

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148360.3 - phase: 0
(301 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
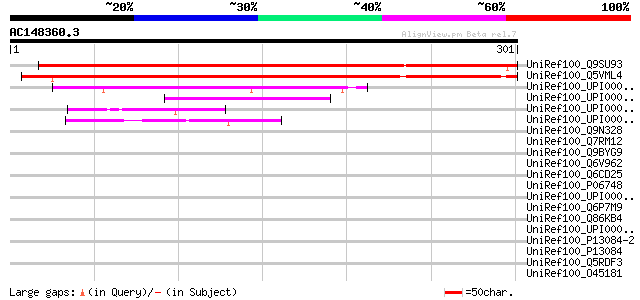
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SU93 Hypothetical protein T16L4.30 [Arabidopsis thal... 384 e-105
UniRef100_Q5VML4 Hypothetical protein P0529C07.1 [Oryza sativa] 362 5e-99
UniRef100_UPI000030936D UPI000030936D UniRef100 entry 107 5e-22
UniRef100_UPI0000274E6A UPI0000274E6A UniRef100 entry 60 5e-08
UniRef100_UPI000032ABFB UPI000032ABFB UniRef100 entry 57 8e-07
UniRef100_UPI00002E7C04 UPI00002E7C04 UniRef100 entry 52 2e-05
UniRef100_Q9N328 Hypothetical protein Y59E9AL.6 [Caenorhabditis ... 45 0.002
UniRef100_Q7RM12 SNF2 family N-terminal domain, putative [Plasmo... 45 0.002
UniRef100_Q9BYG9 Nucleophosmin/B23.2 [Homo sapiens] 45 0.002
UniRef100_Q6V962 Nucleophosmin [Homo sapiens] 45 0.002
UniRef100_Q6CD25 Similar to tr|Q04031 Saccharomyces cerevisiae D... 45 0.002
UniRef100_P06748 Nucleophosmin [Homo sapiens] 45 0.002
UniRef100_UPI000046B225 UPI000046B225 UniRef100 entry 45 0.003
UniRef100_Q6P7M9 Hypothetical protein MGC75666 [Xenopus tropicalis] 44 0.004
UniRef100_Q86KB4 Similar to Y55B1BR.3.p [Caenorhabditis elegans]... 44 0.005
UniRef100_UPI0000452EB9 UPI0000452EB9 UniRef100 entry 44 0.007
UniRef100_P13084-2 Splice isoform B23.2 of P13084 [Rattus norveg... 44 0.007
UniRef100_P13084 Nucleophosmin [Rattus norvegicus] 44 0.007
UniRef100_Q5RDF3 Hypothetical protein DKFZp469O2211 [Pongo pygma... 43 0.009
UniRef100_O45181 Hypothetical protein K07H8.10 [Caenorhabditis e... 43 0.009
>UniRef100_Q9SU93 Hypothetical protein T16L4.30 [Arabidopsis thaliana]
Length = 306
Score = 384 bits (987), Expect = e-105
Identities = 185/286 (64%), Positives = 228/286 (79%), Gaps = 3/286 (1%)
Query: 18 IPLSHCAKKPVGIARKEDVPYIKCQVCEILAKQLYQQVQSKKAEISPKKISEYQIIEIAE 77
+P+S AKKP RKEDVPYIKCQVCE L+ +L+Q V+ K+ +ISPKKISEY+IIEIAE
Sbjct: 22 LPVSDAAKKPSSTPRKEDVPYIKCQVCEKLSSRLHQLVKEKQQQISPKKISEYEIIEIAE 81
Query: 78 NVCNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAE 137
NVCNLKK EADW+L+IDIVEK D L L E EG CNS+CKT+E ACQ+V+GYSDTDVAE
Sbjct: 82 NVCNLKKEEADWMLKIDIVEKGDNLVLVEQQEEGMCNSKCKTIENACQKVIGYSDTDVAE 141
Query: 138 YLYSSKPDIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEG 197
Y+Y SKPD+ SL N+LCKDL+ AC+ KPPPVPKDR PGEPFVAK SK+AEM+K+L+SM+G
Sbjct: 142 YIYKSKPDLVSLVNHLCKDLTDACSKKPPPVPKDRVPGEPFVAKPSKDAEMDKILRSMQG 201
Query: 198 MPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQ 257
MPGAPGMK+YSR+D+ K N G E++D D+DEDEEED+ FP LGK+LK KE++ + K+
Sbjct: 202 MPGAPGMKVYSREDIEKGNIGNEDDDGDDDEDEEEDD-KFPKNLGKVLKEKESKTEELKK 260
Query: 258 KIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK--KNSKSEL 301
I K LK+HA KVSN ++RWWKG ++SSK K+ KSEL
Sbjct: 261 TITKEFKKKGEALKRHAQKVSNRVRRWWKGLGSSSSKKPKSGKSEL 306
>UniRef100_Q5VML4 Hypothetical protein P0529C07.1 [Oryza sativa]
Length = 306
Score = 362 bits (930), Expect = 5e-99
Identities = 178/303 (58%), Positives = 225/303 (73%), Gaps = 14/303 (4%)
Query: 8 LVVSVVIASWIPLSHCA--------KKPVGIARKEDVPYIKCQVCEILAKQLYQQVQSKK 59
+ ++V +A+ + L H A KK ARKED+PYI+CQVCE +A+++ QV K+
Sbjct: 9 VAMAVAVAAVVLLLHPAASAAAAGPKKVATAARKEDIPYIRCQVCERIAREISAQVAKKQ 68
Query: 60 AEI-SPKKISEYQIIEIAENVCNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECK 118
+ + KK+ E +IIEIAENVCNLKK EADW+L+IDIVEK D+LEL E D EG CN+ECK
Sbjct: 69 QALPATKKVPEIEIIEIAENVCNLKKQEADWMLKIDIVEKGDKLELVEQDEEGHCNAECK 128
Query: 119 TVERACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPF 178
T+ERACQEVMGY+DTDVAE++Y KP D L +LCKDLS+AC PPPVPKDR PGEPF
Sbjct: 129 TIERACQEVMGYADTDVAEFVYKKKPSADQLVKFLCKDLSEACVVDPPPVPKDRVPGEPF 188
Query: 179 VAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFP 238
AK SK+AEM+++LKSMEG+PGAP MKMYSRDDLMK NFG + +DDD+DEDE++ DFP
Sbjct: 189 AAKPSKDAEMDRILKSMEGIPGAPSMKMYSRDDLMKNNFGVDGDDDDDDEDEDD---DFP 245
Query: 239 SKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSKKNSK 298
LGK+ K K + K D KQ++ K I DT LK H KVS +++WW+GKK S K+SK
Sbjct: 246 KNLGKVFKDKGSPKKDLKQQVVKQIKDTGKKLKGHVNKVSKVVKKWWQGKKKPS--KSSK 303
Query: 299 SEL 301
+EL
Sbjct: 304 TEL 306
>UniRef100_UPI000030936D UPI000030936D UniRef100 entry
Length = 254
Score = 107 bits (266), Expect = 5e-22
Identities = 57/202 (28%), Positives = 102/202 (50%), Gaps = 19/202 (9%)
Query: 26 KPVGIARKEDVPYIKCQVCEILAKQLYQQ----------VQSKKAEISPKKISEYQIIEI 75
KP K D+ YI+C VC + + Y + + K+ + + E + E
Sbjct: 7 KPQAELLKSDMKYIRCGVCRKMVEIAYDKSSELLEKRFAFKKKRKNEATEFDGEGAVQEF 66
Query: 76 AENVCNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDV 135
E +CN K E +W+ +D+V++ ++L+L + + G+C EC+T+E C++V+ +D +
Sbjct: 67 VEKMCNPVKTEGEWVGMLDLVQEGEQLQLAQQPTHGKCLKECRTIEAVCEDVLDKADVEF 126
Query: 136 AEYLYSS---KPDIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLL 192
E LY + K ++ Y+C + C KPPP+ K R E F + +E +M+ +
Sbjct: 127 TEILYKAIQEKSTVEQAQRYICNRAAGVCKKKPPPLLKPRKHDEKFQPMTPEEKQMQDMQ 186
Query: 193 KSME--GMPGAPGMKMYSRDDL 212
+++ GM G MY R+DL
Sbjct: 187 ANLKDTGMSGT----MYKREDL 204
>UniRef100_UPI0000274E6A UPI0000274E6A UniRef100 entry
Length = 103
Score = 60.5 bits (145), Expect = 5e-08
Identities = 29/99 (29%), Positives = 50/99 (50%), Gaps = 1/99 (1%)
Query: 93 IDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYS-DTDVAEYLYSSKPDIDSLTN 151
+D+VE+ +RLEL +G C EC T++ AC ++ + +++E LY+ + L
Sbjct: 4 VDLVEEGERLELVRQSEDGPCGKECATIQLACNGLLEEGWENELSEALYTGSVSVARLRA 63
Query: 152 YLCKDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEK 190
+CK+ S AC PP + R G F + E + +
Sbjct: 64 DVCKEWSSACRKPPPKLDPARAAGPRFRPYTDSERALRE 102
>UniRef100_UPI000032ABFB UPI000032ABFB UniRef100 entry
Length = 124
Score = 56.6 bits (135), Expect = 8e-07
Identities = 31/96 (32%), Positives = 48/96 (49%), Gaps = 4/96 (4%)
Query: 35 DVPYIKCQVCEILAKQLYQQVQSKKAEISPKKISEYQIIEIAENVCNLKKVEADWILRID 94
DV YI+CQ CE L+ + + A+ PK + E ++ + + C E W+ +D
Sbjct: 24 DVKYIRCQACEHFVHALHTAIDNA-AQAKPK-LGEEDVVGLMKASCQPHTEEGAWLRSLD 81
Query: 95 IVE--KADRLELEEHDSEGQCNSECKTVERACQEVM 128
+VE L L +G CN EC+TVE AC ++
Sbjct: 82 LVEDDSTRSLVLVAEAEDGPCNRECRTVELACNALL 117
>UniRef100_UPI00002E7C04 UPI00002E7C04 UniRef100 entry
Length = 194
Score = 52.0 bits (123), Expect = 2e-05
Identities = 32/135 (23%), Positives = 63/135 (45%), Gaps = 18/135 (13%)
Query: 34 EDVPYIKCQVCEILAKQLYQQVQSKKAEISPKKISEYQIIEIAENVCNLKKVEADWILRI 93
+DV ++C VC L K+ + + + + ++S +CN+ W+ +
Sbjct: 41 KDVARVRCDVCRELVKEQQRVLAASSQRKANAEVS----------LCNVDSRAGIWLTAL 90
Query: 94 DIVEKADRLELEEHDSEGQCNSECKTVERACQEVM-------GYSDTDVAEYLYSSKPDI 146
DIV++ DR+ + + S G C EC+TVE AC++V+ + +AE
Sbjct: 91 DIVQQDDRVRVV-YASPGFCRRECRTVEVACKQVVDEIEQEESLEEAVLAETARLRDAGA 149
Query: 147 DSLTNYLCKDLSKAC 161
D++ + +C ++ C
Sbjct: 150 DAIADAVCTKRARVC 164
>UniRef100_Q9N328 Hypothetical protein Y59E9AL.6 [Caenorhabditis elegans]
Length = 214
Score = 45.4 bits (106), Expect = 0.002
Identities = 36/135 (26%), Positives = 55/135 (40%), Gaps = 3/135 (2%)
Query: 166 PPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDD 225
PP D P P A + + E + K E G D + E +D D
Sbjct: 49 PPAAPDAAPAAPDAAAAPADGEKKDGDKKSEMKEGEEKKNEKKEGDEKDEKKDEEKKDGD 108
Query: 226 EDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWW 285
+ +DE++D+ K K K+ +K D K+K + D KK K + +
Sbjct: 109 DKKDEKKDD---DKKDEKKSDKKDEKKDDKKEKDEEKKDDKKEDEKKDDDKKDDKKEAKS 165
Query: 286 KGKKTTSSKKNSKSE 300
K KK+ SKK+ KS+
Sbjct: 166 KSKKSNKSKKSKKSK 180
>UniRef100_Q7RM12 SNF2 family N-terminal domain, putative [Plasmodium yoelii yoelii]
Length = 1350
Score = 45.1 bits (105), Expect = 0.002
Identities = 38/142 (26%), Positives = 65/142 (45%), Gaps = 27/142 (19%)
Query: 187 EMEKLLKSMEGMPGAP------GMKMYSRDDLMKKNFGAE----NEDDDEDEDEEEDEAD 236
E+EK LK++E + G+ MY+ + + + ++ +EDDDED+D+EEDE +
Sbjct: 752 EIEKKLKNLENIFDLTNISLDGGLNMYNNLEKEESDNSSDEDGSDEDDDEDDDDEEDENE 811
Query: 237 FPSKLGKILKSKENEKG---------DWKQKIRKGIVDTSTTLKKHATKVSNHI------ 281
G L ++ E G K+K +K ++ S+T+KK H+
Sbjct: 812 SEGGDGNNLDNQIGEDGIVDENXKKKKKKKKKKKKKLNKSSTIKKFLKNNKKHMIFLDLG 871
Query: 282 --QRWWKGKKTTSSKKNSKSEL 301
+ WK T +K N K +
Sbjct: 872 ERKSKWKVMNTACTKTNKKKAI 893
>UniRef100_Q9BYG9 Nucleophosmin/B23.2 [Homo sapiens]
Length = 259
Score = 45.1 bits (105), Expect = 0.002
Identities = 59/224 (26%), Positives = 89/224 (39%), Gaps = 30/224 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE---DEEEDEADFPSKLGKILKSKEN 250
S+ G APG S+ K A+ +DDD+DE DE++D+ DF ++
Sbjct: 137 SISGKRSAPGGG--SKVPQKKVKLAADEDDDDDDEEDDDEDDDDDDF-----------DD 183
Query: 251 EKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
E+ + K ++K I DT K+A K SN + K T SK
Sbjct: 184 EEAEEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 223
>UniRef100_Q6V962 Nucleophosmin [Homo sapiens]
Length = 294
Score = 45.1 bits (105), Expect = 0.002
Identities = 59/224 (26%), Positives = 89/224 (39%), Gaps = 30/224 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE---DEEEDEADFPSKLGKILKSKEN 250
S+ G APG S+ K A+ +DDD+DE DE++D+ DF ++
Sbjct: 137 SISGKRSAPGGG--SKVPQKKVKLAADEDDDDDDEEDDDEDDDDDDF-----------DD 183
Query: 251 EKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
E+ + K ++K I DT K+A K SN + K T SK
Sbjct: 184 EEAEEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 223
>UniRef100_Q6CD25 Similar to tr|Q04031 Saccharomyces cerevisiae D9461. 3P [Yarrowia
lipolytica]
Length = 262
Score = 45.1 bits (105), Expect = 0.002
Identities = 35/111 (31%), Positives = 55/111 (49%), Gaps = 14/111 (12%)
Query: 181 KSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSK 240
K +E E ++ K+M+ + A +S DD +K+ + +E +DE+EDEEEDE K
Sbjct: 78 KGDQEKERARMQKAMKDIERARS-GYFSTDD--EKDSDSGDESEDEEEDEEEDEEKTLKK 134
Query: 241 LGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTT 291
K + E++ DW+ +D LKK K + +GKKTT
Sbjct: 135 SKKSSSTDEDDFSDWEG------IDKPGVLKKQKKKYVD-----GQGKKTT 174
>UniRef100_P06748 Nucleophosmin [Homo sapiens]
Length = 294
Score = 45.1 bits (105), Expect = 0.002
Identities = 59/224 (26%), Positives = 89/224 (39%), Gaps = 30/224 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE---DEEEDEADFPSKLGKILKSKEN 250
S+ G APG S+ K A+ +DDD+DE DE++D+ DF ++
Sbjct: 137 SISGKRSAPGGG--SKVPQKKVKLAADEDDDDDDEEDDDEDDDDDDF-----------DD 183
Query: 251 EKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
E+ + K ++K I DT K+A K SN + K T SK
Sbjct: 184 EEAEEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 223
>UniRef100_UPI000046B225 UPI000046B225 UniRef100 entry
Length = 714
Score = 44.7 bits (104), Expect = 0.003
Identities = 35/131 (26%), Positives = 62/131 (46%), Gaps = 19/131 (14%)
Query: 187 EMEKLLKSMEGMPGAP------GMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSK 240
E+EK LK++E + G+ MY+ + + + ++ + DED+D EEDE + +
Sbjct: 132 EIEKKLKNLENIFDLTNISLDGGLNMYNNLEKEESSDSSDEDGSDEDDDGEEDENESEGE 191
Query: 241 LGKILKSKENEKGD-----WKQKIRKGIVDTSTTLKKHATKVSNHI--------QRWWKG 287
G L ++ E G K+K +K ++ S+T+KK H+ + WK
Sbjct: 192 DGNNLDNQIGEDGIVDEPLKKKKKKKKKLNKSSTIKKFLKNNKKHMIFLDLGERKSKWKV 251
Query: 288 KKTTSSKKNSK 298
T+ +K N K
Sbjct: 252 MNTSCTKTNKK 262
>UniRef100_Q6P7M9 Hypothetical protein MGC75666 [Xenopus tropicalis]
Length = 212
Score = 44.3 bits (103), Expect = 0.004
Identities = 26/68 (38%), Positives = 35/68 (51%), Gaps = 1/68 (1%)
Query: 169 PKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE 228
PKD+ P E AK ++ E + +G G K+ R KK E +DDDEDE
Sbjct: 139 PKDKLPYEQKAAKLKEKYEKDVAAYKAKGKSDV-GKKVPGRPTGSKKKAEPEEDDDDEDE 197
Query: 229 DEEEDEAD 236
DEE++E D
Sbjct: 198 DEEDEEDD 205
>UniRef100_Q86KB4 Similar to Y55B1BR.3.p [Caenorhabditis elegans] [Dictyostelium
discoideum]
Length = 727
Score = 43.9 bits (102), Expect = 0.005
Identities = 49/212 (23%), Positives = 84/212 (39%), Gaps = 8/212 (3%)
Query: 98 KADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCKDL 157
K R + EE + S+ E A Q+ + T + KPD S ++
Sbjct: 135 KQSRAKKEEIKVDDAAKSDNDDKESADQKKFKKNITPSKKVTPIEKPDKKSTKKKKDQEE 194
Query: 158 SKACNTKPPPVPKDRTPGEPFVAKSSK-------EAEMEKLLKSMEGMPGAPGMKMYSRD 210
+ N P + T +P A SSK E E E+ + + P K +
Sbjct: 195 EEEENNNEESTPSEPTASKPKKAPSSKSKSKKDKEEEEEEEEEEEDEEEKKPKKKSTPKK 254
Query: 211 DLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEK-GDWKQKIRKGIVDTSTT 269
D E E+++EDE ++ K K N+K D +++ ++ V+ +T
Sbjct: 255 DSKSSKKKEEEEEEEEDETPKKSTEKETKKKPPAATKKSNKKLKDDEEEEKEEKVEKTTK 314
Query: 270 LKKHATKVSNHIQRWWKGKKTTSSKKNSKSEL 301
+KK + KV + + K +TS K +K E+
Sbjct: 315 VKKSSFKVPTAVTPNKEIKSSTSKKTPNKKEI 346
Score = 35.4 bits (80), Expect = 1.9
Identities = 38/149 (25%), Positives = 61/149 (40%), Gaps = 29/149 (19%)
Query: 159 KACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFG 218
K +TK PV K ++K KE E KL S + P +KK
Sbjct: 350 KKTSTKKIPVKK--------ISKDDKEEET-KLSSSKKKTP-------------VKKTSK 387
Query: 219 AENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHAT--- 275
E E+++ED DEEE + + + K K E++ + +K +TT K +
Sbjct: 388 EEEEEEEEDNDEEEQSSKKKTPVKKTSKIVEDDDEEETSPSKKKTPTKNTTPSKRKSVEP 447
Query: 276 ----KVSNHIQRWWKGKKTTSSKKNSKSE 300
K ++ KKT ++K+ KS+
Sbjct: 448 KPKEKEEEDEEKESPKKKTKKAEKSPKSD 476
Score = 35.0 bits (79), Expect = 2.4
Identities = 48/206 (23%), Positives = 81/206 (39%), Gaps = 26/206 (12%)
Query: 49 KQLYQQVQSKKAEISPKKISEYQIIEIAENVCNLKKVEADWILRIDIVEKADRLELEEHD 108
K++ ++ ++ +I KKIS+ E + + KK + K + E EE +
Sbjct: 344 KEIEEEKKTSTKKIPVKKISKDDKEEETKLSSSKKKTP------VKKTSKEEEEEEEEDN 397
Query: 109 SEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCKDLSKACNTKPPPV 168
E + +S+ KT + +++ D D E S K T SK + +P P
Sbjct: 398 DEEEQSSKKKTPVKKTSKIV--EDDDEEETSPSKKKTPTKNTTP-----SKRKSVEPKPK 450
Query: 169 PKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE 228
K+ E K S + + +K KS + G K R + +++D D+
Sbjct: 451 EKEEEDEE----KESPKKKTKKAEKSPKSDTTRSGKKTPKR---------TKQDENDSDD 497
Query: 229 DEEEDEADFPSKLGKILKSKENEKGD 254
E EDE K K K E+ D
Sbjct: 498 SEVEDEKSSKGKKKKGGSKKHQEESD 523
>UniRef100_UPI0000452EB9 UPI0000452EB9 UniRef100 entry
Length = 1604
Score = 43.5 bits (101), Expect = 0.007
Identities = 54/207 (26%), Positives = 90/207 (43%), Gaps = 22/207 (10%)
Query: 97 EKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSKPD--IDSLTNYLC 154
EK E + DS+ + NSE K ++ G +A+ SK + + L + +
Sbjct: 895 EKTKIKEDSKIDSDSEENSESKE-----KKSTGLLAKTIAKAKAKSKSNSVLSKLPSKVN 949
Query: 155 KDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMK 214
K+ SKA N + KD + + K E + + K +P P + S+ + K
Sbjct: 950 KETSKAENDLENKIEKDDESNKKEIKK-----EEDSIAKLKAKVPVKPSPLLKSKSEKEK 1004
Query: 215 KNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIR-KGIVDTSTTLKKH 273
+ + ED DE++ E+ED+ K + KE E+G K+K + K D S + K
Sbjct: 1005 E----KEEDKDEEKKEKEDKEKKEKLKEKEEEGKEKEEGKEKEKEKDKDEKDKSKSKIKE 1060
Query: 274 ATKVSNHIQRWWKGKKTTSSKKNSKSE 300
+ K + + + SSKK SK E
Sbjct: 1061 SQK-----EEKLEETEKDSSKKESKDE 1082
>UniRef100_P13084-2 Splice isoform B23.2 of P13084 [Rattus norvegicus]
Length = 257
Score = 43.5 bits (101), Expect = 0.007
Identities = 54/227 (23%), Positives = 87/227 (37%), Gaps = 30/227 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEDEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKG 253
M G APG + + +K + +DD++DED+E+DE D + E+
Sbjct: 137 GMSGKRSAPG----GGNKVPQKKVKLDEDDDEDDEDDEDDEDDDDDDF-------DEEET 185
Query: 254 DWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSKKNSKSE 300
+ K ++K + DT K+A K + + GK S SK +
Sbjct: 186 EEKVPVKKSVRDTPA---KNAQKSNQN------GKDLKPSTPRSKGQ 223
>UniRef100_P13084 Nucleophosmin [Rattus norvegicus]
Length = 292
Score = 43.5 bits (101), Expect = 0.007
Identities = 54/227 (23%), Positives = 87/227 (37%), Gaps = 30/227 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEDEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKG 253
M G APG + + +K + +DD++DED+E+DE D + E+
Sbjct: 137 GMSGKRSAPG----GGNKVPQKKVKLDEDDDEDDEDDEDDEDDDDDDF-------DEEET 185
Query: 254 DWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSKKNSKSE 300
+ K ++K + DT K+A K + + GK S SK +
Sbjct: 186 EEKVPVKKSVRDTPA---KNAQKSNQN------GKDLKPSTPRSKGQ 223
>UniRef100_Q5RDF3 Hypothetical protein DKFZp469O2211 [Pongo pygmaeus]
Length = 313
Score = 43.1 bits (100), Expect = 0.009
Identities = 57/221 (25%), Positives = 87/221 (38%), Gaps = 23/221 (10%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKG 253
S+ G APG +K A +EDDD+D+DE++D+ D ++E+
Sbjct: 137 SISGKRSAPGGGSKVPQKKVKL---AADEDDDDDDDEDDDDEDDDD------DDFDDEEA 187
Query: 254 DWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
+ K ++K I DT K+A K SN + K T SK
Sbjct: 188 EEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 224
>UniRef100_O45181 Hypothetical protein K07H8.10 [Caenorhabditis elegans]
Length = 798
Score = 43.1 bits (100), Expect = 0.009
Identities = 39/151 (25%), Positives = 67/151 (43%), Gaps = 17/151 (11%)
Query: 96 VEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCK 155
V+K+ +E +E D E + E + E +E +T+ + K + S+ + K
Sbjct: 281 VKKSVAVEEDEEDEEDE--EEIEMDEEDEEEEEDEEETEPVAKTPAVKSSLKSVDSTKGK 338
Query: 156 DLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKK 215
++ + P V K + P P AK+SK A A + + DD +
Sbjct: 339 QVNLSKVATPTTVDK-KAPATPHPAKTSKNA--------------AGDVPKLNFDDDSDE 383
Query: 216 NFGAENEDDDEDEDEEEDEADFPSKLGKILK 246
+ E ++EDE+EEE+E D P+K +LK
Sbjct: 384 EDEEDEEVEEEDEEEEEEEEDVPAKKAPVLK 414
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.311 0.129 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 525,238,690
Number of Sequences: 2790947
Number of extensions: 23308294
Number of successful extensions: 244922
Number of sequences better than 10.0: 1539
Number of HSP's better than 10.0 without gapping: 490
Number of HSP's successfully gapped in prelim test: 1151
Number of HSP's that attempted gapping in prelim test: 223353
Number of HSP's gapped (non-prelim): 7422
length of query: 301
length of database: 848,049,833
effective HSP length: 127
effective length of query: 174
effective length of database: 493,599,564
effective search space: 85886324136
effective search space used: 85886324136
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 74 (33.1 bits)
Medicago: description of AC148360.3


