

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148289.16 + phase: 0
(200 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
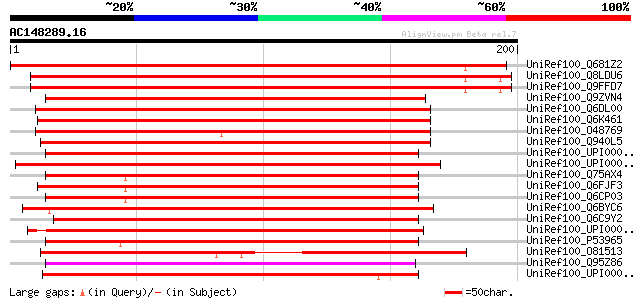
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q681Z2 Hypothetical protein At3g02800 [Arabidopsis tha... 279 3e-74
UniRef100_Q8LDU6 Hypothetical protein [Arabidopsis thaliana] 273 1e-72
UniRef100_Q9FFD7 Similarity to tyrosine phosphatase [Arabidopsis... 273 2e-72
UniRef100_Q9ZVN4 T7A14.14 protein [Arabidopsis thaliana] 204 1e-51
UniRef100_Q6DL00 Tyrosine-specific protein phosphatase protein [... 204 1e-51
UniRef100_Q6K461 Putative tyrosine specific protein phosphatase ... 202 5e-51
UniRef100_O48769 Hypothetical protein At2g32960 [Arabidopsis tha... 194 1e-48
UniRef100_Q940L5 Hypothetical protein At4g03960/T24M8_4 [Arabido... 192 6e-48
UniRef100_UPI00003C1F27 UPI00003C1F27 UniRef100 entry 170 2e-41
UniRef100_UPI000042EE51 UPI000042EE51 UniRef100 entry 164 2e-39
UniRef100_Q75AX4 ADL204Wp [Ashbya gossypii] 163 3e-39
UniRef100_Q6FJF3 Similar to sp|P53965 Saccharomyces cerevisiae Y... 163 3e-39
UniRef100_Q6CP03 Similarities with sp|P53965 Saccharomyces cerev... 162 4e-39
UniRef100_Q6BYC6 Similar to CA3369|IPF4672 Candida albicans IPF4... 162 5e-39
UniRef100_Q6C9Y2 Similar to sp|P53965 Saccharomyces cerevisiae Y... 161 1e-38
UniRef100_UPI00004302F1 UPI00004302F1 UniRef100 entry 159 3e-38
UniRef100_P53965 Hypothetical 32.8 kDa protein in NCE3-HHT2 inte... 156 3e-37
UniRef100_O81513 T24M8.4 protein [Arabidopsis thaliana] 147 2e-34
UniRef100_Q95Z86 Hypothetical protein L3291.05 [Leishmania major] 138 1e-31
UniRef100_UPI00003C1ED0 UPI00003C1ED0 UniRef100 entry 137 1e-31
>UniRef100_Q681Z2 Hypothetical protein At3g02800 [Arabidopsis thaliana]
Length = 203
Score = 279 bits (713), Expect = 3e-74
Identities = 131/197 (66%), Positives = 160/197 (80%), Gaps = 1/197 (0%)
Query: 1 MIVEVENIDEDDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPE 60
M + +E D + DVL PP NFSMVED IYRS P+P +F FL+TLNLRSIIYLCPEPYPE
Sbjct: 1 MCLIMETDDHNGDVLAPPSNFSMVEDGIYRSGFPRPENFSFLKTLNLRSIIYLCPEPYPE 60
Query: 61 ENLDFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTG 120
ENL FL+ NI+L+QFGIEGKT+ P +D++++ALKVLVDVRNHPIL+HCK+GKHRTG
Sbjct: 61 ENLKFLEANNIKLYQFGIEGKTDPPTPMPKDTVLDALKVLVDVRNHPILIHCKRGKHRTG 120
Query: 121 CLVGCFRKLQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLRQCLYSIIYQYQG-A 179
CLVGC RK+Q+WCLSS EEYQ+ AG+K R DL FIE FD+VSLRQCL SI+YQY G
Sbjct: 121 CLVGCLRKVQSWCLSSVLEEYQKNAGLKWRQRDLNFIEAFDIVSLRQCLLSIMYQYHGYG 180
Query: 180 SKKRRLMYQDENIQKPR 196
K+RRL Y++EN++ P+
Sbjct: 181 FKRRRLAYEEENVKTPK 197
>UniRef100_Q8LDU6 Hypothetical protein [Arabidopsis thaliana]
Length = 204
Score = 273 bits (699), Expect = 1e-72
Identities = 131/195 (67%), Positives = 156/195 (79%), Gaps = 5/195 (2%)
Query: 9 DEDDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKE 68
D D +VLIPPPNFSMVED IYRS P+ +F FL TLNLRSIIYLCPEPYPE+NL L
Sbjct: 8 DNDGEVLIPPPNFSMVEDGIYRSGFPELENFGFLSTLNLRSIIYLCPEPYPEDNLKSLAS 67
Query: 69 QNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRK 128
NI+LFQFGIEGKT+ P +D+++ AL+VLVDVRNHPIL+HCK+GKHRTGCLVGC RK
Sbjct: 68 NNIKLFQFGIEGKTDPPTPMPKDTVLSALRVLVDVRNHPILIHCKRGKHRTGCLVGCLRK 127
Query: 129 LQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLRQCLYSIIYQYQG-ASKKRRLMY 187
+QNWCLSS EEYQ+ AG+K R DL FIE FD++ L+QCLYSIIYQY G K+R+L+Y
Sbjct: 128 VQNWCLSSVLEEYQKCAGLKWRQRDLRFIEDFDVLRLKQCLYSIIYQYNGYGLKRRKLLY 187
Query: 188 QDENI----QKPRLT 198
Q+EN+ QKP+ T
Sbjct: 188 QEENVVQEQQKPQAT 202
>UniRef100_Q9FFD7 Similarity to tyrosine phosphatase [Arabidopsis thaliana]
Length = 204
Score = 273 bits (698), Expect = 2e-72
Identities = 131/195 (67%), Positives = 156/195 (79%), Gaps = 5/195 (2%)
Query: 9 DEDDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKE 68
D D +VLIPPPNFSMVED IYRS P+ +F FL TLNLRSIIYLCPEPYPE+NL L
Sbjct: 8 DNDGEVLIPPPNFSMVEDEIYRSGFPELENFGFLSTLNLRSIIYLCPEPYPEDNLKSLAS 67
Query: 69 QNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRK 128
NI+LFQFGIEGKT+ P +D+++ AL+VLVDVRNHPIL+HCK+GKHRTGCLVGC RK
Sbjct: 68 NNIKLFQFGIEGKTDPPTPMPKDTVLSALRVLVDVRNHPILIHCKRGKHRTGCLVGCLRK 127
Query: 129 LQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLRQCLYSIIYQYQG-ASKKRRLMY 187
+QNWCLSS EEYQ+ AG+K R DL FIE FD++ L+QCLYSIIYQY G K+R+L+Y
Sbjct: 128 VQNWCLSSVLEEYQKCAGLKWRQRDLRFIEDFDVLRLKQCLYSIIYQYNGYGLKRRKLLY 187
Query: 188 QDENI----QKPRLT 198
Q+EN+ QKP+ T
Sbjct: 188 QEENVVQEQQKPQAT 202
>UniRef100_Q9ZVN4 T7A14.14 protein [Arabidopsis thaliana]
Length = 215
Score = 204 bits (519), Expect = 1e-51
Identities = 97/150 (64%), Positives = 114/150 (75%)
Query: 15 LIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLF 74
LIPP NFSMV++ I+RS P ++F FLQTL LRSIIYLCPEPYPE NL FLK IRLF
Sbjct: 53 LIPPLNFSMVDNGIFRSGFPDSANFSFLQTLGLRSIIYLCPEPYPESNLQFLKSNGIRLF 112
Query: 75 QFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNWCL 134
QFGIEG E + I ALKVL+D +NHP+L+HCK+GKHRTGCLVGC RKLQ WCL
Sbjct: 113 QFGIEGNKEPFVNIPDHKIRMALKVLLDEKNHPVLIHCKRGKHRTGCLVGCLRKLQKWCL 172
Query: 135 SSAFEEYQRFAGVKSRAADLTFIERFDLVS 164
+S F+EYQRFA K+R +D F+E FD+ S
Sbjct: 173 TSIFDEYQRFAAAKARVSDQRFMEIFDVSS 202
>UniRef100_Q6DL00 Tyrosine-specific protein phosphatase protein [Oryza sativa]
Length = 225
Score = 204 bits (518), Expect = 1e-51
Identities = 92/156 (58%), Positives = 121/156 (76%)
Query: 11 DDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQN 70
++ L+PP NF+MV+D I+RS P ++F FL++LNLRSI+YLCPEPYPE N +FL +
Sbjct: 63 EEATLVPPLNFAMVDDGIFRSGFPAAANFRFLKSLNLRSIVYLCPEPYPETNAEFLAKNG 122
Query: 71 IRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQ 130
I+L QFGIEG+ E + D I EALKV++DV+N P+L+HCK+GKHRTGC+VGC RKLQ
Sbjct: 123 IKLHQFGIEGRKEPFVNIPDDKIREALKVVLDVKNQPLLIHCKRGKHRTGCVVGCLRKLQ 182
Query: 131 NWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLR 166
WCLSS F+EYQRFA K+R+ D F+E FD+ SL+
Sbjct: 183 KWCLSSVFDEYQRFAAAKARSTDQRFMELFDISSLK 218
>UniRef100_Q6K461 Putative tyrosine specific protein phosphatase protein [Oryza
sativa]
Length = 222
Score = 202 bits (513), Expect = 5e-51
Identities = 90/155 (58%), Positives = 119/155 (76%)
Query: 12 DDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNI 71
+ +L+PP NF+MV+ +YRS P S+ PF+++L LRS++ LCPEPYPE N +FL+ I
Sbjct: 60 EGLLVPPLNFAMVDHGVYRSGFPDISNLPFVESLRLRSVLCLCPEPYPEANQEFLRAHGI 119
Query: 72 RLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQN 131
RLFQFGI+G E + D I EALKV++DV NHP+L+HCK+GKHRTGC+VGC RKLQ
Sbjct: 120 RLFQFGIDGSKEPFVNIPEDRIREALKVVLDVANHPVLIHCKRGKHRTGCVVGCLRKLQR 179
Query: 132 WCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLR 166
WCL+S F+EYQRFA K+R +DL F+E FD+ SL+
Sbjct: 180 WCLTSIFDEYQRFAAAKARVSDLRFMELFDISSLK 214
>UniRef100_O48769 Hypothetical protein At2g32960 [Arabidopsis thaliana]
Length = 218
Score = 194 bits (492), Expect = 1e-48
Identities = 92/158 (58%), Positives = 116/158 (73%), Gaps = 2/158 (1%)
Query: 11 DDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQN 70
D+ LIPP NFSMV++ I+RS P ++F F++TL LRSII LCPEPYPE N+ FLK
Sbjct: 50 DELNLIPPLNFSMVDNGIFRSGFPDSANFSFIKTLGLRSIISLCPEPYPENNMQFLKSNG 109
Query: 71 IRLFQFGIEGKT--EVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRK 128
I LFQFGIEG E + L I EALKVL+D +NHP+L+HCK+GKHRTGCLVGC RK
Sbjct: 110 ISLFQFGIEGSKSKEPFVDILDQKIREALKVLLDEKNHPLLIHCKRGKHRTGCLVGCMRK 169
Query: 129 LQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLR 166
LQ WC++S +EY+RFA K+R +D F+E FD+ L+
Sbjct: 170 LQKWCITSILDEYKRFAAAKARVSDQRFLESFDVSGLK 207
>UniRef100_Q940L5 Hypothetical protein At4g03960/T24M8_4 [Arabidopsis thaliana]
Length = 198
Score = 192 bits (487), Expect = 6e-48
Identities = 88/154 (57%), Positives = 115/154 (74%)
Query: 13 DVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIR 72
++ +PP NF+MV++ I+RS P+P SF FLQ+L L+SIIYLCPE YPE N +F K I+
Sbjct: 27 ELFVPPLNFAMVDNGIFRSGFPEPVSFSFLQSLRLKSIIYLCPEAYPEVNREFAKSNGIQ 86
Query: 73 LFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNW 132
+FQFGIE E + + I EAL+VL+D NHP+L+HCK GKHRTGCLVGC RK+Q W
Sbjct: 87 VFQFGIERCKEPFVNIPDEVIREALQVLLDTENHPVLIHCKSGKHRTGCLVGCVRKIQRW 146
Query: 133 CLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLR 166
CLSS F+EYQRFA K+R +D F+E FD+ +L+
Sbjct: 147 CLSSIFDEYQRFAAAKARISDQRFMELFDISNLK 180
>UniRef100_UPI00003C1F27 UPI00003C1F27 UniRef100 entry
Length = 569
Score = 170 bits (430), Expect = 2e-41
Identities = 80/147 (54%), Positives = 100/147 (67%)
Query: 15 LIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLF 74
L+PP NF+MV +YRSS PK FPFL+TL LRS++ L E YPE N FL + I F
Sbjct: 361 LLPPDNFAMVNSHVYRSSFPKKKHFPFLRTLGLRSVLTLILEEYPETNSTFLDQNGITFF 420
Query: 75 QFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNWCL 134
QFGI G E + D I AL ++D RNHPIL+HC +GKHRTGCL+GC RKLQ W L
Sbjct: 421 QFGIPGNKEPFVSIPTDKITSALVTILDRRNHPILIHCNKGKHRTGCLIGCLRKLQQWSL 480
Query: 135 SSAFEEYQRFAGVKSRAADLTFIERFD 161
++ F+EY+RF+ KSR+ D FIE +D
Sbjct: 481 TTIFDEYRRFSWPKSRSMDQEFIELYD 507
>UniRef100_UPI000042EE51 UPI000042EE51 UniRef100 entry
Length = 281
Score = 164 bits (414), Expect = 2e-39
Identities = 80/168 (47%), Positives = 110/168 (64%)
Query: 3 VEVENIDEDDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEEN 62
+E EN + D L PP NF+ V + IYRSS P+P++F FL+ L L+SI+ L PE YP
Sbjct: 104 LEYENCLDYDKPLTPPENFAPVINKIYRSSFPQPNNFAFLKKLKLKSILCLIPEDYPHLQ 163
Query: 63 LDFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCL 122
+F+K +NI+LFQ G+ G E + D I EA+K++++ N PIL+HC +GKHRTGCL
Sbjct: 164 QEFIKNENIKLFQLGMSGNKEPFVKISADLITEAVKIVLNPENQPILIHCNRGKHRTGCL 223
Query: 123 VGCFRKLQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLRQCLY 170
VG RKLQNW L+ F+EY++FA K R D FIE +D + + Y
Sbjct: 224 VGVIRKLQNWSLTLIFDEYRKFACPKERPMDQQFIELYDDTEILEYCY 271
>UniRef100_Q75AX4 ADL204Wp [Ashbya gossypii]
Length = 217
Score = 163 bits (412), Expect = 3e-39
Identities = 79/148 (53%), Positives = 102/148 (68%), Gaps = 1/148 (0%)
Query: 15 LIPPPNFSMVEDCIYRSSLPKPSSFPFLQT-LNLRSIIYLCPEPYPEENLDFLKEQNIRL 73
++PP NFS V IYRSS P+P +F FLQ + LRSI+ L PE YP EN +F++ I+L
Sbjct: 52 VVPPENFSPVVGEIYRSSFPRPENFAFLQERVRLRSILVLIPEEYPPENQEFVERAGIQL 111
Query: 74 FQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNWC 133
FQ G+ G E + RD + AL + +D NHPIL+HC +GKHRTGCLVGC RKLQNW
Sbjct: 112 FQVGMSGNKEPFVNIPRDVLTRALAIALDPANHPILIHCNRGKHRTGCLVGCIRKLQNWS 171
Query: 134 LSSAFEEYQRFAGVKSRAADLTFIERFD 161
L+ F+EY+RFA K+RA D FIE ++
Sbjct: 172 LTMIFDEYRRFAFPKARAMDQQFIEMYE 199
>UniRef100_Q6FJF3 Similar to sp|P53965 Saccharomyces cerevisiae YNL032w SIW14
[Candida glabrata]
Length = 280
Score = 163 bits (412), Expect = 3e-39
Identities = 79/151 (52%), Positives = 106/151 (69%), Gaps = 1/151 (0%)
Query: 12 DDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQT-LNLRSIIYLCPEPYPEENLDFLKEQN 70
D + PP NFS V IYRSS P+ +F FLQ L L+SI+ L PE YP+ENLDF+++ N
Sbjct: 112 DSEVTPPENFSHVVGEIYRSSFPRTENFAFLQKRLKLKSILVLIPEEYPQENLDFMEKAN 171
Query: 71 IRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQ 130
I+LFQ G+ G E + D + +AL+V+++ N PIL+HC +GKHRTGCL+GC RKLQ
Sbjct: 172 IKLFQVGMSGNKEPFVNIPSDLLTKALEVVLNPENQPILIHCNRGKHRTGCLIGCIRKLQ 231
Query: 131 NWCLSSAFEEYQRFAGVKSRAADLTFIERFD 161
+W L+ F+EY+RFA K+RA D FIE +D
Sbjct: 232 SWSLTMIFDEYRRFAFPKARALDQQFIEMYD 262
>UniRef100_Q6CP03 Similarities with sp|P53965 Saccharomyces cerevisiae YNL032w SIW14
[Kluyveromyces lactis]
Length = 274
Score = 162 bits (411), Expect = 4e-39
Identities = 80/148 (54%), Positives = 104/148 (70%), Gaps = 1/148 (0%)
Query: 15 LIPPPNFSMVEDCIYRSSLPKPSSFPFLQT-LNLRSIIYLCPEPYPEENLDFLKEQNIRL 73
+IPP NFS V IYRSS P+P +F FL+ L L+SI+ L PE YP EN+ F++E I+L
Sbjct: 109 VIPPENFSHVCGEIYRSSFPRPENFEFLRDRLKLKSILVLIPEEYPAENMKFMEETGIKL 168
Query: 74 FQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNWC 133
FQ G+ G E + D + +AL+V+++ NHPIL+HC +GKHRTGCLVGC RKLQNW
Sbjct: 169 FQVGMSGNKEPFVNIPSDLLTKALEVVLNPENHPILIHCNRGKHRTGCLVGCIRKLQNWS 228
Query: 134 LSSAFEEYQRFAGVKSRAADLTFIERFD 161
L+ F+EY+RFA K RA D FIE +D
Sbjct: 229 LTMIFDEYRRFAFPKVRALDQQFIELYD 256
>UniRef100_Q6BYC6 Similar to CA3369|IPF4672 Candida albicans IPF4672 unknown Function
[Debaryomyces hansenii]
Length = 270
Score = 162 bits (410), Expect = 5e-39
Identities = 79/164 (48%), Positives = 113/164 (68%), Gaps = 2/164 (1%)
Query: 6 ENIDEDDDV--LIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENL 63
E+ D D++ L PP NF+ V + IYRSS P+PS+FPF++ L L+SI+ L PE YPEE+
Sbjct: 94 ESGDFADEIPQLTPPENFAPVINKIYRSSFPQPSNFPFVKKLKLKSILCLIPEDYPEEHE 153
Query: 64 DFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLV 123
FL+++NI+LFQ G+ G E + + I EA+K++++ N PIL+HC +GKHRTGCLV
Sbjct: 154 QFLEKENIKLFQLGMSGNKEPFVKISHNLITEAIKIVLNPANQPILIHCNRGKHRTGCLV 213
Query: 124 GCFRKLQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLRQ 167
G R+LQ W L+ F+EY++FA K R D FIE ++ V L +
Sbjct: 214 GVLRRLQKWSLTIIFDEYRKFAAPKERPMDQQFIELYNEVELEE 257
>UniRef100_Q6C9Y2 Similar to sp|P53965 Saccharomyces cerevisiae YNL032w SIW14
[Yarrowia lipolytica]
Length = 290
Score = 161 bits (407), Expect = 1e-38
Identities = 74/144 (51%), Positives = 100/144 (69%)
Query: 18 PPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLFQFG 77
P NFS+V IYRSS P+P +F +L+ L L+SI+ L PE YP+ENL F+KE NI+ FQ G
Sbjct: 129 PENFSIVVGQIYRSSFPRPENFEYLKRLKLKSILVLIPEIYPDENLQFMKENNIQFFQVG 188
Query: 78 IEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNWCLSSA 137
+ G E + D I AL++ ++ NHP+L+HC +GKHRTGCL GC R+LQ+W L+
Sbjct: 189 MSGNKEPFVHVPHDVITRALEIAINPANHPLLIHCNRGKHRTGCLSGCIRRLQDWSLTMI 248
Query: 138 FEEYQRFAGVKSRAADLTFIERFD 161
F+EY+RFA K+R D FIE +D
Sbjct: 249 FDEYRRFAYPKARPLDQQFIELYD 272
>UniRef100_UPI00004302F1 UPI00004302F1 UniRef100 entry
Length = 279
Score = 159 bits (403), Expect = 3e-38
Identities = 74/156 (47%), Positives = 106/156 (67%), Gaps = 3/156 (1%)
Query: 8 IDEDDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLK 67
++ED L+PP NF++V +YR PK +F F++TL L++++ L E YP+ NL++ +
Sbjct: 75 VEED---LVPPENFALVSSGVYRCGFPKKRNFKFMETLRLKTVLTLVLEEYPKANLEWCQ 131
Query: 68 EQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFR 127
Q+I+ QFGI G E D I AL ++D RNHPIL+HC +GKHRTGCL+GC R
Sbjct: 132 SQDIQFMQFGIPGNKEPFDNIPEDVICAALVAILDRRNHPILIHCNKGKHRTGCLIGCIR 191
Query: 128 KLQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLV 163
+LQ W L+S F+EY+RF+ KSRA D FI+ FD++
Sbjct: 192 RLQAWSLTSIFDEYRRFSAPKSRAVDQQFIDLFDIM 227
>UniRef100_P53965 Hypothetical 32.8 kDa protein in NCE3-HHT2 intergenic region
[Saccharomyces cerevisiae]
Length = 281
Score = 156 bits (394), Expect = 3e-37
Identities = 76/148 (51%), Positives = 103/148 (69%), Gaps = 1/148 (0%)
Query: 15 LIPPPNFSMVEDCIYRSSLPKPSSFPFL-QTLNLRSIIYLCPEPYPEENLDFLKEQNIRL 73
+IPP NFS V IYRSS P+ +F FL + L L+SI+ L PE YP+ENL+FLK I+L
Sbjct: 116 VIPPENFSHVVGEIYRSSFPRQENFSFLHERLKLKSILVLIPEEYPQENLNFLKLTGIKL 175
Query: 74 FQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNWC 133
+Q G+ G E + + +AL+++++ N PIL+HC +GKHRTGCL+GC RKLQNW
Sbjct: 176 YQVGMSGNKEPFVNIPSHLLTKALEIVLNPANQPILIHCNRGKHRTGCLIGCIRKLQNWS 235
Query: 134 LSSAFEEYQRFAGVKSRAADLTFIERFD 161
L+ F+EY+RFA K+RA D FIE +D
Sbjct: 236 LTMIFDEYRRFAFPKARALDQQFIEMYD 263
>UniRef100_O81513 T24M8.4 protein [Arabidopsis thaliana]
Length = 233
Score = 147 bits (371), Expect = 2e-34
Identities = 79/172 (45%), Positives = 106/172 (60%), Gaps = 22/172 (12%)
Query: 13 DVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIR 72
++ +PP NF+MV++ I+RS P+P SF FLQ+L L+SIIYLCPE YPE N +F K I+
Sbjct: 27 ELFVPPLNFAMVDNGIFRSGFPEPVSFSFLQSLRLKSIIYLCPEAYPEVNREFAKSNGIQ 86
Query: 73 LFQFGIEG-KTEVSLPALR---DSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRK 128
+FQFGIE K + P + + I EAL +HRTGCLVGC RK
Sbjct: 87 VFQFGIERCKVRLVEPFVNIPDEVIREAL------------------QHRTGCLVGCVRK 128
Query: 129 LQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLRQCLYSIIYQYQGAS 180
+Q WCLSS F+EYQRFA K+R +D F+E FD+ +L+ S +G +
Sbjct: 129 IQRWCLSSIFDEYQRFAAAKARISDQRFMELFDISNLKHTPLSFSCSKRGVN 180
>UniRef100_Q95Z86 Hypothetical protein L3291.05 [Leishmania major]
Length = 300
Score = 138 bits (347), Expect = 1e-31
Identities = 67/146 (45%), Positives = 86/146 (58%)
Query: 15 LIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLF 74
L+P NFSMV +YRS P ++ FL L LRSI+YLCPE Y E NL F +E I +
Sbjct: 11 LVPSINFSMVCPGVYRSGYPTKKNYSFLCALRLRSILYLCPEDYAESNLKFCEENGIHVL 70
Query: 75 QFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNWCL 134
+F EG E + L + D RN P+L+HC +GKHRTG +V C R LQ W L
Sbjct: 71 RFPTEGNKEPFCDVSEPLMHRILSAICDTRNLPLLIHCNKGKHRTGTVVACLRHLQGWSL 130
Query: 135 SSAFEEYQRFAGVKSRAADLTFIERF 160
S FEEY+RFA K+R D ++E +
Sbjct: 131 VSIFEEYKRFASDKARVGDQQYVELY 156
>UniRef100_UPI00003C1ED0 UPI00003C1ED0 UniRef100 entry
Length = 158
Score = 137 bits (346), Expect = 1e-31
Identities = 70/149 (46%), Positives = 93/149 (61%), Gaps = 1/149 (0%)
Query: 14 VLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRL 73
+L+PPPN+ MVE+ YRS P +FPFL+ L L+S+I+L PE LDF +QNI L
Sbjct: 1 MLVPPPNYGMVEENFYRSGQPDQLNFPFLEKLGLKSVIWLAPEEPEPGFLDFCVDQNIEL 60
Query: 74 FQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNWC 133
G+ T P + +++AL +LV +P+LV C G+HRTG +VGCFRKLQ W
Sbjct: 61 HHLGVLYSTNAWDPITEEVVLQALHLLVQPATYPVLVMCNLGRHRTGTVVGCFRKLQRWN 120
Query: 134 LSSAFEEYQRF-AGVKSRAADLTFIERFD 161
LS+ EEY+RF G K R + FIE FD
Sbjct: 121 LSAILEEYRRFVGGQKYRILNEQFIELFD 149
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.141 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 329,412,875
Number of Sequences: 2790947
Number of extensions: 12955320
Number of successful extensions: 28693
Number of sequences better than 10.0: 264
Number of HSP's better than 10.0 without gapping: 138
Number of HSP's successfully gapped in prelim test: 127
Number of HSP's that attempted gapping in prelim test: 28441
Number of HSP's gapped (non-prelim): 316
length of query: 200
length of database: 848,049,833
effective HSP length: 121
effective length of query: 79
effective length of database: 510,345,246
effective search space: 40317274434
effective search space used: 40317274434
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 71 (32.0 bits)
Medicago: description of AC148289.16


