIVSRHRSP---VIPEYFRCPISLELMKD 263
Query: 177 PVIVATGQ-------TYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCI 229
PVIV+TGQ TYERS IQ+W+D G+ TCPK+Q+ L H LTPNYVL+SL++ WC
Sbjct: 264 PVIVSTGQLNFSTLQTYERSSIQKWLDAGHKTCPKSQETLLHAGLTPNYVLKSLIALWCE 323
Query: 230 EHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKR 289
+ IE P + + K GS D + +L+ KL+ + E+ RAA E+R L+KR
Sbjct: 324 SNGIELPQNQGSCRTTKIGGSSSSDC-DRTFVLSLLEKLANGTTEQQRAAAGELRLLAKR 382
Query: 290 STDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQ 349
+ DNR+ IAEAGAIP+LV LL+S D TQE++VT++LNLSI E NKG I+ AGAI IV+
Sbjct: 383 NVDNRVCIAEAGAIPLLVELLSSPDPRTQEHSVTALLNLSINEGNKGAIVDAGAITDIVE 442
Query: 350 VLRAGTMEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFN 409
VL+ G+MEARENAAATLFSLS+ DENK+ IGA+GAI AL+ LL+ G+ RGKKDAATA+FN
Sbjct: 443 VLKNGSMEARENAAATLFSLSVIDENKVAIGAAGAIQALISLLEEGTRRGKKDAATAIFN 502
Query: 410 LCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPV 469
LCIYQGNK RA++ GI+ L +L D+ MVDEAL I+++L+++QE K +I +A +IPV
Sbjct: 503 LCIYQGNKSRAVKGGIVDPLTRLLKDAGGGMVDEALAILAILSTNQEGKTAIAEAESIPV 562
Query: 470 LIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATS 529
L++++RTG PRN+ENAAAIL LC + + L+ +GA + L EL GT+RAKRKA S
Sbjct: 563 LVEIIRTGSPRNRENAAAILWYLCIGNIERLNVAREVGADVALKELTENGTDRAKRKAAS 622
Query: 530 LLEHLRKLQ 538
LLE +++ +
Sbjct: 623 LLELIQQTE 631
>UniRef100_Q9ZV31 Expressed protein [Arabidopsis thaliana]
Length = 654
Score = 416 bits (1068), Expect = e-114
Identities = 244/544 (44%), Positives = 348/544 (63%), Gaps = 18/544 (3%)
Query: 6 KIIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGF------MIS 59
+++ +FQ+VT LE+ LS +PY++L+IS+E+KEQV+LV QLRR+ K G +
Sbjct: 96 QVMVKFQKVTSLLEQALSIIPYENLEISDELKEQVELVLVQLRRSLGKRGGDVYDDELYK 155
Query: 60 KMPSFDSSQPLAQEISQV--LGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPFGAGS 117
+ S S + E V + + + + E+L+ L + S F S
Sbjct: 156 DVLSLYSGRGSVMESDMVRRVAEKLQLMTITDLTQESLALLDMVSSSGGDDPGESFEKMS 215
Query: 118 RLERTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDA--IVIPEDFLCPISLELMR 175
+ + + + ++ AP + +LP+ + D ++ PE+F CPISLELM
Sbjct: 216 MVLKKIKDFVQT-YNPNLDDAP----LRLKSSLPKSRDDDRDMLIPPEEFRCPISLELMT 270
Query: 176 DPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQ 235
DPVIV++GQTYER I++W++ G+ TCPKTQ+ L +TPNYVLRSL++QWC + IE
Sbjct: 271 DPVIVSSGQTYERECIKKWLEGGHLTCPKTQETLTSDIMTPNYVLRSLIAQWCESNGIEP 330
Query: 236 PTGLTNGK-IKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNR 294
P + K+ S + IE L+ KL+ + E+ R+A EIR L+K++ NR
Sbjct: 331 PKRPNISQPSSKASSSSSAPDDEHNKIEELLLKLTSQQPEDRRSAAGEIRLLAKQNNHNR 390
Query: 295 ILIAEAGAIPVLVSLLT-SEDVMTQENAVTSILNLSIYENNKGLIMLA-GAIPSIVQVLR 352
+ IA +GAIP+LV+LLT S D TQE+AVTSILNLSI + NKG I+ + GA+P IV VL+
Sbjct: 391 VAIAASGAIPLLVNLLTISNDSRTQEHAVTSILNLSICQENKGKIVYSSGAVPGIVHVLQ 450
Query: 353 AGTMEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCI 412
G+MEARENAAATLFSLS+ DENK+ IGA+GAI LV LL GS RGKKDAATALFNLCI
Sbjct: 451 KGSMEARENAAATLFSLSVIDENKVTIGAAGAIPPLVTLLSEGSQRGKKDAATALFNLCI 510
Query: 413 YQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLID 472
+QGNKG+A+RAG++ L+ +LT+ MVDE+L+I+++L+SH + K + A +PVL+D
Sbjct: 511 FQGNKGKAVRAGLVPVLMRLLTEPESGMVDESLSILAILSSHPDGKSEVGAADAVPVLVD 570
Query: 473 LLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLE 532
+R+G PRNKEN+AA+L+ LC + +L +LG + L E+A GT+R KRKA LL
Sbjct: 571 FIRSGSPRNKENSAAVLVHLCSWNQQHLIEAQKLGIMDLLIEMAENGTDRGKRKAAQLLN 630
Query: 533 HLRK 536
+
Sbjct: 631 RFSR 634
>UniRef100_Q94C99 Hypothetical protein At2g28830 [Arabidopsis thaliana]
Length = 654
Score = 414 bits (1065), Expect = e-114
Identities = 243/544 (44%), Positives = 348/544 (63%), Gaps = 18/544 (3%)
Query: 6 KIIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGF------MIS 59
+++ +FQ+VT LE+ LS +PY++L+IS+E+KEQV+LV QLRR+ K G +
Sbjct: 96 QVMVKFQKVTSLLEQALSIIPYENLEISDELKEQVELVLVQLRRSLGKRGGDVYDDELYK 155
Query: 60 KMPSFDSSQPLAQEISQV--LGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPFGAGS 117
+ S S + E V + + + + E+L+ L + S F S
Sbjct: 156 DVLSLYSGRGSVMESDMVRRVAEKLQLMTITDLTQESLALLDMVSSSGGDDPGESFEKMS 215
Query: 118 RLERTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDA--IVIPEDFLCPISLELMR 175
+ + + + ++ AP + +LP+ + D ++ PE+F CPISLELM
Sbjct: 216 MVLKKIKDFVQT-YNPNLDDAP----LRLKSSLPKSRDDDRDMLIPPEEFRCPISLELMT 270
Query: 176 DPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQ 235
DPVIV++GQTYER I++W++ G+ TCPKTQ+ L +TPNYVLRSL++QWC + IE
Sbjct: 271 DPVIVSSGQTYERECIKKWLEGGHLTCPKTQETLTSDIMTPNYVLRSLIAQWCESNGIEP 330
Query: 236 PTGLTNGK-IKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNR 294
P + K+ S + IE L+ KL+ + E+ ++A EIR L+K++ NR
Sbjct: 331 PKRPNISQPSSKASSSSSAPDDEHNKIEELLLKLTSQQPEDRKSAAGEIRLLAKQNNHNR 390
Query: 295 ILIAEAGAIPVLVSLLT-SEDVMTQENAVTSILNLSIYENNKGLIMLA-GAIPSIVQVLR 352
+ IA +GAIP+LV+LLT S D TQE+AVTSILNLSI + NKG I+ + GA+P IV VL+
Sbjct: 391 VAIAASGAIPLLVNLLTISNDSRTQEHAVTSILNLSICQENKGKIVYSSGAVPGIVHVLQ 450
Query: 353 AGTMEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCI 412
G+MEARENAAATLFSLS+ DENK+ IGA+GAI LV LL GS RGKKDAATALFNLCI
Sbjct: 451 KGSMEARENAAATLFSLSVIDENKVTIGAAGAIPPLVTLLSEGSQRGKKDAATALFNLCI 510
Query: 413 YQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLID 472
+QGNKG+A+RAG++ L+ +LT+ MVDE+L+I+++L+SH + K + A +PVL+D
Sbjct: 511 FQGNKGKAVRAGLVPVLMRLLTEPESGMVDESLSILAILSSHPDGKSEVGAADAVPVLVD 570
Query: 473 LLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLE 532
+R+G PRNKEN+AA+L+ LC + +L +LG + L E+A GT+R KRKA LL
Sbjct: 571 FIRSGSPRNKENSAAVLVHLCSWNQQHLIEAQKLGIMDLLIEMAENGTDRGKRKAAQLLN 630
Query: 533 HLRK 536
+
Sbjct: 631 RFSR 634
>UniRef100_Q9FII3 Arm repeat containing protein [Arabidopsis thaliana]
Length = 656
Score = 365 bits (936), Expect = 2e-99
Identities = 216/547 (39%), Positives = 323/547 (58%), Gaps = 44/547 (8%)
Query: 10 QFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRA-----TDKYGFMISKMPSF 64
+F + KL ++L P+D+L IS + K+++D + QL++A T + M F
Sbjct: 138 RFHSIYEKLNRVLVKAPFDELMISGDAKDEIDSLCKQLKKAKRRTDTQDIELAVDMMVVF 197
Query: 65 DSSQP------LAQEISQVLG-QSVSGLHKQHSCPENLSE----LGSIPKSNEGKSCNPF 113
+ P + + +++ L Q++ L + ++L + L K + + N F
Sbjct: 198 SKTDPRNADSAIIERLAKKLELQTIDDLKTETIAIQSLIQDKGGLNIETKQHIIELLNKF 257
Query: 114 GAGSRLERTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLEL 173
LE T ++ Q + + K ++++P +FLCPI+LE+
Sbjct: 258 KKLQGLEATDILY---------------QPVINKA----ITKSTSLILPHEFLCPITLEI 298
Query: 174 MRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNI 233
M DPVI+ATGQTYE+ IQ+W D G+ TCPKT+Q+L HL+L PN+ L++L+ QWC ++N
Sbjct: 299 MLDPVIIATGQTYEKESIQKWFDAGHKTCPKTRQELDHLSLAPNFALKNLIMQWCEKNNF 358
Query: 234 EQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDN 293
KI + + S + LV LS +EE R +V ++R L++ + +N
Sbjct: 359 ---------KIPEKEVSPDSQNEQKDEVSLLVEALSSSQLEEQRRSVKQMRLLARENPEN 409
Query: 294 RILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRA 353
R+LIA AGAIP+LV LL+ D QENAVT++LNLSI E NK LI GAIP+I+++L
Sbjct: 410 RVLIANAGAIPLLVQLLSYPDSGIQENAVTTLLNLSIDEVNKKLISNEGAIPNIIEILEN 469
Query: 354 GTMEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIY 413
G EAREN+AA LFSLS+ DENK+ IG S I LVDLLQ+G+ RGKKDA TALFNL +
Sbjct: 470 GNREARENSAAALFSLSMLDENKVTIGLSNGIPPLVDLLQHGTLRGKKDALTALFNLSLN 529
Query: 414 QGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDL 473
NKGRAI AGI+ LLN+L D + M+DEAL+I+ +LASH E + +I + S I L++
Sbjct: 530 SANKGRAIDAGIVQPLLNLLKDKNLGMIDEALSILLLLASHPEGRQAIGQLSFIETLVEF 589
Query: 474 LRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEH 533
+R G P+NKE A ++LL L ++ + + G L E+ +GT RA+RKA +L++
Sbjct: 590 IRQGTPKNKECATSVLLELGSNNSSFILAALQFGVYEYLVEITTSGTNRAQRKANALIQL 649
Query: 534 LRKLQQL 540
+ K +Q+
Sbjct: 650 ISKSEQI 656
>UniRef100_Q681N2 Arm repeat containing protein [Arabidopsis thaliana]
Length = 660
Score = 362 bits (929), Expect = 2e-98
Identities = 215/547 (39%), Positives = 322/547 (58%), Gaps = 44/547 (8%)
Query: 10 QFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRA-----TDKYGFMISKMPSF 64
+F + KL ++L P+D+L IS + K+++D + QL++A T + M F
Sbjct: 142 RFHSIYEKLNRVLVKAPFDELMISGDAKDEIDSLCKQLKKAKRRTDTQDIELAVDMMVVF 201
Query: 65 DSSQP------LAQEISQVLG-QSVSGLHKQHSCPENLSE----LGSIPKSNEGKSCNPF 113
+ P + + +++ L Q++ L + ++L + L K + + N F
Sbjct: 202 SKTDPRNADSAIIERLAKKLELQTIDDLKTETIAIQSLIQDKGGLNIETKQHIIELLNKF 261
Query: 114 GAGSRLERTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLEL 173
LE T ++ Q + + K ++++P +FLCPI+L +
Sbjct: 262 KKLQGLEATDILY---------------QPVINKA----ITKSTSLILPHEFLCPITLGI 302
Query: 174 MRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNI 233
M DPVI+ATGQTYE+ IQ+W D G+ TCPKT+Q+L HL+L PN+ L++L+ QWC ++N
Sbjct: 303 MLDPVIIATGQTYEKESIQKWFDAGHKTCPKTRQELDHLSLAPNFALKNLIMQWCEKNNF 362
Query: 234 EQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDN 293
KI + + S + LV LS +EE R +V ++R L++ + +N
Sbjct: 363 ---------KIPEKEVSPDSQNEQKDEVSLLVEALSSSQLEEQRRSVKQMRLLARENPEN 413
Query: 294 RILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRA 353
R+LIA AGAIP+LV LL+ D QENAVT++LNLSI E NK LI GAIP+I+++L
Sbjct: 414 RVLIANAGAIPLLVQLLSYPDSGIQENAVTTLLNLSIDEVNKKLISNEGAIPNIIEILEN 473
Query: 354 GTMEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIY 413
G EAREN+AA LFSLS+ DENK+ IG S I LVDLLQ+G+ RGKKDA TALFNL +
Sbjct: 474 GNREARENSAAALFSLSMLDENKVTIGLSNGIPPLVDLLQHGTLRGKKDALTALFNLSLN 533
Query: 414 QGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDL 473
NKGRAI AGI+ LLN+L D + M+DEAL+I+ +LASH E + +I + S I L++
Sbjct: 534 SANKGRAIDAGIVQPLLNLLKDKNLGMIDEALSILLLLASHPEGRQAIGQLSFIETLVEF 593
Query: 474 LRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEH 533
+R G P+NKE A ++LL L ++ + + G L E+ +GT RA+RKA +L++
Sbjct: 594 IRQGTPKNKECATSVLLELGSNNSSFILAALQFGVYEYLVEITTSGTNRAQRKANALIQL 653
Query: 534 LRKLQQL 540
+ K +Q+
Sbjct: 654 ISKSEQI 660
>UniRef100_Q6Z250 Putative Avr9/Cf-9 rapidly elicited protein [Oryza sativa]
Length = 642
Score = 360 bits (924), Expect = 6e-98
Identities = 219/532 (41%), Positives = 310/532 (58%), Gaps = 16/532 (3%)
Query: 10 QFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRA-----TDKYGFMISKMPSF 64
+F+ V K+ L +PY +L IS+EVKEQV+L+ QL R T + M
Sbjct: 120 RFRAVYEKMNSALDGMPYSELAISDEVKEQVELMNAQLTRCKKRADTQDIELSMDLMVIL 179
Query: 65 DSSQPLAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPFGAGSRLERTRS 124
D+ + +L + L Q + + +E +I K ++ G + T+
Sbjct: 180 DNKEGERNADRAILERLAKKLELQ-TLADLRAETMAIKKLISERN------GQSGDSTKQ 232
Query: 125 IHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELMRDPVIVATGQ 184
I + E + + K +++IP DFLCPI+L +MRDPVIVATGQ
Sbjct: 233 IIELLNKFKEVAGVDEKNVLGEVSVTKSLDKCPSLMIPNDFLCPITLAIMRDPVIVATGQ 292
Query: 185 TYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLTNGKI 244
TYER IQ+W+D G TCPKT+Q+L H++L PNY L++L+ +WC ++ +E
Sbjct: 293 TYERRSIQKWLDSGERTCPKTRQRLSHMSLAPNYALKNLILEWCDKNKVELQKREPEPVA 352
Query: 245 KKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIP 304
++ D R I +LV +S ++ R AV IR LSK +NR LIA++G IP
Sbjct: 353 EQDDEHQRGAED----IPSLVEGMSSIHLDVQRKAVKRIRMLSKECPENRTLIADSGGIP 408
Query: 305 VLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAA 364
L+ LL D QEN VTS+LNLSI E+NK I GA+P I+++LR G+ EA+EN+AA
Sbjct: 409 ALIGLLACPDKKVQENTVTSLLNLSIDESNKRHITKGGALPLIIEILRNGSAEAQENSAA 468
Query: 365 TLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAG 424
TLFSLS+ DENK+ IG G I+ LV+LLQNGS RGKKDAATA+FNL + Q NK RA +AG
Sbjct: 469 TLFSLSMIDENKLTIGRLGGIAPLVELLQNGSIRGKKDAATAIFNLVLNQQNKVRATQAG 528
Query: 425 IITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKEN 484
I+ ALL ++ D + +MVDEAL+I +L+S+ I I L+ L++ G P+NKE
Sbjct: 529 IVPALLKIIDDKALNMVDEALSIFLLLSSNAACCGEIGTTPFIEKLVRLIKDGTPKNKEC 588
Query: 485 AAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRK 536
A ++LL L ++ L R G LS++A+ GT RA+RKATSL++ RK
Sbjct: 589 ALSVLLELGSKNKPLLVHALRFGLHEDLSKIAKNGTSRAQRKATSLIQLARK 640
>UniRef100_Q5Z7P7 Putative cell death-related protein SPL11 [Oryza sativa]
Length = 621
Score = 342 bits (877), Expect = 2e-92
Identities = 214/543 (39%), Positives = 313/543 (57%), Gaps = 39/543 (7%)
Query: 4 AKKIIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDK------YGFM 57
++ ++ +F+ V K+ L +PY +L +S+EVKEQV+L+ QL++ + K
Sbjct: 106 SEAVMGRFRGVYEKMNMALEGMPYAELGVSDEVKEQVELISAQLKKRSKKRTETQDMELA 165
Query: 58 ISKMPSFDSSQPLAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKS-NEGKSCNPFGAG 116
+ M S + A + + ++ + S + +E +I K N+ +S +
Sbjct: 166 MDLMMILQSKEQDANNADRPILDRLAKRLQLQSLADLRAETMAIKKLINDHQSDSTNQIV 225
Query: 117 SRLERTRSIHASSEVSFSIKTAPESQEISGSGNLPE-VKKPDAIVIPEDFLCPISLELMR 175
L R ++I E + I G +P+ ++K +++IP DFLCPISLE+M
Sbjct: 226 DLLHRLKAIAGVDE-----------KNILGDVFIPKYLEKCPSLMIPNDFLCPISLEIMT 274
Query: 176 DPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQ 235
DP TYER IQ+W+D G TCPKTQQ L HL+L PNY L++L+ QWC ++ +E
Sbjct: 275 DP-------TYERRSIQKWLDAGQRTCPKTQQPLGHLSLAPNYALKNLIMQWCDKNKVE- 326
Query: 236 PTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRI 295
I D I TLV+ LS +++ R AV +IR+LSK + +NR+
Sbjct: 327 --------IHSGDPPPEPPEDPKVVIPTLVKDLSSPNLDVQRKAVKKIRTLSKENPENRL 378
Query: 296 LIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGT 355
L+ + IP L+ LL D QEN VTS+LNLSI E NK LI GAIP I+ VLR G+
Sbjct: 379 LVTDNAGIPALIGLLPYPDKKMQENTVTSLLNLSIDEANKLLIARGGAIPLIIDVLRNGS 438
Query: 356 MEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQG 415
+E +EN+AA LFSLS+ DENK+ IG G I LVDLLQNG+ RGKKDA+TA+FNL + G
Sbjct: 439 VEGQENSAAALFSLSMVDENKVAIGTLGGIPPLVDLLQNGTVRGKKDASTAIFNLMLNNG 498
Query: 416 NKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLR 475
NK RAI AGI+ LL +L D +MVDEAL+I +LAS+ + + + L+ +++
Sbjct: 499 NKLRAIEAGILPTLLKLLDDKKAAMVDEALSIFLLLASNPTCRGEVGTEHFVEKLVQIIK 558
Query: 476 TGLPRNKENAAAILLALCKRDTDNLSCISRLGAVI--PLSELARTGTERAKRKATSLLEH 533
G P+NKE A ++LL L ++N LG + L+++A+ GT RA+RKA SL++
Sbjct: 559 EGTPKNKECAVSVLLEL--GSSNNALMAHALGFDLHDHLADIAKNGTSRAQRKANSLIQL 616
Query: 534 LRK 536
RK
Sbjct: 617 ARK 619
>UniRef100_Q9C7R6 Arm repeat-containing protein, putative; 6839-9028 [Arabidopsis
thaliana]
Length = 729
Score = 279 bits (714), Expect = 1e-73
Identities = 201/545 (36%), Positives = 299/545 (53%), Gaps = 31/545 (5%)
Query: 11 FQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRAT---DKYGFMI-----SKMP 62
F + ++ LL LP +DL +S++++EQ++L++ Q R+A DK + S +
Sbjct: 143 FHDLNQEISTLLDVLPVNDLGLSDDIREQIELLQRQSRKARLYIDKNDESLRESFYSFLD 202
Query: 63 SFDSSQ-PLAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEG------KSCNPFGA 115
F++ + P + ++ + + G+ SC + L +++G N F A
Sbjct: 203 GFENGKIPSSVDLRMFFVEKL-GIRDSKSCRSEIEFLEEQIVNHDGDLEPTGSVINGFVA 261
Query: 116 GSRLERTRSIHASSE-VSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELM 174
+R R + + + I+ P+ G + + I +P+DF+CPISL+LM
Sbjct: 262 ITRYCRFLLFGFEEDGMEWWIENNPKKPR---KGFVAQEIGDTFITVPKDFVCPISLDLM 318
Query: 175 RDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIE 234
DPVI++TGQTY+R+ I RWI+ G+ TCPKT Q L + PN L++L+ QWC I
Sbjct: 319 TDPVIISTGQTYDRNSIARWIEEGHCTCPKTGQMLMDSRIVPNRALKNLIVQWCTASGIS 378
Query: 235 QPTGLT---NGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRST 291
+ T N + + V + A + L++ L+ S A EIR L+K
Sbjct: 379 YESEFTDSPNESFASALPTKAAVEANKATVSILIKYLADGSQAAQTVAAREIRLLAKTGK 438
Query: 292 DNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAG-AIPSIVQV 350
+NR IAEAGAIP L LLTSE+ + QEN+VT++LNLSIYE NK IM G + SIV V
Sbjct: 439 ENRAYIAEAGAIPHLCRLLTSENAIAQENSVTAMLNLSIYEKNKSRIMEEGDCLESIVSV 498
Query: 351 LRAG-TMEARENAAATLFSLSLADE-NKIIIGASGAISALVDLLQNGSPRGKKDAATALF 408
L +G T+EA+ENAAATLFSLS E K I + AL LLQNG+PRGKKDA TAL+
Sbjct: 499 LVSGLTVEAQENAAATLFSLSAVHEYKKRIAIVDQCVEALALLLQNGTPRGKKDAVTALY 558
Query: 409 NLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKA-STI 467
NL + N R I G +++L+ L ++ + +EA +++L +I K S +
Sbjct: 559 NLSTHPDNCSRMIEGGGVSSLVGAL--KNEGVAEEAAGALALLVRQSLGAEAIGKEDSAV 616
Query: 468 PVLIDLLRTGLPRNKENAAAILLALCKRDTDNLS-CISRLGAVIPLSE-LARTGTERAKR 525
L+ ++R G PR KENA A LL LC+ ++ + R A+ L + L TGT+RA+R
Sbjct: 617 AGLMGMMRCGTPRGKENAVAALLELCRSGGAAVAEKVLRAPAIAGLLQTLLFTGTKRARR 676
Query: 526 KATSL 530
KA SL
Sbjct: 677 KAASL 681
>UniRef100_Q84QD7 Avr9/Cf-9 rapidly elicited protein 276 [Nicotiana tabacum]
Length = 726
Score = 276 bits (707), Expect = 8e-73
Identities = 208/549 (37%), Positives = 293/549 (52%), Gaps = 39/549 (7%)
Query: 11 FQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRA---TDKYGFMI-----SKMP 62
F + ++ LL P +L + E+V EQV+L++ Q R++ DK+ M+ S +
Sbjct: 138 FHDLNQEISTLLDVFPLKELTLPEDVMEQVELLQKQARKSMLFVDKHDEMLRLKLFSFLN 197
Query: 63 SFDSS--QPLAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEG------KSCNPFG 114
F++ AQ S + + V + SC + L ++EG N F
Sbjct: 198 EFENGGIPGSAQLYSFFVDKLV--ICNPRSCRVEIEFLEEQIVNHEGDIEPTASVLNGFV 255
Query: 115 AGSRLERTRSI-HASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLEL 173
A R R +V + + ++ G + + +I +P+DF CPISL+L
Sbjct: 256 ALIRYCRFLLFGFEEDDVGLGVGKHKKQKK----GLISQEIADTSISVPKDFCCPISLDL 311
Query: 174 MRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNI 233
MRDPVIV+TGQTY+R+ I RW++ G+ TCPKT Q L H L PN LR+L+ QWC H I
Sbjct: 312 MRDPVIVSTGQTYDRASISRWMEEGHCTCPKTGQLLDHTRLVPNRALRNLIMQWCAAHKI 371
Query: 234 EQPTGLTNGKIKKSDG----SFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKR 289
+S G S V + A L+++L+ + A EIR L+K
Sbjct: 372 PYDNMEGGDPCVESFGAASPSKAAVEANRATTALLIKQLANGTQIAKTIAAREIRLLAKT 431
Query: 290 STDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIM-LAGAIPSIV 348
+NR IAEAGAIP L +LL+S D + QEN+VT++LNLSI++ NKG IM G + +V
Sbjct: 432 GKENRAYIAEAGAIPHLKNLLSSPDAVAQENSVTAMLNLSIFDKNKGRIMDEVGCLTLVV 491
Query: 349 QVLRAG-TMEARENAAATLFSLS-LADENKIIIGASGAISALVDLLQNGSPRGKKDAATA 406
VL G T EARENAAATLFSLS + D K I GA+ AL LL+ GSPRGKKDA TA
Sbjct: 492 GVLIFGHTTEARENAAATLFSLSAVHDYKKQIAKEDGAVEALAGLLREGSPRGKKDAVTA 551
Query: 407 LFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSI-VKAS 465
LFNL + N R I G ITAL+ L S+ + +EA ++++ ++ +
Sbjct: 552 LFNLSTHTENCARMIELGAITALVGAL--GSEGVAEEAAGALALIVRQPIGAAAVGNEEM 609
Query: 466 TIPVLIDLLRTGLPRNKENAAAILLALCK----RDTDNLSCISRLGAVIPLSELARTGTE 521
+ LI ++R G PR KENA A LL LC+ T+ + L ++ L L TGT+
Sbjct: 610 AVAGLIGMMRCGTPRGKENAVAALLELCRGGGAAATERVLKAPALASL--LQTLLFTGTK 667
Query: 522 RAKRKATSL 530
RA+RKA SL
Sbjct: 668 RARRKAASL 676
>UniRef100_Q6K5K9 Avr9/Cf-9 rapidly elicited protein-like [Oryza sativa]
Length = 467
Score = 273 bits (698), Expect = 9e-72
Identities = 158/394 (40%), Positives = 242/394 (61%), Gaps = 28/394 (7%)
Query: 157 DAIVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTP 216
+A +PE FLCPIS E+MRDPV++A+GQTY+R +IQ W+ GN TCP+TQQ L + L P
Sbjct: 79 EAAAVPEQFLCPISSEIMRDPVVLASGQTYDRRFIQEWLSAGNRTCPQTQQVLSNTILIP 138
Query: 217 NYVLRSLVSQWCIEHNI------EQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSC 270
N+++RS+++QWC E+ I Q L +KS D R S
Sbjct: 139 NHLVRSMIAQWCTENGIALSPLENQEEDLVTNNERKSFSELFD------------RISSS 186
Query: 271 RSVEESRAAVAEIRSLSKRSTDNRILIAE-AGAIPVLVSLLTSEDVMTQ----ENAVTSI 325
++ E R A+ ++R L+KR++ R +I E +I ++S +++ ++ + E+ VT+I
Sbjct: 187 SNISEKRQAIKDLRLLTKRNSSFRAVIGENPDSISQMISAVSNPELESNSEVLEDTVTTI 246
Query: 326 LNLSIYENNKGLI-MLAGAIPSIVQVLRAGTMEARENAAATLFSLSLADENKIIIGASGA 384
LNLSI+E+NK +I AI ++ L++GTMEAR NAAA +FSLS D NK IG SGA
Sbjct: 247 LNLSIHESNKKIIGDDTKAITFLISALQSGTMEARSNAAAAIFSLSALDSNKAKIGESGA 306
Query: 385 ISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEA 444
+ LVDLL++GS KKDAA+A+F+LC NK RA ++G+I +L ++D +S+ DE+
Sbjct: 307 MRPLVDLLEHGSMTAKKDAASAIFSLCKLHENKSRATKSGVIDVVLKAISD--ESLTDES 364
Query: 445 LTIMSVLASHQEAKVSIVKASTIPVLIDLLRTG-LPRNKENAAAILLALCKRDTDNL-SC 502
LTI+++L+S E I + +P ++ +++ RNKENA A+L ++C D L
Sbjct: 365 LTILALLSSDHETVEEIGETGGVPCMLHIIKDDQCKRNKENAVAVLFSICMYDRTKLREV 424
Query: 503 ISRLGAVIPLSELARTGTERAKRKATSLLEHLRK 536
+ L+ LA+ GT RA+RKA +L+ L++
Sbjct: 425 VEDENLNGSLAWLAQNGTSRARRKAAGILDKLKR 458
Score = 50.1 bits (118), Expect = 2e-04
Identities = 50/188 (26%), Positives = 87/188 (45%), Gaps = 10/188 (5%)
Query: 360 ENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGR 419
EN A L L +E+ + + S L D + + S +K A L + + R
Sbjct: 152 ENGIA-LSPLENQEEDLVTNNERKSFSELFDRISSSSNISEKRQAIKDLRLLTKRNSSFR 210
Query: 420 AIR-------AGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVK-ASTIPVLI 471
A+ + +I+A+ N +S+ ++++ +T + L+ H+ K I I LI
Sbjct: 211 AVIGENPDSISQMISAVSNPELESNSEVLEDTVTTILNLSIHESNKKIIGDDTKAITFLI 270
Query: 472 DLLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLL 531
L++G + NAAA + +L D+ N + I GA+ PL +L G+ AK+ A S +
Sbjct: 271 SALQSGTMEARSNAAAAIFSLSALDS-NKAKIGESGAMRPLVDLLEHGSMTAKKDAASAI 329
Query: 532 EHLRKLQQ 539
L KL +
Sbjct: 330 FSLCKLHE 337
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.315 0.131 0.359
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 808,743,926
Number of Sequences: 2790947
Number of extensions: 31878962
Number of successful extensions: 92787
Number of sequences better than 10.0: 1062
Number of HSP's better than 10.0 without gapping: 373
Number of HSP's successfully gapped in prelim test: 694
Number of HSP's that attempted gapping in prelim test: 89491
Number of HSP's gapped (non-prelim): 2497
length of query: 540
length of database: 848,049,833
effective HSP length: 132
effective length of query: 408
effective length of database: 479,644,829
effective search space: 195695090232
effective search space used: 195695090232
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 77 (34.3 bits)
Medicago: description of AC148469.7


