

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148404.10 + phase: 0
(400 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
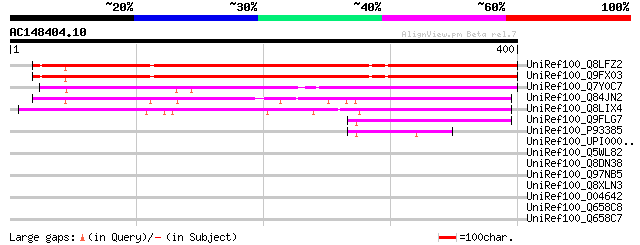
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8LFZ2 Hypothetical protein [Arabidopsis thaliana] 288 2e-76
UniRef100_Q9FX03 T3F24.7 protein [Arabidopsis thaliana] 286 9e-76
UniRef100_Q7Y0C7 Hypothetical protein OSJNBa0079B15.24 [Oryza sa... 192 1e-47
UniRef100_Q84JN2 Hypothetical protein At5g64510 [Arabidopsis tha... 114 6e-24
UniRef100_Q8LIX4 Hypothetical protein OSJNBb0053G03.20 [Oryza sa... 101 3e-20
UniRef100_Q9FLG7 Arabidopsis thaliana genomic DNA, chromosome 5,... 64 6e-09
UniRef100_P93385 Nicotiana tabacum ORF [Nicotiana tabacum] 48 6e-04
UniRef100_UPI0000318DC0 UPI0000318DC0 UniRef100 entry 40 0.15
UniRef100_Q5WL82 Succinate-semialdehyde dehydrogenase [Bacillus ... 37 0.75
UniRef100_Q8DN38 Choline-binding protein [Streptococcus pneumoniae] 37 0.99
UniRef100_Q97NB5 Choline binding protein PcpA [Streptococcus pne... 37 0.99
UniRef100_Q8XLN3 Hypothetical protein CPE1008 [Clostridium perfr... 35 3.7
UniRef100_O04642 Hypothetical protein F2P16.15 [Arabidopsis thal... 35 3.7
UniRef100_Q658C8 Hypothetical protein P0436E04.12-1 [Oryza sativa] 34 8.3
UniRef100_Q658C7 Hypothetical protein P0436E04.12-2 [Oryza sativa] 34 8.3
>UniRef100_Q8LFZ2 Hypothetical protein [Arabidopsis thaliana]
Length = 395
Score = 288 bits (737), Expect = 2e-76
Identities = 167/387 (43%), Positives = 240/387 (61%), Gaps = 11/387 (2%)
Query: 19 FLQLFTAFSNASSSSSSTHSNITH--IFQDILKAISSRQKWDLNDVRVFNFDVAKIRFGT 76
F+Q+ T + A S SNIT I QD+LK IS +QKW+L +VR +V KIR GT
Sbjct: 15 FIQVLT-LAVALDPSQPDESNITATPILQDVLKEISVKQKWNLEEVRFSKLEVKKIRIGT 73
Query: 77 SQNYLFRIGSSKNNFTVKFSDEISSWNHNKFTTTPKPDLASLVDQLSSIAFLDY-IKLEG 135
S+ + RI K+ F F DE++ W + +L LV +++S LD + L+G
Sbjct: 74 SRRFEIRIRLGKSRFVFIFPDEVTDWRRSGGGRDV--ELQELVREVNSSKVLDPPLVLKG 131
Query: 136 PFELRVHESHHLSLSLPMNVSYNGLKHIIVGKGITVEVRRAREISFYYQSDLDLQRNGSV 195
PFELRV LSLSLPMN+S++GLK ++V +GI+VE+R A+ +S ++ S
Sbjct: 132 PFELRVDGDDRLSLSLPMNISHSGLKRVLVSEGISVEIREAQAVSLFHSSHRRYAATVDP 191
Query: 196 ICSNQKNEFWPFLQSMCVPLIPIRIIGSASLIAYVARNPYVQIGTALISEDAVELLPEKC 255
+ Q + W F S+CVPL PI+IIGSASL+A+ N QI T+ +S++A+ L EKC
Sbjct: 192 VNIKQGSSLWSFWGSVCVPLPPIQIIGSASLVAFRTSNATTQIKTSYLSDEAIHLYAEKC 251
Query: 256 YHGC-VFRKQACPVASLNLRLILLEKILRSLLGHKILQDRLSGLIKANIKAYAGVKFPLE 314
Y+ +R+ P L L++ LEK+L S LG+ Q S + A +KA V+F LE
Sbjct: 252 YYKAHTYRQHRFPNDLLGLKIHKLEKVLNS-LGNGTRQTVSS--VTAKLKASGMVRFQLE 308
Query: 315 LERDVG-NNATLSTLPDWRTRPSVERVWFEVMARVEDSRLKPLSIKKVKPFIESDSVSWA 373
+ER +G N + +S WRT+P +ERVWFEV A++E +LK + ++KV PFIE D+ +W+
Sbjct: 309 IERSIGKNESVISKKVAWRTKPKIERVWFEVTAKIEGDKLKAVRLRKVVPFIEVDTEAWS 368
Query: 374 NLMSNLSYTKLRPVLLPPEALTLDVKW 400
+LMSN+S+TK +L+P EALTLDVKW
Sbjct: 369 SLMSNMSFTKFPSLLVPQEALTLDVKW 395
>UniRef100_Q9FX03 T3F24.7 protein [Arabidopsis thaliana]
Length = 395
Score = 286 bits (731), Expect = 9e-76
Identities = 166/387 (42%), Positives = 241/387 (61%), Gaps = 11/387 (2%)
Query: 19 FLQLFTAFSNASSSSSSTHSNITH--IFQDILKAISSRQKWDLNDVRVFNFDVAKIRFGT 76
F+Q+ T + A S SNIT I QD+LK IS +QKW+L +VR +V KIR GT
Sbjct: 15 FIQVLT-LAVALDPSQPDESNITATPILQDVLKEISVKQKWNLEEVRFSKLEVKKIRIGT 73
Query: 77 SQNYLFRIGSSKNNFTVKFSDEISSWNHNKFTTTPKPDLASLVDQLSSIAFLDY-IKLEG 135
S+ + RI K+ F F DEI+ W + + +L LV +++S LD + L+G
Sbjct: 74 SRRFEIRIRLGKSRFVFIFPDEITDWRRSGGGSDV--ELQELVREVNSSKVLDPPLVLKG 131
Query: 136 PFELRVHESHHLSLSLPMNVSYNGLKHIIVGKGITVEVRRAREISFYYQSDLDLQRNGSV 195
PFEL V + LSLSLPMN+S++GLK ++V +GI+VE+R A+ +S ++ S
Sbjct: 132 PFELLVDGNDRLSLSLPMNISHSGLKRVLVSEGISVEIREAQAVSLFHSSHRRYAATVDP 191
Query: 196 ICSNQKNEFWPFLQSMCVPLIPIRIIGSASLIAYVARNPYVQIGTALISEDAVELLPEKC 255
+ + + W F S+CVPL PI+IIGSASL+A+ N QI T+ +S++A+ L EKC
Sbjct: 192 VNIKEGSSLWSFWGSVCVPLPPIQIIGSASLVAFRTSNATTQIKTSYLSDEAIHLYAEKC 251
Query: 256 YHGC-VFRKQACPVASLNLRLILLEKILRSLLGHKILQDRLSGLIKANIKAYAGVKFPLE 314
Y+ +R+ P L L++ LEK+L S LG+ Q S + A +KA V+F LE
Sbjct: 252 YYKAHTYRQHRFPNDLLGLKIHKLEKVLNS-LGNGTRQTVSS--VTAKLKASGMVRFQLE 308
Query: 315 LERDVG-NNATLSTLPDWRTRPSVERVWFEVMARVEDSRLKPLSIKKVKPFIESDSVSWA 373
+ER +G N + +S WRT+P +ERVWFEV A++E +LK + ++KV PFIE D+ +W+
Sbjct: 309 IERSIGKNESVISKKVAWRTKPKIERVWFEVTAKIEGDKLKAVRLRKVVPFIEVDTEAWS 368
Query: 374 NLMSNLSYTKLRPVLLPPEALTLDVKW 400
+LMSN+S+TK +L+P EALTLDVKW
Sbjct: 369 SLMSNMSFTKFPSLLVPQEALTLDVKW 395
>UniRef100_Q7Y0C7 Hypothetical protein OSJNBa0079B15.24 [Oryza sativa]
Length = 425
Score = 192 bits (489), Expect = 1e-47
Identities = 132/399 (33%), Positives = 206/399 (51%), Gaps = 27/399 (6%)
Query: 24 TAFSNASSSSSSTHSNITHI------FQDILKAISSRQKWDLN-DVRVFNFDVAKIRFGT 76
TA SNAS S+ S + ++++ A+S + WD + +VRV+ D R G
Sbjct: 32 TASSNASLSTFSDSDPEPELAREPTFLEEVIDAVSEKYDWDPDAEVRVWPLDADAARVGA 91
Query: 77 SQNYLFRIGSSKNNFTVKFSDEISSWNHNKFTTTPKPDLASLVDQLSSIAFLDY------ 130
Q Y FR + + V+ +DE W H + D +D + L +
Sbjct: 92 VQRYEFRARAGSASALVRLADESVEWRHPAAPAVEEVDGPDGLDVVPGDGALGFRPGVRD 151
Query: 131 IKLEGPFELRVH---ESHHLSLSLPM-NVSYNGLKHIIVGKGITVEVRRAREISFYYQSD 186
+ L GP E+RV + + L LP N +Y GLK +IV G+ ++V A+++ F +
Sbjct: 152 VDLVGPVEVRVASGGDGGSIELQLPSRNATYAGLKRVIVAAGVALKVIGAQKVIFTHPHS 211
Query: 187 LDLQRNGSVICSNQK-NEFWPFLQSMCVPLIPIRIIGSASLIAYVARNPYVQIGTALISE 245
+ L NGS++ SN + WP + C P++ + ++GS ++ N +G S
Sbjct: 212 IGLLTNGSLLASNNDPSRIWPLSYATCAPILQVSVVGSVMIVV----NESNVLGRRR-SH 266
Query: 246 DAVELLPEKCYHGCVFRK-QACPVASLNLRLILLEKILRSLLGHKILQDRLSGLIKANIK 304
D VELL EKC R C S++ RL L+KIL++ +K + I+A +
Sbjct: 267 DTVELLSEKCEVDVANRLISVCVFCSISSRLPRLDKILKTWFSNKTQDSKSMQFIQAKVT 326
Query: 305 AYAGVKFPLELERDVGNNATL-STLPDWRTRPSVERVWFEVMARVEDS-RLKPLSIKKVK 362
+ +KF LELERD+ + + W+T P V+RV +V+A+VE+ RLK +S+KKVK
Sbjct: 327 SIPLIKFRLELERDITEEDGIWENISKWKTVPMVQRVALDVVAKVEEEGRLKAMSVKKVK 386
Query: 363 -PFIESDSVSWANLMSNLSYTKLRPVLLPPEALTLDVKW 400
P+ D+ SW++L SN+S+TK +LPPE LTLDVKW
Sbjct: 387 KPYPVVDASSWSSLTSNISFTKFMSFVLPPEPLTLDVKW 425
>UniRef100_Q84JN2 Hypothetical protein At5g64510 [Arabidopsis thaliana]
Length = 424
Score = 114 bits (284), Expect = 6e-24
Identities = 104/419 (24%), Positives = 188/419 (44%), Gaps = 48/419 (11%)
Query: 19 FLQLFTAFSNASSSSSSTHSNITH--IFQDILKAISSRQKWDLNDVRVFNFDVAKIRFGT 76
FL LFT F ++S+ SS +N + D+ ++I + +V+V FD+ G
Sbjct: 13 FLILFTIFPSSSALISSPDANPPYPKAISDLKESIVKGLGFQSEEVKVSGFDIRDALVGH 72
Query: 77 SQNYLFRIGSSKNNFTVKFSDEISSWNHNKFTT--TPKPDLASLVDQLSSIAFLDYI--- 131
S +Y F + K ++++ W + +P LV + D +
Sbjct: 73 SVSYEFDLEIDNKVLPFKLLEDVNRWEYVDLPIFQVEQPSENGLVPMRNKKTSSDDVLPV 132
Query: 132 ----KLEGPFELRVHESHHLSLSLPMNVSYNGLKHIIVGKGITVEVRRAREISFYYQSDL 187
+L GP EL + +++++ LSLP +V LK +I+ G V V+ AR +S + DL
Sbjct: 133 LAPFQLSGPMELWIQDANNMRLSLPYDVDAGVLKKVILADGAVVTVKGARSVSLRHPIDL 192
Query: 188 DLQRNGSVICSNQKNEFWPFLQSMC-----------VPLIPIRIIGSASLIAYVARNPYV 236
L N S NEF L S+ P++ +RI+G SL A +++P
Sbjct: 193 PLPLNQS------SNEFASGLLSLAEQLRRASTDQESPVLSLRIVGPTSL-ASTSQSPDN 245
Query: 237 QIGTALISEDAVEL------------LPEKCYHGCVFRKQ---ACPVASL---NLRLILL 278
++ ++ VEL + + ++ P+ S+ N L+
Sbjct: 246 KLKLKRLAPGLVELSSMSKDKRSLSTIGANAMTTVLTPREFTTMWPITSINGSNANLLGF 305
Query: 279 EKILRSLLGHKILQDRLSGLIKANIKAYAGVKFPLELERDVGN-NATLSTLPDWRTRPSV 337
EK+L S+LG K + ++KA + A +K +E+ + + + P+WRT+P
Sbjct: 306 EKLLTSVLGPKAQEKGSFKVLKAKVAAQTFMKIGFGIEKKLKEADVEGLSFPEWRTKPET 365
Query: 338 ERVWFEVMARVEDSRLKPLSIKKVKPFIESDSVSWANLMSNLSYTKLRPVLLPPEALTL 396
R+ FEV+A+V+ + P ++ +V P D+++ + N++ +KL + PP TL
Sbjct: 366 MRMHFEVLAKVDGENVIPENVMRVDPIPLEDTIAQNVITGNVTMSKLPIIESPPSPFTL 424
>UniRef100_Q8LIX4 Hypothetical protein OSJNBb0053G03.20 [Oryza sativa]
Length = 422
Score = 101 bits (252), Expect = 3e-20
Identities = 102/416 (24%), Positives = 179/416 (42%), Gaps = 28/416 (6%)
Query: 8 RLHQLPFFIIFFLQLFTAFSNASSSSSSTHSNITHIFQDILKAISSRQKWDLNDVRVFNF 67
R+ LP ++ + +SSS++ + D+ +AI + +++V F
Sbjct: 8 RILALPLALLLAVGSSPGLVRQASSSAAAKPPVPKAISDLREAIVKGLGFQSEELKVSGF 67
Query: 68 DVAKIRFGTSQNYLFRIGSSKNNFTVKFSDEISSWNHNK---FTTTPKPDLASLVD---- 120
DV G + Y F I + V+ ++++ W+ F + D +L +
Sbjct: 68 DVRDALVGQAVAYEFDIEVGRKVVPVRLLEDVNRWDFVDLPIFRSQADADDTALAEIRRG 127
Query: 121 QLSSIAF---LDYIKLEGPFELRVHESHHLSLSLPMNVSYNGLKHIIVGKGITVEVRRAR 177
+ AF L +L GP EL + + + L+LP +V LK +++ G V V+ A+
Sbjct: 128 KSGKRAFDPTLPPFQLAGPMELWIQDGDDVRLALPHDVEAGTLKKVVLSDGAVVTVKGAK 187
Query: 178 EISFYYQSDLDLQRNGSVICSNQKN------EFWPFLQSMCVPLIPIRIIGSASLIAYVA 231
+S +L L N + + +S PL+ +RI G SL + +
Sbjct: 188 AVSLRLPLELPLPLNRTTYKGRLSSLISIAQTLRGAARSNQKPLLSLRIEGPTSLSSTPS 247
Query: 232 RNPYVQI-------GTALISEDAVELLPEKCYHGCVFRKQACPVASLNLR---LILLEKI 281
+P ++ G +S A+ + + G P+ SLN L E++
Sbjct: 248 MSPNDRLKLKRLAPGQVELSSRAIPAVTDDDGDGS-HAAGLWPLLSLNGSDGSLQGFEEL 306
Query: 282 LRSLLGHKILQDRLSGLIKANIKAYAGVKFPLELERDVGNN-ATLSTLPDWRTRPSVERV 340
L S+LG K + L+KA A VK +E+ + + S P+W+T+P R
Sbjct: 307 LASVLGKKAGEKGTFKLLKARASAQTYVKMGFAVEKRIADGEVNWSNFPEWKTKPKKLRA 366
Query: 341 WFEVMARVEDSRLKPLSIKKVKPFIESDSVSWANLMSNLSYTKLRPVLLPPEALTL 396
+EV+ARVE + P I +V+PF +++S + L N+S +K V PP TL
Sbjct: 367 HYEVLARVEGGQAIPERIAQVQPFEADEAMSESVLTGNVSMSKTEVVHPPPVYFTL 422
>UniRef100_Q9FLG7 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MUB3
[Arabidopsis thaliana]
Length = 163
Score = 64.3 bits (155), Expect = 6e-09
Identities = 38/134 (28%), Positives = 71/134 (52%), Gaps = 4/134 (2%)
Query: 267 PVASLN---LRLILLEKILRSLLGHKILQDRLSGLIKANIKAYAGVKFPLELERDVGN-N 322
P+ S+N L+ EK+L S+LG K + ++KA + A +K +E+ + +
Sbjct: 30 PITSINGSNANLLGFEKLLTSVLGPKAQEKGSFKVLKAKVAAQTFMKIGFGIEKKLKEAD 89
Query: 323 ATLSTLPDWRTRPSVERVWFEVMARVEDSRLKPLSIKKVKPFIESDSVSWANLMSNLSYT 382
+ P+WRT+P R+ FEV+A+V+ + P ++ +V P D+++ + N++ +
Sbjct: 90 VEGLSFPEWRTKPETMRMHFEVLAKVDGENVIPENVMRVDPIPLEDTIAQNVITGNVTMS 149
Query: 383 KLRPVLLPPEALTL 396
KL + PP TL
Sbjct: 150 KLPIIESPPSPFTL 163
>UniRef100_P93385 Nicotiana tabacum ORF [Nicotiana tabacum]
Length = 126
Score = 47.8 bits (112), Expect = 6e-04
Identities = 30/88 (34%), Positives = 48/88 (54%), Gaps = 5/88 (5%)
Query: 267 PVASLN---LRLILLEKILRSLLGHKILQDRLSGLIKANIKAYAGVKFPLELERDV--GN 321
PV S+N L+ E +L ++LG K Q L+KA++ A VK +E+ + G+
Sbjct: 27 PVTSVNGSNSNLLGFEALLTNVLGPKASQKGSFKLLKADVSAQTFVKIGFGVEKKLKEGD 86
Query: 322 NATLSTLPDWRTRPSVERVWFEVMARVE 349
P+WRT+P R+ FEV+A+V+
Sbjct: 87 GFNWEGYPEWRTKPESVRMHFEVLAKVD 114
>UniRef100_UPI0000318DC0 UPI0000318DC0 UniRef100 entry
Length = 569
Score = 39.7 bits (91), Expect = 0.15
Identities = 34/148 (22%), Positives = 63/148 (41%), Gaps = 8/148 (5%)
Query: 29 ASSSSSSTHSNITHIFQDILKAISSRQKWDLNDVRVFNF-DVAKIRFGTSQNYLFRIGSS 87
A+ +S H I QD+L+ S Q+W L D+ + + + L+R+ +
Sbjct: 187 ANKYCTSAHKLIVATAQDVLEVYSDWQRWVLGDLPFSHMVRILDVYLVEGYKVLYRVAIA 246
Query: 88 KNNFTVKFSDEISSWNHNKFTTTPKPDLASLVDQLSSIAFLDYIKLEGPFELRVHESHHL 147
F K + + + T K D+ + V ++S+ D + LE F +R+ +
Sbjct: 247 LLKFYRKHKAGLQGSQQQQDSATVKADIRAFVQDIASVITPDKL-LEKAFSIRLFSRKEI 305
Query: 148 SLSLPMNVSYNGLKHIIVGKGITVEVRR 175
+L N + + KGITV+ +R
Sbjct: 306 TLLQLTN------ERSLQQKGITVKQKR 327
>UniRef100_Q5WL82 Succinate-semialdehyde dehydrogenase [Bacillus clausii]
Length = 472
Score = 37.4 bits (85), Expect = 0.75
Identities = 19/59 (32%), Positives = 35/59 (59%), Gaps = 6/59 (10%)
Query: 322 NATLSTLPDWRTRPSVER-----VWFEVMARVEDSRLKPLSIKKVKPFIES-DSVSWAN 374
+A + LP+WR + + ER W+E++ R E+ K +++++ KPF E+ V +AN
Sbjct: 45 DAAAAALPEWRAKTAAERGAYLQKWYELIQREEEEIAKLMTLEQGKPFAEALGEVRYAN 103
>UniRef100_Q8DN38 Choline-binding protein [Streptococcus pneumoniae]
Length = 690
Score = 37.0 bits (84), Expect = 0.99
Identities = 56/228 (24%), Positives = 98/228 (42%), Gaps = 22/228 (9%)
Query: 6 SLRLHQLPFFIIFFLQLFT--AFSNASSSSSST-HSNITHIFQDILKAISSRQKWDLNDV 62
S +L +L F L+L + AF+N S+ T ++ + ++ + +S + D+ +
Sbjct: 196 SQKLKKLTFSSSSKLELISHEAFANLSNLEKLTLPKSVKTLGSNLFRLTTSLKHVDVEEG 255
Query: 63 RVFNFDVAKIRFGTSQNYLFRIGSSKNNF---TVKFSDEISSWNHNKFTTTPKPDLASLV 119
V + F + L S KN+ T K + E++S++ NK + K +L +
Sbjct: 256 NESFASVDGVLFSKDKTQLIYYPSQKNDESYKTPKETKELASYSFNKNSYLKKLELNEGL 315
Query: 120 DQLSSIAFLDYIKLEG---PFELRVHES-------HHLSLSLPMNVSYNGLKHIIVG--K 167
+++ + AF D IKLE P L E L LP NV G KH++ G K
Sbjct: 316 EKIGTFAFADAIKLEEISLPNSLETIERLAFYGNLELKELILPDNVKNFG-KHVMNGLPK 374
Query: 168 GITVEVRRAREI-SFYYQSDLDLQRNGSVICSNQKNEFWPFLQSMCVP 214
+T+ + SF+ LD + + N+ EF + +P
Sbjct: 375 FLTLSGNNINSLPSFFLSGVLDSLK--EIHIKNKSTEFSVKKDTFAIP 420
>UniRef100_Q97NB5 Choline binding protein PcpA [Streptococcus pneumoniae]
Length = 621
Score = 37.0 bits (84), Expect = 0.99
Identities = 56/229 (24%), Positives = 97/229 (41%), Gaps = 23/229 (10%)
Query: 6 SLRLHQLPFFIIFFLQLFT--AFSNASSSSSST-HSNITHIFQDILKAISSRQKWDLNDV 62
S +L +L F L+L + AF+N S+ T ++ + ++ + +S + D+ +
Sbjct: 186 SQKLKKLTFSSSSKLELISHEAFANLSNLEKLTLPKSVKTLGSNLFRLTTSLKHVDVEEG 245
Query: 63 RVFNFDVAKIRFGTSQNYLFRIGSSKNNF---TVKFSDEISSWNHNKFTTTPKPDLASLV 119
V + F + L S KN+ T K + E++S++ NK + K +L +
Sbjct: 246 NESFASVDGVLFSKDKTQLIYYPSQKNDESYKTPKETKELASYSFNKNSYLKKLELNEGL 305
Query: 120 DQLSSIAFLDYIKLEG---PFELRVHES-------HHLSLSLPMNVSYNGLKHIIVG--- 166
+++ + AF D IKLE P L E L LP NV G KH++ G
Sbjct: 306 EKIGTFAFADAIKLEEISLPNSLETIERLAFYGNLELKELILPDNVKNFG-KHVMNGLPK 364
Query: 167 -KGITVEVRRAREISFYYQSDLDLQRNGSVICSNQKNEFWPFLQSMCVP 214
K +T+ SF+ LD + + N+ EF + +P
Sbjct: 365 LKSLTIGNNINSLPSFFLSGVLDSLK--EIHIKNKSTEFSVKKDTFAIP 411
>UniRef100_Q8XLN3 Hypothetical protein CPE1008 [Clostridium perfringens]
Length = 287
Score = 35.0 bits (79), Expect = 3.7
Identities = 19/78 (24%), Positives = 36/78 (45%), Gaps = 1/78 (1%)
Query: 93 VKFSDEISSWNHNKFTTTPKPDLASLVDQLSSIAFLDYIKLEGPFELRVHE-SHHLSLSL 151
+K + + WNH + D+ ++++ +I LD E E + +++ L
Sbjct: 61 IKGYETLIRWNHETLGSISASDIVKTLEEIDAIKDLDLYVFESTCEFQSRLIQNNIELQC 120
Query: 152 PMNVSYNGLKHIIVGKGI 169
+N+S N LKH VG +
Sbjct: 121 SVNISINTLKHRGVGSNL 138
>UniRef100_O04642 Hypothetical protein F2P16.15 [Arabidopsis thaliana]
Length = 559
Score = 35.0 bits (79), Expect = 3.7
Identities = 35/158 (22%), Positives = 69/158 (43%), Gaps = 24/158 (15%)
Query: 16 IIFFLQLFTAFSNASSSSSSTHSNITHIFQDILKAISSRQKWDLNDVRVFNFDVAKIRFG 75
+ FL + F++ S + + + FQD L I + Q+ + K F
Sbjct: 286 LTIFLLVLIGFAHVSGILKEIVAVLINCFQDFLPLIHASQR------------INKESFD 333
Query: 76 TSQNYLFRIGSSKNNFTVKFSDEISSWNHNKFTTTPKPDLASLVDQLSSIAFLDYIKLEG 135
++ L IG + +KFS + H K+ +P+ + ++D+ IA + KL G
Sbjct: 334 CIRHILRSIG-----YAIKFSIRRHTQRHTKWLPSPEENTLMILDR--DIASMLSKKLLG 386
Query: 136 PFELRVHESHHLSLSLPMNVSYNGLKHIIVGKGITVEV 173
F L +H + S+ + +++N L ++ G+ E+
Sbjct: 387 SFPL----NHENNFSVEV-MNHNSLLFLLFNNGVLTEI 419
>UniRef100_Q658C8 Hypothetical protein P0436E04.12-1 [Oryza sativa]
Length = 555
Score = 33.9 bits (76), Expect = 8.3
Identities = 33/119 (27%), Positives = 47/119 (38%), Gaps = 22/119 (18%)
Query: 14 FFIIFFLQL---FTAFSNASSSSSSTHSNITHIFQDILKAISSRQKWDLNDVRVFN---- 66
FF FF L TA ++ S+SS IT I + WD ++ N
Sbjct: 9 FFFFFFFILPASLTATASTSTSSCPDGWQITPALDKCFIYIPTPLSWDRSEALCRNNFTA 68
Query: 67 ----------FDVAKIRFGTSQNYLFRIGSSKNN----FTVKFSDEISSWNHNKFTTTP 111
++AK G S + + +G +NN F K+SD+ SSWN F P
Sbjct: 69 HLAALSSLQDLNLAKSLCGPSPSGCW-VGGHRNNTASAFAWKWSDDSSSWNDTAFPADP 126
>UniRef100_Q658C7 Hypothetical protein P0436E04.12-2 [Oryza sativa]
Length = 554
Score = 33.9 bits (76), Expect = 8.3
Identities = 33/119 (27%), Positives = 47/119 (38%), Gaps = 22/119 (18%)
Query: 14 FFIIFFLQL---FTAFSNASSSSSSTHSNITHIFQDILKAISSRQKWDLNDVRVFN---- 66
FF FF L TA ++ S+SS IT I + WD ++ N
Sbjct: 9 FFFFFFFILPASLTATASTSTSSCPDGWQITPALDKCFIYIPTPLSWDRSEALCRNNFTA 68
Query: 67 ----------FDVAKIRFGTSQNYLFRIGSSKNN----FTVKFSDEISSWNHNKFTTTP 111
++AK G S + + +G +NN F K+SD+ SSWN F P
Sbjct: 69 HLAALSSLQDLNLAKSLCGPSPSGCW-VGGHRNNTASAFAWKWSDDSSSWNDTAFPADP 126
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 638,391,960
Number of Sequences: 2790947
Number of extensions: 25718941
Number of successful extensions: 60398
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 60367
Number of HSP's gapped (non-prelim): 20
length of query: 400
length of database: 848,049,833
effective HSP length: 129
effective length of query: 271
effective length of database: 488,017,670
effective search space: 132252788570
effective search space used: 132252788570
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC148404.10


