

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148405.4 - phase: 0
(609 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
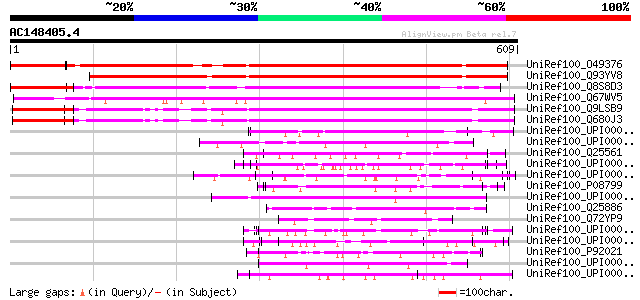
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O49376 Hypothetical protein F10N7.30 [Arabidopsis thal... 441 e-122
UniRef100_Q93YV8 Hypothetical protein At4g32160 [Arabidopsis tha... 438 e-121
UniRef100_Q8S8D3 Hypothetical protein At2g25350 [Arabidopsis tha... 371 e-101
UniRef100_Q67WV5 Phox (PX) domain-containing protein-like [Oryza... 355 2e-96
UniRef100_Q9LSB9 Emb|CAA16573.1 [Arabidopsis thaliana] 291 4e-77
UniRef100_Q680J3 Hypothetical protein At3g15920 [Arabidopsis tha... 291 4e-77
UniRef100_UPI0000338083 UPI0000338083 UniRef100 entry 71 8e-11
UniRef100_UPI00003AD06F UPI00003AD06F UniRef100 entry 67 2e-09
UniRef100_Q25561 Myosin II heavy chain [Naegleria fowleri] 67 2e-09
UniRef100_UPI0000437337 UPI0000437337 UniRef100 entry 66 3e-09
UniRef100_UPI0000432992 UPI0000432992 UniRef100 entry 65 4e-09
UniRef100_P08799 Myosin II heavy chain, non muscle [Dictyosteliu... 64 1e-08
UniRef100_UPI000042CCB9 UPI000042CCB9 UniRef100 entry 64 2e-08
UniRef100_Q25886 Liver stage antigen-1 [Plasmodium falciparum] 64 2e-08
UniRef100_Q72YP9 S-layer homology domain protein [Bacillus cereus] 63 3e-08
UniRef100_UPI000049A29E UPI000049A29E UniRef100 entry 62 4e-08
UniRef100_UPI000042F5AA UPI000042F5AA UniRef100 entry 62 4e-08
UniRef100_P92021 Hypothetical protein T10G3.5 [Caenorhabditis el... 62 4e-08
UniRef100_UPI0000437813 UPI0000437813 UniRef100 entry 62 5e-08
UniRef100_UPI000036865B UPI000036865B UniRef100 entry 62 5e-08
>UniRef100_O49376 Hypothetical protein F10N7.30 [Arabidopsis thaliana]
Length = 731
Score = 441 bits (1135), Expect = e-122
Identities = 268/537 (49%), Positives = 345/537 (63%), Gaps = 39/537 (7%)
Query: 69 SAFQDASQQNSVKDPDSNNRDYSVQSS--LHSGLSLT-AGSSSVASDYGSDTPYDASEVG 125
SA QD Q S DSNN S SS +H LSL AG S++ SDYGSDT Y+ SEVG
Sbjct: 192 SACQDVDQNAS----DSNNDRSSTSSSPMVHPSLSLFHAGGSTLTSDYGSDTAYETSEVG 247
Query: 126 TPRIGQDDNSEVGTDDLTLDEDTT--NPIEKLVKYGISNIDEGLFMGQTILEQLEGLPRH 183
+P +GQDD SE+GT+DLTLDED T NPIEKLV + +SNIDEGL M +TILEQLE P+H
Sbjct: 248 SPSVGQDDISEIGTEDLTLDEDLTLTNPIEKLVNFSMSNIDEGLSMSETILEQLEDFPKH 307
Query: 184 KVNARHGNNVTGKDRSNGNSYDSSLLGNNTMELFSQPGHAKVLGHIRKLSNESVGSDGSS 243
KV +R+ NN+ GKD NGN+ L NN L S+P + S SV D +
Sbjct: 308 KVRSRYVNNILGKDVYNGNASKGVFLANNGSRLLSEP----------EPSTHSVMHDRND 357
Query: 244 IRGSDISNFGIPNSSGDGSVDLPGSASVSRETDIVGHTKLKSSGDAQLVLPLDQRNKLNR 303
++ D +DL VS T +V + + + G AQ+VLPL+ RNKLNR
Sbjct: 358 ------------SAERDSHLDLRQGPGVSLGTGLVCNPERQ--GSAQIVLPLELRNKLNR 403
Query: 304 ILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAIL 363
IL RLV AKTDMEDLI RLNQEIA KD+L KV DLE ELET+KQ++KENL+QAI+
Sbjct: 404 ILLATNERLVNAKTDMEDLIARLNQEIAVKDYLNKKVNDLEGELETTKQRSKENLEQAIM 463
Query: 364 IERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAARE 423
ERERF QMQWDMEELR+KS EMEMKLKS G+ + T +S + + VL ++LDA ++
Sbjct: 464 SERERFNQMQWDMEELRQKSYEMEMKLKSREDGSSHAEPTVQSTISEKHVLSKELDARKQ 523
Query: 424 QLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHERE 483
QLE LS++Y ELEAKSKAD+KVLVKEVKSLR S EL+KEL+ S+ +K AEKLL ER+
Sbjct: 524 QLEDLSRRYEELEAKSKADMKVLVKEVKSLRRSHVELEKELTHSLTDKTNAEKLLQEERK 583
Query: 484 KSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQID 543
E A +K L C +L +L+E ++ L + S++ +D L SDDQI+
Sbjct: 584 LLENTVAARKKLLSDCRILHDRLKEYNLNLSMDGNGNFVDDSTTISDVLRLLSISDDQIE 643
Query: 544 ILLAEVENLEK-DAGSAASNVDKTNDI-KDGVICGDEMRKIISDLFLDNVRLRKQTN 598
E + L D +AA ++DKT + + I DE+RKI++++F++N +LRKQ N
Sbjct: 644 ----EAQLLSGFDENAAAEDIDKTLSMDTETRIMEDELRKILANIFVENAKLRKQVN 696
Score = 139 bits (351), Expect = 2e-31
Identities = 57/67 (85%), Positives = 62/67 (92%)
Query: 1 MQRRSPPKHRHDGTSPLPLGMDWSPAPRKWNGRDTEWPHNHRTGWSYCVIIPSWVFVPKS 60
MQRRSPPKHRHDGTSPLPLGMDWSP PRKWNGRDT WPH+ RTGWSYCV IPSW+ +PKS
Sbjct: 2 MQRRSPPKHRHDGTSPLPLGMDWSPPPRKWNGRDTVWPHDPRTGWSYCVTIPSWIVLPKS 61
Query: 61 KNSDPIV 67
+NSDP+V
Sbjct: 62 RNSDPVV 68
>UniRef100_Q93YV8 Hypothetical protein At4g32160 [Arabidopsis thaliana]
Length = 534
Score = 438 bits (1127), Expect = e-121
Identities = 259/509 (50%), Positives = 337/509 (65%), Gaps = 17/509 (3%)
Query: 96 LHSGLSLT-AGSSSVASDYGSDTPYDASEVGTPRIGQDDNSEVGTDDLTLDEDTT--NPI 152
+H LSL AG S++ SDYGSDT Y+ SEVG+P +GQDD SE+GT+DLTLDED T NPI
Sbjct: 2 VHPSLSLFHAGGSTLTSDYGSDTAYETSEVGSPSVGQDDISEIGTEDLTLDEDLTLTNPI 61
Query: 153 EKLVKYGISNIDEGLFMGQTILEQLEGLPRHKVNARHGNNVTGKDRSNGNSYDSSLLGNN 212
EKLV + +SNIDEGL M +TILEQLE P+HKV +R+ NN+ GKD NGN+ L NN
Sbjct: 62 EKLVNFSMSNIDEGLSMSETILEQLEDFPKHKVRSRYVNNILGKDVYNGNASKGVFLANN 121
Query: 213 TMELFSQPGHAK-VLGHIRKLSNESVGSDGSSIRGSDISNFGIPNSSGDGSVDLPGSASV 271
L S+P + + H R S E ++ S G+ SS D +DL V
Sbjct: 122 GSRLLSEPEPSTHSVMHDRNDSAERF-----ALHTGQTSTSGLLISSRDSHLDLRQGPGV 176
Query: 272 SRETDIVGHTKLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIA 331
S T +V + + + G AQ+VLPL+ RNKLNRIL RLV AKTDMEDLI RLNQEIA
Sbjct: 177 SLGTGLVCNPERQ--GSAQIVLPLELRNKLNRILLATNERLVNAKTDMEDLIARLNQEIA 234
Query: 332 AKDFLTTKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLK 391
KD+L KV DLE ELET+KQ++KENL+QAI+ ERERF QMQWDMEELR+KS EMEMKLK
Sbjct: 235 VKDYLNKKVNDLEGELETTKQRSKENLEQAIMSERERFNQMQWDMEELRQKSYEMEMKLK 294
Query: 392 SESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVK 451
S G+ + T +S + + VL ++LDA ++QLE LS++Y ELEAKSKAD+KVLVKEVK
Sbjct: 295 SREDGSSHAEPTVQSTISEKHVLSKELDARKQQLEDLSRRYEELEAKSKADMKVLVKEVK 354
Query: 452 SLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSV 511
SLR S EL+KEL+ S+ +K AEKLL ER+ E A +K L C +L +L+E ++
Sbjct: 355 SLRRSHVELEKELTHSLTDKTNAEKLLQEERKLLENTVAARKKLLSDCRILHDRLKEYNL 414
Query: 512 ELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEK-DAGSAASNVDKTNDI- 569
L + S++ +D L SDDQI+ E + L D +AA ++DKT +
Sbjct: 415 NLSMDGNGNFVDDSTTISDVLRLLSISDDQIE----EAQLLSGFDEKAAAEDIDKTLSMD 470
Query: 570 KDGVICGDEMRKIISDLFLDNVRLRKQTN 598
+ I DE+RKI++++F++N +LRKQ N
Sbjct: 471 TETRIMEDELRKILANIFVENAKLRKQVN 499
>UniRef100_Q8S8D3 Hypothetical protein At2g25350 [Arabidopsis thaliana]
Length = 643
Score = 371 bits (952), Expect = e-101
Identities = 228/521 (43%), Positives = 316/521 (59%), Gaps = 42/521 (8%)
Query: 69 SAFQDASQQNSVKDPDSNNRDYSVQSSLHSGLSLTAGSSSVASDYGSDTPYDASEVGTPR 128
SA Q Q S + D N+ S+ S +H + GSS ++SDYGSDT Y+ SE+G+
Sbjct: 165 SACQVVDQNASDSNDDGNST--SLSSLVHPN---SGGSSLLSSDYGSDTAYETSELGSAS 219
Query: 129 IGQDDNSEVGTDDLTLDEDTTNPIEKLVKYGISNIDEGLFMGQTILEQLEGLPRHKVNAR 188
+GQDD SE T DLTLDED TNP EKLVK+ +SNIDEGL M QTI+EQLE P+H+V+
Sbjct: 220 LGQDDVSETDTGDLTLDEDLTNPTEKLVKFSMSNIDEGLSMSQTIIEQLEDFPKHRVHLG 279
Query: 189 HGNNVTGKDRSNGNSYDSSLLGNNTMELFSQPGHAKVLGHIRKLSNESVGSDGSSIRGSD 248
+ N++T + NG + NN + S+ + + H RKLS ES +DG S+ +
Sbjct: 280 YANDITETNSYNGKASKGVFRANNDLRCRSESETSHSVMHDRKLSLES--ADGVSLLAGE 337
Query: 249 ISNFGIPNSSGDGSVDLPGSASVSRETDIVGHTKLKSSGDAQLVLPLDQRNKLNRILSTM 308
S I +S +D+ SV L+ G+ ++VLPL +KLNRIL TM
Sbjct: 338 TSTSSILSSIVHSQLDVNHDISVGN---------LEIPGNGRIVLPLKMHSKLNRILLTM 388
Query: 309 QRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERER 368
RL+ +KTDMEDLI RLNQE A K++L KV DLEVELET+KQ+NKENL+QA++ ER+
Sbjct: 389 NERLLNSKTDMEDLIARLNQETAVKEYLNRKVDDLEVELETTKQRNKENLEQALMTERQS 448
Query: 369 FTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEIL 428
T+MQWDMEELR+K+ EME+KLKS+ G+ + S + ++ LLQ++DA ++QLE L
Sbjct: 449 VTKMQWDMEELRQKTFEMELKLKSKEDGSSDSKTSGNSTISESHELLQEMDATKQQLEDL 508
Query: 429 SKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQA 488
S++Y ELEAKSKADIKVLV+EVKSLR S E++KEL+ S+ EK + EKLL ER E
Sbjct: 509 SRRYVELEAKSKADIKVLVREVKSLRRSHMEMEKELTRSLTEKSDTEKLLQQERIIVENT 568
Query: 489 EIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAE 548
A R+ C +L +L+ + L ++ +S+++ D L+E
Sbjct: 569 LEARRRLYSDCEILHDRLKVNNTNLSMDE-------------------SSNNRED--LSE 607
Query: 549 VENLEKDAGSAASNVDKTNDIKDGVICGDEMRKIISDLFLD 589
V N +D A + D DE+RKI++D++ D
Sbjct: 608 VSNALQDQIEAQLLLG-----FDETASEDELRKIMADMYED 643
Score = 134 bits (337), Expect = 8e-30
Identities = 58/76 (76%), Positives = 61/76 (79%)
Query: 1 MQRRSPPKHRHDGTSPLPLGMDWSPAPRKWNGRDTEWPHNHRTGWSYCVIIPSWVFVPKS 60
MQRRSPPKHRHDG SPLPLGMDWSP PR WNGRDT WPH+ RTGWSYCV IPSW + KS
Sbjct: 1 MQRRSPPKHRHDGASPLPLGMDWSPPPRNWNGRDTIWPHDFRTGWSYCVTIPSWTLLSKS 60
Query: 61 KNSDPIVMSAFQDASQ 76
KNSDPIV Q + Q
Sbjct: 61 KNSDPIVFYRVQVSVQ 76
>UniRef100_Q67WV5 Phox (PX) domain-containing protein-like [Oryza sativa]
Length = 710
Score = 355 bits (911), Expect = 2e-96
Identities = 242/682 (35%), Positives = 353/682 (51%), Gaps = 93/682 (13%)
Query: 5 SPPKHRHDGTSPLPLGMDWSPAPRKWNGRDTEWPHNHRTGWSYCVIIPSWVFVPKSKNSD 64
SPPKHRHDGTS LP GMDWSP P++W GR+T WPHN +TGWSYCV+IPSW
Sbjct: 16 SPPKHRHDGTSGLPFGMDWSPPPKRWEGRNTIWPHNPQTGWSYCVMIPSW---------- 65
Query: 65 PIVMSAFQDASQQNSVKDPDSNNRDYSVQSSLHSGLSLTAGSSSVASDY---GSDTPYDA 121
I + A+ N +K +QS G+S + G SD+ SD +
Sbjct: 66 -IAQTPEASATADNFLKSIVFYRIHVGIQSP--EGISSSHGVLRRFSDFLKLSSDLKQEF 122
Query: 122 SEVGTPRIGQDDNSEVGTDDLTLDEDTTNPIEKLVKYGISNIDEGLFMGQTILEQLEGLP 181
G P L E+ N +E+ ++ +S+I+ +LE
Sbjct: 123 PRKGIPPAPPKHAFSRINSSRVLLEERRNALEEWMQKLLSDIELSRSAPVAAFLELEAAA 182
Query: 182 RH----------------KVNA-------RHGNNVTGKDRSNGNSY-----DSSLLGNNT 213
R K +A HG+ V + +++ + GN
Sbjct: 183 RSYYQDWNQRPSEVGSSAKSSADSSPRPDEHGSGVLSESSQMNSAFAHGNGPTGATGNGM 242
Query: 214 M--ELFSQPG-HAKVLGHIRK---------------------LSNESVGSDGSSIRGSDI 249
+ + QP A + + RK +S E S+ R
Sbjct: 243 LGESILDQPNERASSMSNHRKKNHVFLEHGVRNGSIDTYKGVVSEEDHDSNPGHARKDSA 302
Query: 250 SNFGIPNSSGDGS-VDLPGSAS------VSRETDIVGHT----------KLKSSGDAQLV 292
+ G SS GS + +PG +S V + I GH+ +L DAQ++
Sbjct: 303 ESIGSDLSSLRGSELSVPGVSSSLWDGPVDLPSGIDGHSQTEQFTGLDMQLLYDMDAQII 362
Query: 293 LPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQ 352
LP DQR KL R+L +M RR VTAKTDMEDLI RLNQE+A K++L TKVKDLEVELE +K+
Sbjct: 363 LPADQRPKLTRLLISMDRRQVTAKTDMEDLIARLNQEVAVKEYLATKVKDLEVELEATKK 422
Query: 353 KNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQND 412
K+KE L QA+L ERE+ TQ+QWD +EL RK EME LK E K + + +
Sbjct: 423 KDKEILHQAVLTEREKITQLQWDKDELYRKYSEMESNLKIEQNEKTRVQSEKTTASGEKE 482
Query: 413 VLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKY 472
+LL++L+ R+++E L + GE EAKSKADIKVLVKEVKSLR+SQ E+KK L++ +EK
Sbjct: 483 MLLEELETKRKEVESLQQHIGEFEAKSKADIKVLVKEVKSLRNSQKEMKKVLNQYHEEKT 542
Query: 473 EAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAF 532
E E+++ E+++S +A + K L +C LL ++LQEC+ + +++D + SS DA
Sbjct: 543 ELERIVNREKQRSTRARFSREKILHECRLLRERLQECTAKFVADEQDTMTIDLSSLPDAL 602
Query: 533 NKLKTSDDQIDILLAEVENLEKDAGSAASNVDKTNDIK--------DGVICGDEMRKIIS 584
+ + TSD++I +L+AE + L +D +S+ +D K D + +E K++S
Sbjct: 603 DLVTTSDNRIRLLVAEAQLLSRDDEQGSSDDGDNSDGKSSVTMSSEDAYVTDEETTKMLS 662
Query: 585 DLFLDNVRLRKQTNRVTRHMLS 606
DL +DN +LR + N + R+ ++
Sbjct: 663 DLLIDNAQLRLRLNSLIRNAVN 684
>UniRef100_Q9LSB9 Emb|CAA16573.1 [Arabidopsis thaliana]
Length = 755
Score = 291 bits (745), Expect = 4e-77
Identities = 190/546 (34%), Positives = 314/546 (56%), Gaps = 52/546 (9%)
Query: 67 VMSAFQDASQQNSVKDPDSNNRDYSVQSSLHSGLSLTAGSSSVASDYGSDTPYDASEVGT 126
V S F D Q+ D + S L + +S GSSSV D+ +D+ + S T
Sbjct: 223 VRSYFNDEYQETEDTSGD-------IPSLLPTTISDVPGSSSVTIDHDNDSADETSNAST 275
Query: 127 PRIGQDDNSEVGTDDLTLDEDTTNPIEKLVKYGISNIDEGLFMGQTILEQLEGLPRHKVN 186
+ + + + + + T ++ T+ E + +YG+ +D+ F E++E +++
Sbjct: 276 MKHDEANLKNLFSRNSTAVDNVTDWHELITEYGL--LDQSSFQ-----EKVE-----RLS 323
Query: 187 ARHGNNVTGKDRSNGNSYDSSLLGNNTMELFSQPGHAKVLGHIRKLSNESVGSDGSSIRG 246
+ +G+ TG G S S G ++ G RK ++ SI+
Sbjct: 324 STNGDAATGTVTRGGIS--------------SGVGIQRLDGSDRKFQELTI----ESIKK 365
Query: 247 SDISNFGI-----PNSSGDGSVDLPGSASVSRETDIVGHTKLKSSGDAQLVLPLDQRNKL 301
+ +S+F P+ G++D+ G A + + G T+ + D +V ++R+KL
Sbjct: 366 THVSDFEASTAVEPDLVNQGAMDIHGEAHGNMYGAVGGDTETQK--DLAIVFQSEERHKL 423
Query: 302 NRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQA 361
R++ T+++RL TAK D EDLI RLNQE+A + FL+TKV+DLEVELET+++ K+ +++
Sbjct: 424 KRVIDTLKQRLETAKADTEDLISRLNQELAVRQFLSTKVRDLEVELETTRESCKQGMEKT 483
Query: 362 ILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAA 421
+L E+ERFTQ+QWDMEELR++ +EME L S + ES+VQ+N +LLQ ++
Sbjct: 484 VLDEKERFTQIQWDMEELRKQCMEMESFLNSIKDEKTHIETANESLVQENQMLLQQINDI 543
Query: 422 REQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHE 481
RE E K++ ELE K+KA++KVLVKEVKSLR++Q++L++ELS +KEK E E+++ E
Sbjct: 544 RENFENFHKEHEELEVKAKAELKVLVKEVKSLRTTQSDLRQELSGIMKEKLEMERIVQRE 603
Query: 482 REKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQ 541
+++ E A+ A +K L +C +L +LQEC+V+ E+E + + SSS ++A L TSD++
Sbjct: 604 KDREETAKNADKKLLHECDVLQNRLQECNVKFDIEEEGKLIMDSSSLSEAIELLATSDNR 663
Query: 542 IDILLAEVENLEKDAGSAASNVDKTNDIKDGVICGDEM-RKIISDLFLDNVRLRKQTNRV 600
I +L+AE + L ++ V+K G D++ RK+++++ +DN RLRKQ N V
Sbjct: 664 IGLLIAETQLLSEE-------VEKLKLTSGGHRGTDDLVRKMLTEVLIDNARLRKQVNSV 716
Query: 601 TRHMLS 606
R LS
Sbjct: 717 LRCSLS 722
Score = 111 bits (278), Expect = 5e-23
Identities = 49/73 (67%), Positives = 53/73 (72%)
Query: 4 RSPPKHRHDGTSPLPLGMDWSPAPRKWNGRDTEWPHNHRTGWSYCVIIPSWVFVPKSKNS 63
R+ P HRHDG SPLPLGMDWS PR GRDT WPH+HRTGWSYCV +PSWV +PKS S
Sbjct: 64 RASPPHRHDGRSPLPLGMDWSSPPRHLEGRDTVWPHDHRTGWSYCVTVPSWVDLPKSSVS 123
Query: 64 DPIVMSAFQDASQ 76
DP V Q A Q
Sbjct: 124 DPAVFYRVQVAIQ 136
>UniRef100_Q680J3 Hypothetical protein At3g15920 [Arabidopsis thaliana]
Length = 755
Score = 291 bits (745), Expect = 4e-77
Identities = 190/546 (34%), Positives = 314/546 (56%), Gaps = 52/546 (9%)
Query: 67 VMSAFQDASQQNSVKDPDSNNRDYSVQSSLHSGLSLTAGSSSVASDYGSDTPYDASEVGT 126
V S F D Q+ D + S L + +S GSSSV D+ +D+ + S T
Sbjct: 223 VRSYFNDEYQETEDTSGD-------IPSLLPTTISDVPGSSSVTIDHDNDSADETSNAST 275
Query: 127 PRIGQDDNSEVGTDDLTLDEDTTNPIEKLVKYGISNIDEGLFMGQTILEQLEGLPRHKVN 186
+ + + + + + T ++ T+ E + +YG+ +D+ F E++E +++
Sbjct: 276 MKHDEANLKNLFSRNSTAVDNVTDWHELITEYGL--LDQSSFQ-----EKVE-----RLS 323
Query: 187 ARHGNNVTGKDRSNGNSYDSSLLGNNTMELFSQPGHAKVLGHIRKLSNESVGSDGSSIRG 246
+ +G+ TG G S S G ++ G RK ++ SI+
Sbjct: 324 STNGDAATGTVTRGGIS--------------SGVGIQRLDGSDRKFQELTI----ESIKK 365
Query: 247 SDISNFGI-----PNSSGDGSVDLPGSASVSRETDIVGHTKLKSSGDAQLVLPLDQRNKL 301
+ +S+F P+ G++D+ G A + + G T+ + D +V ++R+KL
Sbjct: 366 THVSDFEASTAVEPDLVNQGAMDIHGEAHGNMYGAVGGDTETQK--DLAIVFQSEERHKL 423
Query: 302 NRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQA 361
R++ T+++RL TAK D EDLI RLNQE+A + FL+TKV+DLEVELET+++ K+ +++
Sbjct: 424 KRVIDTLKQRLETAKADTEDLISRLNQELAVRQFLSTKVRDLEVELETTRESCKQGMEKT 483
Query: 362 ILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAA 421
+L E+ERFTQ+QWDMEELR++ +EME L S + ES+VQ+N +LLQ ++
Sbjct: 484 VLDEKERFTQIQWDMEELRKQCMEMESFLNSIKDEKTHIETANESLVQENQMLLQQINDI 543
Query: 422 REQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHE 481
RE E K++ ELE K+KA++KVLVKEVKSLR++Q++L++ELS +KEK E E+++ E
Sbjct: 544 RENFENFHKEHEELEVKAKAELKVLVKEVKSLRTTQSDLRQELSGIMKEKLEMERIVQRE 603
Query: 482 REKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQ 541
+++ E A+ A +K L +C +L +LQEC+V+ E+E + + SSS ++A L TSD++
Sbjct: 604 KDREETAKNADKKLLHECDVLQNRLQECNVKFDIEEEGKLIMDSSSLSEAIELLATSDNR 663
Query: 542 IDILLAEVENLEKDAGSAASNVDKTNDIKDGVICGDEM-RKIISDLFLDNVRLRKQTNRV 600
I +L+AE + L ++ V+K G D++ RK+++++ +DN RLRKQ N V
Sbjct: 664 IGLLIAETQLLSEE-------VEKLKLTSGGHRGTDDLVRKMLTEVLIDNARLRKQVNSV 716
Query: 601 TRHMLS 606
R LS
Sbjct: 717 LRCSLS 722
Score = 111 bits (278), Expect = 5e-23
Identities = 49/73 (67%), Positives = 53/73 (72%)
Query: 4 RSPPKHRHDGTSPLPLGMDWSPAPRKWNGRDTEWPHNHRTGWSYCVIIPSWVFVPKSKNS 63
R+ P HRHDG SPLPLGMDWS PR GRDT WPH+HRTGWSYCV +PSWV +PKS S
Sbjct: 64 RASPPHRHDGRSPLPLGMDWSSPPRHLEGRDTVWPHDHRTGWSYCVTVPSWVDLPKSSVS 123
Query: 64 DPIVMSAFQDASQ 76
DP V Q A Q
Sbjct: 124 DPAVFYRVQVAIQ 136
>UniRef100_UPI0000338083 UPI0000338083 UniRef100 entry
Length = 623
Score = 71.2 bits (173), Expect = 8e-11
Identities = 78/345 (22%), Positives = 161/345 (46%), Gaps = 53/345 (15%)
Query: 288 DAQLVLPLDQRNKLNRILSTMQRRLVTAKTD-MEDLIVRLNQEI----AAKDFLTTKVKD 342
+ + +L L +R++ +L T + R K + +E+ V +E+ + + +T +V +
Sbjct: 34 ERERLLKLQERHEKRTVLLTNELRQTKQKLESVEETFVHAKEELQRANSKNETMTLRVDE 93
Query: 343 LEVELETSKQKNKENLQQAILIERERF-----------TQMQWDMEELRRKSLEMEMKLK 391
L+V L K+ KEN + +RERF T Q D+ + +K E+++K+
Sbjct: 94 LQVLL---KELKKENKDLDVKYQRERFKRKHAREDLDHTNCQNDI--VTKKVEELQVKIL 148
Query: 392 SESGGN--VGQDLTKESIVQQNDVLLQDLDAAREQ--LEILSKQYGELEAKSKADIKVLV 447
N + + KE + + + AA Q LE SK Y LE+K + D
Sbjct: 149 DVLTNNKCLEEKYEKEKSTHMHIIAEMEQQAAEHQARLEQASKDYQSLESKYQNDTSDYT 208
Query: 448 KEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQL- 506
K+++ L + +L+ +L ++++ ++ ER + E+ + +++ +++ L LK++
Sbjct: 209 KKIQELDRDKEKLQAQLEQALEGNNHLKEKYQDERAQQERTQGEFQQNVKELQLQLKKVE 268
Query: 507 -QECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDK 565
++ SVE Y +E T+ K + +++++ + +++N AG + V+K
Sbjct: 269 KEKQSVEEKYHNEKDTY-------------KRTKEELELEIYKLQNQLMQAGEESKRVEK 315
Query: 566 TNDIKDGVICGDEM---RKIISDL--FLDNVRLRKQTNRVTRHML 605
DEM RKI+ +L D ++L ++ N+ R L
Sbjct: 316 EYQ--------DEMEINRKIMKELKQLQDKLQLAEEDNKFLREKL 352
Score = 52.8 bits (125), Expect = 3e-05
Identities = 63/293 (21%), Positives = 130/293 (43%), Gaps = 43/293 (14%)
Query: 290 QLVLPLDQRNKLNRILSTM-QRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVE-- 346
+L + L + K N+ L QR K EDL + D +T KV++L+V+
Sbjct: 93 ELQVLLKELKKENKDLDVKYQRERFKRKHAREDL----DHTNCQNDIVTKKVEELQVKIL 148
Query: 347 --------LETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNV 398
LE +K K I ++ + Q +E+ + +E K ++++
Sbjct: 149 DVLTNNKCLEEKYEKEKSTHMHIIAEMEQQAAEHQARLEQASKDYQSLESKYQNDTS--- 205
Query: 399 GQDLTK--ESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSS 456
D TK + + + + L L+ A E L ++Y + A+ + + VK L+
Sbjct: 206 --DYTKKIQELDRDKEKLQAQLEQALEGNNHLKEKYQDERAQQERTQGEFQQNVKELQLQ 263
Query: 457 QTELKKELSESIKEKYEAEK-------------------LLLHEREKSEQAEIAWRKQLE 497
+++KE +S++EKY EK L+ E+S++ E ++ ++E
Sbjct: 264 LKKVEKE-KQSVEEKYHNEKDTYKRTKEELELEIYKLQNQLMQAGEESKRVEKEYQDEME 322
Query: 498 KCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVE 550
++K+L++ +L +ED FL+ + +K K++ ++++ +AE++
Sbjct: 323 INRKIMKELKQLQDKLQLAEEDNKFLREKLGIEK-SKHKSTREELNQDIAELQ 374
Score = 37.7 bits (86), Expect = 0.99
Identities = 41/219 (18%), Positives = 95/219 (42%), Gaps = 39/219 (17%)
Query: 295 LDQRNKLNRILSTMQRRLVTAKTD----MEDLIVRLNQEIAAKDFLTTKVKDLEVEL--- 347
++ K+ + L +Q +L A+ D E L + ++ + ++ L + +L+++L
Sbjct: 321 MEINRKIMKELKQLQDKLQLAEEDNKFLREKLGIEKSKHKSTREELNQDIAELQLQLKLA 380
Query: 348 ETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESI 407
E K + ++ Q + RE ++Q + + +++ +E K + E + ++
Sbjct: 381 EDDKTRLEDKHQNEVTQVRE---ELQAPLRQAEKENRCVEQKYQQELSKHRATSEELDTA 437
Query: 408 VQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSES 467
++N VLL L+ Q+EI +Q ++ L++++ +LK++L
Sbjct: 438 RRENHVLLGKLEELEHQMEIRDQQ------------------IQQLQTNEKKLKEDLHTH 479
Query: 468 IKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQL 506
H +S++ R QL CG+ ++ L
Sbjct: 480 -----------QHAEAESKRCVEKLRDQLHGCGVFMRYL 507
>UniRef100_UPI00003AD06F UPI00003AD06F UniRef100 entry
Length = 1248
Score = 66.6 bits (161), Expect = 2e-09
Identities = 84/358 (23%), Positives = 156/358 (43%), Gaps = 46/358 (12%)
Query: 229 IRKLSNESVGSDGSSIRGSD---ISNFGIP--NSSGDGSVDLPGSASVSRETDI------ 277
IR +NE +G + SD I N G P +S DLP S S E ++
Sbjct: 506 IRSAANEQTSKEGINRAMSDQSIIHNLGHPVVDSCRQHHYDLPVSPVKSCERELHLLRRA 565
Query: 278 -VGHTKLKSSGDA--QLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKD 334
V L S+ D QL + ++ K M+ ++ KT E + +L QE
Sbjct: 566 QVAEEDLSSARDVIQQLTSTVSEKEK------EMETKIAELKTRYEKELSQLGQE---NY 616
Query: 335 FLTTKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEE----LRRKSLEMEMKL 390
L +K++ +E E SK+K + + + E+ +Q+Q +ME+ ++ E E
Sbjct: 617 VLQSKLRSME---ELSKEKRWIHHTATVPVCEEKLSQIQKEMEDQEVIIQGYQQENERLY 673
Query: 391 KSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLE---ILSKQYGELEAKSKADIKVLV 447
K + +E + ++N L+ +L A RE++E I S+ E E L+
Sbjct: 674 KQMKELQIQNKKNEEQMYKENQCLMSELIALREKVERTNIQSQIVQESEPTRNQSFTELI 733
Query: 448 KEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQ 507
E+++ R +T+L++E+ ++K E L +++ + A++ + K ++
Sbjct: 734 SELRAARKEETKLQEEIRRLKQDKQALELDLGQAKKERDLAKVQITSTSSEKSYEFKVME 793
Query: 508 ECSVE--------LPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAG 557
E + L + E++ L ++ +LK + ++I+ L EVE L +AG
Sbjct: 794 ESYKQEILHLKRRLQWYAENQDLLDKDAA-----RLKDAREEIEKLKQEVERLRAEAG 846
Score = 43.9 bits (102), Expect = 0.014
Identities = 40/173 (23%), Positives = 84/173 (48%), Gaps = 15/173 (8%)
Query: 334 DFLTTKVKDLEVELETSKQKNKENL---QQAILIERERFTQMQWDMEELRRKSLEMEM-K 389
DFL ++K LE ELE + K +L +Q + ++ Q ++E+L + E K
Sbjct: 913 DFLERRIKKLETELEGKDDEAKTSLRAMEQQFQKIKMQYEQRLAELEQLLAYKWKSESPK 972
Query: 390 LKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKE 449
L S+ ++ +L +++ + + +++L + ++E L Q +L+ +SK K+
Sbjct: 973 LNSDKANSIELELQLQNLKTTHQITVENL---KTEIENLKSQNSQLKLRSK-------KD 1022
Query: 450 VKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLL 502
K L+S+ ++K+ ++ K E L+ RE + + + Q E+ +L
Sbjct: 1023 NKDLQSTDWQMKQGKTKEKLLKLNQE-LITKNREIQDLTKTVEKLQKERMAML 1074
Score = 41.2 bits (95), Expect = 0.090
Identities = 55/232 (23%), Positives = 107/232 (45%), Gaps = 23/232 (9%)
Query: 286 SGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEV 345
S ++QL L + K N+ L + ++ KT ++ +++LNQE+ K+ +++DL
Sbjct: 1010 SQNSQLKL---RSKKDNKDLQSTDWQMKQGKT--KEKLLKLNQELITKN---REIQDLTK 1061
Query: 346 ELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVG----QD 401
+E +++ L L R + + E L++ ++ E + S S +G
Sbjct: 1062 TVEKLQKERMAMLSDNNL--RNKTDNKENCQEILKKNTMATEKRNSSHSEPFIGIFNNDK 1119
Query: 402 LTKESIVQQNDVL--LQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTE 459
+ + ++VL LQ+ +E+LE LS + + KS+A + ++ ++ E
Sbjct: 1120 MYQPHNFSDSNVLEVLQENARLKEELEKLSLEMNQQRVKSQATLAYSENNIRRIQEDTAE 1179
Query: 460 LKKELSESIKEKYEAEKLL---LHEREKSEQAEIAWRKQLEKCGLLLKQLQE 508
L S + E EK+L E S+ AE+ R ++ +L+K LQE
Sbjct: 1180 YVAALKAS--HQREVEKILSQYTKEHSASKVAELNGRISTQE--ILIKHLQE 1227
>UniRef100_Q25561 Myosin II heavy chain [Naegleria fowleri]
Length = 746
Score = 66.6 bits (161), Expect = 2e-09
Identities = 78/337 (23%), Positives = 149/337 (44%), Gaps = 50/337 (14%)
Query: 282 KLKSSGDAQLVLPL----DQRNKLNRILSTMQRRLVTAKTDMEDL----------IVRLN 327
KL S +L L L D++N+L L ++ L K +DL I +L
Sbjct: 20 KLSESSKDELTLQLNKTNDEKNELVNKLKKAEKDLKNLKKSKDDLQAEKDDSDNRIRKLE 79
Query: 328 QEIAAKDFLTT----KVKDLEVELETSKQKNKENLQQAILIE------RERFTQMQWDME 377
Q++ K+ L+ ++ DLE E T + + K + ++ ++R Q+Q D+E
Sbjct: 80 QDLREKEQLSENLAKRIADLENEARTKEAQKKSTEMELSSVKDDLNRTKQRAEQLQSDLE 139
Query: 378 ELRRKSLEMEMKLKSESGGNVGQD--------------LTKESIVQQNDVLLQDLDAARE 423
R ++ E+E L GG D + + +N+ L ++L+ +
Sbjct: 140 AQRERANELENLLSDTEGGKNQLDSQFKQLQNELQNERTNLQKMKSENERLQRELEEMKR 199
Query: 424 QLEILSKQYGELEAKSKA---DIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLH 480
L + L++K K+ I+ L +++ RSS+T+L K+ S+ KE K L
Sbjct: 200 SLSDKQNESTSLDSKVKSLEDKIRELTALLETERSSKTDLDKKRSKMDKEV----KRLAQ 255
Query: 481 EREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDD 540
+ +++EQA ++ +KQL E ++ + DR ++++ N LK D
Sbjct: 256 QLQETEQALKGETQKKNDADNRVKQL-ESELQGVKSERDRLNKDLNNTSGDMNGLKRQLD 314
Query: 541 QIDILLA----EVENLEKDAGSAASNVDKTNDIKDGV 573
+ + L+A E++ L+KD + ++T + D +
Sbjct: 315 ESNNLVAKLKAEIQKLQKDLSDHHGDREETEEQLDAL 351
Score = 51.2 bits (121), Expect = 9e-05
Identities = 73/335 (21%), Positives = 142/335 (41%), Gaps = 60/335 (17%)
Query: 306 STMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIE 365
S + R +++ ++L ++LN+ K+ L K+K E +L+ K K+K++LQ
Sbjct: 13 SEIDRLKKLSESSKDELTLQLNKTNDEKNELVNKLKKAEKDLKNLK-KSKDDLQAEKDDS 71
Query: 366 RERFTQMQWDMEELRRKS---------LEMEMKLKSESGGNVGQDL---------TKESI 407
R +++ D+ E + S LE E + K + +L TK+
Sbjct: 72 DNRIRKLEQDLREKEQLSENLAKRIADLENEARTKEAQKKSTEMELSSVKDDLNRTKQRA 131
Query: 408 VQ-QNDV------------LLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLR 454
Q Q+D+ LL D + + QL+ KQ + +++ + E + L+
Sbjct: 132 EQLQSDLEAQRERANELENLLSDTEGGKNQLDSQFKQLQNELQNERTNLQKMKSENERLQ 191
Query: 455 SSQTELKKELSE-------------SIKEKY-EAEKLLLHEREKSEQAEIAWRKQLEKCG 500
E+K+ LS+ S+++K E LL ER + K ++
Sbjct: 192 RELEEMKRSLSDKQNESTSLDSKVKSLEDKIRELTALLETERSSKTDLDKKRSKMDKEVK 251
Query: 501 LLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAA 560
L +QLQE L E + + DA N++K + ++ + +E + L KD + +
Sbjct: 252 RLAQQLQETEQALKGETQKK--------NDADNRVKQLESELQGVKSERDRLNKDLNNTS 303
Query: 561 SNVDKTNDIKDGVICGDEMRKIISDLFLDNVRLRK 595
++ N +K + DE +++ L + +L+K
Sbjct: 304 GDM---NGLKRQL---DESNNLVAKLKAEIQKLQK 332
Score = 35.4 bits (80), Expect = 4.9
Identities = 54/253 (21%), Positives = 109/253 (42%), Gaps = 29/253 (11%)
Query: 261 GSVDLPGSASVSRETDIVGHTKLKSSGDAQLVLPLDQRNKLNRI---LSTMQRRLVTA-- 315
G VD + RE ++ +LK+ + +D NK + L+ M+ RL T
Sbjct: 455 GGVDSAEVEKLRREYEMQ-LAQLKARVEEVTQQRVDVENKKRSVEMDLTEMKTRLQTEER 513
Query: 316 -KTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNK---ENLQQAILIERE---- 367
+ +E + E L + +DL EL +K +++ + L+Q +L ER
Sbjct: 514 LRKKVEQQKKSVEMECDELRELAEEAEDLRDELNRTKLEHQALIQQLRQDLLQERHSRAS 573
Query: 368 ----------RFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKE--SIVQQNDVLL 415
++Q D+E+ R K E +LK + + DL + ++
Sbjct: 574 AEESATRQKREIEELQQDLEQERAKLDEAARRLKQQYENEI-LDLNNQIAQAKKERSAAS 632
Query: 416 QDLDAAREQLEILSKQYGELEAKSKADIKV-LVKEVKSLRSSQTELKKELSESIKEKYEA 474
+D+ A L +++ E EA++K D++ L K + + Q++ + + S+ K + E
Sbjct: 633 RDMKKADRDLREYQRRFQE-EARAKQDLEQRLTKVERENKLLQSQSQSDASKYQKAEQEK 691
Query: 475 EKLLLHEREKSEQ 487
++L R++ ++
Sbjct: 692 QRLEAENRQQKDK 704
>UniRef100_UPI0000437337 UPI0000437337 UniRef100 entry
Length = 1701
Score = 66.2 bits (160), Expect = 3e-09
Identities = 67/294 (22%), Positives = 147/294 (49%), Gaps = 20/294 (6%)
Query: 290 QLVLPLD-QRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELE 348
Q L +D QRN+++ +RL + D+E+ +L + + + K++ ++ EL
Sbjct: 1043 QFKLKIDHQRNEMSAAFEKEHKRLENMENDLENQRAKLAESLNDAAKMEQKLQQMKNELN 1102
Query: 349 TSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEME-----MKLKSESGGNVGQDLT 403
K++ E ++Q + ER + Q + EE+R+++ E+E ++ K+E + QD
Sbjct: 1103 KEKEQ-VEEMKQMVKTERVKIEQ---EKEEMRKQTQEIEDGHEKLQKKTEIADKLLQDAN 1158
Query: 404 KESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKV-LVKEVKSLRSSQTELKK 462
+ + L+++L +EQLEI SK E E K ++K L ++++SLR + ++
Sbjct: 1159 MTTEQNEYRKLIEELQREKEQLEI-SKNQIEQEKKDLQNMKSNLERQLESLRHEKANVED 1217
Query: 463 ELSESIKEKYEAEKLLLHEREKSEQAEIAWRK-QLEKCGLLLKQLQECSVELPYEDEDRT 521
E + K + E++++ R++ E E W + + EK K+++E + L E+
Sbjct: 1218 EKNALEKMRSESKEIQNAIRKEREDLENCWVEIEGEK-----KRMEEETRRLEMHREEIK 1272
Query: 522 FLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKTND-IKDGVI 574
+ S N +++ +++I+ + E+ ++ + G ++K + ++D I
Sbjct: 1273 KVDSELQKSREN-IESREEKINNKMEELLQMKVEMGRTKEEIEKESQRLQDATI 1325
Score = 62.4 bits (150), Expect = 4e-08
Identities = 75/346 (21%), Positives = 151/346 (42%), Gaps = 45/346 (13%)
Query: 271 VSRETDIVGHTKLKSSGDAQLVLPLD-----QRNKLNRILSTMQRRLVTAKTDMEDLIVR 325
+ RE D + KL++ D Q V + R+K+N ++ R+ + D+ +
Sbjct: 386 LEREADEIEKIKLETQHDRQRVEEMTAQIQKDRDKINNLIEETNRKDMVLNEKNRDIEKK 445
Query: 326 LNQEIAAKDFLTTKVKDLE------VELETSKQKNKENLQQAILIERERFTQMQ----WD 375
+ + KD L + DLE +++ +K KEN I ERE +M +
Sbjct: 446 IKSIQSDKDMLEKEKHDLEKTRSELYKVKEDLEKQKENTLAQIQKEREDLEKMNENITRE 505
Query: 376 MEELRRKSLEMEMK------LKSESGGNVGQDLTKESIV-----QQNDVLLQDLDAAR-- 422
M E++ + +M K LK+E N+ Q+L KE + Q D +LD +
Sbjct: 506 MHEIKHQEEQMNQKQDELDQLKTEI-QNLQQELEKEKEIIMKARSQLDRRQSELDKQQTN 564
Query: 423 --EQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLH 480
+ +E + + +L+ K K +++ +E++ R + + +K L E +K ++++
Sbjct: 565 MNDIMETMKNERKQLD-KDKEEMEEQKQEMEKERDNMDQSRKSLDEDLKMMKLQKQVIEE 623
Query: 481 EREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDD 540
E+ ++ ++A + L+E + + ED ++ +K++ +D
Sbjct: 624 EKNNLDEMKVANESSMADLQKEKSNLEEMKENISKQTED---IEKEK-----DKIRLRED 675
Query: 541 QIDILLAEVENLEKDAGSAASNVDKTND--IKDGVICGDEMRKIIS 584
+++ L AE+ + + SN+++ IKD D KIIS
Sbjct: 676 ELEQLQAEIHKQQSETEIEKSNIERERAAIIKD---VEDLQSKIIS 718
Score = 57.8 bits (138), Expect = 9e-07
Identities = 74/345 (21%), Positives = 157/345 (45%), Gaps = 45/345 (13%)
Query: 282 KLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVK 341
K+K + Q L + K L M + +++ ++NQ+ D L T+++
Sbjct: 4 KVKEDLEKQKENTLAEIQKEREDLEKMNENITREMHEIKHQEEQMNQKQDELDQLKTEIQ 63
Query: 342 DLEVELETSK--------------------QKNKENLQQAILIERERFTQMQWDMEELRR 381
+L+ ELE K Q N ++ + + ER++ + + +MEE ++
Sbjct: 64 NLQQELEKEKEIIMKDRSQLDLRQSELDKQQTNMNDIMETMKNERKQLDKDKEEMEE-QK 122
Query: 382 KSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEI-LSKQYGEL----- 435
+ +E E +S ++ +D K + +Q + ++ EQ++I L ++ E+
Sbjct: 123 QEMEKERDNMDQSRKSLDEDQKKMKLQKQ---MFEEEKNKLEQMKIELEREADEISKIKE 179
Query: 436 EAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQ 495
E ++K K L K++K ++ L+K++ E K K E K+ L ERE E +I Q
Sbjct: 180 ETQNKRQRKSLDKDLKMMK-----LQKQVFEEEKNKLEQMKIEL-EREADEIRKIKEETQ 233
Query: 496 LEKCGLLLKQLQECSVELPYEDEDRTFL--QSSSSTDAFNKLKTSDDQIDI-LLAEVENL 552
E+ + L++ + EL E E T L + + +K+K +++ + L E NL
Sbjct: 234 NER-----QSLEKMTEELKKEKESFTHLAEDTKKEQEILDKVKVANESLMADLQKEKSNL 288
Query: 553 EKDAGSAASNVDKTNDIKDGV-ICGDEMRKIISDLFLDNVRLRKQ 596
E+ + + + + K+ + + DE+ ++ ++++ + K+
Sbjct: 289 EEMRENISKQTEDVENKKENLRLREDELWQLQAEIYKQQREIEKE 333
Score = 53.1 bits (126), Expect = 2e-05
Identities = 72/330 (21%), Positives = 150/330 (44%), Gaps = 47/330 (14%)
Query: 297 QRNKLNRILST-MQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNK 355
Q +K I+ +Q + K ++ RL++EI K VKDLE+E+ K++ +
Sbjct: 1337 QEDKFEDIIDIKIQEKEREIKIQLDKETKRLSEEIELK------VKDLEMEMADMKRQKQ 1390
Query: 356 ENLQQAILIERERFTQMQWDMEELRRKSLEM---------EMKLKSESGGNVGQDLTKES 406
E L+E+E+ +++ + +EL + +++ EMK K+E + + KE
Sbjct: 1391 EIEDTKGLLEKEK-QELKQEKKELEDQMMDLTQETKIKIIEMKTKTEP-----EKIKKEK 1444
Query: 407 IVQQNDVLL-------QDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTE 459
++ +V+ ++LD + QLE + + + K D K++ +E + L ++E
Sbjct: 1445 EKEEEEVMRAKVEMKKEELDQIKSQLERVRSEIDHEQKKLNDDKKMIEQEKEDLEKMKSE 1504
Query: 460 LKKELSESIKEKYEAEKLLLH---EREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYE 516
+ K+ + +E+ E + + ER E ++ +K + + K ++ EL E
Sbjct: 1505 IMKQRQQMEEERSELDNKIKQTDLERHDIENSKEIVQKLMVEVEEQRKDIRLQKEELDIE 1564
Query: 517 -----DEDRTFLQSSSSTDAFN-KLKTSDDQI---DILLAEVE-NLEKDAGSAASNVDKT 566
DE +Q+ + N ++K D++I L E+E +L K+ S +++T
Sbjct: 1565 RQKIADEQGLVVQNKAKLQNENERIKEMDEEIKKEKETLKEMEAHLRKEKEEMRSVIEET 1624
Query: 567 NDIKDGVICGDEMRKIISDLFLDNVRLRKQ 596
+ +++ K+ +D+ N L Q
Sbjct: 1625 QRRQK-----EDLEKMSTDVNKQNQDLMNQ 1649
Score = 50.8 bits (120), Expect = 1e-04
Identities = 67/302 (22%), Positives = 133/302 (43%), Gaps = 35/302 (11%)
Query: 297 QRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSK----- 351
Q+ +N I+ TM+ D E++ + + +D + K L+ +L+ K
Sbjct: 561 QQTNMNDIMETMKNERKQLDKDKEEMEEQKQEMEKERDNMDQSRKSLDEDLKMMKLQKQV 620
Query: 352 -QKNKENLQQAILIERERFTQMQWD---MEELRRKSLEMEMKLKSESGG-----NVGQDL 402
++ K NL + + +Q + +EE++ + ++ E + + L
Sbjct: 621 IEEEKNNLDEMKVANESSMADLQKEKSNLEEMKENISKQTEDIEKEKDKIRLREDELEQL 680
Query: 403 TKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKA---DIKVLVKEVKSLRSSQTE 459
E QQ++ ++ + RE+ I+ K +L++K + D + L + ++ + + E
Sbjct: 681 QAEIHKQQSETEIEKSNIERERAAII-KDVEDLQSKIISLDRDAESLKLDREAFENEKEE 739
Query: 460 LKKELSESIKEKYEAEKLLL---HEREKSEQAEIAW-------RKQLEKCGLLLKQLQEC 509
LK+ +E +E E EK+ L HER++ E+ + RKQL+K +++++ Q+
Sbjct: 740 LKQMKTELEREADEIEKIKLETQHERQRVEEMTADFMETMNNERKQLDKNKVMIEE-QKQ 798
Query: 510 SVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKTNDI 569
+E +D D QS S D LK Q +L E LE+ D+ + I
Sbjct: 799 EMEKKRDDMD----QSRKSLD--EDLKMMKAQKQVLEEEKNKLEQMKIGLEREADEISKI 852
Query: 570 KD 571
K+
Sbjct: 853 KE 854
Score = 47.4 bits (111), Expect = 0.001
Identities = 56/277 (20%), Positives = 121/277 (43%), Gaps = 23/277 (8%)
Query: 282 KLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVK 341
K+K + Q L Q K L M + +++ ++NQ+ D L T+++
Sbjct: 472 KVKEDLEKQKENTLAQIQKEREDLEKMNENITREMHEIKHQEEQMNQKQDELDQLKTEIQ 531
Query: 342 DLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQD 401
+L+ ELE K KE + +A +Q+ EL ++ M +++ D
Sbjct: 532 NLQQELE----KEKEIIMKA-------RSQLDRRQSELDKQQTNMNDIMETMKNERKQLD 580
Query: 402 LTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELK 461
KE + +Q Q+++ R+ ++ K E K +V+ +E +L + +
Sbjct: 581 KDKEEMEEQK----QEMEKERDNMDQSRKSLDEDLKMMKLQKQVIEEEKNNLDEMKVANE 636
Query: 462 KELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRT 521
+++ KEK E++ + +++E E K+ +K L +L++ E+ + +
Sbjct: 637 SSMADLQKEKSNLEEMKENISKQTEDIE----KEKDKIRLREDELEQLQAEIHKQQSETE 692
Query: 522 FLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGS 558
+S+ + +K +D L +++ +L++DA S
Sbjct: 693 IEKSNIERERAAIIKDVED----LQSKIISLDRDAES 725
Score = 47.0 bits (110), Expect = 0.002
Identities = 61/296 (20%), Positives = 130/296 (43%), Gaps = 57/296 (19%)
Query: 299 NKLNRILSTMQRRLVTAKTDMEDLIV------RLNQEIAAKDFLTTKVKDLEVELETSK- 351
++L + ++ R ME+L+ R +EI + + +++D ++LE +K
Sbjct: 1276 SELQKSRENIESREEKINNKMEELLQMKVEMGRTKEEIEKE---SQRLQDATIDLENAKA 1332
Query: 352 --QKNKENLQQAILI---ERERFTQMQWDME--------ELRRKSLEMEM----KLKSES 394
Q+ ++ + I I E+ER ++Q D E EL+ K LEMEM + K E
Sbjct: 1333 EIQRQEDKFEDIIDIKIQEKEREIKIQLDKETKRLSEEIELKVKDLEMEMADMKRQKQEI 1392
Query: 395 GGNVG------QDLTKE---------SIVQQNDVLLQDLDAAREQLEILSKQYGELEAKS 439
G Q+L +E + Q+ + + ++ E +I ++ E E
Sbjct: 1393 EDTKGLLEKEKQELKQEKKELEDQMMDLTQETKIKIIEMKTKTEPEKIKKEKEKEEEEVM 1452
Query: 440 KADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKC 499
+A +++ +E+ ++S ++ E+ K+ + +K++ E+E E+ + KQ ++
Sbjct: 1453 RAKVEMKKEELDQIKSQLERVRSEIDHEQKKLNDDKKMIEQEKEDLEKMKSEIMKQRQQ- 1511
Query: 500 GLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKD 555
+ ++ E ++ D +R + ++ S + + L+ EVE KD
Sbjct: 1512 --MEEERSELDNKIKQTDLER------------HDIENSKEIVQKLMVEVEEQRKD 1553
Score = 45.1 bits (105), Expect = 0.006
Identities = 60/304 (19%), Positives = 136/304 (44%), Gaps = 23/304 (7%)
Query: 300 KLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQ 359
K RIL M+ + D+E+ + L T+V+ + E++ K N E+ +
Sbjct: 904 KEKRILEEMRENISKQIEDIENEKEKSKLREDELKKLQTEVQKQQSEIDMEKT-NIESER 962
Query: 360 QAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLD 419
A++ E++ M EL++KS ++EM++K ++ E + ND L D
Sbjct: 963 AAMIREKQNM------MTELKKKSEDVEMQMK---------EILTEKELLHNDRNLLKKD 1007
Query: 420 AAREQLEILSKQYGELEAKS---KADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEK 476
Q +++ GEL+ + ++ + L+ E + + E+S + +++++ +
Sbjct: 1008 VEDLQQKLMKLFRGELQRQRDEVESSRQALIMEKDQFKLKIDHQRNEMSAAFEKEHKRLE 1067
Query: 477 LLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLK 536
+ ++ E +++A++A + L + ++LQ+ EL E E ++ T+ K++
Sbjct: 1068 NMENDLE-NQRAKLA--ESLNDAAKMEQKLQQMKNELNKEKEQVEEMKQMVKTERV-KIE 1123
Query: 537 TSDDQIDILLAEVENLEKDAGSAASNVDKTNDIKDGVICGDEMRKIISDLFLDNVRLRKQ 596
+++ E+E+ + DK + +E RK+I +L + +L
Sbjct: 1124 QEKEEMRKQTQEIEDGHEKLQKKTEIADKLLQDANMTTEQNEYRKLIEELQREKEQLEIS 1183
Query: 597 TNRV 600
N++
Sbjct: 1184 KNQI 1187
Score = 42.4 bits (98), Expect = 0.040
Identities = 59/261 (22%), Positives = 125/261 (47%), Gaps = 29/261 (11%)
Query: 308 MQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQ---QAILI 364
+Q + + ++E + + E AA + V+DL+ ++ S ++ E+L+ +A
Sbjct: 319 LQAEIYKQQREIEKEKINIESERAA---IIKNVEDLQHKI-ISLDRDSESLKLDREAFEN 374
Query: 365 ERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQ 424
ERE QM+ EL R++ E+E K+K E+ + ++ + + + +D D
Sbjct: 375 EREVLKQMK---TELEREADEIE-KIKLETQHD------RQRVEEMTAQIQKDRDKINNL 424
Query: 425 LEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREK 484
+E +++ L K++ DI+ K++KS++S + L+KE + K + E K+ ++
Sbjct: 425 IEETNRKDMVLNEKNR-DIE---KKIKSIQSDKDMLEKEKHDLEKTRSELYKVKEDLEKQ 480
Query: 485 SEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDI 544
E +K+ E + + + E+ +++E Q + D ++LKT +I
Sbjct: 481 KENTLAQIQKEREDLEKMNENITREMHEIKHQEE-----QMNQKQDELDQLKT---EIQN 532
Query: 545 LLAEVENLEKDAGSAASNVDK 565
L E+E ++ A S +D+
Sbjct: 533 LQQELEKEKEIIMKARSQLDR 553
>UniRef100_UPI0000432992 UPI0000432992 UniRef100 entry
Length = 1058
Score = 65.5 bits (158), Expect = 4e-09
Identities = 103/404 (25%), Positives = 175/404 (42%), Gaps = 50/404 (12%)
Query: 221 GHAKVLGHIRKLSNESVGSDGSSI-----RGSDISNFGIPNSSGDGSVDLPGSASVSRET 275
G K L + K NE S ++ R +SN + N + L + SV RE
Sbjct: 80 GQKKALEDLNKQLNEDNESMKRTMGNLEARIDSLSN-ELSNVERERDALLDENESVKREL 138
Query: 276 D--IVGHTKLKSS---GDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEI 330
+ + + LK+ D QL +RN+L R TM+ T K +++ L L +
Sbjct: 139 ERTLTENENLKTELDKADEQLDKLKTERNELQRNFDTMKLENETLKENVKALKDDLEESK 198
Query: 331 AAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKL 390
D + V D + E K LQQ + + +++ + ++LR +S E+E KL
Sbjct: 199 REVDEMKA-VGDALKDKEELKDAEFRELQQNMQNLKTENGELKKENDDLRTRSSELEHKL 257
Query: 391 KSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELE---AKSKADIKVLV 447
NV ++L K + +N L +D ++LE K+ +L+ + K + V
Sbjct: 258 D-----NVKKELDK--VESENADLRAKIDNLEKELEKDKKEIEQLKLEISSLKDALDKCV 310
Query: 448 KEVKSLRSSQTELKKE--------LSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKC 499
E++ L+ +LKKE L E++ K A+ L E + E+ K +
Sbjct: 311 DEMEKLKVENEKLKKEGMKVEATWLEENVNLK--AKNTELEENLANTVNELD--KMRSEN 366
Query: 500 GLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSA 559
LL +L EL E +++ TDA KL + + L AEV+ L+K+ SA
Sbjct: 367 ADLLSELNRLKQELESELQEKL-------TDASKKLDEAKTEDSDLRAEVDRLKKELESA 419
Query: 560 ASNVD----KTNDIKDGV-ICGDEMRKIISDLFLDNVRLRKQTN 598
+D + N +K+G+ C +EM K+ + +N L+ Q +
Sbjct: 420 GKEIDQLKAEMNSLKNGLNKCVEEMEKLTN----ENSELKSQVH 459
Score = 55.1 bits (131), Expect = 6e-06
Identities = 70/319 (21%), Positives = 146/319 (44%), Gaps = 22/319 (6%)
Query: 296 DQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNK 355
D +LN +M+R + + ++ L L+ +D L + + ++ ELE + +N
Sbjct: 87 DLNKQLNEDNESMKRTMGNLEARIDSLSNELSNVERERDALLDENESVKRELERTLTEN- 145
Query: 356 ENLQQAILIERERFTQMQWDMEELRRK--SLEMEMKLKSESGGNVGQDLTKESIVQQNDV 413
ENL+ + E+ +++ + EL+R ++++E + E+ + DL E ++ D
Sbjct: 146 ENLKTELDKADEQLDKLKTERNELQRNFDTMKLENETLKENVKALKDDL--EESKREVDE 203
Query: 414 LLQDLDAAREQLEILSKQYGELEAKS---KADIKVLVKEVKSLRSSQTELKKELSESIKE 470
+ DA +++ E+ ++ EL+ K + L KE LR+ +EL+ +L KE
Sbjct: 204 MKAVGDALKDKEELKDAEFRELQQNMQNLKTENGELKKENDDLRTRSSELEHKLDNVKKE 263
Query: 471 KYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTD 530
+ E R K + E K+LEK K++++ +E+ + D
Sbjct: 264 LDKVESENADLRAKIDNLE----KELEKDK---KEIEQLKLEISSLKD-----ALDKCVD 311
Query: 531 AFNKLKTSDDQI--DILLAEVENLEKDAGSAASNVDKTNDIKDGVICGDEMRKIISDLFL 588
KLK ++++ + + E LE++ A N + ++ + V D+MR +DL
Sbjct: 312 EMEKLKVENEKLKKEGMKVEATWLEENVNLKAKNTELEENLANTVNELDKMRSENADLLS 371
Query: 589 DNVRLRKQTNRVTRHMLSN 607
+ RL+++ + L++
Sbjct: 372 ELNRLKQELESELQEKLTD 390
Score = 53.1 bits (126), Expect = 2e-05
Identities = 67/311 (21%), Positives = 141/311 (44%), Gaps = 21/311 (6%)
Query: 296 DQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNK 355
D+ +KL ++ M+R L T+ + L +++ + + KVK LE +L ++ K
Sbjct: 758 DENDKLQTQINEMKRSLDKMGTENDRLKREVDESRKKLEDMEAKVKSLENQL-SNLSAEK 816
Query: 356 ENLQQAILIERERFTQMQWDMEELR--RKSLEMEMKLKSESGGNVGQDLTKESIVQQNDV 413
E L + + RE ++ ++E+ + ++E E E + ++L K +ND
Sbjct: 817 EELVKELYRTREDLNNLRNELEKQTGVKDTMEKESTNLKEELKALKEELNKTR--DENDR 874
Query: 414 LLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYE 473
L + D ++ L+KQ L+ +S A++K ++++L EL KEL+ +
Sbjct: 875 LKNENDKLNAEIARLNKQLDALKDES-ANLK---NDIENLNERNAELSKELAVAKDNLMG 930
Query: 474 AEKLLLHEREKSEQAE---IAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTD 530
E L + +++++ + I +++ L +QL+E EL + L+S++
Sbjct: 931 METRLSNLKKENDDMKNKIITLEDSIQEVDDLKRQLKEAKKELDKPSPELDTLKSTN--- 987
Query: 531 AFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDK-TNDIKDGVICGDEMRKIISDLFLD 589
K D +D E NL+ D + ++ + ++ D D R+ + L D
Sbjct: 988 -----KKLQDDLDNARNESLNLKNDLDNLQNDYNNLQTELADVKEERDTFRERAAALEKD 1042
Query: 590 NVRLRKQTNRV 600
VR++++ +
Sbjct: 1043 LVRVKRENEEL 1053
Score = 50.1 bits (118), Expect = 2e-04
Identities = 71/294 (24%), Positives = 125/294 (42%), Gaps = 26/294 (8%)
Query: 316 KTDMEDLIVRLNQ----EI----AAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERE 367
K D+E ++ LN+ EI D T + L+ +L++ K +N + Q ++R
Sbjct: 714 KADLEKVLKDLNECQRVEIDRLRTTLDAEKTAAEKLKSDLQSCKDENDKLQTQINEMKRS 773
Query: 368 ------RFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAA 421
+++ +++E R+K +ME K+KS KE +V++ +DL+
Sbjct: 774 LDKMGTENDRLKREVDESRKKLEDMEAKVKSLENQLSNLSAEKEELVKELYRTREDLNNL 833
Query: 422 REQLEILSKQYGELEAKS---KADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLL 478
R +LE + +E +S K ++K L +E+ R LK E + E K L
Sbjct: 834 RNELEKQTGVKDTMEKESTNLKEELKALKEELNKTRDENDRLKNENDKLNAEIARLNKQL 893
Query: 479 LHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTS 538
+++S + E+ L K+L L E R + D NK+ T
Sbjct: 894 DALKDESANLKNDIENLNERNAELSKELAVAKDNL-MGMETRLSNLKKENDDMKNKIITL 952
Query: 539 DDQIDILLAEVENLEKDAGSAASNVDKTNDIKDGVICGDEMRKIISDLFLDNVR 592
+D I EV++L++ A +DK + D + +K+ D LDN R
Sbjct: 953 EDSIQ----EVDDLKRQLKEAKKELDKPSPELDTL--KSTNKKLQDD--LDNAR 998
Score = 48.1 bits (113), Expect = 7e-04
Identities = 64/293 (21%), Positives = 125/293 (41%), Gaps = 54/293 (18%)
Query: 296 DQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNK 355
D ++LNR+ ++ L TD +L++ L +V L+ ELE S K
Sbjct: 368 DLLSELNRLKQELESELQEKLTDASK---KLDEAKTEDSDLRAEVDRLKKELE-SAGKEI 423
Query: 356 ENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKS--ESGGNVGQDLTKESIVQQNDV 413
+ L+ + + + +ME+L ++ E++ ++ G ++ +LT ++ +N
Sbjct: 424 DQLKAEMNSLKNGLNKCVEEMEKLTNENSELKSQVHGLRGEGDSLASELT--NVKDENSA 481
Query: 414 LLQDLDAAREQL-------EILSKQYGELEA------------------------KSKAD 442
L + D +QL E L KQ ELE+ K K +
Sbjct: 482 LKDEKDQLNKQLAENKTENERLKKQNDELESENTKIKKELESCKNENNNLKEENNKLKEE 541
Query: 443 IKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLL 502
++ L +++KSL +L++EL E+ + E L R ++E+++ + +L
Sbjct: 542 LEKLGEQLKSLNDETNKLRRELKEAEDKIQILEPQLSRARSENEKSQ-------NELAVL 594
Query: 503 LKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKD 555
+ E V+L E D T ++ N LK +DQ+ L +++N ++
Sbjct: 595 RNEANELKVKLDREMLDNTNMR--------NALKILEDQVLDLNKKLDNCREE 639
Score = 47.8 bits (112), Expect = 0.001
Identities = 60/288 (20%), Positives = 121/288 (41%), Gaps = 23/288 (7%)
Query: 298 RNKLNRILSTMQRRLV------TAKTDMEDLIVRLNQEI----AAKDFLTTKVKDLEVEL 347
RN+ N + + R ++ A +ED ++ LN+++ D L + KDL+ +L
Sbjct: 595 RNEANELKVKLDREMLDNTNMRNALKILEDQVLDLNKKLDNCREENDALKEENKDLKTKL 654
Query: 348 ETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESI 407
+ Q NL+ +E +Q +E+L++K + E ++ + +L E +
Sbjct: 655 SDTGQVVL-NLKTECDNLKEDIASLQKTIEQLKQKIADQEAEIDHWKVEHCKFELDNEKL 713
Query: 408 VQQNDVLLQDLDAAR----EQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKE 463
+ +L+DL+ + ++L K K+D++ E L++ E+K+
Sbjct: 714 KADLEKVLKDLNECQRVEIDRLRTTLDAEKTAAEKLKSDLQSCKDENDKLQTQINEMKRS 773
Query: 464 LSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFL 523
L + E ++ + R+K E E + + L + +E EL ED L
Sbjct: 774 LDKMGTENDRLKREVDESRKKLEDMEAKVKSLENQLSNLSAEKEELVKELYRTREDLNNL 833
Query: 524 QSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKTNDIKD 571
+ N+L+ D + E NL+++ + ++KT D D
Sbjct: 834 R--------NELEKQTGVKDTMEKESTNLKEELKALKEELNKTRDEND 873
>UniRef100_P08799 Myosin II heavy chain, non muscle [Dictyostelium discoideum]
Length = 2116
Score = 64.3 bits (155), Expect = 1e-08
Identities = 68/279 (24%), Positives = 127/279 (45%), Gaps = 41/279 (14%)
Query: 308 MQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERE 367
+++ V +++++DL VRL+ E K L + K LE EL + +Q+A+ E
Sbjct: 1001 LEKTRVRLQSELDDLTVRLDSETKDKSELLRQKKKLEEEL--------KQVQEALAAETA 1052
Query: 368 RFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEI 427
+ ++L+ + E+ K SE + +K+++ Q + +LD ++ +
Sbjct: 1053 AKLAQEAANKKLQGEYTELNEKFNSEVTARSNVEKSKKTLESQLVAVNNELDEEKKNRDA 1112
Query: 428 LSKQYGELEA----------------KSKADIKVLVK-EVKSLRSSQTELKKELS--ESI 468
L K+ L+A KS D+KV + ++++LR+ +EL+ ++ E I
Sbjct: 1113 LEKKKKALDAMLEEMKDQLESTGGEKKSLYDLKVKQESDMEALRNQISELQSTIAKLEKI 1172
Query: 469 KEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSS 528
K E E L ++EQ + +EK Q+ VEL ED+ + +++
Sbjct: 1173 KSTLEGEVARLQGELEAEQLA---KSNVEK--------QKKKVELDLEDKSAQLAEETAA 1221
Query: 529 TDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKTN 567
A +KLK +Q L+EV+ +A + N D TN
Sbjct: 1222 KQALDKLKKKLEQ---ELSEVQTQLSEANNKNVNSDSTN 1257
Score = 58.9 bits (141), Expect = 4e-07
Identities = 59/281 (20%), Positives = 125/281 (43%), Gaps = 33/281 (11%)
Query: 305 LSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILI 364
L+ ++++ + +++++ +L EI AKD L + LEVELE + + +E
Sbjct: 1666 LNASEKKIKSLVAEVDEVKEQLEDEILAKDKLVKAKRALEVELEEVRDQLEE-------- 1717
Query: 365 ERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQ 424
E + ++++ L + +++ K +E N D K+ + D L + L+ +++
Sbjct: 1718 EEDSRSELEDSKRRLTTEVEDIKKKYDAEVEQNTKLDEAKKKLTDDVDTLKKQLEDEKKK 1777
Query: 425 LEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREK 484
L + LE++++ + L EVK+ ++ + KK + KY+ L E
Sbjct: 1778 LNESERAKKRLESENEDFLAKLDAEVKNRSRAEKDRKKYEKDLKDTKYK----LNDEAAT 1833
Query: 485 SEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDI 544
Q EI K ++ L +L++ + + +T A KT + +ID
Sbjct: 1834 KTQTEIGAAKLEDQIDELRSKLEQ---------------EQAKATQADKSKKTLEGEIDN 1878
Query: 545 LLAEVENLEKDAGSAASNVDKTNDIKDGVICGDEMRKIISD 585
L A++E D G ++K +G + +E+R+ + +
Sbjct: 1879 LRAQIE----DEGKIKMRLEKEKRALEGEL--EELRETVEE 1913
Score = 51.2 bits (121), Expect = 9e-05
Identities = 77/330 (23%), Positives = 136/330 (40%), Gaps = 43/330 (13%)
Query: 298 RNKLNRILSTMQRRLVTA--------------KTDMEDLIVRLNQEIAAKDFLTTKVKDL 343
+ KL + LS +Q +L A +T +L + L E AK L K L
Sbjct: 1229 KKKLEQELSEVQTQLSEANNKNVNSDSTNKHLETSFNNLKLELEAEQKAKQALEKKRLGL 1288
Query: 344 EVEL-------ETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGG 396
E EL E K++ + N ++ + +E+E EE+ K E K K ES
Sbjct: 1289 ESELKHVNEQLEEEKKQKESNEKRKVDLEKEVSELKDQIEEEVASKKAVTEAKNKKES-- 1346
Query: 397 NVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSS 456
+ I +Q ++ D + EQL+ L + EL ++ L + +S + +
Sbjct: 1347 ------ELDEIKRQYADVVSSRDKSVEQLKTLQAKNEELRNTAEEAEGQLDRAERSKKKA 1400
Query: 457 QTELK---KELSESIKEKYEAEKLLL-----HEREKSE--QAEIAWRKQLEKCGLLLKQL 506
+ +L+ K L E +K +AEK + + KSE A+ +Q + L ++L
Sbjct: 1401 EFDLEEAVKNLEEETAKKVKAEKAMKKAETDYRSTKSELDDAKNVSSEQYVQIKRLNEEL 1460
Query: 507 QECSVELPYEDE--DRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVD 564
E L DE + ++ A LK D + A+ E K+ + ++
Sbjct: 1461 SELRSVLEEADERCNSAIKAKKTAESALESLKDEIDAANNAKAKAERKSKELEVRVAELE 1520
Query: 565 KTNDIKDGVICGDEMRKIISDLFLDNVRLR 594
++ + K G + + +RK D +D++R R
Sbjct: 1521 ESLEDKSGTVNVEFIRK--KDAEIDDLRAR 1548
Score = 47.0 bits (110), Expect = 0.002
Identities = 66/282 (23%), Positives = 132/282 (46%), Gaps = 30/282 (10%)
Query: 283 LKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKD 342
+K DA++ +Q KL+ + + T K +ED +LN+ AK L ++ +D
Sbjct: 1739 IKKKYDAEV----EQNTKLDEAKKKLTDDVDTLKKQLEDEKKKLNESERAKKRLESENED 1794
Query: 343 LEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRK-SLEMEMKLKSESGGNVGQD 401
+L+ ++ KN+ ++ +R+++ + D+++ + K + E K ++E G +D
Sbjct: 1795 FLAKLD-AEVKNRSRAEK----DRKKYEK---DLKDTKYKLNDEAATKTQTEIGAAKLED 1846
Query: 402 LTKE--SIVQQNDVLLQDLDAAREQLE-ILSKQYGELEAKSKADIKVLVKEVKSLRSSQT 458
E S ++Q D +++ LE + ++E + K ++ L KE ++L
Sbjct: 1847 QIDELRSKLEQEQAKATQADKSKKTLEGEIDNLRAQIEDEGKIKMR-LEKEKRALEGELE 1905
Query: 459 ELKKELSESIKEKYEAE------KLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVE 512
EL++ + E+ K EAE +L L + ++ Q EI ++ E LQ VE
Sbjct: 1906 ELRETVEEAEDSKSEAEQSKRLVELELEDARRNLQKEIDAKEIAEDA---KSNLQREIVE 1962
Query: 513 LPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEK 554
E+ +S + T++ K + +ID L A+V+ +K
Sbjct: 1963 AKGRLEE----ESIARTNSDRSRKRLEAEIDALTAQVDAEQK 2000
Score = 44.7 bits (104), Expect = 0.008
Identities = 63/284 (22%), Positives = 120/284 (42%), Gaps = 25/284 (8%)
Query: 294 PLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLE---VELETS 350
PL +R + + +R ++ K+++ D + KD L +KD E ++L+
Sbjct: 819 PLLKRRNFEKEIKEKEREILELKSNLTDSTTQ-------KDKLEKSLKDTESNVLDLQRQ 871
Query: 351 KQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNV-GQDLTKESIVQ 409
+ KE L+ + + + + Q E+R + +E E+ K + N+ Q + E V+
Sbjct: 872 LKAEKETLKA--MYDSKDALEAQKRELEIRVEDMESELDEKKLALENLQNQKRSVEEKVR 929
Query: 410 QNDVLLQDLDAAREQLEILSKQYGE-LEAKSKAD------IKVLVKEVKSLRSSQTELKK 462
+ LQ+ R LE L K+Y E LE + + I L K L+ EL +
Sbjct: 930 DLEEELQEEQKLRNTLEKLKKKYEEELEEMKRVNDGQSDTISRLEKIKDELQKEVEELTE 989
Query: 463 ELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTF 522
SE K+K EK + + + + + + + LL+Q ++ EL E
Sbjct: 990 SFSEESKDKGVLEKTRVRLQSELDDLTVRLDSETKDKSELLRQKKKLEEELKQVQE---- 1045
Query: 523 LQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKT 566
++ T A + ++ ++ E+ +A SNV+K+
Sbjct: 1046 -ALAAETAAKLAQEAANKKLQGEYTELNEKFNSEVTARSNVEKS 1088
Score = 42.4 bits (98), Expect = 0.040
Identities = 68/309 (22%), Positives = 132/309 (42%), Gaps = 36/309 (11%)
Query: 296 DQRNKLNRILSTM-QRRLVTAKTD--MEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQ 352
D + KLN +T Q + AK + +++L +L QE A K LE E++ +
Sbjct: 1822 DTKYKLNDEAATKTQTEIGAAKLEDQIDELRSKLEQEQAKATQADKSKKTLEGEIDNLRA 1881
Query: 353 KNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQND 412
+ ++ + + +E+E+ ++ ++EELR E E S+S + L + +
Sbjct: 1882 QIEDEGKIKMRLEKEK-RALEGELEELRETVEEAE---DSKSEAEQSKRLVELELEDARR 1937
Query: 413 VLLQDLDAA-----------REQLEILSKQYGELEAKSKADI--KVLVKEVKSLRSSQTE 459
L +++DA RE +E + E A++ +D K L E+ +L +
Sbjct: 1938 NLQKEIDAKEIAEDAKSNLQREIVEAKGRLEEESIARTNSDRSRKRLEAEIDALTAQVDA 1997
Query: 460 LKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDED 519
+K ++ IKE + E L R+K ++E K+ L + E E +
Sbjct: 1998 EQKAKNQQIKENKKIETELKEYRKKFGESEKTKTKEFLVVEKLETDYKRAKKEAADEQQQ 2057
Query: 520 RTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKT-NDIKDGVICGDE 578
R ++ N L+ +I +L ++ L++D DKT +++ E
Sbjct: 2058 RLTVE--------NDLRKHLSEISLLKDAIDKLQRDH-------DKTKRELETETASKIE 2102
Query: 579 MRKIISDLF 587
M++ ++D F
Sbjct: 2103 MQRKMADFF 2111
Score = 39.7 bits (91), Expect = 0.26
Identities = 50/242 (20%), Positives = 114/242 (46%), Gaps = 11/242 (4%)
Query: 318 DMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDME 377
+++DL RL++E ++ K+ + + K +E ++ + I+R + +++ D+
Sbjct: 1541 EIDDLRARLDRETESRIKSDEDKKNTRKQFADLEAKVEEAQREVVTIDRLK-KKLESDII 1599
Query: 378 ELRRK-SLEMEMKLKSE-SGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGEL 435
+L + E + ++K E S + Q L + ++ D + ++ + + + +L
Sbjct: 1600 DLSTQLDTETKSRIKIEKSKKKLEQTLAERRAAEEGSSKAADEEIRKQVWQEVDELRAQL 1659
Query: 436 EAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQ 495
+++ +A + K++KSL + E+K++L + I K +KL+ +R + E R Q
Sbjct: 1660 DSE-RAALNASEKKIKSLVAEVDEVKEQLEDEILAK---DKLVKAKRALEVELEEV-RDQ 1714
Query: 496 LEKCGLLLKQLQECSVELPYEDED---RTFLQSSSSTDAFNKLKTSDDQIDILLAEVENL 552
LE+ +L++ L E ED + + +T K D +D L ++E+
Sbjct: 1715 LEEEEDSRSELEDSKRRLTTEVEDIKKKYDAEVEQNTKLDEAKKKLTDDVDTLKKQLEDE 1774
Query: 553 EK 554
+K
Sbjct: 1775 KK 1776
>UniRef100_UPI000042CCB9 UPI000042CCB9 UniRef100 entry
Length = 658
Score = 63.5 bits (153), Expect = 2e-08
Identities = 77/335 (22%), Positives = 148/335 (43%), Gaps = 8/335 (2%)
Query: 243 SIRGSDISNFGIPNSSGDGSVDLPGSASVSRE-TDIVGHTKLKSSGDAQLVLPLDQRNKL 301
SI+ ++N + + D SV S+ E TD + K K+ Q+V ++ K
Sbjct: 155 SIKKLKLANQDLEENIRDYSVKSEESSMKLEELTDFLKTHKFKTID--QVVSKYEEVVKE 212
Query: 302 NRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQA 361
+ T +T +L+V + AAK L + ++++EL K+++K ++
Sbjct: 213 LEVAKAGLENSHTWETKYNELVVSHEEANAAKKELLKETNEVKIELNMLKRQHKLEIESK 272
Query: 362 ILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAA 421
+E +QM + ++ +E K+++ N + E++ + V +
Sbjct: 273 NATIKELKSQMSASKQSFAQEISRLEDKIETLRLENESNETVSENVNEDGKVDFDEFSKL 332
Query: 422 REQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKK---ELSESIKEKYEAEKLL 478
E IL KQY + KA L +V +L SS LKK +L++ + + + L
Sbjct: 333 SEAHRILQKQYLSSQENWKAIESNLSLKVDNLSSSVESLKKSKHKLTQDLAKVNNSMHLQ 392
Query: 479 LHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTS 538
L E EK EQ +I +++ + L ++ + VE+ + E + + +K+++
Sbjct: 393 LKEMEKLEQDKIKLQEERNQLQLTVEMKENEIVEITEKFEKFKGIYNQERAALNSKIQSL 452
Query: 539 DDQI-DILLAEVENLEKDAGSAASNVDKTNDIKDG 572
+D+I + E NL +D S +S N+IK G
Sbjct: 453 NDKIKEEKRPERLNLIRD-NSVSSGFSWDNEIKLG 486
>UniRef100_Q25886 Liver stage antigen-1 [Plasmodium falciparum]
Length = 493
Score = 63.5 bits (153), Expect = 2e-08
Identities = 75/276 (27%), Positives = 132/276 (47%), Gaps = 29/276 (10%)
Query: 309 QRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSK--QKNKENLQQAILIER 366
Q RL AK +++ L QE AK+ L + DLE E + Q+ + +L+Q L +
Sbjct: 43 QERL--AKEKLQEQQSDLEQERRAKEKLQEQQSDLEQERRAKEKLQEQQSDLEQDRL-AK 99
Query: 367 ERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLE 426
E+ + Q D+E+ RR + KL+ + + KE + +Q DL+ R E
Sbjct: 100 EKLQEQQSDLEQERR----AKEKLQEQQSDLEQERRAKEKLQEQQ----SDLEQERLAKE 151
Query: 427 ILSKQYGELEAKSKADIKVLVK--EVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREK 484
L +Q +LE + +A K+ + +++ R ++ +L+++ S+ +E+ EKL +R+
Sbjct: 152 KLQEQQSDLEQERRAKEKLQEQQSDLEQERRAKEKLQEQQSDLEQERRAKEKLQEQQRD- 210
Query: 485 SEQAEIAWRKQLEK--------CGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLK 536
EQ + +K LE+ L +L+ ++ELP E+E ++ SS N+
Sbjct: 211 LEQRKADTKKNLERKKEHGDILAEDLYGRLEIPAIELPSENERGYYIPHQSSLPQDNR-G 269
Query: 537 TSDDQIDILLAEVENLEKDAGSAASNVDKTNDIKDG 572
S D +I + E N E S +NV+ DI G
Sbjct: 270 NSRDSKEISIIEKTNRE----SITTNVEGRRDIHKG 301
>UniRef100_Q72YP9 S-layer homology domain protein [Bacillus cereus]
Length = 939
Score = 62.8 bits (151), Expect = 3e-08
Identities = 58/216 (26%), Positives = 109/216 (49%), Gaps = 29/216 (13%)
Query: 324 VRLNQEIAAKDFLTTKV---KDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELR 380
++ +E+ KD + K ++ +V E +QK KE L +++ +++ EEL+
Sbjct: 139 LKSQEELKEKDGVEVKENNGQEEKVHEEFEEQKKKEEL-------KKQQDELRKQQEELK 191
Query: 381 RKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYG---ELEA 437
++ LE+E ++K E ++ K+ + L Q + A+ +LE+ K+ ELE
Sbjct: 192 KQQLELEQRIKQELERKQQEEQAKQELE-----LKQKEEQAKRELELKQKEEQVKQELEL 246
Query: 438 KSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLE 497
K K + VK+ L+ + ++K+EL KE+ E ++L L ++E+ E+ E+ +++ E
Sbjct: 247 KQKEE---QVKQELELKQKEEQVKQELELKQKEEQEKQELELKQKEEQEKQELELKQKEE 303
Query: 498 --KCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDA 531
K L LKQ +E E +R Q+ SS A
Sbjct: 304 QKKQELELKQKEE------QEKGERELKQAVSSIQA 333
Score = 47.4 bits (111), Expect = 0.001
Identities = 62/259 (23%), Positives = 120/259 (45%), Gaps = 36/259 (13%)
Query: 306 STMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIE 365
S +Q++L AK ++ ++ + + ++L ++++ KQK +E L+QA
Sbjct: 48 SELQKQLEAAK--------EREMKLEKREEILKQQEELFIKIDDLKQKKEELLEQAGE-H 98
Query: 366 RERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQL 425
+ ++ ++ EL+++ E+E K + N + T E + Q ++ +D +E
Sbjct: 99 NVQIEEVYQELNELKKQQEELEEKNPLQVKSNDQEKKTNE-LKSQEELKEKDGVEVKENN 157
Query: 426 EILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKK---ELSESIKEKYEAEKLLLHER 482
K + E E + K + L K+ LR Q ELKK EL + IK++ E +
Sbjct: 158 GQEEKVHEEFEEQKKKE--ELKKQQDELRKQQEELKKQQLELEQRIKQELE-------RK 208
Query: 483 EKSEQA----EIAWRKQLEKCGLLLKQLQE-CSVELPYEDEDRTFLQSSSSTDAFNKLKT 537
++ EQA E+ +++ K L LKQ +E EL + ++ Q +LK
Sbjct: 209 QQEEQAKQELELKQKEEQAKRELELKQKEEQVKQELELKQKEEQVKQEL-------ELKQ 261
Query: 538 SDDQI--DILLAEVENLEK 554
++Q+ ++ L + E EK
Sbjct: 262 KEEQVKQELELKQKEEQEK 280
Score = 40.8 bits (94), Expect = 0.12
Identities = 43/173 (24%), Positives = 85/173 (48%), Gaps = 15/173 (8%)
Query: 345 VELETSKQKN-KENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLT 403
+E + Q N K LQ+ + +ER +++ EE+ ++ E+ +K+ DL
Sbjct: 36 IEAVSHMQSNEKSELQKQLEAAKEREMKLE-KREEILKQQEELFIKI---------DDLK 85
Query: 404 --KESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLR-SSQTEL 460
KE +++Q ++ ++L L KQ ELE K+ +K +E K+ SQ EL
Sbjct: 86 QKKEELLEQAGEHNVQIEEVYQELNELKKQQEELEEKNPLQVKSNDQEKKTNELKSQEEL 145
Query: 461 KKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVEL 513
K++ +KE E+ +HE + ++ + +KQ ++ ++L++ +EL
Sbjct: 146 KEKDGVEVKENNGQEE-KVHEEFEEQKKKEELKKQQDELRKQQEELKKQQLEL 197
Score = 37.7 bits (86), Expect = 0.99
Identities = 37/175 (21%), Positives = 85/175 (48%), Gaps = 21/175 (12%)
Query: 297 QRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKE 356
++ +L + ++++ K +L R+ QE+ K ++LE+ KQK ++
Sbjct: 172 KKEELKKQQDELRKQQEELKKQQLELEQRIKQELERKQQEEQAKQELEL-----KQKEEQ 226
Query: 357 NLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQ 416
++ L ++E + + ++++ + + ++ E++LK KE V+Q L Q
Sbjct: 227 AKRELELKQKEEQVKQELELKQ-KEEQVKQELELKQ-----------KEEQVKQELELKQ 274
Query: 417 DLDAAREQLEILSK---QYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESI 468
+ +++LE+ K + ELE K K + K E+K + Q + ++EL +++
Sbjct: 275 KEEQEKQELELKQKEEQEKQELELKQKEEQKKQELELKQ-KEEQEKGERELKQAV 328
>UniRef100_UPI000049A29E UPI000049A29E UniRef100 entry
Length = 1813
Score = 62.4 bits (150), Expect = 4e-08
Identities = 66/289 (22%), Positives = 136/289 (46%), Gaps = 39/289 (13%)
Query: 299 NKLNRILSTMQR------RLVTAKTDMEDLIVRLNQEIAA----KDFLTTKVKDLEVELE 348
N+LN+ Q+ +L+T ++ D I +LN+E+ K+ + ++ ++ E
Sbjct: 742 NELNQTKDEKQKIEDEKSKLITELSNGNDGISKLNEELTQTKQEKENVLNELNQIKNEFA 801
Query: 349 TSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDL--TKES 406
+ K++N + E E + +EL +K+ E+ KL+ E G N+ +L TK+
Sbjct: 802 SFKEQNTQK-------ENELKDENNKVQQELEQKNNEVS-KLEEEKG-NISNELSNTKQE 852
Query: 407 IVQQNDVLLQDLDAAREQLEILSKQYGELEA-KSKADIKVLVKEVKSLRSSQTELKKELS 465
+ Q+ ++ E+ L +Q ++E KSK L+ E+ + ++L +EL+
Sbjct: 853 LEQKKQEIITITQEKEEKENELKEQVKKIEEEKSK-----LITELSNGSDGISKLNEELT 907
Query: 466 ESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQS 525
++ +EK E +K L E+EK E+ E LK+++E EL E++++T +
Sbjct: 908 QTKQEKEEIQKALEEEKEKLERIETE-----------LKEIKEAKQELE-EEKNKTIEEK 955
Query: 526 SSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKTNDIKDGVI 574
++ N+ K +++ E E + + S + + K+ +I
Sbjct: 956 TNLQQELNENKKIVEELTQTKQEKEEINNELNSIKEEKKRIEEEKNQII 1004
Score = 60.1 bits (144), Expect = 2e-07
Identities = 73/307 (23%), Positives = 138/307 (44%), Gaps = 40/307 (13%)
Query: 300 KLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKD-------LEVELETSKQ 352
K I+ + + K + E+L LNQ K L T + + L E+ET
Sbjct: 1330 KNQEIICDNNKEIAKFKEEQENLQKELNQIKEEKSKLITDLSNGNDGLSKLNEEIETIN- 1388
Query: 353 KNKENLQQAILIERERFTQMQWDME----ELRRKSLEMEMKLKSESGGNVG-----QDLT 403
K KE +++ + +E ++Q ++E EL + E E + + GN G +DL
Sbjct: 1389 KEKEGIRKELESLKEENNKIQDELEQKNQELSKVKEEKEKLIHDLTNGNDGINQLNEDLN 1448
Query: 404 -----KESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQT 458
KE + ++N L +++ + + E LS + K +K + +EV +++ +
Sbjct: 1449 QIKNDKEELTEKNVQLQNEINKLKSENEELSNNL----SFEKEGLKQVNEEVNAIKEERD 1504
Query: 459 ELKKELSESIKEKYEAEKLL-LHEREKSEQ-AEIAWRKQL--EKCGLL---LKQLQECSV 511
EL K++ + +EK + E+ L + E +EQ A+I K+ ++C L LK+LQ
Sbjct: 1505 ELVKQIKKIEEEKRKVEEELNFNGSEVNEQIAQINNEKEQLNQECNELKQNLKELQSKIE 1564
Query: 512 ELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEK-------DAGSAASNVD 564
E+ E E + + ++ D+ I L E+E +EK D ++N +
Sbjct: 1565 EIEQEKESNEIKKKEELQELQEEITEKDNDIKNLKEEIERIEKELQEKEEDMEQMSNNTE 1624
Query: 565 KTNDIKD 571
+ ++K+
Sbjct: 1625 ELEELKN 1631
Score = 55.8 bits (133), Expect = 4e-06
Identities = 66/311 (21%), Positives = 142/311 (45%), Gaps = 42/311 (13%)
Query: 281 TKLKSSGDAQLVLPLDQRNKLNRILSTMQR---RLVTAKTDMEDLIVRLNQEIAAKDFLT 337
T+LK+ D + N+LN++ + T+K + E ++ L+Q K+
Sbjct: 278 TQLKTDND-------QKENELNQVRHEKDEVIEKFNTSKEENEKIMNELSQLKQEKEEKE 330
Query: 338 TKVKDLEVELETSKQK---NKENLQQAILIERERFTQMQWDMEELRRK--SLEMEMKLKS 392
++K+ ++E K K N I E TQ + + EE+ + S++ E K
Sbjct: 331 NELKEQVKKMEEEKSKLITELSNGSDGISKLNEELTQTKQEKEEINNELNSIKEEKKRIE 390
Query: 393 ESGGNVGQDLT-----KESIVQQNDVLLQDLDAARE-------QLEILSKQYGELEAKSK 440
E + + KE I ++ LL++++ +E ++ + + E+E K++
Sbjct: 391 EEKNQIINENKEIKEEKEKIEEEKKELLKEIEKEKEGNNQLQNEINTIQTRMKEIEEKNQ 450
Query: 441 ADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCG 500
I KE+ + Q L+KEL++ +EK + E EK+E ++ +K+ E
Sbjct: 451 EIICDNNKEIAKFKEEQENLQKELNQIKEEKQKT------ENEKNELVDVKTQKENE--- 501
Query: 501 LLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAA 560
L +L+E E ++T +++S + K K ++++ + + E+++++ D +
Sbjct: 502 --LNKLKE---EKEQIFNEKTTIENSLNQIVEEKNKLTEEK-ESIKQELDSIKADNSTKE 555
Query: 561 SNVDKTNDIKD 571
++K N+ K+
Sbjct: 556 LEINKINEEKN 566
Score = 53.1 bits (126), Expect = 2e-05
Identities = 58/285 (20%), Positives = 127/285 (44%), Gaps = 45/285 (15%)
Query: 297 QRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKE 356
+RNK+N T+ L K ++ DL + + EI LE+ KNK+
Sbjct: 1158 ERNKINEEYKTVNEELEKNKKELNDLQTKYDNEI------------LEL------NKNKD 1199
Query: 357 NLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGG--NVGQDLTKESIVQQNDVL 414
L I +E T ++ ++++ + ++ +L + S G + ++LT+ Q+ + +
Sbjct: 1200 ELNSLINNLKEEKTNLEEQVKKMEEEKSKLITELSNGSDGVSKLNEELTQTK--QEKEEI 1257
Query: 415 LQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEA 474
+L++ +E+ + + ++ ++ ++ KE+K + E KKEL + I+++ E
Sbjct: 1258 NNELNSIKEEKKRIEEEKNQIINEN--------KEIKEEKEKIEEEKKELLKEIEKEKEG 1309
Query: 475 EKLLLHE-----------REKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFL 523
L +E EK+++ K++ K + LQ+ + E++ +
Sbjct: 1310 NNQLQNEINTIQTRMKEIEEKNQEIICDNNKEIAKFKEEQENLQK-ELNQIKEEKSKLIT 1368
Query: 524 QSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKTND 568
S+ D +KL +++I+ + E E + K+ S +K D
Sbjct: 1369 DLSNGNDGLSKL---NEEIETINKEKEGIRKELESLKEENNKIQD 1410
Score = 51.6 bits (122), Expect = 7e-05
Identities = 58/315 (18%), Positives = 142/315 (44%), Gaps = 28/315 (8%)
Query: 295 LDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKD----LEVELETS 350
L++ N++ ++ + + + +++D ++ QE+ K+ +K+++ + EL +
Sbjct: 790 LNELNQIKNEFASFKEQNTQKENELKDENNKVQQELEQKNNEVSKLEEEKGNISNELSNT 849
Query: 351 KQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSE-SGGNVGQDLTKESIVQ 409
KQ+ ++ Q+ I I +E+ + + +++E +K E + KL +E S G+ G E + Q
Sbjct: 850 KQELEQKKQEIITITQEK-EEKENELKEQVKKIEEEKSKLITELSNGSDGISKLNEELTQ 908
Query: 410 QNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIK 469
Q+ + ++ LE + K ++ + E+K ++ ++ EL++E +++I+
Sbjct: 909 TK----QEKEEIQKALE-----------EEKEKLERIETELKEIKEAKQELEEEKNKTIE 953
Query: 470 EKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSST 529
EK ++ L+E +K + +++ E+ L ++E + E
Sbjct: 954 EKTNLQQ-ELNENKKIVEELTQTKQEKEEINNELNSIKEEKKRIEEEKNQIINENKEIKE 1012
Query: 530 DAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKTNDIKDGVICGDEMRKIISDLFLD 589
+ ++ +I+ L +E L+ + + +D VI ++D+ L
Sbjct: 1013 ENIKSIEEKTQEINSLTTSIEELKGRLEESKGERIEIEKERDRVI------SELNDIKLQ 1066
Query: 590 NVRLRKQTNRVTRHM 604
N ++KQ M
Sbjct: 1067 NEGMKKQVEEAHNRM 1081
Score = 47.8 bits (112), Expect = 0.001
Identities = 63/305 (20%), Positives = 145/305 (46%), Gaps = 41/305 (13%)
Query: 299 NKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELET---SKQKNK 355
N LN+I+ + +L K ++ + + + + K+ K+ + + +L+ + Q+ K
Sbjct: 521 NSLNQIVEE-KNKLTEEKESIKQELDSIKADNSTKELEINKINEEKNQLQNDYDTVQQEK 579
Query: 356 ENLQQAILIERERFTQMQWDMEELRRKSLEME---MKLKSE-SGGNVGQDLTK-----ES 406
EN+Q+ + + +Q + ++ +++ + ++E KL ++ + GN G LTK +
Sbjct: 580 ENIQKELNQIKIEKSQKEEELNKIKEEKQQVEDEKAKLITDIANGNDG--LTKLNEVIDK 637
Query: 407 IVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSE 466
+ + + + +L+ + + + +S ++ K+K +IK E L ++ L EL++
Sbjct: 638 LKDEKENISNELNQIKNERDNISNEFN----KTKEEIKQKENETIQLNEEKSVLLNELNQ 693
Query: 467 SI--KEKYEAEKLLLHEREKSEQAEIAWRKQL--EKCGLLLKQLQECSVEL--------P 514
K+K E EK ++ + +++E ++ K + + + + QE EL
Sbjct: 694 IKEEKQKIEDEKAVIQQEKENEITKLNEDKTVIENELNQIKTEKQEIENELNQTKDEKQK 753
Query: 515 YEDE-DRTFLQSSSSTDAFNKL-----KTSDDQIDIL--LAEVENLEKDAGSAASNVDKT 566
EDE + + S+ D +KL +T ++ ++L L +++N + A N K
Sbjct: 754 IEDEKSKLITELSNGNDGISKLNEELTQTKQEKENVLNELNQIKN--EFASFKEQNTQKE 811
Query: 567 NDIKD 571
N++KD
Sbjct: 812 NELKD 816
Score = 42.7 bits (99), Expect = 0.031
Identities = 51/277 (18%), Positives = 124/277 (44%), Gaps = 29/277 (10%)
Query: 340 VKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVG 399
+ L ++ +K K+ +Q + ++ + +Q ++EE+++ +E + K +
Sbjct: 1096 INSLNNQITQLNEKEKQMNEQVMALQTQ-LSQSNINLEEVKKDLIESQNKYTQ-----IN 1149
Query: 400 QDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVK---EVKSLRSS 456
++ K+ + Q+ + + ++ E+LE K+ +L+ K +I L K E+ SL ++
Sbjct: 1150 EE--KDCVEQERNKINEEYKTVNEELEKNKKELNDLQTKYDNEILELNKNKDELNSLINN 1207
Query: 457 QTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELP-- 514
E K L E +K+ E + L+ E ++L + ++ +E + EL
Sbjct: 1208 LKEEKTNLEEQVKKMEEEKSKLITELSNGSDGVSKLNEELTQ---TKQEKEEINNELNSI 1264
Query: 515 YEDEDRTFLQSSSSTDAFNKLKTSDDQID----ILLAEVE-------NLEKDAGSAASNV 563
E++ R + + + ++K ++I+ LL E+E L+ + + + +
Sbjct: 1265 KEEKKRIEEEKNQIINENKEIKEEKEKIEEEKKELLKEIEKEKEGNNQLQNEINTIQTRM 1324
Query: 564 DKTNDIKDGVICGDEMRKIISDLFLDNVRLRKQTNRV 600
+ + +IC + K I+ + L+K+ N++
Sbjct: 1325 KEIEEKNQEIIC--DNNKEIAKFKEEQENLQKELNQI 1359
Score = 40.8 bits (94), Expect = 0.12
Identities = 61/299 (20%), Positives = 121/299 (40%), Gaps = 47/299 (15%)
Query: 298 RNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAA----KDFLTTKVKDLEVELETSKQK 353
+N++N++ S + + E L ++N+E+ A +D L ++K +E E +++
Sbjct: 1465 QNEINKLKSENEELSNNLSFEKEGL-KQVNEEVNAIKEERDELVKQIKKIEEEKRKVEEE 1523
Query: 354 -----------------NKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGG 396
KE L Q ++ ++Q +EE+ ++ E+K K E
Sbjct: 1524 LNFNGSEVNEQIAQINNEKEQLNQECNELKQNLKELQSKIEEIEQEKESNEIKKKEEL-- 1581
Query: 397 NVGQDLTKESIVQQNDVL------------LQDLDAAREQLEILSKQYGELEAKSKADIK 444
Q+L +E + ND+ LQ+ + EQ+ +++ EL+ K +
Sbjct: 1582 ---QELQEEITEKDNDIKNLKEEIERIEKELQEKEEDMEQMSNNTEELEELKNKLTETQR 1638
Query: 445 VLVKEVKSLRSSQTELKK-------ELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLE 497
+L +E K S E ++ EL E + ++ + + E+ + K
Sbjct: 1639 LLEEEKKEKESISNEFEETKEQVLVELQRVNNEMNKMNEIKQEDENEKEELQEHINKLKS 1698
Query: 498 KCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDA 556
+ +QL+E S +L +E + S NK D+ D +L +E L D+
Sbjct: 1699 QIERENEQLKEVS-KLKWELSELKTENESMKQMIMNKKSLLDNTDDFILMSLEYLYFDS 1756
>UniRef100_UPI000042F5AA UPI000042F5AA UniRef100 entry
Length = 1880
Score = 62.4 bits (150), Expect = 4e-08
Identities = 71/309 (22%), Positives = 140/309 (44%), Gaps = 42/309 (13%)
Query: 303 RILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAI 362
R S L AK++ E+ V+ +E+ + LT+K+ +LE EL+ +Q++K+N
Sbjct: 960 RSASVAYNDLKKAKSESEEETVKAKEEL---ETLTSKIDNLEKELK--EQQSKKN----- 1009
Query: 363 LIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAR 422
E Q+Q + K E+E +LKS N + I QN L+Q L+
Sbjct: 1010 ----ELEGQLQNITDSTNEKFKELEDELKSIKKSN-------KEISSQNSELIQKLEKTE 1058
Query: 423 EQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHER 482
+ L+ ++ +L+A++K++I L E+ SL+S E ++ S + K E L + +
Sbjct: 1059 KDLQAKDEEIDKLKAETKSNIDNLNSEISSLQSKLKEAEESHSST---KDEHSSLSENLK 1115
Query: 483 EKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQ----------SSSSTDAF 532
+ E+ E + K +++ ++ + E+ + + T LQ S D
Sbjct: 1116 KLKEEYENTKTSMIAKLSAKIEEHKKATDEIETKTKHITDLQEEHAKQKSQFESERNDIK 1175
Query: 533 NKLKTSDDQIDILLAEVENLEKDA----GSAASNVDKTNDIKDGVICGDEMRKI----IS 584
+ L ++ ++ ++ NLEK+ + +K +D++ V ++ K I
Sbjct: 1176 SNLDEANKELSDNREKLSNLEKEKTELNNKLKTQEEKISDLETSVAISEDKSKSLKHDIE 1235
Query: 585 DLFLDNVRL 593
DL + ++L
Sbjct: 1236 DLKREKIKL 1244
Score = 54.3 bits (129), Expect = 1e-05
Identities = 54/244 (22%), Positives = 116/244 (47%), Gaps = 36/244 (14%)
Query: 324 VRLNQEIAAKDFLTTKVKDLEVELETSKQKNK--------ENLQQAILIERERFTQMQWD 375
++ N + A K+ + K +E E ++ NK +L+ ++ I ++ ++ D
Sbjct: 1174 IKSNLDEANKELSDNREKLSNLEKEKTELNNKLKTQEEKISDLETSVAISEDKSKSLKHD 1233
Query: 376 MEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGEL 435
+E+L+R+ +++E LK +E++ ++ +EQL++++ + EL
Sbjct: 1234 IEDLKREKIKLETTLKE----------NEETMFEK-----------KEQLQVVNDKCKEL 1272
Query: 436 EAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQ 495
EA K + KE+ L +L+ S+ E+ + L+ + +SE+ I +Q
Sbjct: 1273 EACLKKLTETKEKEINDL---IRKLEAAKSDHDTERKKLSLLIEDTKSESEKNVIKLNEQ 1329
Query: 496 LEKC-GLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKL---KTSDDQIDILLAEVEN 551
+EK G K++++ +L + D ++++ K KT+ + +D L EVEN
Sbjct: 1330 IEKLKGEREKEVRDIQSQLAAKTTDWEKIKTTLDKVLKEKSDLEKTNKESVDTLKKEVEN 1389
Query: 552 LEKD 555
L+K+
Sbjct: 1390 LKKE 1393
Score = 51.6 bits (122), Expect = 7e-05
Identities = 70/329 (21%), Positives = 135/329 (40%), Gaps = 41/329 (12%)
Query: 301 LNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELE------------ 348
L + +S ++ + T ++L +L + + D T +++ E+EL+
Sbjct: 1390 LKKEISLLEDQKKDDTTKYKELAAQLETKTSNLDSTTMELEKTELELKKVRNELTEATSE 1449
Query: 349 -TSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKS--------ESGGNVG 399
T Q N ++L + I + T+ D+E + E++ LKS E+ N
Sbjct: 1450 LTKLQDNNQSLTEEIEKTKAALTKSSKDLEVCGNQKSELQDSLKSVKSELKNFENKYNQE 1509
Query: 400 QDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTE 459
K+ I ++ ++ ++++ + K+ L S+ IK ++KSL S
Sbjct: 1510 TTSLKDEIEEKQKEIVTLQTELKDRISEVEKERAMLSENSETVIKEYSDKIKSLESKINS 1569
Query: 460 LKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDED 519
+K+ S+ I E + L + K Q + + QLE LK+L + S+E + E
Sbjct: 1570 IKENHSKEITTHNEQKTSLKQDIAKLSQDHESAQTQLEDKENQLKEL-KASLE-KHNTES 1627
Query: 520 RTFLQ---------SSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKTNDIK 570
T ++ S + +LKTS D + E + L+ S ++K
Sbjct: 1628 ATSIEEKNNQIKELSETIKSLKTELKTSGDALKQSQKEYKTLKTKNSDTESKLEKQL--- 1684
Query: 571 DGVICGDEMRKIISDLFLDNVRLRKQTNR 599
+E+ K+ SDL + +L+ T R
Sbjct: 1685 ------EELEKVKSDLQTADEKLKGITER 1707
Score = 48.1 bits (113), Expect = 7e-04
Identities = 62/242 (25%), Positives = 108/242 (44%), Gaps = 36/242 (14%)
Query: 281 TKLKSSGDAQL----------VLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEI 330
T+LK+SGDA D +KL + L +++ +T E L +EI
Sbjct: 1650 TELKTSGDALKQSQKEYKTLKTKNSDTESKLEKQLEELEKVKSDLQTADEKLKGITEREI 1709
Query: 331 AAKDFLTT-------KVKDLEVELETSKQKNKENLQQAILI-----ERERFTQMQWDMEE 378
A K L T +L +T K KE + L E E + Q D+ E
Sbjct: 1710 ALKSELETVKNSGLSTTSELAALTKTVKSLEKEKEELQFLSGNKSKELEDYIQKHSDISE 1769
Query: 379 LRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQ---YGEL 435
+ K+L E+K K++ + + LT+ L DL + +++LE Q + L
Sbjct: 1770 -KLKALTDELKEKTKQFDDSKKKLTE---------LENDLTSTKKELETEKTQTSKFKNL 1819
Query: 436 EAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQ 495
E + +I L KE++ L++ + KKELSE + K E+E +L ++ + +++ + +
Sbjct: 1820 EERKDKEIVKLNKELELLKNDNSGAKKELSEKV-SKLESEIEILSKKLEDKKSVMKQHDE 1878
Query: 496 LE 497
L+
Sbjct: 1879 LK 1880
Score = 46.6 bits (109), Expect = 0.002
Identities = 62/273 (22%), Positives = 113/273 (40%), Gaps = 55/273 (20%)
Query: 336 LTTKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESG 395
L TKV +L EL+T K+ + N ++ + +E+L S ++E KL+ +
Sbjct: 733 LDTKVLNLTKELQTEKENAESNDKE-----------LNEKIEKLTNLSTKLETKLEDKE- 780
Query: 396 GNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRS 455
Q+L +E + L++++ + A S IK KE +++
Sbjct: 781 --------------------QELAKIQEDHKSLNEKF-LVTANSLCGIKARTKESETISG 819
Query: 456 SQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPY 515
++EL E++K+ +E L +EK + E A +K + + + L
Sbjct: 820 PD---QQELQEALKKGNTSESTLKQLKEKLDSTEQAKKKLEDGINNMTRDL--------- 867
Query: 516 EDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKT-NDIKDGVI 574
F S ++A ++K + + L E EN +KD +N++K+ N+ K +
Sbjct: 868 ------FHLKKSKSEAETQIKQREREFKNLTYEFENTKKDYELQINNLNKSNNEFKQKI- 920
Query: 575 CGDEMRKIISDLFLDNVRLRKQTNRVTRHMLSN 607
+E+ K I L DN KQ R N
Sbjct: 921 --NELSKKIESLTEDNKFNAKQLEEKLRDTEEN 951
Score = 46.2 bits (108), Expect = 0.003
Identities = 56/256 (21%), Positives = 112/256 (42%), Gaps = 23/256 (8%)
Query: 314 TAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERERFTQMQ 373
T+++ ++ L +L+ AK L + ++ +L K+ E Q ERE F +
Sbjct: 834 TSESTLKQLKEKLDSTEQAKKKLEDGINNMTRDLFHLKKSKSEAETQIKQRERE-FKNLT 892
Query: 374 WDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYG 433
++ E + K E+++ ++S Q + + L + +++ E + +KQ
Sbjct: 893 YEFENTK-KDYELQINNLNKSNNEFKQKINE---------LSKKIESLTEDNKFNAKQLE 942
Query: 434 ELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWR 493
E ++ + + L+ +++S + +LKK SES +E +A++ L K + E +
Sbjct: 943 EKLRDTEENNEHLMDKLRSASVAYNDLKKAKSESEEETVKAKEELETLTSKIDNLEKELK 1002
Query: 494 KQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDI----LLAEV 549
+Q K L QLQ + D T + D +K S+ +I L+ ++
Sbjct: 1003 EQQSKKNELEGQLQNIT--------DSTNEKFKELEDELKSIKKSNKEISSQNSELIQKL 1054
Query: 550 ENLEKDAGSAASNVDK 565
E EKD + +DK
Sbjct: 1055 EKTEKDLQAKDEEIDK 1070
Score = 40.0 bits (92), Expect = 0.20
Identities = 58/279 (20%), Positives = 113/279 (39%), Gaps = 25/279 (8%)
Query: 339 KVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNV 398
++ DL +LE +K + ++ L+ + ++ + ++ +L E KLK E V
Sbjct: 1286 EINDLIRKLEAAKSDHDTERKKLSLLIEDTKSESEKNVIKLN----EQIEKLKGEREKEV 1341
Query: 399 GQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSL----- 453
I Q D + + L+ + K+ +LE +K + L KEV++L
Sbjct: 1342 ------RDIQSQLAAKTTDWEKIKTTLDKVLKEKSDLEKTNKESVDTLKKEVENLKKEIS 1395
Query: 454 -----RSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQE 508
+ T KEL+ ++ K E EK+E R +L + L +LQ+
Sbjct: 1396 LLEDQKKDDTTKYKELAAQLETKTSNLDSTTMELEKTELELKKVRNELTEATSELTKLQD 1455
Query: 509 CSVELPYEDEDRTFLQSSSSTD---AFNKLKTSDDQIDILLAEVENLEKDAGSAASNV-- 563
+ L E E + SS D N+ D + + +E++N E +++
Sbjct: 1456 NNQSLTEEIEKTKAALTKSSKDLEVCGNQKSELQDSLKSVKSELKNFENKYNQETTSLKD 1515
Query: 564 DKTNDIKDGVICGDEMRKIISDLFLDNVRLRKQTNRVTR 602
+ K+ V E++ IS++ + L + + V +
Sbjct: 1516 EIEEKQKEIVTLQTELKDRISEVEKERAMLSENSETVIK 1554
>UniRef100_P92021 Hypothetical protein T10G3.5 [Caenorhabditis elegans]
Length = 1205
Score = 62.4 bits (150), Expect = 4e-08
Identities = 80/315 (25%), Positives = 143/315 (45%), Gaps = 40/315 (12%)
Query: 285 SSGDAQLVLPLDQ----RNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIA-AKDFLTTK 339
S+G+ + ++Q + KL L T + A ++E I L +++ A+ T K
Sbjct: 502 SAGEGGAKMAIEQLEQEKVKLTNELQTSSEKTKKASGELEAKISELEKKLRDAEASRTDK 561
Query: 340 VKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKS---ESGG 396
+ + E E+ ++K E + I + ERF +M+ +MEE R+K+ + +KLK S
Sbjct: 562 EQKWKQEKESFERKLAE-AEDEIKRKGERFVEMEKEMEEERQKATDRTLKLKDALVNSEK 620
Query: 397 NV----GQDLTKESIVQQNDVLLQD----LDAAREQLEILSKQYGELEAK-SKADIKVLV 447
N+ + +E IV++ D L++ ++ A ++LE K+ ELEA S D V
Sbjct: 621 NLETIKKESEDREKIVREKDAHLEENKKRIEDAVQKLEEAEKRARELEASVSSRDTTVST 680
Query: 448 KEVKSLRSSQTELKKELSES---IKE-KYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLL 503
KE S +ELK +L+ES I+E K + EK+ EK ++ E + +K
Sbjct: 681 KE-----SELSELKGKLTESNSFIEELKVQVEKVSNEISEKQQEVENLMAEMRDKEAHWK 735
Query: 504 KQLQECSVELPYEDED-------------RTFLQSSSSTDAFNKLKTSDDQIDILLAEVE 550
+ E ++ ED + + +S + N+L + Q++ L EVE
Sbjct: 736 TKRDEFEAQMLRNQEDNEEASSTLKSVQEQLMKEKETSGEEKNQLISVKSQLEELKTEVE 795
Query: 551 NLEKDAGSAASNVDK 565
L + ++K
Sbjct: 796 RLIRSEEEKTQEIEK 810
Score = 48.9 bits (115), Expect = 4e-04
Identities = 73/278 (26%), Positives = 132/278 (47%), Gaps = 29/278 (10%)
Query: 300 KLNRILSTMQRRLVTAKTDMEDLIVRLNQEI----AAKDFLTTKVKDLEVELETSKQKNK 355
++ L ++ + A ED I + Q+I A K +T + +LE E S Q+++
Sbjct: 118 RIKEELDNIKSVMAIASEVTEDEIPYMAQQIQVLTADKGMVTRQFLELEKE---SGQQSR 174
Query: 356 ENLQQAILIERERFTQMQWDMEELRRKSLEM-EMKLKSESGGNVGQDLTKESIVQQNDVL 414
E LQQ +++ER M +L++ S+ M E+ +SESG +DL +E V ++DV+
Sbjct: 175 E-LQQ---VKQERGDLMA----KLKQMSVTMREITDESESGKVEMEDLKRELKVVKSDVV 226
Query: 415 LQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEA 474
+++ +R LE + Q S+ D+ VL E+ + + + +E IKE +
Sbjct: 227 RYEIEVSR--LEKMLDQ-----RPSEDDVNVLRTELVNAQKLMDAISQEKDIEIKEHLNS 279
Query: 475 EKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQ----ECSVELPYEDEDRTFLQSSSSTD 530
+ L EREK K++ + +KQLQ S EL +E L++ +
Sbjct: 280 IRNLSMEREKQHIVNENLEKKIGEGEETVKQLQISYDAQSEELKQRNERVVQLEARIEEN 339
Query: 531 AFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKTND 568
F +L + + L +V+ +DA SN++ +N+
Sbjct: 340 VF-ELSENKQNVKRLEDKVQE-SQDALQMLSNINGSNE 375
Score = 42.0 bits (97), Expect = 0.053
Identities = 65/318 (20%), Positives = 135/318 (42%), Gaps = 42/318 (13%)
Query: 270 SVSRETDIVGHTKLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQE 329
++S+E DI L S + + +R K + + +++++ + ++ L + + +
Sbjct: 264 AISQEKDIEIKEHLNSIRNLSM-----EREKQHIVNENLEKKIGEGEETVKQLQISYDAQ 318
Query: 330 IAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIE---RERFTQMQWDMEELRRKSLEM 386
+V LE +E + + EN Q +E +E +Q + + + E
Sbjct: 319 SEELKQRNERVVQLEARIEENVFELSENKQNVKRLEDKVQESQDALQM-LSNINGSNEEQ 377
Query: 387 EMKLKSESGGNVGQDLTKESIVQQNDVL----LQDLDAAREQLEILSKQYGELEAKSKAD 442
+ L S+ N + E++ ++ + L+ L+ A L G L K ++
Sbjct: 378 MISLNSKFERNTAERKRIEAVFEEKVTVQGERLKTLEMANLDLTNELASMGSLLDKERSL 437
Query: 443 IKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLL 502
++ KE+ SS +LK++L+ES K K ++ E E A++ +E L
Sbjct: 438 LEEKNKEISERDSSINDLKEKLAESEK------KATKYKNELKEHADL-----VENLTLQ 486
Query: 503 LKQLQECSVELPYE---------------DEDRTFLQSSSSTDAFNKLKTSDDQIDILLA 547
L +LQE S +L + ++++ L + T + K K + +++ ++
Sbjct: 487 LNKLQENSKDLMEKISAGEGGAKMAIEQLEQEKVKLTNELQTSS-EKTKKASGELEAKIS 545
Query: 548 EVENLEKDAGSAASNVDK 565
E+E +DA AS DK
Sbjct: 546 ELEKKLRDA--EASRTDK 561
>UniRef100_UPI0000437813 UPI0000437813 UniRef100 entry
Length = 1361
Score = 62.0 bits (149), Expect = 5e-08
Identities = 64/266 (24%), Positives = 121/266 (45%), Gaps = 19/266 (7%)
Query: 300 KLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNK---- 355
+L I S++ + V + EDL ++ + L+ K+K ++ LE + ++K
Sbjct: 627 ELKHIKSSLSKAEVEKRQLQEDLTTLEKEKNKQEIDLSFKLKAVQQSLEQEEAEHKTTKA 686
Query: 356 -----ENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQ 410
+ Q+I + E M+ + E R ++E +L N D + +
Sbjct: 687 RLADNNKINQSIEAKSETLKDMEHKLLEERSAKQQLENRLMQLEKENSVLDCDYKQAKHE 746
Query: 411 NDVLLQDLDAAREQLEILS---KQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSES 467
L + EQ+E+L+ +Q + + + D+KV +E+ SLRSS+ +LK+EL+
Sbjct: 747 LQELRSLKENLTEQVEVLNVRVQQETQRKTLCQGDLKVQRQEINSLRSSEQQLKQELNHL 806
Query: 468 IKEKYEAEKL---LLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQ 524
++ K EK L ERE+SE+ + QLE K + EL E +++ L
Sbjct: 807 LELKLTLEKQNQELSKEREESEKQLKEMKDQLEAEQYFTKLYKTQIRELKEESDEKVKLY 866
Query: 525 SSSSTDAFNKLKTSDDQIDILLAEVE 550
DA +++ ++ D L +++E
Sbjct: 867 K----DAQQRIEDLQEERDSLASQLE 888
Score = 35.8 bits (81), Expect = 3.8
Identities = 47/234 (20%), Positives = 98/234 (41%), Gaps = 36/234 (15%)
Query: 306 STMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIE 365
S +++ + + +++D+I R Q++A KD +++ L + E
Sbjct: 909 SDLEKEKIMKELEIKDMIARHRQDLAEKDGTINSLEESNRTLTVDVAN--------LASE 960
Query: 366 RERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQL 425
+E +++ K E E ++KS + ++ E +Q L + A +
Sbjct: 961 KEELNNKLKHIQQKLEKIKEEEKQMKSLT-------VSYEKQIQVEKTL--KIQAINKLA 1011
Query: 426 EILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKS 485
E++ K G + DI+ KE++ L+ K++L+ SI + ++RE +
Sbjct: 1012 EVMKKTDGRPHQINNTDIRRKEKEIRQLQLQLRTEKEKLNHSI---------IKYQREIN 1062
Query: 486 E-QAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTS 538
+ QA IA Q+ E + L +D D L+S ++ + + L T+
Sbjct: 1063 DMQAMIAEESQVR---------LEMQMSLDSKDSDIERLRSQLTSLSIHSLDTT 1107
>UniRef100_UPI000036865B UPI000036865B UniRef100 entry
Length = 3157
Score = 62.0 bits (149), Expect = 5e-08
Identities = 93/403 (23%), Positives = 175/403 (43%), Gaps = 79/403 (19%)
Query: 274 ETDIVGHTKLKSSGDAQLVLPLDQRNKLNRILSTMQR---RLVTAKTDMEDLIVRLNQEI 330
E ++ T+L + + + L + L L ++++ L K ++E+ I +LN+E
Sbjct: 2049 EAEVKEKTELLQTLSSDVSELLKDKTHLQEKLQSLEKDSQALSFTKCELENQIAQLNKE- 2107
Query: 331 AAKDFLTTKVKDLEVELETS--KQKNKENLQQAILIERERFT-----------QMQWDME 377
K+ L + + L+ L S ++ N +A L+E+ F Q++ +E
Sbjct: 2108 --KELLVKESESLQARLSESDYEKLNVSKALEAALVEKGEFALRLSSTQEEVHQLRRGIE 2165
Query: 378 ELR-------RKSLEMEMKLKSESGGNVG-----QDLTKESIVQQNDVLLQDLDA--ARE 423
+LR +K L + +LK N ++L +E + + + L LDA ++
Sbjct: 2166 KLRVRIEADEKKQLHVAEELKERERENDSLKDKVENLERELQMSEENQELVILDAENSKA 2225
Query: 424 QLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHER- 482
++E L Q E+ A+S +KV ++ +LRS + L K++ E + E +KLL +
Sbjct: 2226 EVETLKTQIEEM-ARS---LKVFELDLVTLRSEKENLTKQIQEKQGQLSELDKLLSSFKS 2281
Query: 483 --EKSEQAEIAWRKQ----LEKCGLLLKQLQEC----------------SVELPYEDED- 519
E+ EQAEI +++ +E LK+L E S++ P E+E
Sbjct: 2282 LLEEKEQAEIQIKEESKTAVEMLQNQLKELNEAVAALCGDQEIMKATEQSLDPPIEEEHQ 2341
Query: 520 --------RTFLQSSSSTD--AFNKLKTSDDQIDILLAEVENLEKDAGSAASNVD----- 564
R L++ +LK S+ D+L VENLE++ A +N +
Sbjct: 2342 LRNSIEKLRARLEADEKKQLCVLQQLKESEHHADLLKGRVENLERELEIARTNQEHAAHE 2401
Query: 565 ---KTNDIKDGVICGDEMRKIISDLFLDNVRLRKQTNRVTRHM 604
+++ + M + + DL LD V +R + +T +
Sbjct: 2402 AENSKGEVEALKAKIEGMTQSLRDLELDLVTIRSEKENLTNEL 2444
Score = 51.2 bits (121), Expect = 9e-05
Identities = 48/223 (21%), Positives = 104/223 (46%), Gaps = 22/223 (9%)
Query: 289 AQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEI--AAKDFLTTKVKDLEVE 346
A+L L Q ++ +L +L + K +E+ + Q++ A + F +++K+ E+
Sbjct: 419 ARLTQELQQAKNMHNVLQAELDKLTSVKQQLENNLEEFKQKLCRAEQAFQASQIKENELR 478
Query: 347 LETSKQKNKENLQQAILIERER-FTQMQWDMEELRR-----KSLEMEMKLKSESGGNVGQ 400
+ K + NL ++ ++ R ++ +++ +++ ++ EMK K+ S + +
Sbjct: 479 RSMEEMKKENNLLKSQSEQKAREVCHLEAELKNIKQCLNQSQNFAEEMKAKNTSQETMLR 538
Query: 401 DLTKESIVQQNDVLLQDLDAAREQLE--------ILSKQYGELE------AKSKADIKVL 446
DL ++ Q+N + L+ L A LE +L K+ +E +K++ + K L
Sbjct: 539 DLQEKINQQENSLTLEKLKLAVADLEKQRDCSQDLLKKREHHIEQLNDKLSKTEKESKAL 598
Query: 447 VKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAE 489
+ ++ + ELK+E + K E EKLL + E +
Sbjct: 599 LSALELKKKEYEELKEEKTLFSCWKSENEKLLTQMESEKENLQ 641
Score = 36.2 bits (82), Expect = 2.9
Identities = 67/268 (25%), Positives = 102/268 (38%), Gaps = 45/268 (16%)
Query: 337 TTKVKDLEVELETSKQ---KNKENLQQAILIERERFTQMQWDMEE----LRRKSLEMEMK 389
T+ ++ + ELE K+ K + + ER + ++E LR + + +
Sbjct: 1104 TSLCENRKNELEQLKEAFAKEHQEFLTKLAFAEERNQNLMLELETVQQALRSEMTDNQNN 1163
Query: 390 LKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKE 449
KSE GG + +T + + + DL EQL + K E + I+ VKE
Sbjct: 1164 SKSEPGGLKQEIMTLKEEQNKMQKEVNDLLQENEQLMKVMKTKHECQNLESEPIRNSVKE 1223
Query: 450 VKSLRSSQTELKKELSESIKE----KYEAE-----------KLLLHEREKSE---QAEI- 490
+S R +Q K ++ +KE Y A+ +L L E EK + Q E+
Sbjct: 1224 RESER-NQCNFKPQMDLEVKEISLDSYNAQLVQLEAMLRSKELKLQESEKEKECLQHELQ 1282
Query: 491 AWRKQLEKCGLLLKQLQE------CSV-----------ELPYEDEDRTFLQSSSSTDAFN 533
R LE L Q QE C + EL D LQ S T N
Sbjct: 1283 TIRGDLESSNLQDMQSQEISGLKDCEIDAEEKYISGPHELSTSQNDNAHLQCSLQT-TMN 1341
Query: 534 KLKTSDDQIDILLAEVENLEKDAGSAAS 561
KL + +IL AE L + + S
Sbjct: 1342 KLNELEKICEILQAEKYELVTELNDSRS 1369
Score = 34.7 bits (78), Expect = 8.4
Identities = 52/251 (20%), Positives = 107/251 (41%), Gaps = 47/251 (18%)
Query: 371 QMQWDMEELRRKSLEMEMKLKSESGGNVGQ-----------DLTKESIVQQNDVLLQDLD 419
Q++ +ELR K E+E++L+ GQ + K ++++ V L+
Sbjct: 282 QLKAQNQELRNKINELELRLQGHEKEMKGQVNKFQELQLQLEKAKVELIEKEKV----LN 337
Query: 420 AAREQLEILSKQYGELEAKSKA---DIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEK 476
R++L + QY + K A +K L +++ R + + L + IKEK + +
Sbjct: 338 KCRDELVRTTAQYDQASTKYTALEQKLKKLTEDLSCQRQNAESARCSLEQKIKEKEKEFQ 397
Query: 477 LLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLK 536
L +++S Q L+ QEC + + R + + + N L+
Sbjct: 398 EELSRQQRSFQT-------LD---------QEC-----IQMKARLTQELQQAKNMHNVLQ 436
Query: 537 TSDDQIDILLAEVE-NLE--KDAGSAASNVDKTNDIKDGVICGDEMRKIISDLFLDNVRL 593
D++ + ++E NLE K A + + IK+ +E+R+ + ++ +N L
Sbjct: 437 AELDKLTSVKQQLENNLEEFKQKLCRAEQAFQASQIKE-----NELRRSMEEMKKENNLL 491
Query: 594 RKQTNRVTRHM 604
+ Q+ + R +
Sbjct: 492 KSQSEQKAREV 502
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.310 0.129 0.356
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 986,146,669
Number of Sequences: 2790947
Number of extensions: 43038621
Number of successful extensions: 191009
Number of sequences better than 10.0: 8145
Number of HSP's better than 10.0 without gapping: 252
Number of HSP's successfully gapped in prelim test: 8160
Number of HSP's that attempted gapping in prelim test: 167158
Number of HSP's gapped (non-prelim): 21788
length of query: 609
length of database: 848,049,833
effective HSP length: 133
effective length of query: 476
effective length of database: 476,853,882
effective search space: 226982447832
effective search space used: 226982447832
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 78 (34.7 bits)
Medicago: description of AC148405.4


