

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148405.1 + phase: 0
(149 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
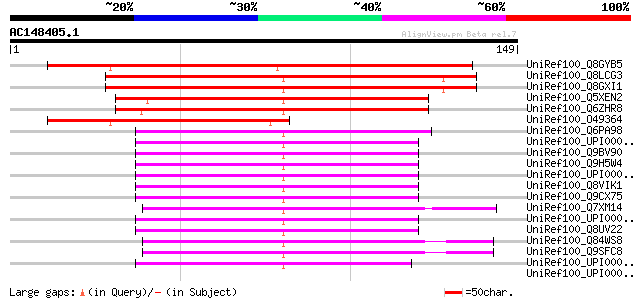
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8GYB5 Hypothetical protein At4g32270/F10M6_90 [Arabid... 144 5e-34
UniRef100_Q8LCG3 Hypothetical protein [Arabidopsis thaliana] 119 2e-26
UniRef100_Q8GXI1 Hypothetical protein At5g25340/F18G18_80 [Arabi... 118 2e-26
UniRef100_Q5XEN2 At1g80060 [Arabidopsis thaliana] 93 1e-18
UniRef100_Q6ZHR8 Hypothetical protein OJ1145_F01.17 [Oryza sativa] 80 9e-15
UniRef100_O49364 Hypothetical protein F10M6.90 [Arabidopsis thal... 70 1e-11
UniRef100_Q6PA98 MGC68520 protein [Xenopus laevis] 61 4e-09
UniRef100_UPI0000360C06 UPI0000360C06 UniRef100 entry 60 1e-08
UniRef100_Q9BV90 U11/U12 snRNP 25K protein [Homo sapiens] 60 1e-08
UniRef100_Q9H5W4 Hypothetical protein FLJ22940 [Homo sapiens] 60 1e-08
UniRef100_UPI00001D0334 UPI00001D0334 UniRef100 entry 59 2e-08
UniRef100_Q8VIK1 C16ORF33 [Mus musculus] 59 2e-08
UniRef100_Q9CX75 Mus musculus 17 days embryo head cDNA, RIKEN fu... 59 2e-08
UniRef100_Q7XM14 OSJNBa0084K01.17 protein [Oryza sativa] 58 5e-08
UniRef100_UPI00002DAC25 UPI00002DAC25 UniRef100 entry 57 6e-08
UniRef100_Q8UV22 C16ORF33 [Sphoeroides nephelus] 57 8e-08
UniRef100_Q84WS8 Hypothetical protein At3g07860 [Arabidopsis tha... 55 2e-07
UniRef100_Q9SFC8 F17A17.20 protein [Arabidopsis thaliana] 55 2e-07
UniRef100_UPI0000247CD5 UPI0000247CD5 UniRef100 entry 54 5e-07
UniRef100_UPI00003633B9 UPI00003633B9 UniRef100 entry 33 1.3
>UniRef100_Q8GYB5 Hypothetical protein At4g32270/F10M6_90 [Arabidopsis thaliana]
Length = 239
Score = 144 bits (362), Expect = 5e-34
Identities = 73/134 (54%), Positives = 94/134 (69%), Gaps = 9/134 (6%)
Query: 12 PRRSLSIQLSSMIIVDC-----TLPYDKLPSQNLTLTVLKLDSSSFQVEVAKTATVAVLK 66
PRRSL+ LS + ++D + Y+++P + + LTVLKLD SSF ++V KTATV LK
Sbjct: 5 PRRSLAASLSPLRLIDGLPRRRSFNYNQMPEEPIKLTVLKLDGSSFGIQVLKTATVGELK 64
Query: 67 QAVEAAFRHKP----EKISWPLVWGQFCLCYEGQKLVTETDYLRDYGIKDGDQLRFIRHV 122
AVEAAF H P KISWP VWGQFCL YE ++L+ E++YL ++GIKDGDQLRFIRH+
Sbjct: 65 MAVEAAFSHLPISGPGKISWPHVWGQFCLSYEDKRLINESEYLTEFGIKDGDQLRFIRHI 124
Query: 123 SNVCNFQRKPKKRT 136
SN C K K +T
Sbjct: 125 SNYCMLMVKHKSKT 138
>UniRef100_Q8LCG3 Hypothetical protein [Arabidopsis thaliana]
Length = 208
Score = 119 bits (297), Expect = 2e-26
Identities = 60/117 (51%), Positives = 79/117 (67%), Gaps = 8/117 (6%)
Query: 29 TLPYDKLPSQNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK----ISWPL 84
T YDKLP++ + L+VLKLD SSF V V +ATV LK A+E AF H P+K ISW
Sbjct: 8 TFSYDKLPNEPIRLSVLKLDGSSFDVYVLTSATVGDLKVAIETAFSHVPKKGPSKISWSH 67
Query: 85 VWGQFCLCYEGQKLVTETDYLRDYGIKDGDQLRFIRHVSNVC----NFQRKPKKRTV 137
VWG FCLC+ GQKL+T+TD + +YG+KDGD++RF HVS + RK K++ +
Sbjct: 68 VWGHFCLCFGGQKLITDTDCIGNYGMKDGDEVRFKNHVSGNAELSKGYSRKSKRKNL 124
>UniRef100_Q8GXI1 Hypothetical protein At5g25340/F18G18_80 [Arabidopsis thaliana]
Length = 208
Score = 118 bits (296), Expect = 2e-26
Identities = 60/117 (51%), Positives = 79/117 (67%), Gaps = 8/117 (6%)
Query: 29 TLPYDKLPSQNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK----ISWPL 84
T YDKLP++ + L+VLKLD SSF V V +ATV LK A+E AF H P+K ISW
Sbjct: 8 TFSYDKLPNEPIRLSVLKLDGSSFDVYVLTSATVGDLKVAIETAFSHVPKKGPSKISWSH 67
Query: 85 VWGQFCLCYEGQKLVTETDYLRDYGIKDGDQLRFIRHVSNVC----NFQRKPKKRTV 137
VWG FCLC+ GQKL+T+TD + +YG+KDGD++RF HVS + RK K++ +
Sbjct: 68 VWGHFCLCFGGQKLITDTDCIGNYGMKDGDEVRFKNHVSGNAVLSKGYSRKSKQKNL 124
>UniRef100_Q5XEN2 At1g80060 [Arabidopsis thaliana]
Length = 243
Score = 93.2 bits (230), Expect = 1e-18
Identities = 50/97 (51%), Positives = 63/97 (64%), Gaps = 5/97 (5%)
Query: 32 YDKLPSQN-LTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK----ISWPLVW 86
Y KLP Q + L+V+KL+ S F VEVAK +VA LK+AVE F P + ISW VW
Sbjct: 44 YLKLPPQGRIKLSVVKLNGSLFDVEVAKDCSVAELKRAVEQVFTISPLEGHGMISWSHVW 103
Query: 87 GQFCLCYEGQKLVTETDYLRDYGIKDGDQLRFIRHVS 123
G FCLCY Q+LV + +R G+ DGDQL F+RH+S
Sbjct: 104 GHFCLCYRDQRLVNDKTSIRYLGLNDGDQLHFVRHLS 140
>UniRef100_Q6ZHR8 Hypothetical protein OJ1145_F01.17 [Oryza sativa]
Length = 324
Score = 80.1 bits (196), Expect = 9e-15
Identities = 42/98 (42%), Positives = 61/98 (61%), Gaps = 6/98 (6%)
Query: 32 YDKLPS-QNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK-----ISWPLV 85
Y +LP + L LTV KLD S F V++A++A V LK A+E F ++ ISW V
Sbjct: 105 YRRLPEPRRLRLTVRKLDDSYFDVQIARSAAVWELKAAIEDVFAALYDETDNKAISWQHV 164
Query: 86 WGQFCLCYEGQKLVTETDYLRDYGIKDGDQLRFIRHVS 123
W FCLC++ +KL + LR +GI+DGD++ F +H+S
Sbjct: 165 WSHFCLCFKDEKLTDDKATLRAFGIRDGDEVHFAQHLS 202
>UniRef100_O49364 Hypothetical protein F10M6.90 [Arabidopsis thaliana]
Length = 364
Score = 69.7 bits (169), Expect = 1e-11
Identities = 40/80 (50%), Positives = 52/80 (65%), Gaps = 9/80 (11%)
Query: 12 PRRSLSIQLSSMIIVDC-----TLPYDKLPSQNLTLTVLKLDSSSFQVEVAKTATVAVLK 66
PRRSL+ LS + ++D + Y+++P + + LTVLKLD SSF ++V KTATV LK
Sbjct: 195 PRRSLAASLSPLRLIDGLPRRRSFNYNQMPEEPIKLTVLKLDGSSFGIQVLKTATVGELK 254
Query: 67 QAVEAAFRH----KPEKISW 82
AVEAAF H P KISW
Sbjct: 255 MAVEAAFSHLPISGPGKISW 274
>UniRef100_Q6PA98 MGC68520 protein [Xenopus laevis]
Length = 168
Score = 61.2 bits (147), Expect = 4e-09
Identities = 34/93 (36%), Positives = 52/93 (55%), Gaps = 6/93 (6%)
Query: 38 QNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK------ISWPLVWGQFCL 91
Q +T+ V K D V V + ATV LK+A++ + K ++ ISW VW + L
Sbjct: 75 QAMTVRVCKEDEEIMPVVVVQNATVLDLKRAIQRYIQLKHQREGGIQHISWKYVWRTYHL 134
Query: 92 CYEGQKLVTETDYLRDYGIKDGDQLRFIRHVSN 124
Y G+KL E LR+YGIK+ D++ F++ + N
Sbjct: 135 SYSGEKLDDEAKSLREYGIKNRDEVVFVKKLKN 167
>UniRef100_UPI0000360C06 UPI0000360C06 UniRef100 entry
Length = 140
Score = 59.7 bits (143), Expect = 1e-08
Identities = 32/89 (35%), Positives = 50/89 (55%), Gaps = 6/89 (6%)
Query: 38 QNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK------ISWPLVWGQFCL 91
Q +T+ VLK D + V + ATV LK+A+ K ++ +SW VW F L
Sbjct: 47 QAMTVKVLKADGEVMPIVVVQNATVLDLKKAIRRFMELKQQREGGVKCVSWRYVWRTFNL 106
Query: 92 CYEGQKLVTETDYLRDYGIKDGDQLRFIR 120
++G+KL + LRDYGIK+ D++ F++
Sbjct: 107 VFQGEKLEDDRMRLRDYGIKNRDEVTFMK 135
>UniRef100_Q9BV90 U11/U12 snRNP 25K protein [Homo sapiens]
Length = 132
Score = 59.7 bits (143), Expect = 1e-08
Identities = 33/89 (37%), Positives = 51/89 (57%), Gaps = 6/89 (6%)
Query: 38 QNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK------ISWPLVWGQFCL 91
Q +T+ V K+D V V ++ATV LK+A++ + K E+ ISW VW + L
Sbjct: 39 QAMTVRVCKMDGEVMPVVVVQSATVLDLKKAIQRYVQLKQEREGGIQHISWSYVWRTYHL 98
Query: 92 CYEGQKLVTETDYLRDYGIKDGDQLRFIR 120
G+KL + LRDYGI++ D++ FI+
Sbjct: 99 TSAGEKLTEDRKKLRDYGIRNRDEVSFIK 127
>UniRef100_Q9H5W4 Hypothetical protein FLJ22940 [Homo sapiens]
Length = 132
Score = 59.7 bits (143), Expect = 1e-08
Identities = 33/89 (37%), Positives = 51/89 (57%), Gaps = 6/89 (6%)
Query: 38 QNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK------ISWPLVWGQFCL 91
Q +T+ V K+D V V ++ATV LK+A++ + K E+ ISW VW + L
Sbjct: 39 QAMTVRVCKMDGEVMPVVVVQSATVLDLKKAIQRYVQLKQEREGGIQHISWSYVWRTYHL 98
Query: 92 CYEGQKLVTETDYLRDYGIKDGDQLRFIR 120
G+KL + LRDYGI++ D++ FI+
Sbjct: 99 TSAGEKLAEDRKKLRDYGIRNRDEVSFIK 127
>UniRef100_UPI00001D0334 UPI00001D0334 UniRef100 entry
Length = 182
Score = 59.3 bits (142), Expect = 2e-08
Identities = 33/89 (37%), Positives = 50/89 (56%), Gaps = 6/89 (6%)
Query: 38 QNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK------ISWPLVWGQFCL 91
Q +T+ V K+D V V + ATV LK+A++ + K E+ ISW VW + L
Sbjct: 89 QAMTVRVCKMDGEVMPVVVVQNATVLDLKKAIQRYVQLKQEREGGVQHISWSYVWRTYHL 148
Query: 92 CYEGQKLVTETDYLRDYGIKDGDQLRFIR 120
G+KL + LRDYGI++ D++ FI+
Sbjct: 149 TSAGEKLTEDRKKLRDYGIRNRDEVSFIK 177
>UniRef100_Q8VIK1 C16ORF33 [Mus musculus]
Length = 123
Score = 59.3 bits (142), Expect = 2e-08
Identities = 33/89 (37%), Positives = 50/89 (56%), Gaps = 6/89 (6%)
Query: 38 QNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK------ISWPLVWGQFCL 91
Q +T+ V K+D V V + ATV LK+A++ + K E+ ISW VW + L
Sbjct: 30 QAMTVRVCKMDGEVMPVVVVQNATVLDLKKAIQRYVQLKQEREGGVQHISWSYVWRTYHL 89
Query: 92 CYEGQKLVTETDYLRDYGIKDGDQLRFIR 120
G+KL + LRDYGI++ D++ FI+
Sbjct: 90 TSAGEKLTEDRKKLRDYGIRNRDEVSFIK 118
>UniRef100_Q9CX75 Mus musculus 17 days embryo head cDNA, RIKEN full-length enriched
library, clone:3300001G02 product:POLYMERASE (RNA) III
(DNA DIRECTED) POLYPEPTIDE K (12.3 kDa) (MINUS-99
PROTEIN) (HYPOTHETICAL 15.3 kDa PROTEIN) homolog [Mus
musculus]
Length = 123
Score = 59.3 bits (142), Expect = 2e-08
Identities = 33/89 (37%), Positives = 50/89 (56%), Gaps = 6/89 (6%)
Query: 38 QNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK------ISWPLVWGQFCL 91
Q +T+ V K+D V V + ATV LK+A++ + K E+ ISW VW + L
Sbjct: 30 QAMTVRVCKMDGEVMPVVVVQNATVLDLKKAIQRYVQLKQEREGGVQHISWSYVWRTYHL 89
Query: 92 CYEGQKLVTETDYLRDYGIKDGDQLRFIR 120
G+KL + LRDYGI++ D++ FI+
Sbjct: 90 TSAGEKLTEDRKKLRDYGIRNRDEVSFIK 118
>UniRef100_Q7XM14 OSJNBa0084K01.17 protein [Oryza sativa]
Length = 187
Score = 57.8 bits (138), Expect = 5e-08
Identities = 35/110 (31%), Positives = 55/110 (49%), Gaps = 8/110 (7%)
Query: 40 LTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK------ISWPLVWGQFCLCY 93
+ LTV+KLD +SF V + TATV LK A+ ++ ISW +W +CL +
Sbjct: 75 MRLTVVKLDGTSFDVAMLNTATVKDLKMAIRKKTDEIEQEKMGHRHISWKHIWDNYCLTH 134
Query: 94 EGQKLVTETDYLRDYGIKDGDQLRFIRHVSNVCNFQRKPKKRTVNLKQHG 143
+ +KL+ + L GI + ++ F HV + RK +R + HG
Sbjct: 135 QNEKLIDDNSVLSSNGICNNSKVYFSPHV--MSRVYRKHSRRRKHRFFHG 182
>UniRef100_UPI00002DAC25 UPI00002DAC25 UniRef100 entry
Length = 188
Score = 57.4 bits (137), Expect = 6e-08
Identities = 30/89 (33%), Positives = 50/89 (55%), Gaps = 6/89 (6%)
Query: 38 QNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK------ISWPLVWGQFCL 91
Q +T+ VLK D + V + ATV LK+A+ K ++ +SW VW + L
Sbjct: 95 QAMTVRVLKADGEVMPIVVVQNATVLDLKKAIRRFMELKQQREGGVKYVSWRYVWRTYHL 154
Query: 92 CYEGQKLVTETDYLRDYGIKDGDQLRFIR 120
++G+KL + L+DYGIK+ D++ F++
Sbjct: 155 VFQGEKLEDDRMRLKDYGIKNRDEVTFMK 183
>UniRef100_Q8UV22 C16ORF33 [Sphoeroides nephelus]
Length = 109
Score = 57.0 bits (136), Expect = 8e-08
Identities = 30/89 (33%), Positives = 50/89 (55%), Gaps = 6/89 (6%)
Query: 38 QNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK------ISWPLVWGQFCL 91
Q +T+ VLK D + V + ATV LK+A+ K ++ +SW VW + L
Sbjct: 16 QAMTVRVLKADGEVMPIVVVQNATVLDLKKAIRRFMELKQQREGGVKYVSWRYVWRTYHL 75
Query: 92 CYEGQKLVTETDYLRDYGIKDGDQLRFIR 120
++G+KL + L+DYGIK+ D++ F++
Sbjct: 76 VFQGEKLEGDRMRLKDYGIKNRDEVTFMK 104
>UniRef100_Q84WS8 Hypothetical protein At3g07860 [Arabidopsis thaliana]
Length = 165
Score = 55.5 bits (132), Expect = 2e-07
Identities = 34/109 (31%), Positives = 54/109 (49%), Gaps = 12/109 (11%)
Query: 40 LTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK------ISWPLVWGQFCLCY 93
+ L+V+KLD SS V V +AT+ LK ++ + ISW VW FCL
Sbjct: 52 MRLSVVKLDGSSLDVAVMNSATLKDLKLLIKKKVNEMEQANMGHRHISWKHVWSNFCLSC 111
Query: 94 EGQKLVTETDYLRDYGIKDGDQLRFIRHVSNVCNFQRKPKKRTVNLKQH 142
+KL+ + L+D GI++ Q+ F+ +V +K + R K+H
Sbjct: 112 NNEKLLDDNAVLQDVGIRNNSQVTFMPYV------MKKGRGRHSKRKKH 154
>UniRef100_Q9SFC8 F17A17.20 protein [Arabidopsis thaliana]
Length = 232
Score = 55.5 bits (132), Expect = 2e-07
Identities = 34/109 (31%), Positives = 54/109 (49%), Gaps = 12/109 (11%)
Query: 40 LTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK------ISWPLVWGQFCLCY 93
+ L+V+KLD SS V V +AT+ LK ++ + ISW VW FCL
Sbjct: 52 MRLSVVKLDGSSLDVAVMNSATLKDLKLLIKKKVNEMEQANMGHRHISWKHVWSNFCLSC 111
Query: 94 EGQKLVTETDYLRDYGIKDGDQLRFIRHVSNVCNFQRKPKKRTVNLKQH 142
+KL+ + L+D GI++ Q+ F+ +V +K + R K+H
Sbjct: 112 NNEKLLDDNAVLQDVGIRNNSQVTFMPYV------MKKGRGRHSKRKKH 154
>UniRef100_UPI0000247CD5 UPI0000247CD5 UniRef100 entry
Length = 145
Score = 54.3 bits (129), Expect = 5e-07
Identities = 28/87 (32%), Positives = 47/87 (53%), Gaps = 6/87 (6%)
Query: 38 QNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK------ISWPLVWGQFCL 91
Q +T+ V K D + V ++ATV LK+A+ K ++ +SW VW F L
Sbjct: 52 QAMTVRVCKADGEVMPIVVVQSATVLDLKKAIRRYMELKQQREGGVKHVSWKYVWRTFHL 111
Query: 92 CYEGQKLVTETDYLRDYGIKDGDQLRF 118
+ G+KL + L+DYG+++ D++ F
Sbjct: 112 VFNGEKLEDDRRKLKDYGVRNRDEVTF 138
>UniRef100_UPI00003633B9 UPI00003633B9 UniRef100 entry
Length = 538
Score = 33.1 bits (74), Expect = 1.3
Identities = 20/83 (24%), Positives = 40/83 (48%), Gaps = 10/83 (12%)
Query: 45 LKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEKISWPLVWGQFCLCYEGQKLVTETDY 104
+K ++ +A+ A+V+ K+ + F+ K + Q L + G K++ + D
Sbjct: 5 VKTPKDKEEIAIAEDASVSQFKEEISRRFKAKQD---------QLVLIFAG-KILKDGDS 54
Query: 105 LRDYGIKDGDQLRFIRHVSNVCN 127
L +GIKDG + + ++ CN
Sbjct: 55 LSQHGIKDGLTVHLVIKTAHNCN 77
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.133 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 218,464,101
Number of Sequences: 2790947
Number of extensions: 7602868
Number of successful extensions: 22446
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 25
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 22405
Number of HSP's gapped (non-prelim): 38
length of query: 149
length of database: 848,049,833
effective HSP length: 125
effective length of query: 24
effective length of database: 499,181,458
effective search space: 11980354992
effective search space used: 11980354992
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC148405.1


