

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148345.17 + phase: 0
(528 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
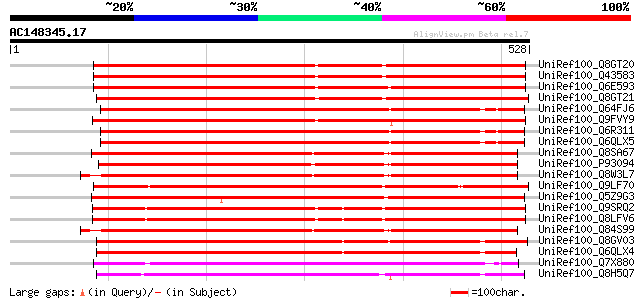
Sequences producing significant alignments: (bits) Value
UniRef100_Q8GT20 Benzoyl coenzyme A: benzyl alcohol benzoyl tran... 536 e-151
UniRef100_Q43583 Hsr201 protein [Nicotiana tabacum] 533 e-150
UniRef100_Q6E593 Benzoyl coenzyme A: benzyl alcohol benzoyl tran... 532 e-150
UniRef100_Q8GT21 Benzoyl coenzyme A: benzyl alcohol benzoyl tran... 511 e-143
UniRef100_Q64FJ6 Alcohol acyl transferase [Malus domestica] 451 e-125
UniRef100_Q9FVY9 Putative hypersensitivity-related (Hsr)protein ... 450 e-125
UniRef100_Q6R311 Alcohol acyl transferase [Malus domestica] 441 e-122
UniRef100_Q6QLX5 Alcohol acyl transferase [Pyrus communis] 439 e-122
UniRef100_Q8SA67 Putative acyltransferase [Cucumis melo] 422 e-117
UniRef100_P93094 Hypothetical protein [Cucumis melo] 420 e-116
UniRef100_Q8W3L7 Alcohol acetyltransferase [Cucumis melo] 419 e-115
UniRef100_Q9LF70 Hypothetical protein K10A8_20 [Arabidopsis thal... 412 e-113
UniRef100_Q5Z9G3 Putative benzoyl coenzyme A, benzyl alcohol ben... 403 e-111
UniRef100_Q9SRQ2 Putative hypersensitivity-related gene [Arabido... 400 e-110
UniRef100_Q8LFV6 Putative hypersensitivity-related gene [Arabido... 399 e-109
UniRef100_Q84S99 Alcohol acyltransferase [Cucumis melo] 385 e-105
UniRef100_Q8GV03 Acyltransferase 2 [Capsicum chinense] 364 3e-99
UniRef100_Q6QLX4 Alcohol acyl transferase [Lycopersicon esculentum] 353 7e-96
UniRef100_Q7X880 OSJNBa0043L24.8 protein [Oryza sativa] 343 7e-93
UniRef100_Q8H5Q7 Putative benzoyl coenzyme A [Oryza sativa] 343 9e-93
>UniRef100_Q8GT20 Benzoyl coenzyme A: benzyl alcohol benzoyl transferase [Nicotiana
tabacum]
Length = 460
Score = 536 bits (1380), Expect = e-151
Identities = 269/442 (60%), Positives = 333/442 (74%), Gaps = 8/442 (1%)
Query: 86 SAPLLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPV 145
S+ L+FTVRR +PEL+ PA PTPRE+K LSDIDDQEGLRF IP++ Y + SM KDPV
Sbjct: 6 SSELVFTVRRQKPELIAPAKPTPREIKFLSDIDDQEGLRFQIPVIQFYHKDSSMGRKDPV 65
Query: 146 KVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPL 205
KV++ A+++ LV+YYPFAGR+REG GRKLMVDCTGEG+MFVEA+ADVTL++FGD L PP
Sbjct: 66 KVIKKAIAETLVFYYPFAGRLREGNGRKLMVDCTGEGIMFVEADADVTLEQFGDELQPPF 125
Query: 206 PCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEM 265
PC EELLYDVP S +++ P+ LIQVTRL+CG FI + LNHTMSD GL FM A EM
Sbjct: 126 PCLEELLYDVPDSAGVLNCPLLLIQVTRLRCGGFIFALRLNHTMSDAPGLVQFMTAVGEM 185
Query: 266 ARGAHKPSIQPVWNREILMARDPPHITCNHHEYEQI-FSSNTIKEEDTTTLVHQSFFFRT 324
ARGA PSI PVW RE+L AR+PP +TC HHEY+++ + TI D +VH+SFFF
Sbjct: 186 ARGASAPSILPVWCRELLNARNPPQVTCTHHEYDEVRDTKGTIIPLD--DMVHKSFFFGP 243
Query: 325 SDIVVLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNS 384
S++ LR VP HLR C+TF+L+T+ W CRT +L+ + ++E+R +CIVNARSRF N
Sbjct: 244 SEVSALRRFVPHHLRKCSTFELLTAVLWRCRTMSLKPDPEEEVRALCIVNARSRF---NP 300
Query: 385 PL-VGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERC 443
PL GYYGN FA+P AVTTA KL N LGYA+ELV+K K++VTEEYM SVADLMV+K R
Sbjct: 301 PLPTGYYGNAFAFPVAVTTAAKLSKNPLGYALELVKKTKSDVTEEYMKSVADLMVLKGRP 360
Query: 444 LFTTIRSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPG-AAYIIPHKNVEGEEGF 502
FT +R+ +VSDVTR EVDFGWG+AVYGG AK G G PG A++ IP KN +GE G
Sbjct: 361 HFTVVRTFLVSDVTRGGFGEVDFGWGKAVYGGPAKGGVGAIPGVASFYIPFKNKKGENGI 420
Query: 503 ILLVCLSFEAMKRFAKELDEML 524
++ +CL AM+ F KELD ML
Sbjct: 421 VVPICLPGFAMETFVKELDGML 442
>UniRef100_Q43583 Hsr201 protein [Nicotiana tabacum]
Length = 460
Score = 533 bits (1373), Expect = e-150
Identities = 268/442 (60%), Positives = 331/442 (74%), Gaps = 8/442 (1%)
Query: 86 SAPLLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPV 145
S+ L+FTVRR +PEL+ PA PTPRE K LSDIDDQEGLRF IP++ Y + SM KDPV
Sbjct: 6 SSELVFTVRRQKPELIAPAKPTPRETKFLSDIDDQEGLRFQIPVIQFYHKDSSMGRKDPV 65
Query: 146 KVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPL 205
KV++ A+++ LV+YYPFAGR+REG GRKLMVDCTGEG+MFVEA+ADVTL++FGD L PP
Sbjct: 66 KVIKKAIAETLVFYYPFAGRLREGNGRKLMVDCTGEGIMFVEADADVTLEQFGDELQPPF 125
Query: 206 PCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEM 265
PC EELLYDVP S +++ P+ LIQVTRL+CG FI + LNHTMSD GL FM A EM
Sbjct: 126 PCLEELLYDVPDSAGVLNCPLLLIQVTRLRCGGFIFALRLNHTMSDAPGLVQFMTAVGEM 185
Query: 266 ARGAHKPSIQPVWNREILMARDPPHITCNHHEYEQI-FSSNTIKEEDTTTLVHQSFFFRT 324
ARG PSI PVW RE+L AR+PP +TC HHEY+++ + TI D +VH+SFFF
Sbjct: 186 ARGGSAPSILPVWCRELLNARNPPQVTCTHHEYDEVRDTKGTIIPLD--DMVHKSFFFGP 243
Query: 325 SDIVVLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNS 384
S++ LR VP HLR C+TF+L+T+ W CRT +L+ + ++E+R +CIVNARSRF N
Sbjct: 244 SEVSALRRFVPHHLRKCSTFELLTAVLWRCRTMSLKPDPEEEVRALCIVNARSRF---NP 300
Query: 385 PL-VGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERC 443
PL GYYGN FA+P AVTTA KL N LGYA+ELV+K K++VTEEYM SVADLMV+K R
Sbjct: 301 PLPTGYYGNAFAFPVAVTTAAKLSKNPLGYALELVKKTKSDVTEEYMKSVADLMVLKGRP 360
Query: 444 LFTTIRSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPG-AAYIIPHKNVEGEEGF 502
FT +R+ +VSDVTR EVDFGWG+AVYGG AK G G PG A++ IP KN +GE G
Sbjct: 361 HFTVVRTFLVSDVTRGGFGEVDFGWGKAVYGGPAKGGVGAIPGVASFYIPFKNKKGENGI 420
Query: 503 ILLVCLSFEAMKRFAKELDEML 524
++ +CL AM+ F KELD ML
Sbjct: 421 VVPICLPGFAMETFVKELDGML 442
>UniRef100_Q6E593 Benzoyl coenzyme A: benzyl alcohol benzoyl transferase [Petunia
hybrida]
Length = 460
Score = 532 bits (1371), Expect = e-150
Identities = 265/441 (60%), Positives = 331/441 (74%), Gaps = 6/441 (1%)
Query: 86 SAPLLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPV 145
S+ L+FTVRR +PEL+ PA PTPRE K LSDIDDQEGLRF IP++ YR + SM KDPV
Sbjct: 6 SSELVFTVRRQEPELIAPAKPTPRETKFLSDIDDQEGLRFQIPVINFYRKDSSMGGKDPV 65
Query: 146 KVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPL 205
+V++ A+++ LV+YYPFAGR+REG RKLMVDCTGEGVMFVEA ADVTL+EFGD L PP
Sbjct: 66 EVIKKAIAETLVFYYPFAGRLREGNDRKLMVDCTGEGVMFVEANADVTLEEFGDELQPPF 125
Query: 206 PCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEM 265
PC EELLYDVPGS ++ P+ LIQVTRL+CG FI + LNHTMSD GL FM A EM
Sbjct: 126 PCLEELLYDVPGSAGVLHCPLLLIQVTRLRCGGFIFALRLNHTMSDAPGLVQFMTAVGEM 185
Query: 266 ARGAHKPSIQPVWNREILMARDPPHITCNHHEYEQI-FSSNTIKEEDTTTLVHQSFFFRT 324
ARGA PS PVW RE+L AR+PP +TC HHEYE++ + T+ D +VH+SFFF
Sbjct: 186 ARGATAPSTLPVWCRELLNARNPPQVTCTHHEYEEVPDTKGTLIPLD--DMVHRSFFFGP 243
Query: 325 SDIVVLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNS 384
+++ LR VP HL +C+TF+++T+ W CRT +++ + ++E+R++CIVNARSRFN
Sbjct: 244 TEVSALRRFVPPHLHNCSTFEVLTAALWRCRTISIKPDPEEEVRVLCIVNARSRFNPQLP 303
Query: 385 PLVGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCL 444
GYYGN FA+P AVTTA KLC N LGYA+ELV+K K++VTEEYM SVADLMVIK R
Sbjct: 304 S--GYYGNAFAFPVAVTTAEKLCKNPLGYALELVKKTKSDVTEEYMKSVADLMVIKGRPH 361
Query: 445 FTTIRSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPG-AAYIIPHKNVEGEEGFI 503
FT +R+ +VSDVTR EVDFGWG+AVYGG AK G G PG A++ IP +N +GE G +
Sbjct: 362 FTVVRTYLVSDVTRAGFGEVDFGWGKAVYGGPAKGGVGAIPGVASFYIPFRNKKGENGIV 421
Query: 504 LLVCLSFEAMKRFAKELDEML 524
+ +CL AM++F KELD ML
Sbjct: 422 VPICLPGFAMEKFVKELDSML 442
>UniRef100_Q8GT21 Benzoyl coenzyme A: benzyl alcohol benzoyl transferase [Clarkia
breweri]
Length = 456
Score = 511 bits (1317), Expect = e-143
Identities = 262/444 (59%), Positives = 330/444 (74%), Gaps = 11/444 (2%)
Query: 89 LLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHE-PSMKEKDPVKV 147
L F V R +PEL+ PA TP E K LSD++DQEGLRF IP++ Y+H SM+E+DPV+V
Sbjct: 7 LSFEVCRRKPELIRPAKQTPHEFKKLSDVEDQEGLRFQIPVIQFYKHNNESMQERDPVQV 66
Query: 148 LRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPC 207
+R +++ALVYYYPFAGR+RE GRKL+V+CTGEGVMF+EA+ADVTL++FGDAL PP PC
Sbjct: 67 IREGIARALVYYYPFAGRLREVDGRKLVVECTGEGVMFIEADADVTLEQFGDALQPPFPC 126
Query: 208 FEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMAR 267
F++LL+DVPGS I+D P+ LIQVTRLKCGSFI + LNHTM+D AG+ LFM A EMAR
Sbjct: 127 FDQLLFDVPGSGGILDSPLLLIQVTRLKCGSFIFALRLNHTMADAAGIVLFMKAVGEMAR 186
Query: 268 GAHKPSIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFFFRTSDI 327
GA PS PVW+R IL AR PP +T NH EYE++ + +D L H+SFFF +++I
Sbjct: 187 GAATPSTLPVWDRHILNARVPPQVTFNHREYEEVKGTIFTPFDD---LAHRSFFFGSTEI 243
Query: 328 VVLRLLVPFHLRHC-TTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPL 386
+R +P HLR C TT +++T+C W CRT A++ D+E+RM+CIVNARS+F N PL
Sbjct: 244 SAMRKQIPPHLRSCSTTIEVLTACLWRCRTLAIKPNPDEEVRMICIVNARSKF---NPPL 300
Query: 387 V-GYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCLF 445
GYYGN FA PAAVTTAGKLC N LG+A+EL+RK K EVTEEYMHSVADLMV R F
Sbjct: 301 PDGYYGNAFAIPAAVTTAGKLCNNPLGFALELIRKAKREVTEEYMHSVADLMVATGRPHF 360
Query: 446 TTIRSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPG-AAYIIPHKNVEGEEGFIL 504
T + + +VSDVTR EVDFGWGEAVYGG AK G G PG ++ IP +N +GE+G +L
Sbjct: 361 TVVNTYLVSDVTRAGFGEVDFGWGEAVYGGPAKGGVGVIPGVTSFYIPLRNRQGEKGIVL 420
Query: 505 LVCLSFEAMKRFAKELDEML-GKQ 527
+CL AM+ FA+ L+ L GK+
Sbjct: 421 PICLPSAAMEIFAEALNNTLNGKE 444
>UniRef100_Q64FJ6 Alcohol acyl transferase [Malus domestica]
Length = 455
Score = 451 bits (1160), Expect = e-125
Identities = 231/436 (52%), Positives = 303/436 (68%), Gaps = 11/436 (2%)
Query: 93 VRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSM-KEKDPVKVLRHA 151
V+R QPEL+ PA TP+E K LSDIDDQE LR IP++ Y+ PS+ K ++PVK +R A
Sbjct: 9 VKRLQPELITPAKSTPQETKFLSDIDDQESLRVQIPIIMCYKDNPSLNKNRNPVKAIREA 68
Query: 152 LSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCFEEL 211
LS+ALVYYYP AGR+REG RKL+VDC GEG++FVEA ADVTL++ GD + PP P EE
Sbjct: 69 LSRALVYYYPLAGRLREGPNRKLVVDCNGEGILFVEASADVTLEQLGDKILPPCPLLEEF 128
Query: 212 LYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGAHK 271
LY+ PGS+ IID P+ LIQVT L CG FIL + LNHTM D AGL LF+ A AEMARGAH
Sbjct: 129 LYNFPGSDGIIDCPLLLIQVTCLTCGGFILALRLNHTMCDAAGLLLFLTAIAEMARGAHA 188
Query: 272 PSIQPVWNREILMARDPPHITCNHHEYEQIF--SSNTIKEEDTTTLVHQSFFFRTSDIVV 329
PSI PVW RE+L ARDPP ITC HHEYE + S + + + +V +SF+F ++ V
Sbjct: 189 PSILPVWERELLFARDPPRITCAHHEYEDVIGHSDGSYASSNQSNMVQRSFYFGAKEMRV 248
Query: 330 LRLLVPFHL-RHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPLVG 388
LR +P HL C+TFDLIT+C W CRT AL + + +R+ CIVNAR + N PL G
Sbjct: 249 LRKQIPPHLISTCSTFDLITACLWKCRTLALNINPKEAVRVSCIVNARGKHNNVRLPL-G 307
Query: 389 YYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCLFTTI 448
YYGN FA+PAA++ A LC N LGYA+ELV+K KA + EEY+ SVADL+V++ R +++
Sbjct: 308 YYGNAFAFPAAISKAEPLCKNPLGYALELVKKAKATMNEEYLRSVADLLVLRGRPQYSST 367
Query: 449 RS-CVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPGAAYIIPHKNVEGEEGFILLVC 507
S +VSD TR+ +V+FGWG+ V+ G K+ ++ + HKN E+G ++ +C
Sbjct: 368 GSYLIVSDNTRVGFGDVNFGWGQPVFAGPVKA----LDLISFYVQHKN-NTEDGILVPMC 422
Query: 508 LSFEAMKRFAKELDEM 523
L AM+RF +EL+ +
Sbjct: 423 LPSSAMERFQQELERI 438
>UniRef100_Q9FVY9 Putative hypersensitivity-related (Hsr)protein [Oryza sativa]
Length = 553
Score = 450 bits (1158), Expect = e-125
Identities = 235/448 (52%), Positives = 305/448 (67%), Gaps = 10/448 (2%)
Query: 85 TSAPLLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDP 144
T+A L FTVRR ELV PA PTPRE+K LSDIDDQ+GLRF+IP++ YR +M +DP
Sbjct: 5 TAAALKFTVRRKPAELVAPAGPTPRELKKLSDIDDQDGLRFHIPVIQFYRRSAAMGGRDP 64
Query: 145 VKVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPP 204
V+R A+++ALV YYPFAGR+RE GRKL VDCTGEGV+F+EA+ADV L+ FG AL PP
Sbjct: 65 APVIRAAVARALVSYYPFAGRLRELEGRKLAVDCTGEGVLFIEADADVRLEHFGGALQPP 124
Query: 205 LPCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAE 264
PC EEL++DVPGS ++ P+ L QVTRL CG FIL V L+HTM+D GL F+ A AE
Sbjct: 125 FPCLEELVFDVPGSSEVLGSPLLLFQVTRLACGGFILAVRLHHTMADAQGLVQFLGAVAE 184
Query: 265 MARG--AHKPSIQPVWNREILMARDPPHITCNHHEYEQI-FSSNTIKEEDTTTLVHQSFF 321
MARG A PS+ PVW RE+L AR PP H EY+++ + TI D + H+SFF
Sbjct: 185 MARGGAAAAPSVAPVWGREMLEARSPPRPAFAHREYDEVPDTKGTIIPLD--DMAHRSFF 242
Query: 322 FRTSDIVVLRL-LVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFN 380
F ++ +R L P TTF+++T C W CRT AL + D+ +RM+CIVNAR
Sbjct: 243 FGAREVAAVRSHLAPGIRERATTFEVLTGCLWRCRTAALAPDDDEVMRMICIVNARGGGK 302
Query: 381 ANNSPLV---GYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLM 437
+ + GYYGN FA+P AV TAG+L LGYAVELVR K EV+ EYM SVADLM
Sbjct: 303 SGGGAGMIPEGYYGNAFAFPVAVATAGELRARPLGYAVELVRAAKGEVSVEYMRSVADLM 362
Query: 438 VIKERCLFTTIRSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPG-AAYIIPHKNV 496
V + R FT +R+ +VSDVT+ ++DFGWG+ YGG AK G G PG A+++IP KN
Sbjct: 363 VQRGRPHFTVVRAYLVSDVTKAGFGDLDFGWGKPAYGGPAKGGVGAIPGVASFLIPFKNA 422
Query: 497 EGEEGFILLVCLSFEAMKRFAKELDEML 524
+GE+G ++ +CL AM +F +E+ +++
Sbjct: 423 KGEDGIVVPMCLPGPAMDKFVEEMGKLM 450
>UniRef100_Q6R311 Alcohol acyl transferase [Malus domestica]
Length = 459
Score = 441 bits (1135), Expect = e-122
Identities = 227/436 (52%), Positives = 301/436 (68%), Gaps = 11/436 (2%)
Query: 93 VRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSM-KEKDPVKVLRHA 151
V+R Q EL+ PA PT +E K LSDIDDQEGLRF +P++ Y+ PS+ K +PVKV+R A
Sbjct: 9 VKRLQLELITPAKPTLQEAKFLSDIDDQEGLRFQVPVIMCYKDNPSLNKNCNPVKVIREA 68
Query: 152 LSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCFEEL 211
LS+ALVYYYP AGR++EG RKLMVDC GEG++FVEA ADVTL++ GD + PP P EE
Sbjct: 69 LSRALVYYYPLAGRLKEGPNRKLMVDCNGEGILFVEASADVTLEQLGDKILPPCPLLEEF 128
Query: 212 LYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGAHK 271
L++ PGS+ II P+ L+QVT L CG FIL + +NHTM D GL LF+ A AEMARGAH
Sbjct: 129 LFNFPGSDGIIGCPLLLVQVTCLTCGGFILALRVNHTMCDAPGLLLFLTAIAEMARGAHA 188
Query: 272 PSIQPVWNREILMARDPPHITCNHHEYEQIF--SSNTIKEEDTTTLVHQSFFFRTSDIVV 329
PSI PVW RE+L +RDPP ITC HHEYE + S + + +V +SF+F ++ V
Sbjct: 189 PSILPVWERELLFSRDPPRITCAHHEYEDVIDHSDGLYASSNQSNMVQRSFYFGAKEMRV 248
Query: 330 LRLLVPFHL-RHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPLVG 388
LR +P HL C+TFDLIT+C W CRT AL + + +R+ CIVNAR + N PL G
Sbjct: 249 LRKQIPPHLISTCSTFDLITACLWKCRTLALNINPKEAVRVSCIVNARGKHNNVRLPL-G 307
Query: 389 YYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCLFTTI 448
YYGN FA+PAA++ A LC N LGYA+ELV+K KA + EEY+ SVADL+V++ R +++
Sbjct: 308 YYGNAFAFPAAISKAEPLCKNPLGYALELVKKAKATMNEEYLRSVADLLVLRGRPQYSST 367
Query: 449 RS-CVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPGAAYIIPHKNVEGEEGFILLVC 507
S +VSD TR +V+FGWG+ V+ G AK+ ++ + HKN E+G ++ +C
Sbjct: 368 GSYLIVSDNTRAGFGDVNFGWGQPVFAGPAKA----LDLISFYVQHKN-NTEDGILVPMC 422
Query: 508 LSFEAMKRFAKELDEM 523
L AM+RF +EL+ +
Sbjct: 423 LPSSAMERFQQELERI 438
>UniRef100_Q6QLX5 Alcohol acyl transferase [Pyrus communis]
Length = 442
Score = 439 bits (1129), Expect = e-122
Identities = 224/436 (51%), Positives = 302/436 (68%), Gaps = 11/436 (2%)
Query: 93 VRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSM-KEKDPVKVLRHA 151
V+R QPEL+ PA PTP+E K LSDIDDQEGLRF +P++ Y+ PS+ K ++P+KV++ A
Sbjct: 9 VKRLQPELITPAKPTPQETKFLSDIDDQEGLRFQLPVIMCYKDNPSLNKNRNPIKVIKEA 68
Query: 152 LSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCFEEL 211
LS+ALVYYYP AGR+REG RKLMV+C GEG++FVEA ADVTL++ GD + PP P EE
Sbjct: 69 LSRALVYYYPLAGRLREGPNRKLMVNCNGEGILFVEASADVTLEQLGDKILPPCPLLEEF 128
Query: 212 LYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGAHK 271
L++ PGS+ II P+ L+QVT L CG FIL + LNHTM D GL +F+ A EM RGA
Sbjct: 129 LFNFPGSDGIIGCPLLLVQVTCLTCGGFILALRLNHTMCDATGLLMFLTAITEMGRGADA 188
Query: 272 PSIQPVWNREILMARDPPHITCNHHEYEQIF--SSNTIKEEDTTTLVHQSFFFRTSDIVV 329
PSI PVW RE+L ARDPP ITC H+EYE + S + + + +V +SF+F ++ V
Sbjct: 189 PSILPVWERELLFARDPPRITCAHYEYEDVIDHSDGSYAFSNQSNMVQRSFYFGAKEMRV 248
Query: 330 LRLLVPFHL-RHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPLVG 388
LR +P HL C+TFDLIT+C W CRT L++ +R+ CIVNAR + N + PL G
Sbjct: 249 LRKQIPPHLISTCSTFDLITACLWKCRTLVLKINPKQAVRVSCIVNARGKHNNVHIPL-G 307
Query: 389 YYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCLFTTI 448
YYGN FA+PAAV+ A LC N LGYA+ELV+K KA + EEY+ SVADL+V++ R +++
Sbjct: 308 YYGNAFAFPAAVSKAEPLCKNPLGYALELVKKAKATMNEEYLRSVADLLVLRGRPQYSST 367
Query: 449 RS-CVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPGAAYIIPHKNVEGEEGFILLVC 507
S +VSD TR +V+FGWG+ V+ G AK+ ++ + HKN E+G ++ +C
Sbjct: 368 GSYLIVSDNTRAGFGDVNFGWGQPVFAGPAKA----LDLISFYVQHKN-NIEDGILVPMC 422
Query: 508 LSFEAMKRFAKELDEM 523
L AM+RF +EL+ +
Sbjct: 423 LPSSAMERFQQELERI 438
>UniRef100_Q8SA67 Putative acyltransferase [Cucumis melo]
Length = 461
Score = 422 bits (1086), Expect = e-117
Identities = 209/435 (48%), Positives = 295/435 (67%), Gaps = 7/435 (1%)
Query: 84 MTSAPLLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKD 143
M + F VR+ QPEL+ PA PTP E K LSD+DDQ+ LRF +P++ IY H PS++ +D
Sbjct: 4 MQTIDFSFQVRKCQPELIAPANPTPYEFKQLSDVDDQQSLRFQLPLVNIYHHNPSLEGRD 63
Query: 144 PVKVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHP 203
PVKV++ A+++ALV+YYP AGR+REG GRKL V+CTGEG++F+EA+ADV+L++F D L
Sbjct: 64 PVKVIKEAIAKALVFYYPLAGRLREGPGRKLFVECTGEGILFIEADADVSLEQFRDTLPY 123
Query: 204 PLPCFE-ELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAW 262
L E ++++ S+ +++ P+ LIQVTRLKCG FI ++ +HTM+DG G+ FM A
Sbjct: 124 SLSSMENNIIHNSLNSDGVLNSPLLLIQVTRLKCGGFIFGIHFDHTMADGFGIAQFMKAI 183
Query: 263 AEMARGAHKPSIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFFF 322
AE+ARGA PSI PVW R +L ARDPP IT H+EY+Q+ + + ++ + FFF
Sbjct: 184 AEIARGAFAPSILPVWQRALLTARDPPRITVRHYEYDQVVDTKSTL-IPANNMIDRLFFF 242
Query: 323 RTSDIVVLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNAN 382
I LR +P HL C++F+++ + W RT A QL+ ++E+R +C+VN RS+ +
Sbjct: 243 TQRQISTLRQTLPAHLHDCSSFEVLAAYVWRLRTIAFQLKPEEEVRFLCVVNLRSKIDI- 301
Query: 383 NSPLVGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKER 442
PL G+YGN +PA +TT KLCGN LGYAV+L+RK KA+ T+EY+ S+ D MVIK R
Sbjct: 302 --PL-GFYGNAIVFPAVITTVAKLCGNPLGYAVDLIRKAKAKATKEYIKSMVDFMVIKGR 358
Query: 443 CLFTTIRSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPG-AAYIIPHKNVEGEEG 501
FT I ++SD+TRI VDFGWG+A++GG G G G +Y I N GE+G
Sbjct: 359 PRFTEIGPFMMSDITRIGFENVDFGWGKAIFGGPIIGGCGIIRGMISYSIAFMNRNGEKG 418
Query: 502 FILLVCLSFEAMKRF 516
++ +CL AM+RF
Sbjct: 419 IVVPLCLPPPAMERF 433
>UniRef100_P93094 Hypothetical protein [Cucumis melo]
Length = 455
Score = 420 bits (1080), Expect = e-116
Identities = 214/430 (49%), Positives = 291/430 (66%), Gaps = 11/430 (2%)
Query: 91 FTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVKVLRH 150
F VR+ QPEL+ PA PTP E K LSD+DDQ+ LR +P + IY H PS++ +DPVKV++
Sbjct: 4 FHVRKCQPELIAPANPTPYEFKQLSDVDDQQSLRLQLPFVNIYPHNPSLEGRDPVKVIKE 63
Query: 151 ALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCFE- 209
A+ +ALV+YYP AGR+REG GRKL V+CTGEG++F+EA+ADV+L+EF D L L +
Sbjct: 64 AIGKALVFYYPLAGRLREGPGRKLFVECTGEGILFIEADADVSLEEFWDTLPYSLSSMQN 123
Query: 210 ELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGA 269
++++ S+ +++ P+ LIQVTRLKCG FI + NHTM+DG G+ FM A AE+ARGA
Sbjct: 124 NIIHNALNSDEVLNSPLLLIQVTRLKCGGFIFGLCFNHTMADGFGIVQFMKATAEIARGA 183
Query: 270 HKPSIQPVWNREILMARDPPHITCNHHEYEQI--FSSNTIKEEDTTTLVHQSFFFRTSDI 327
PSI PVW R +L ARDPP IT H+EY+Q+ S I + + Q FFF I
Sbjct: 184 FAPSILPVWQRALLTARDPPRITFRHYEYDQVVDMKSGLI---PVNSKIDQLFFFSQLQI 240
Query: 328 VVLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPLV 387
LR +P HL C +F+++T+ W RT ALQ + ++E+R +C++N RS+ + PL
Sbjct: 241 STLRQTLPAHLHDCPSFEVLTAYVWRLRTIALQFKPEEEVRFLCVMNLRSKIDI---PL- 296
Query: 388 GYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCLFTT 447
GYYGN PA +TTA KLCGN LGYAV+L+RK KA+ T EY+ S DLMVIK R FT
Sbjct: 297 GYYGNAVVVPAVITTAAKLCGNPLGYAVDLIRKAKAKATMEYIKSTVDLMVIKGRPYFTV 356
Query: 448 IRSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPG-AAYIIPHKNVEGEEGFILLV 506
+ S ++SD+TRI + VDFGWG+A++GG +GA G ++ +P N GE+G L +
Sbjct: 357 VGSFMMSDLTRIGVENVDFGWGKAIFGGPTTTGARITRGLVSFCVPFMNRNGEKGTALSL 416
Query: 507 CLSFEAMKRF 516
CL AM+RF
Sbjct: 417 CLPPPAMERF 426
>UniRef100_Q8W3L7 Alcohol acetyltransferase [Cucumis melo]
Length = 461
Score = 419 bits (1076), Expect = e-115
Identities = 212/446 (47%), Positives = 295/446 (65%), Gaps = 17/446 (3%)
Query: 73 THTTLIFSYHIMTSAPLLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFI 132
T T+ FS+H VR+ QPEL+ PA PTP E K LSD+DDQ+ LR +P + I
Sbjct: 3 TMQTIDFSFH----------VRKCQPELIAPANPTPYEFKQLSDVDDQQSLRLQLPFVNI 52
Query: 133 YRHEPSMKEKDPVKVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADV 192
Y H PS++ +DPVKV++ A+ +ALV+YYP AGR+REG GRKL V+CTGEG++F+EA+ADV
Sbjct: 53 YPHNPSLEGRDPVKVIKEAIGKALVFYYPLAGRLREGPGRKLFVECTGEGILFIEADADV 112
Query: 193 TLDEFGDALHPPLPCFE-ELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSD 251
+L+EF D L L + ++++ S+ +++ P+ LIQVTRLKCG FI + NHTM+D
Sbjct: 113 SLEEFWDTLPYSLSSMQNNIIHNALNSDEVLNSPLLLIQVTRLKCGGFIFGLCFNHTMAD 172
Query: 252 GAGLKLFMNAWAEMARGAHKPSIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEED 311
G G+ FM A AE+ARGA PSI PVW R +L ARDPP IT H+EY+Q+ + +
Sbjct: 173 GFGIVQFMKATAEIARGAFAPSILPVWQRALLTARDPPRITVRHYEYDQVVDTKSTL-IP 231
Query: 312 TTTLVHQSFFFRTSDIVVLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMC 371
++ + FFF I LR +P HL C++F+++ + W RT A QL+ ++E+R +C
Sbjct: 232 ANNMIDRLFFFTQRQISTLRQTLPAHLHDCSSFEVLAAYVWRLRTIAFQLKPEEEVRFLC 291
Query: 372 IVNARSRFNANNSPLVGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMH 431
+VN RS+ + PL G+YGN +PA +TT KLCGN LGYAV+L+RK KA+ T+EY+
Sbjct: 292 VVNLRSKIDI---PL-GFYGNAIVFPAVITTIAKLCGNPLGYAVDLIRKAKAKATKEYIK 347
Query: 432 SVADLMVIKERCLFTTIRSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPG-AAYI 490
S+ D MVIK R FT I ++SD+TRI VDFGWG+A++GG G G G +Y
Sbjct: 348 SMVDFMVIKGRPRFTEIGPFMMSDITRIGFENVDFGWGKAIFGGPIIGGCGIIRGMISYS 407
Query: 491 IPHKNVEGEEGFILLVCLSFEAMKRF 516
I N GE+G ++ +CL AM+RF
Sbjct: 408 IAFMNRNGEKGIVVPLCLPPPAMERF 433
>UniRef100_Q9LF70 Hypothetical protein K10A8_20 [Arabidopsis thaliana]
Length = 461
Score = 412 bits (1059), Expect = e-113
Identities = 223/447 (49%), Positives = 297/447 (65%), Gaps = 10/447 (2%)
Query: 86 SAPLLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPV 145
S L F + R +PELV PA PTPRE+K LSDIDDQEGLRF+IP +F YRH P+ DPV
Sbjct: 2 SGSLTFKIYRQKPELVSPAKPTPRELKPLSDIDDQEGLRFHIPTIFFYRHNPTTNS-DPV 60
Query: 146 KVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFG--DALHP 203
V+R AL++ LVYYYPFAGR+REG RKL VDCTGEGV+F+EA+ADVTL EF DAL P
Sbjct: 61 AVIRRALAETLVYYYPFAGRLREGPNRKLAVDCTGEGVLFIEADADVTLVEFEEKDALKP 120
Query: 204 PLPCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWA 263
P PCFEELL++V GS +++ P+ L+QVTRLKCG FI V +NH MSD GL LF+
Sbjct: 121 PFPCFEELLFNVEGSCEMLNTPLMLMQVTRLKCGGFIFAVRINHAMSDAGGLTLFLKTMC 180
Query: 264 EMARGAHKPSIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFFFR 323
E RG H P++ PVW R +L AR +T H EY+++ + T LV +S FF
Sbjct: 181 EFVRGYHAPTVAPVWERHLLSARVLLRVTHAHREYDEMPAIGTELGSRRDNLVGRSLFFG 240
Query: 324 TSDIVVLRLLVPFHL-RHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNAN 382
++ +R L+P +L T +++TS W RT AL+ + D E+R++ IVNARS+
Sbjct: 241 PCEMSAIRRLLPPNLVNSSTNMEMLTSFLWRYRTIALRPDQDKEMRLILIVNARSKL--K 298
Query: 383 NSPLV-GYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKE 441
N PL GYYGN FA+P A+ TA +L L +A+ L+++ K+ VTEEYM S+ADLMVIK
Sbjct: 299 NPPLPRGYYGNAFAFPVAIATANELTKKPLEFALRLIKEAKSSVTEEYMRSLADLMVIKG 358
Query: 442 RCLFTTIRSCVVSDVTRIKLREVDFG-WGEAVYGGAAKSGAGPFPGAAYIIPHKNVEGEE 500
R F++ + +VSDV RI ++DFG WG+ VYGG +G PGA++ + + GE
Sbjct: 359 RPSFSSDGAYLVSDV-RI-FADIDFGIWGKPVYGGIGTAGVEDLPGASFYVSFEKRNGEI 416
Query: 501 GFILLVCLSFEAMKRFAKELDEMLGKQ 527
G ++ VCL +AM+RF +EL+ + Q
Sbjct: 417 GIVVPVCLPEKAMQRFVEELEGVFNGQ 443
>UniRef100_Q5Z9G3 Putative benzoyl coenzyme A, benzyl alcohol benzoyl transferase
[Oryza sativa]
Length = 450
Score = 403 bits (1036), Expect = e-111
Identities = 220/448 (49%), Positives = 283/448 (63%), Gaps = 11/448 (2%)
Query: 84 MTSAPL-LFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEK 142
M S+PL FTVRR +P LV PAAPTPREVK LSDIDD EG+RF + +YR+ P+ K +
Sbjct: 1 MASSPLPAFTVRRGEPVLVTPAAPTPREVKALSDIDDGEGMRFYSSGIHLYRNNPAKKGQ 60
Query: 143 DPVKVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALH 202
DP V+R AL++ALV YYP AGR+RE AGRKL+V+C G+GVMF EA+AD+T D+FGD
Sbjct: 61 DPAMVIREALARALVPYYPLAGRLREEAGRKLVVECAGQGVMFAEADADLTADDFGDVQS 120
Query: 203 PPLPCFEELLYD---VPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFM 259
PP PCFE + + V G E ++ RP+ IQVTRL+CG FI H + D G F
Sbjct: 121 PPFPCFERFILESTTVAGVEPVVGRPLLYIQVTRLRCGGFIFGQRFCHCVVDAPGGMQFE 180
Query: 260 NAWAEMARGAHKPSIQPVWNREILMARDPPHITCNHHEY-EQIFSSNTIKEEDTTTLVHQ 318
A E+ARGA PS+ P W RE+ MARDPP + H EY E ++ + +V
Sbjct: 181 KAVCELARGAAAPSVSPSWGREMFMARDPPRPSYPHLEYREPAGGADRLLATPPEDMVRV 240
Query: 319 SFFFRTSDIVVLRLLVPFHLR-HCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARS 377
FFF +I LR P +R C+ F+L+ +C W RT AL +E+R+ IVNAR
Sbjct: 241 PFFFGPREIAGLRQHAPASVRGACSRFELVAACIWRSRTAALGYAPGEEVRLSFIVNARG 300
Query: 378 RFNANNSPL-VGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADL 436
R + PL G+YGN FAY A TTAG+LCG LGYA+ LV+K K+ VT EY+ SVADL
Sbjct: 301 RADV---PLPEGFYGNAFAYSVAATTAGELCGGDLGYALGLVKKAKSAVTYEYLQSVADL 357
Query: 437 MVIKERCLFTTIRSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPGAA-YIIPHKN 495
MV+ R LF R+ +VSDV+ + VDFGWGEAVYGG AK G GP G Y KN
Sbjct: 358 MVVAGRPLFALSRTYIVSDVSHAGFKSVDFGWGEAVYGGPAKGGEGPLLGVTNYFSRSKN 417
Query: 496 VEGEEGFILLVCLSFEAMKRFAKELDEM 523
+GE+ ++ +CL +AM +F E+ +
Sbjct: 418 GKGEQSVVVPICLPKDAMDKFQLEVQAL 445
>UniRef100_Q9SRQ2 Putative hypersensitivity-related gene [Arabidopsis thaliana]
Length = 454
Score = 400 bits (1029), Expect = e-110
Identities = 214/445 (48%), Positives = 291/445 (65%), Gaps = 11/445 (2%)
Query: 85 TSAPLLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDP 144
T+ L F V R Q ELV PA PTPRE+K LSDIDDQ+GLRF IP++F YR S + DP
Sbjct: 11 TTTGLSFKVHRQQRELVTPAKPTPRELKPLSDIDDQQGLRFQIPVIFFYRPNLS-SDLDP 69
Query: 145 VKVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEF--GDALH 202
V+V++ AL+ ALVYYYPFAGR+RE + RKL VDCTGEGV+F+EAEADV L E DAL
Sbjct: 70 VQVIKKALADALVYYYPFAGRLRELSNRKLAVDCTGEGVLFIEAEADVALAELEEADALL 129
Query: 203 PPLPCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAW 262
PP P EELL+DV GS +++ P+ L+QVTRLKC FI + NHTM+DGAGL LF+ +
Sbjct: 130 PPFPFLEELLFDVEGSSDVLNTPLLLVQVTRLKCCGFIFALRFNHTMTDGAGLSLFLKSL 189
Query: 263 AEMARGAHKPSIQPVWNREIL-MARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFF 321
E+A G H PS+ PVWNR +L ++ +T H EY+ + + LV +SFF
Sbjct: 190 CELACGLHAPSVPPVWNRHLLTVSASEARVTHTHREYDDQVGIDVVATGH--PLVSRSFF 247
Query: 322 FRTSDIVVLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNA 381
FR +I +R L+P L H T+F+ ++S W CRT AL + + E+R+ CI+N+RS+
Sbjct: 248 FRAEEISAIRKLLPPDL-HNTSFEALSSFLWRCRTIALNPDPNTEMRLTCIINSRSKL-- 304
Query: 382 NNSPL-VGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIK 440
N PL GYYGN F PAA+ TA L L +A+ L+++ K+ VTE+Y+ SV LM +
Sbjct: 305 RNPPLEPGYYGNVFVIPAAIATARDLIEKPLEFALRLIQETKSSVTEDYVRSVTALMATR 364
Query: 441 ERCLFTTIRSCVVSDVTRIKLREVDFG-WGEAVYGGAAKSGAGPFPGAAYIIPHKNVEGE 499
R +F + ++SD+ L ++DFG WG+ VYGG AK+G FPG ++ +P KN +GE
Sbjct: 365 GRPMFVASGNYIISDLRHFDLGKIDFGPWGKPVYGGTAKAGIALFPGVSFYVPFKNKKGE 424
Query: 500 EGFILLVCLSFEAMKRFAKELDEML 524
G ++ + L AM+ F EL+ +L
Sbjct: 425 TGTVVAISLPVRAMETFVAELNGVL 449
>UniRef100_Q8LFV6 Putative hypersensitivity-related gene [Arabidopsis thaliana]
Length = 454
Score = 399 bits (1025), Expect = e-109
Identities = 213/445 (47%), Positives = 290/445 (64%), Gaps = 11/445 (2%)
Query: 85 TSAPLLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDP 144
T+ L F V R Q ELV PA PTPRE+K LSDIDDQ+GLRF IP++F YR S + DP
Sbjct: 11 TTTGLSFKVHRQQRELVTPAKPTPRELKPLSDIDDQQGLRFQIPVIFFYRPNLS-SDLDP 69
Query: 145 VKVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEF--GDALH 202
V+V++ AL+ ALVYYYPFAGR+RE + RKL VDCTGEGV+F+EAEADV L E DAL
Sbjct: 70 VQVIKKALADALVYYYPFAGRLRELSNRKLAVDCTGEGVLFIEAEADVALAELEEADALL 129
Query: 203 PPLPCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAW 262
PP P EELL+DV GS +++ P+ L+QVTRLKC FI + NHTM+DGAGL LF+ +
Sbjct: 130 PPFPFLEELLFDVEGSSDVLNTPLLLVQVTRLKCCGFIFALRFNHTMTDGAGLSLFLKSL 189
Query: 263 AEMARGAHKPSIQPVWNREIL-MARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFF 321
E+A G H PS+ PVWNR +L ++ +T H EY+ + + LV +SFF
Sbjct: 190 CELACGLHAPSVPPVWNRHLLTVSASEARVTHTHREYDDQVGIDVVATGH--PLVSRSFF 247
Query: 322 FRTSDIVVLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNA 381
FR +I +R L+P L H T+F+ ++S W CRT AL + + E+R+ CI+N+RS+
Sbjct: 248 FRAEEISAIRKLLPPDL-HNTSFEALSSFLWRCRTIALNPDPNTEMRLTCIINSRSKL-- 304
Query: 382 NNSPL-VGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIK 440
N PL GYYGN F PAA+ TA L L + + L+++ K+ VTE+Y+ SV LM +
Sbjct: 305 RNPPLEPGYYGNVFVIPAAIATARDLIEKPLEFVLRLIQETKSSVTEDYVRSVTALMATR 364
Query: 441 ERCLFTTIRSCVVSDVTRIKLREVDFG-WGEAVYGGAAKSGAGPFPGAAYIIPHKNVEGE 499
R +F + ++SD+ L ++DFG WG+ VYGG AK+G FPG ++ +P KN +GE
Sbjct: 365 GRPMFVASGNYIISDLRHFDLGKIDFGPWGKPVYGGTAKAGIALFPGVSFYVPFKNKKGE 424
Query: 500 EGFILLVCLSFEAMKRFAKELDEML 524
G ++ + L AM+ F EL+ +L
Sbjct: 425 TGTVVAISLPVRAMETFVAELNGVL 449
>UniRef100_Q84S99 Alcohol acyltransferase [Cucumis melo]
Length = 461
Score = 385 bits (990), Expect = e-105
Identities = 199/446 (44%), Positives = 279/446 (61%), Gaps = 17/446 (3%)
Query: 73 THTTLIFSYHIMTSAPLLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFI 132
T T+ FS+H VR+ QPEL+ PA PTP E K LSD+DDQ+ LR +P + I
Sbjct: 3 TMQTIDFSFH----------VRKCQPELIAPANPTPYEFKQLSDVDDQQSLRLQLPFVNI 52
Query: 133 YRHEPSMKEKDPVKVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADV 192
Y H PS++ +DPVKV++ A+ +ALV YYP AGR+REG GRKL V+CTGEG++F+EA+ADV
Sbjct: 53 YPHNPSLEGRDPVKVIKEAIGKALVLYYPLAGRLREGPGRKLFVECTGEGILFIEADADV 112
Query: 193 TLDEFGDALHPPLPCFE-ELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSD 251
+L++F D L L E ++++ S+ +++ + LIQVTRLKCG FI + NHTM+D
Sbjct: 113 SLEQFRDTLPYSLSSMENNIIHNALNSDGVLNSQLLLIQVTRLKCGGFIFGIRFNHTMAD 172
Query: 252 GAGLKLFMNAWAEMARGAHKPSIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEED 311
G G+ FM A AE+ARGA PSI PVW R +L AR IT H+EY+Q+ + +
Sbjct: 173 GFGIAQFMKAIAEIARGAFAPSILPVWQRALLTARYLSEITVRHYEYDQVVDTKSTL-IP 231
Query: 312 TTTLVHQSFFFRTSDIVVLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMC 371
++ + FFF I LR +P H C++F+++ C W RT QL+ ++++R +C
Sbjct: 232 ANNMIDRLFFFSQLQISTLRQTLPAHRHDCSSFEVLADCVWRLRTIGFQLKPEEDVRFLC 291
Query: 372 IVNARSRFNANNSPLVGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMH 431
+VN RS+ + PL GYYGN +PA +TTA KLCGN LGYAV+L+RK KA+ T
Sbjct: 292 VVNLRSKIDI---PL-GYYGNAVVFPAVITTAAKLCGNPLGYAVDLIRKAKAKATTHLTS 347
Query: 432 SVADLMVIKERCLFTTIRSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPG-AAYI 490
+ D MVIK R FT + ++SD+TRI VDFGWG+A++GG G G G +Y
Sbjct: 348 PMVDFMVIKGRPRFTELMRFMMSDITRIGFENVDFGWGKAIFGGPIIGGCGIIRGMISYS 407
Query: 491 IPHKNVEGEEGFILLVCLSFEAMKRF 516
I N + ++ +CL M+ F
Sbjct: 408 IAFMNRHNKNSLVVPLCLPPPDMESF 433
>UniRef100_Q8GV03 Acyltransferase 2 [Capsicum chinense]
Length = 453
Score = 364 bits (935), Expect = 3e-99
Identities = 189/438 (43%), Positives = 274/438 (62%), Gaps = 6/438 (1%)
Query: 89 LLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVKVL 148
+ +++ +P+LV P+ TPRE K LSDIDDQ R IP++ Y++ M+ KDP K++
Sbjct: 11 MAISIKHHKPKLVVPSIVTPRETKHLSDIDDQGSARLQIPILMFYKYNSLMEGKDPAKLI 70
Query: 149 RHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCF 208
+ LS+ L +YYP AGR+ EG RKLMV+C EGV+FVEA+A+V L++ GD++ PP P
Sbjct: 71 KDGLSKTLSFYYPLAGRLIEGPNRKLMVNCNSEGVLFVEADANVELEKLGDSIKPPCPYL 130
Query: 209 EELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARG 268
+ LL++VPGS+ II P+ L+QVTR CG F + + LNHTM DG GL +F+NA +E+ +G
Sbjct: 131 DLLLHNVPGSDGIIGCPLLLVQVTRFSCGGFAVGLRLNHTMMDGYGLNMFLNALSELIQG 190
Query: 269 AHKPSIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFFFRTSDIV 328
A PSI PVW R +L AR P ITC HHE+++ S E L+ +SFFF ++
Sbjct: 191 ASAPSILPVWERHLLSARSLPSITCTHHEFDEQIESKIAWESMEDKLIQKSFFFGKKEME 250
Query: 329 VLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPLVG 388
++ V + T F+L+ + W CRT AL L ++ +R+ ++N R + P G
Sbjct: 251 AIKNQVSPNC-ESTKFELLMAFLWKCRTIALGLHPEEIVRLTYLINIRGKLQKFELP-AG 308
Query: 389 YYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCLFTTI 448
YYGN F P AV+ AG LC NSL YAVELV+K+K + EEY+ S+ DLM IK R T
Sbjct: 309 YYGNAFVTPTAVSKAGLLCSNSLTYAVELVKKVKDHMNEEYIKSLIDLMAIKGRPELTKS 368
Query: 449 RSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPGAAYIIPHKNVEGEEGFILLVCL 508
+ +VSD I L EV FGWG ++GG AK+ ++ +P KN +G++G ++ + L
Sbjct: 369 WNFIVSDNRSIGLDEVGFGWGRPIFGGVAKA----ISFISFGVPVKNDKGDKGILIAISL 424
Query: 509 SFEAMKRFAKELDEMLGK 526
AMK+F + + ++ K
Sbjct: 425 PPMAMKKFEEVVYKVSSK 442
>UniRef100_Q6QLX4 Alcohol acyl transferase [Lycopersicon esculentum]
Length = 442
Score = 353 bits (906), Expect = 7e-96
Identities = 180/427 (42%), Positives = 265/427 (61%), Gaps = 5/427 (1%)
Query: 89 LLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVKVL 148
L ++ +P+LV P++ T E K LS+IDDQ +R IP++ Y++ SMK KD K++
Sbjct: 5 LPISINYHKPKLVVPSSVTSHETKRLSEIDDQGFIRLQIPILMFYKYNSSMKGKDLAKII 64
Query: 149 RHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCF 208
+ LS+ LV+YYP AGR+ EG +KLMV+C GEGV+F+E +A++ L++ G+++ PP P
Sbjct: 65 KDGLSKTLVFYYPLAGRLIEGPNKKLMVNCNGEGVLFIEGDANIELEKLGESIKPPCPYL 124
Query: 209 EELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARG 268
+ LL++V GS+ II P+ LIQVTR CG F + NHTM D G K+F+NA +E+ +G
Sbjct: 125 DLLLHNVHGSDGIIGSPLLLIQVTRFTCGGFAVGFRFNHTMMDAYGFKMFLNALSELIQG 184
Query: 269 AHKPSIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFFFRTSDIV 328
A PSI PVW R +L AR P ITC HHE+++ S E L+ QSFFF ++
Sbjct: 185 ASTPSILPVWERHLLSARSSPSITCIHHEFDEEIESKIAWESMEDKLIQQSFFFGNEEME 244
Query: 329 VLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPLVG 388
V++ VP + CT F+L+ + W CRT AL L +D+ +R+ ++N R + + N +G
Sbjct: 245 VIKNQVPPNY-ECTKFELLMAFLWKCRTIALNLHSDEIVRLTYVINIRGKKSLNIELPIG 303
Query: 389 YYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCLFTTI 448
YYGN F P V+ AG LC N + YAVEL++K+K + EEY+ S+ DLMV K R T
Sbjct: 304 YYGNAFITPVVVSKAGLLCSNPVTYAVELIKKVKDHINEEYIKSLIDLMVTKGRPELTKS 363
Query: 449 RSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPGAAYIIPHKNVEGEEGFILLVCL 508
+ +VSD I E DFGWG ++GG K+ ++ + KN +GE+G ++ + L
Sbjct: 364 WNFLVSDNRYIGFDEFDFGWGNPIFGGILKA----ISFTSFGVSVKNDKGEKGVLIAISL 419
Query: 509 SFEAMKR 515
AMK+
Sbjct: 420 PPLAMKK 426
>UniRef100_Q7X880 OSJNBa0043L24.8 protein [Oryza sativa]
Length = 449
Score = 343 bits (880), Expect = 7e-93
Identities = 193/435 (44%), Positives = 256/435 (58%), Gaps = 16/435 (3%)
Query: 86 SAPLLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPV 145
S+ L FT RR PELV PA PTPR ++ LSDIDDQ RF +++ YR DP
Sbjct: 4 SSSLAFTARRGDPELVAPAGPTPRGLRRLSDIDDQRSFRFYRSIIYFYRSGGG----DPA 59
Query: 146 KVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPL 205
+V+R AL+ ALV+YYP AGRIRE G KL+VDCTGEGV FVEA+ADV+L+EFGD+L PP+
Sbjct: 60 RVIRGALAAALVHYYPIAGRIRELPGGKLVVDCTGEGVSFVEADADVSLEEFGDSLCPPI 119
Query: 206 PCFEELL-YDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAE 264
PC ELL S ++ DRP+ +QVTRL+CG F+ + H + D AG+ F A E
Sbjct: 120 PCAGELLTLPESNSAVVTDRPLLYVQVTRLRCGGFVFGTQICHNLVDAAGITQFWQAVGE 179
Query: 265 MARGAHKPSIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFFFRT 324
+A+GA PS++PVW RE+L AR PP +H EYE + K LVH+ F F
Sbjct: 180 LAQGAAAPSVRPVWARELLDARHPPRPAYDHPEYEPASDEASDKLRPGDELVHRRFLFGP 239
Query: 325 SDIVVLRLLVPFHL-RHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANN 383
D+ LR +P L C+ F L+++ W CRT AL DE+R M +VN R R +
Sbjct: 240 DDVAALRGQLPARLGPRCSRFLLLSAFTWRCRTAALGYAPGDEVRFMFVVNGRGRGHGGR 299
Query: 384 SPLVGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAE-VTEEYMHSVADLMVIKER 442
G+YGN + A TTAG+LC L AVEL+ +A + + Y S AD +V++ R
Sbjct: 300 PLPEGFYGNALTFGVARTTAGELCSGPLSRAVELIAAARARTMADGYAQSAADAVVLRGR 359
Query: 443 CLFTTIRSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPGAAYIIPHKNVEGEEGF 502
FTT R+ +V+D+T+ L EVD GWG ++GG A + F H G G
Sbjct: 360 RRFTTARTYLVTDLTKSPLHEVDLGWGRPLFGGPATTTLATF--------HMPARG-GGI 410
Query: 503 ILLVCLSFEAMKRFA 517
+ +CL AM+RFA
Sbjct: 411 AVPMCLPPRAMERFA 425
>UniRef100_Q8H5Q7 Putative benzoyl coenzyme A [Oryza sativa]
Length = 451
Score = 343 bits (879), Expect = 9e-93
Identities = 197/443 (44%), Positives = 265/443 (59%), Gaps = 19/443 (4%)
Query: 89 LLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVKVL 148
L F VRR ELV PAAPTPRE K LSD+DD E LR+ +P++F+YR PS DPV +
Sbjct: 3 LSFAVRRRATELVAPAAPTPRETKRLSDVDDPESLRWQVPVVFVYR--PSAAAADPVDTI 60
Query: 149 RHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDA-LHPPLPC 207
R AL+ ALV YYPFAGR+RE GRKL+VDCTGEGVMFVEA+ADV + E A L P PC
Sbjct: 61 RRALAAALVPYYPFAGRLREVEGRKLVVDCTGEGVMFVEADADVRVVELEAAGLRAPFPC 120
Query: 208 FEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMAR 267
++LL+DV GS ++ P+ LIQVTRL CG F+L + LNH M D +G+ FM+A A++AR
Sbjct: 121 MDQLLFDVDGSAAVLGTPLLLIQVTRLLCGGFVLGIRLNHAMCDASGIVQFMDAVADLAR 180
Query: 268 GAHKPSIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEE--DTTTLVHQSFFFRTS 325
GA +P++ P W+RE+L AR PP + + EY ++ +V ++F F
Sbjct: 181 GAREPAVSPAWSRELLDARKPPKLAFHLREYNDFAAAPPAAPSVGALGDMVMRTFSFSPG 240
Query: 326 DIVVLRLLVPFHLR-HCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNS 384
D+ L+ +P HLR T+FD++ S W R +AL+ A ++ R+ IV+ R NN
Sbjct: 241 DVAALKGALPPHLRGRATSFDVLASFVWRARARALETPAGEDARLAIIVSFR-----NNG 295
Query: 385 PL---VGYYGN-CFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIK 440
L GYYGN C A+ SL VE VR+ K V EY+ SVAD +V++
Sbjct: 296 ELRLPRGYYGNVCVPVTVAMPAEALRRRGSLADVVEQVREAKKTVNAEYVRSVADTLVMR 355
Query: 441 ERCLFTTIRSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPGAAYIIPHKNVEGEE 500
R T ++SDV VDFGWGE VYGG + + + G +Y+I KN GE+
Sbjct: 356 GRPAIDTANLLLLSDVRLAGFHRVDFGWGEPVYGGPSHA----WYGVSYLIAVKNGAGED 411
Query: 501 GFILLVCLSFEAMKRFAKELDEM 523
G + V L AM+RF E++ +
Sbjct: 412 GVAVPVVLPAAAMERFTSEIERL 434
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.326 0.140 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 897,156,073
Number of Sequences: 2790947
Number of extensions: 37716761
Number of successful extensions: 98875
Number of sequences better than 10.0: 280
Number of HSP's better than 10.0 without gapping: 181
Number of HSP's successfully gapped in prelim test: 99
Number of HSP's that attempted gapping in prelim test: 97867
Number of HSP's gapped (non-prelim): 361
length of query: 528
length of database: 848,049,833
effective HSP length: 132
effective length of query: 396
effective length of database: 479,644,829
effective search space: 189939352284
effective search space used: 189939352284
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 77 (34.3 bits)
Medicago: description of AC148345.17


