

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148241.11 + phase: 0
(390 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
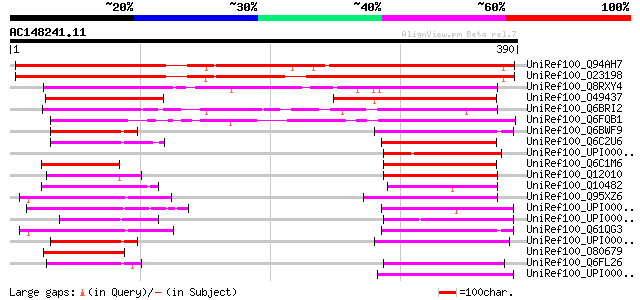
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q94AH7 Hypothetical protein At4g36850 [Arabidopsis tha... 412 e-114
UniRef100_O23198 Hypothetical protein AT4g36850 [Arabidopsis tha... 382 e-105
UniRef100_Q8RXY4 Hypothetical protein At2g41050 [Arabidopsis tha... 215 2e-54
UniRef100_O49437 Hypothetical protein AT4g20100 [Arabidopsis tha... 112 1e-23
UniRef100_Q6BRI2 Similar to CA4714|IPF5912 Candida albicans IPF5... 104 4e-21
UniRef100_Q6FQB1 Similar to tr|Q12010 Saccharomyces cerevisiae Y... 100 7e-20
UniRef100_Q6BWF9 Similar to CA5668|IPF1261 Candida albicans IPF1... 77 6e-13
UniRef100_Q6C2U6 Similar to tr|Q12010 Saccharomyces cerevisiae Y... 75 3e-12
UniRef100_UPI00003C26ED UPI00003C26ED UniRef100 entry 74 7e-12
UniRef100_Q6C1M6 Similar to sp|P38279 Saccharomyces cerevisiae Y... 74 7e-12
UniRef100_Q12010 S.cerevisiae chromosome XV reading frame ORF YO... 71 5e-11
UniRef100_Q10482 Seven transmembrane protein 1 [Schizosaccharomy... 70 1e-10
UniRef100_Q95XZ6 Hypothetical protein Y43H11AL.2 [Caenorhabditis... 69 3e-10
UniRef100_UPI0000438C63 UPI0000438C63 UniRef100 entry 68 4e-10
UniRef100_UPI000042E29B UPI000042E29B UniRef100 entry 67 7e-10
UniRef100_Q61QG3 Hypothetical protein CBG07028 [Caenorhabditis b... 67 7e-10
UniRef100_UPI000042C49A UPI000042C49A UniRef100 entry 66 1e-09
UniRef100_O80679 Hypothetical protein At2g41050 [Arabidopsis tha... 66 1e-09
UniRef100_Q6FL26 Candida glabrata strain CBS138 chromosome L com... 66 1e-09
UniRef100_UPI000021A472 UPI000021A472 UniRef100 entry 65 3e-09
>UniRef100_Q94AH7 Hypothetical protein At4g36850 [Arabidopsis thaliana]
Length = 392
Score = 412 bits (1059), Expect = e-114
Identities = 224/394 (56%), Positives = 271/394 (67%), Gaps = 29/394 (7%)
Query: 5 YCVKENKECVKWVETYFKDCLCNLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGI 64
YC+KE K CV+WVE YF DCLCNL DD+SF+LG+ SL+ WGVAEIPQ+IT FR KSS+G+
Sbjct: 6 YCLKEKKTCVRWVEIYFDDCLCNLNDDVSFALGIASLLCWGVAEIPQVITNFRTKSSNGV 65
Query: 65 SLAFLLTWVAGDICNLVGCLLEPATLPTQFYTALTVINCSFMQALQSYYYCKLYTMITSS 124
SL+FLL WVAGDI NLVGCLLEPATLPTQFYTAL + + +Q+ YY +Y +
Sbjct: 66 SLSFLLAWVAGDIFNLVGCLLEPATLPTQFYTALLYTVSTVVLVIQTIYYDYIYKL---- 121
Query: 125 DGANIVRMLRVNCYFYNCIILQDNEE--EKRPLNPKPSQVYSGIAIPNGTQKEAARGEYY 182
C I Q +EE EKRPL P P + S I+IP G+ K+++R E+Y
Sbjct: 122 ------------CRHRRTKICQKDEEDEEKRPLKP-PKTMGSAISIPGGSYKDSSRREFY 168
Query: 183 YMSARSLAGSATPPSFT-HLRAAKSGPSALEFIHD--SSDDDEASQVTSNIST---TKPW 236
Y SARSLAGS TPP T + R AKSGPSAL +D SSD+DE I+ TKP
Sbjct: 169 YTSARSLAGSGTPPLRTSYFRVAKSGPSALAIDNDGSSSDEDETMSTCPVITAKTITKPR 228
Query: 237 SIPRSVDGRYGTFLATAINLPLKGNSMRYGYIGFTGIKLLKVYVVVHSTYGQYLGWIMAA 296
IPR +GTFLA + +LPL+ S+ Y + +LL +V HS GQ+LGW+MAA
Sbjct: 229 PIPRQAG--FGTFLAASASLPLQAKSLAEKYAHASSRRLLNERIVEHSALGQWLGWLMAA 286
Query: 297 IYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESIKANLPWLLD 356
IY RIPQIWLNIKRGSVEGLNP MF+FAL+AN +YVGSILVRTTE+++IK NLPWLLD
Sbjct: 287 IYMGGRIPQIWLNIKRGSVEGLNPLMFIFALVANATYVGSILVRTTEWDNIKPNLPWLLD 346
Query: 357 ATVCVALDFFIISQYIYYRYFR--SSESSDDGEY 388
A VCV LD FII QYIYY+Y R S ES ++ Y
Sbjct: 347 AIVCVVLDLFIILQYIYYKYCRIKSLESREEDAY 380
>UniRef100_O23198 Hypothetical protein AT4g36850 [Arabidopsis thaliana]
Length = 374
Score = 382 bits (982), Expect = e-105
Identities = 208/389 (53%), Positives = 256/389 (65%), Gaps = 37/389 (9%)
Query: 5 YCVKENKECVKWVETYFKDCLCNLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGI 64
YC+KE K CV+WVE YF DCLCNL DD+SF+LG+ SL+ WGVAEIPQ+IT FR KSS+G+
Sbjct: 6 YCLKEKKTCVRWVEIYFDDCLCNLNDDVSFALGIASLLCWGVAEIPQVITNFRTKSSNGV 65
Query: 65 SLAFLLTWVAGDICNLVGCLLEPATLPTQFYTALTVINCSFMQALQSYYYCKLYTMITSS 124
SL+FLL WVAGDI NLVGCLLEPATLPTQFYTAL + + +Q+ YY +Y +
Sbjct: 66 SLSFLLAWVAGDIFNLVGCLLEPATLPTQFYTALLYTVSTVVLVIQTIYYDYIYKL---- 121
Query: 125 DGANIVRMLRVNCYFYNCIILQDNE--EEKRPLNPKPSQVYSGIAIPNGTQKEAARGEYY 182
C I Q +E EEKRPL P P + S I+IP G+ K+++R E+Y
Sbjct: 122 ------------CRHRRTKICQKDEEDEEKRPLKP-PKTMGSAISIPGGSYKDSSRREFY 168
Query: 183 YMSARSLAGSATPPSFT-HLRAAKSGPSALEFIHDSSDDDEASQVTSNISTTKPWSIPRS 241
Y SARSLAGS TPP T + R AKSG + S + + +
Sbjct: 169 YTSARSLAGSGTPPLRTSYFRVAKSGLRLWQ---------------STMMVRHRTKMRQC 213
Query: 242 VDGRYGTFLATAINLPLKGNSMRYGYIGFTGIKLLKVYVVVHSTYGQYLGWIMAAIYTCS 301
+GTFLA + +LPL+ S+ Y + +LL +V HS GQ+LGW+MAAIY
Sbjct: 214 RRAGFGTFLAASASLPLQAKSLAEKYAHASSRRLLNERIVEHSALGQWLGWLMAAIYMGG 273
Query: 302 RIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESIKANLPWLLDATVCV 361
RIPQIWLNIKRGSVEGLNP MF+FAL+AN +YVGSILVRTTE+++IK NLPWLLDA VCV
Sbjct: 274 RIPQIWLNIKRGSVEGLNPLMFIFALVANATYVGSILVRTTEWDNIKPNLPWLLDAIVCV 333
Query: 362 ALDFFIISQYIYYRYFR--SSESSDDGEY 388
LD FII QYIYY+Y R S ES ++ Y
Sbjct: 334 VLDLFIILQYIYYKYCRIKSLESREEDAY 362
>UniRef100_Q8RXY4 Hypothetical protein At2g41050 [Arabidopsis thaliana]
Length = 376
Score = 215 bits (547), Expect = 2e-54
Identities = 143/367 (38%), Positives = 203/367 (54%), Gaps = 32/367 (8%)
Query: 27 NLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLE 86
+ RD +S SLG++S++SWGVAEIPQI+T + KS+ G+S+ FL TW+ GDI NL+GCL+E
Sbjct: 6 SFRDGLSLSLGIISVISWGVAEIPQIMTNYSEKSTEGLSITFLTTWMIGDIFNLLGCLME 65
Query: 87 PATLPTQFYTALTVINCSFMQALQSYYYCKLYTMITSSDGANIVRMLRVNCYFYNCIILQ 146
PATLPTQFY AL + + +QS YY +Y + + +V R++ I+
Sbjct: 66 PATLPTQFYMALLYTVTTSVLYVQSIYYGHIYPRLKNRRD-QMVEAERIS------NIIS 118
Query: 147 DNEEEKRPLNPKPSQVYSGIAIP----NGTQKEAARG-EYYYMSARSLAGSATPPSFTHL 201
D + R N + G P G+Q+ + G E +Y SARSL+ S TPP+ + L
Sbjct: 119 DVKIPGRWRNSSDTTTCGGQTTPITMIPGSQRTSFTGRELFYTSARSLSSSHTPPAGSVL 178
Query: 202 RAAKSGPSALEFIHDSSDDDEASQVTSNISTTKPWSIPRSVDGRYGTFLATAINLPLKGN 261
+ + + + ++ + +TK SV GTF NL +
Sbjct: 179 AQRMARGYSEPTLEEPLLPEDVTH-----PSTKSLLCVVSVFLFLGTF--NLPNLLSESR 231
Query: 262 SMRYG-----YIGFTGIKLLKV---YVVVH-----STYGQYLGWIMAAIYTCSRIPQIWL 308
+M G ++ KLL+V V H S G +LGW MAAIY R+PQI L
Sbjct: 232 TMALGEGDRVFVVRAARKLLQVTSSNVAEHSGGESSRIGMFLGWAMAAIYMGGRLPQICL 291
Query: 309 NIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESIKANLPWLLDATVCVALDFFII 368
N++RG VEGLNP MF FAL+ N +YV SILV + E+ + NLPWL+DA CV LDF I+
Sbjct: 292 NMRRGHVEGLNPLMFFFALVGNMTYVASILVNSVEWLKLAPNLPWLVDAGGCVVLDFLIL 351
Query: 369 SQYIYYR 375
Q+ ++R
Sbjct: 352 LQFFHFR 358
>UniRef100_O49437 Hypothetical protein AT4g20100 [Arabidopsis thaliana]
Length = 288
Score = 112 bits (281), Expect = 1e-23
Identities = 57/128 (44%), Positives = 82/128 (63%), Gaps = 3/128 (2%)
Query: 250 LATAINLPLKGNSMRYGYIGFTGIKLLKVY---VVVHSTYGQYLGWIMAAIYTCSRIPQI 306
L+ + ++ L+G + G KLL+V + ++ G +LGW MAAIY R+PQI
Sbjct: 140 LSGSRSMDLRGKDRVFVVGGAGARKLLEVSSGNLGENNNIGMWLGWAMAAIYMGGRLPQI 199
Query: 307 WLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESIKANLPWLLDATVCVALDFF 366
+N++RG+VEGLNP MF FA I N +YV SILV + E+ I+ NLPWL+D+ C LDF
Sbjct: 200 CMNVRRGNVEGLNPLMFFFAFIGNVTYVASILVNSVEWSKIEPNLPWLVDSGGCAVLDFL 259
Query: 367 IISQYIYY 374
I+ Q+ Y+
Sbjct: 260 ILLQFFYF 267
Score = 109 bits (273), Expect = 1e-22
Identities = 49/91 (53%), Positives = 68/91 (73%)
Query: 28 LRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLEP 87
+RDD+S SLG++S++SW VAEIPQI+T + KS G+S+ FL TW+ GDI N+VGCL+EP
Sbjct: 2 IRDDLSLSLGIISVISWSVAEIPQIMTNYNQKSIEGVSITFLTTWMLGDIFNVVGCLMEP 61
Query: 88 ATLPTQFYTALTVINCSFMQALQSYYYCKLY 118
A+LP QFYTA+ + + +QS YY +Y
Sbjct: 62 ASLPVQFYTAVLYTLATLVLYVQSIYYGHIY 92
>UniRef100_Q6BRI2 Similar to CA4714|IPF5912 Candida albicans IPF5912 [Debaryomyces
hansenii]
Length = 352
Score = 104 bits (260), Expect = 4e-21
Identities = 96/362 (26%), Positives = 150/362 (40%), Gaps = 55/362 (15%)
Query: 26 CNLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLL 85
C+ IS SL ++SL SW A++PQI +RNKS+ GIS FLL W GD + CLL
Sbjct: 4 CSQFSFISSSLSILSLSSWVWAQLPQIYCNYRNKSAEGISPLFLLLWFMGDFLSFTSCLL 63
Query: 86 EPATLPTQFYTALTVINCSFMQALQSYYYCKLYTMITSSDGANIVRMLRVNCYFYNCIIL 145
L Q Y +L + C+ + YYY Y + NI +
Sbjct: 64 NDVVLKFQVYLSLFFL-CNDVTLCYQYYY---YNSVYPRKYGNIENL------------- 106
Query: 146 QDNEE----EKRPLNPKPSQVYSGIAIPNGTQKEAARGEYYYMSARSLAGSATPPSFTHL 201
D EE ++ +N N + S+ S + + PS T +
Sbjct: 107 -DTEERSKGDETMVNTDIKASEDDPVRANNASVHTTANAIHIRSSNSNSNNKA-PSHTLV 164
Query: 202 RAAKSGPSALEFIHDSSDDDEASQVTSNISTTKPWSIPRSVDGRYGTFLATAI--NLPLK 259
++ S SS+++ S + N K +I G+ L T++ +P+
Sbjct: 165 QSISSSA--------SSNNNSYSSINDNQQYLKTLAI--------GSILNTSVAAAMPVS 208
Query: 260 GNSMRYGYIGFTGIKLLKVYVVVHSTYGQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLN 319
+M + + I G +L W +Y SR PQ++ N KR SV+G++
Sbjct: 209 MGTMADDTLFSSSIN--------KEVLGVFLAWSCTCVYVASRCPQLYKNYKRKSVDGIS 260
Query: 320 PFMFVFALIANTSYVGSILVRTT-EFESIKAN-----LPWLLDATVCVALDFFIISQYIY 373
P +F AL+ N +Y SIL + K+N LP++L ++ V DF Q
Sbjct: 261 PILFGCALVGNLTYTLSILTSCDFVYGDAKSNFFVKELPYILGSSGTVVFDFLYFYQRFL 320
Query: 374 YR 375
Y+
Sbjct: 321 YK 322
>UniRef100_Q6FQB1 Similar to tr|Q12010 Saccharomyces cerevisiae YOL092w [Candida
glabrata]
Length = 309
Score = 100 bits (249), Expect = 7e-20
Identities = 92/366 (25%), Positives = 153/366 (41%), Gaps = 79/366 (21%)
Query: 32 ISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLEPATLP 91
+S G +S+ W V +PQI F KS+ G+SL F++ W+AGD+ NLVG +++ L
Sbjct: 14 VSGIAGSVSIACWVVVFVPQIYENFYRKSADGLSLLFVVLWLAGDVFNLVGAMMQHLLLT 73
Query: 92 TQFYTALTVINCSFMQALQSYYYCKLYTMITSSDGANIVRMLRVNCYFYNCIILQDNEEE 151
M L +YY T++D +L + C +Y DNEE+
Sbjct: 74 --------------MVILAAYY--------TAADV-----ILLIQCLWY------DNEEK 100
Query: 152 KRPLNPKPSQVYSGIAI--------PNGTQKEAARGEYYYMSARSLAGSATPPSFTHLRA 203
P++ P+ + + P T E RG ++RS A
Sbjct: 101 LDPIHFSPANPINENVLQDVFNEHQPLLTSGEPTRGSEAENNSRSQA------------- 147
Query: 204 AKSGPSALEFIHDSSDDDEASQVTSNISTTKPWSIPRSVDGRYGTFLATAINLPLKGNSM 263
I + DE SN+ + G +++ N P
Sbjct: 148 ----------IEALLEADEQKTKRSNLFNDFTIVALVILGGIVSWYVSYCANPPEP---- 193
Query: 264 RYGYIGFTGIKLLKVYVVVHSTYGQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMF 323
+G +++ + Q G++ A +Y SR+PQI LN KR S EG++ F
Sbjct: 194 ---IVGEPDVEM--------NMLAQSFGYLSAVLYLGSRVPQILLNFKRKSCEGISFLFF 242
Query: 324 VFALIANTSYVGSILVRTTEFESIKANLPWLLDATVCVALDFFIISQYIYYRYFRSSESS 383
+FA + NT+++ S+L +T++ + N WL+ ++ + +DF I Q+ Y + E
Sbjct: 243 LFACLGNTTFIISVLAISTDYRYLLVNASWLIGSSGTLVMDFIIFIQFFAYGTSKPIELP 302
Query: 384 DDGEYL 389
D E L
Sbjct: 303 RDEERL 308
>UniRef100_Q6BWF9 Similar to CA5668|IPF1261 Candida albicans IPF1261 unknown function
[Debaryomyces hansenii]
Length = 320
Score = 77.4 bits (189), Expect = 6e-13
Identities = 43/107 (40%), Positives = 58/107 (54%), Gaps = 2/107 (1%)
Query: 281 VVHSTYGQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVR 340
+V+ Q+ GW+ A Y SRIPQI LN KR S EG++ F+FA + N +YV SIL
Sbjct: 200 LVYDPLAQFFGWLCAVFYLGSRIPQILLNYKRKSCEGISFMFFLFACLGNLTYVISILAI 259
Query: 341 TTEFESIKANLPWLLDATVCVALDFFIISQYIYYRYFRSSESSDDGE 387
+ + N WL + +ALDF I Q+ Y SE S+D E
Sbjct: 260 DMSWYYLWVNSSWLAGSLGTLALDFTIFVQFFLYN--EDSEDSEDSE 304
Score = 56.6 bits (135), Expect = 1e-06
Identities = 28/67 (41%), Positives = 41/67 (60%), Gaps = 1/67 (1%)
Query: 32 ISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLEPATLP 91
+S G +SL W + PQI FR KSS G+SL F++ W+AGD+ N++G +L+ LP
Sbjct: 14 VSGITGSISLACWIIVFAPQIYENFRRKSSDGLSLLFIILWLAGDVFNVLGSILQ-GVLP 72
Query: 92 TQFYTAL 98
T A+
Sbjct: 73 TMIILAI 79
>UniRef100_Q6C2U6 Similar to tr|Q12010 Saccharomyces cerevisiae YOL092w [Yarrowia
lipolytica]
Length = 278
Score = 75.1 bits (183), Expect = 3e-12
Identities = 38/88 (43%), Positives = 53/88 (60%)
Query: 287 GQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFES 346
GQ+ GW+ AA Y SR+PQI LN +R S EG++ F+FA + N + V SIL++ T +
Sbjct: 189 GQFFGWLCAAFYLGSRVPQIVLNYERKSCEGISFMFFLFACLGNLTAVASILLKDTSRQY 248
Query: 347 IKANLPWLLDATVCVALDFFIISQYIYY 374
+ N WLL A + LDF I Q+ Y
Sbjct: 249 LIINASWLLGAIGTLFLDFVIFCQFWIY 276
Score = 54.3 bits (129), Expect = 6e-06
Identities = 29/88 (32%), Positives = 49/88 (54%), Gaps = 5/88 (5%)
Query: 32 ISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLEPATLP 91
+S +G +S+ W + PQI F+ +SS G+SL+F++ W+ GDI N++G +L+ +P
Sbjct: 20 LSGIMGCISIACWIIVFTPQIYENFKRQSSEGLSLSFVVIWLIGDIFNVLGAILQ-KIIP 78
Query: 92 TQFYTALTVINCSFMQALQSYYYCKLYT 119
T A+ + LQ C +YT
Sbjct: 79 TMIILAIYYTLADILLLLQ----CLVYT 102
>UniRef100_UPI00003C26ED UPI00003C26ED UniRef100 entry
Length = 353
Score = 73.9 bits (180), Expect = 7e-12
Identities = 36/91 (39%), Positives = 58/91 (63%), Gaps = 1/91 (1%)
Query: 288 QYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESI 347
Q GW+ A +Y SR+PQI N + EGL+ +FVFA+ N +YV SIL+++T+ + +
Sbjct: 239 QVAGWLSAILYLSSRVPQILKN-RTTKCEGLSLALFVFAVAGNLTYVASILLKSTQHDYL 297
Query: 348 KANLPWLLDATVCVALDFFIISQYIYYRYFR 378
+ WL+ + V LDF +++Q+I+YR R
Sbjct: 298 VESFSWLVGSLGTVFLDFIVLAQFIHYRKAR 328
Score = 43.1 bits (100), Expect = 0.013
Identities = 24/79 (30%), Positives = 39/79 (48%), Gaps = 5/79 (6%)
Query: 40 SLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLEPATLPTQFYTALT 99
S++ W + Q+ + +KS+ G+S+ F++ W+AGD N G + L T AL
Sbjct: 66 SIMGW----VSQMHQNYVSKSAEGLSITFVIIWLAGDALNAAGAWFQ-GLLWTMIVLALY 120
Query: 100 VINCSFMQALQSYYYCKLY 118
C + Q +YY K Y
Sbjct: 121 YCTCDCILIFQYWYYRKYY 139
>UniRef100_Q6C1M6 Similar to sp|P38279 Saccharomyces cerevisiae YBR147w [Yarrowia
lipolytica]
Length = 268
Score = 73.9 bits (180), Expect = 7e-12
Identities = 34/87 (39%), Positives = 56/87 (64%)
Query: 288 QYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESI 347
+ LG++ AA+Y +RIPQI N +R SVEGL+ F+F+L+ N +Y G IL+ +TE++ +
Sbjct: 169 EILGYLSAALYLLARIPQIIRNTRRKSVEGLSLLFFIFSLLGNLTYAGYILLFSTEWDYV 228
Query: 348 KANLPWLLDATVCVALDFFIISQYIYY 374
+PWL+ + + D I +Q+ Y
Sbjct: 229 VKYMPWLIGSLGTIVEDLIIFAQFAAY 255
Score = 51.2 bits (121), Expect = 5e-05
Identities = 24/60 (40%), Positives = 36/60 (60%)
Query: 25 LCNLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCL 84
+ LR ++ G SL W V +PQ+I +R +S+ GI++ F+L W+ GDI NL G L
Sbjct: 1 MSTLRTQLAGICGSTSLACWIVVLLPQLIEQWRLQSAEGIAIGFILVWLVGDITNLAGAL 60
>UniRef100_Q12010 S.cerevisiae chromosome XV reading frame ORF YOL092w [Saccharomyces
cerevisiae]
Length = 308
Score = 71.2 bits (173), Expect = 5e-11
Identities = 33/88 (37%), Positives = 56/88 (63%)
Query: 288 QYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESI 347
Q G++ A +Y SRIPQI LN KR S EG++ F+FA + NT+++ S++V + +++ +
Sbjct: 213 QIFGYLSALLYLGSRIPQILLNFKRKSCEGISFLFFLFACLGNTTFIFSVIVISLDWKYL 272
Query: 348 KANLPWLLDATVCVALDFFIISQYIYYR 375
N WL+ + + +DF I SQ+ Y+
Sbjct: 273 IMNASWLVGSIGTLFMDFVIFSQFFIYK 300
Score = 52.4 bits (124), Expect = 2e-05
Identities = 28/77 (36%), Positives = 42/77 (54%), Gaps = 4/77 (5%)
Query: 29 RDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGC----L 84
R +S G +S+ W + +PQI F KSS G+SL F++ W+AGD+ NL+G L
Sbjct: 10 RSTLSGISGSISISCWIIVFVPQIYENFYRKSSDGLSLLFVVLWLAGDVFNLMGAVMQHL 69
Query: 85 LEPATLPTQFYTALTVI 101
L + +YT +I
Sbjct: 70 LSTMIILAAYYTVADII 86
>UniRef100_Q10482 Seven transmembrane protein 1 [Schizosaccharomyces pombe]
Length = 271
Score = 70.1 bits (170), Expect = 1e-10
Identities = 40/87 (45%), Positives = 50/87 (56%), Gaps = 2/87 (2%)
Query: 291 GWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILV--RTTEFESIK 348
G I + +Y C+RIPQI N K S EGL+ FV A + NTSY SILV +
Sbjct: 182 GCISSVLYFCARIPQIIKNHKAKSTEGLSIIFFVLASVGNTSYAFSILVFPASDYLNYTY 241
Query: 349 ANLPWLLDATVCVALDFFIISQYIYYR 375
ANLPW+L A + LD +I Q+I YR
Sbjct: 242 ANLPWILGAFSTIFLDIYIFYQFIKYR 268
Score = 51.6 bits (122), Expect = 4e-05
Identities = 27/90 (30%), Positives = 49/90 (54%), Gaps = 1/90 (1%)
Query: 25 LCNLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCL 84
+ N+ ++S LG +SL W V IPQ++ ++N+S IS FL+ W+ GD N++G +
Sbjct: 10 MANILTELSSFLGALSLGCWVVLLIPQLLENYKNQSGESISDLFLIIWLIGDFFNVLGSI 69
Query: 85 LEPATLPTQFYTALTVINCSFMQALQSYYY 114
+ + +++ S + +Q YYY
Sbjct: 70 YGNVSSTVLVLSFYYIVSDSTL-LMQIYYY 98
Score = 33.9 bits (76), Expect = 8.0
Identities = 21/63 (33%), Positives = 34/63 (53%), Gaps = 3/63 (4%)
Query: 30 DDIS---FSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLE 86
DD++ F+ G +S V + A IPQII + KS+ G+S+ F + G+ L+
Sbjct: 172 DDLNAWPFTAGCISSVLYFCARIPQIIKNHKAKSTEGLSIIFFVLASVGNTSYAFSILVF 231
Query: 87 PAT 89
PA+
Sbjct: 232 PAS 234
>UniRef100_Q95XZ6 Hypothetical protein Y43H11AL.2 [Caenorhabditis elegans]
Length = 311
Score = 68.6 bits (166), Expect = 3e-10
Identities = 38/120 (31%), Positives = 61/120 (50%), Gaps = 4/120 (3%)
Query: 8 KENKEC---VKWVETYFKDCLCNLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGI 64
+E+ C ++W++ F DC+ + F +GL+SL W + PQ+ ++ K G+
Sbjct: 14 EEDANCTQGIQWIKDVFTDCVDTDLKLLGFIIGLISLALWLIPLFPQLWQNYKTKKCEGL 73
Query: 65 SLAFLLTWVAGDICNLVGCLLEPATLPTQFYTALTVINCSFMQALQSYYYCKLYTMITSS 124
SLAFL W+ GD CN++G +L P Q + I + Q YY K+Y T+S
Sbjct: 74 SLAFLFFWLVGDTCNMLGAILTNQQ-PIQKIIGVYYIIQDLVLWTQYGYYLKIYNRPTTS 132
Score = 66.6 bits (161), Expect = 1e-09
Identities = 37/103 (35%), Positives = 53/103 (50%)
Query: 273 IKLLKVYVVVHSTYGQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTS 332
+K+ ++ G +G + A Y RIPQI N + S EGL+ MF + AN +
Sbjct: 187 LKMWPIFTSYTDMLGYIIGSMAAVCYFGGRIPQIIKNYRHSSCEGLSLTMFYIIVAANFT 246
Query: 333 YVGSILVRTTEFESIKANLPWLLDATVCVALDFFIISQYIYYR 375
Y S+L+ TT + + +LPWL + C D IISQY YR
Sbjct: 247 YGISVLLATTSWLYLLRHLPWLAGSLGCCCFDAVIISQYYLYR 289
Score = 34.7 bits (78), Expect = 4.7
Identities = 28/100 (28%), Positives = 43/100 (43%), Gaps = 11/100 (11%)
Query: 287 GQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANT-SYVGSILVRTTEFE 345
G +G I A++ PQ+W N K EGL+ F L+ +T + +G+IL +
Sbjct: 42 GFIIGLISLALWLIPLFPQLWQNYKTKKCEGLSLAFLFFWLVGDTCNMLGAILTNQQPIQ 101
Query: 346 SIKANLPWLLDATVCVALDFFIISQYIYYR--YFRSSESS 383
I + D + +QY YY Y R + SS
Sbjct: 102 KI--------IGVYYIIQDLVLWTQYGYYLKIYNRPTTSS 133
>UniRef100_UPI0000438C63 UPI0000438C63 UniRef100 entry
Length = 295
Score = 68.2 bits (165), Expect = 4e-10
Identities = 36/106 (33%), Positives = 59/106 (54%), Gaps = 5/106 (4%)
Query: 287 GQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTT---- 342
G +G I + +Y CSR+PQI+ N +R S EGL+ F+F ++ NT+Y S+L++
Sbjct: 183 GFVIGSISSVLYLCSRLPQIYTNFQRKSTEGLSYFLFALVILGNTTYGVSVLLKNPDPGQ 242
Query: 343 -EFESIKANLPWLLDATVCVALDFFIISQYIYYRYFRSSESSDDGE 387
E + +LPWL+ + ++LD I Q++ YR + D E
Sbjct: 243 GEASYMVHHLPWLIGSLGTLSLDLVISVQFMMYRKSPQVSADTDEE 288
Score = 60.5 bits (145), Expect = 8e-08
Identities = 45/125 (36%), Positives = 65/125 (52%), Gaps = 5/125 (4%)
Query: 14 VKWVETYFKDCLCNLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKS-SHGISLAFLLTW 72
+ W+ +C + RD S LGL+S+V + V+ IPQ + + + +S+ FLL W
Sbjct: 14 IAWIWYGLGECAQDGRDVASVVLGLLSIVCFMVSSIPQYYSSCKTGNMDSALSIWFLLFW 73
Query: 73 VAGDICNLVGCLLEPATLPTQFYTALTVINCSFMQALQSYYYCKLYTMITSSDGANIVRM 132
+AGD CNL+G L LP Q YTA+ + + L Y Y KL SSD A ++
Sbjct: 74 LAGDSCNLIGSFLAD-QLPLQKYTAVYYVVADLLM-LAMYMYYKLKNK-RSSDRA-LLNA 129
Query: 133 LRVNC 137
L V C
Sbjct: 130 LSVLC 134
>UniRef100_UPI000042E29B UPI000042E29B UniRef100 entry
Length = 256
Score = 67.4 bits (163), Expect = 7e-10
Identities = 39/100 (39%), Positives = 54/100 (54%), Gaps = 1/100 (1%)
Query: 288 QYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESI 347
Q LGW A +Y SRIPQI N K GL+ MF FA+ N +YV SIL+++ I
Sbjct: 150 QALGWASAVLYLGSRIPQIVHNYKTRCA-GLSLAMFFFAISGNITYVLSILLKSLNPRWI 208
Query: 348 KANLPWLLDATVCVALDFFIISQYIYYRYFRSSESSDDGE 387
AN PWL + + V LD F+++Q+ + + GE
Sbjct: 209 LANAPWLAGSGLTVFLDLFVLAQFAIFSWQDERRKKTSGE 248
Score = 51.2 bits (121), Expect = 5e-05
Identities = 27/76 (35%), Positives = 40/76 (52%), Gaps = 1/76 (1%)
Query: 39 MSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLEPATLPTQFYTAL 98
MS+ W V PQ+ ++ KS G+S+ F++ W+ GDI NL G ++ LPT A+
Sbjct: 1 MSIALWVVVYSPQMWENYQLKSGEGLSVPFIILWLLGDITNLFGGMM-AHLLPTVIILAV 59
Query: 99 TVINCSFMQALQSYYY 114
C + Q YYY
Sbjct: 60 YYTICDLVLLFQVYYY 75
>UniRef100_Q61QG3 Hypothetical protein CBG07028 [Caenorhabditis briggsae]
Length = 310
Score = 67.4 bits (163), Expect = 7e-10
Identities = 37/122 (30%), Positives = 61/122 (49%), Gaps = 4/122 (3%)
Query: 8 KENKEC---VKWVETYFKDCLCNLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGI 64
+E+ C ++W++ F DC+ + F +GL+SL W + PQ+ ++ K G+
Sbjct: 14 EEDANCTQGIQWIKNVFTDCVDTDLKLLGFIIGLISLALWLIPLFPQLWQNYKTKKCEGL 73
Query: 65 SLAFLLTWVAGDICNLVGCLLEPATLPTQFYTALTVINCSFMQALQSYYYCKLYTMITSS 124
SLAFL W+ GD CN++G +L P Q + I + Q YY K+Y ++
Sbjct: 74 SLAFLFFWLVGDTCNMLGAILTNQQ-PIQKIIGVYYIFQDLILWAQYGYYMKIYHRPATT 132
Query: 125 DG 126
G
Sbjct: 133 SG 134
Score = 64.3 bits (155), Expect = 6e-09
Identities = 39/101 (38%), Positives = 54/101 (52%), Gaps = 2/101 (1%)
Query: 287 GQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFES 346
G +G + A Y RIPQI N + S EGL+ MF + AN +Y S+L+ TT +
Sbjct: 200 GYLIGSVAALCYFGGRIPQIIKNYQHRSCEGLSLTMFYIIVAANFTYGISVLMATTSWLY 259
Query: 347 IKANLPWLLDATVCVALDFFIISQYIYYRYFRSSESSDDGE 387
+ +LPWL + C D IISQY YR + S++D E
Sbjct: 260 LLRHLPWLAGSLGCCCFDAVIISQYYLYR--PKTPSAEDTE 298
Score = 34.7 bits (78), Expect = 4.7
Identities = 27/100 (27%), Positives = 43/100 (43%), Gaps = 11/100 (11%)
Query: 287 GQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANT-SYVGSILVRTTEFE 345
G +G I A++ PQ+W N K EGL+ F L+ +T + +G+IL +
Sbjct: 42 GFIIGLISLALWLIPLFPQLWQNYKTKKCEGLSLAFLFFWLVGDTCNMLGAILTNQQPIQ 101
Query: 346 SIKANLPWLLDATVCVALDFFIISQYIYYR--YFRSSESS 383
I + D + +QY YY Y R + +S
Sbjct: 102 KI--------IGVYYIFQDLILWAQYGYYMKIYHRPATTS 133
>UniRef100_UPI000042C49A UPI000042C49A UniRef100 entry
Length = 331
Score = 66.2 bits (160), Expect = 1e-09
Identities = 36/104 (34%), Positives = 53/104 (50%)
Query: 281 VVHSTYGQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVR 340
++ Q GW+ A Y SRIPQI LN +R S +G++ F+FA + N +YV SIL
Sbjct: 205 LIFDPLAQIFGWLCAFFYLGSRIPQIVLNYERKSCDGISFMFFLFACLGNLTYVISILSI 264
Query: 341 TTEFESIKANLPWLLDATVCVALDFFIISQYIYYRYFRSSESSD 384
+ + N WL + + LDF I Q+ Y + E S+
Sbjct: 265 DMSWNYLWVNSSWLAGSLGTLGLDFTIFIQFFLYNENKDDEFSE 308
Score = 54.7 bits (130), Expect = 4e-06
Identities = 26/67 (38%), Positives = 41/67 (60%), Gaps = 1/67 (1%)
Query: 32 ISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLEPATLP 91
+S G +S+ W + PQI F+ KSS G+SL F++ W+AGD+ N++G +L+ LP
Sbjct: 16 VSGITGSISIACWIIVFAPQIYENFKRKSSEGLSLTFIVLWLAGDVFNVLGAVLQ-GVLP 74
Query: 92 TQFYTAL 98
T A+
Sbjct: 75 TMIILAV 81
>UniRef100_O80679 Hypothetical protein At2g41050 [Arabidopsis thaliana]
Length = 219
Score = 66.2 bits (160), Expect = 1e-09
Identities = 30/62 (48%), Positives = 43/62 (68%)
Query: 27 NLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLE 86
+ RD +S SLG++S++SWGVAEIPQI+T + KS+ G+S+ FL TW+ G +L
Sbjct: 6 SFRDGLSLSLGIISVISWGVAEIPQIMTNYSEKSTEGLSITFLTTWMIGSARSLSSSHTP 65
Query: 87 PA 88
PA
Sbjct: 66 PA 67
Score = 46.6 bits (109), Expect = 0.001
Identities = 49/150 (32%), Positives = 69/150 (45%), Gaps = 20/150 (13%)
Query: 185 SARSLAGSATPPSFTHLRAAKSGPSALEFIHDSSDDDEASQVTSNISTTKPWSIPRSVDG 244
SARSL+ S TPP+ + L + + + + ++ + +TK SV
Sbjct: 55 SARSLSSSHTPPAGSVLAQRMARGYSEPTLEEPLLPEDVTH-----PSTKSLLCVVSVFL 109
Query: 245 RYGTFLATAINLPLKGNSMRYG-----YIGFTGIKLLKVY---VVVHS-----TYGQYLG 291
GTF NL + +M G ++ KLL+V V HS G +LG
Sbjct: 110 FLGTF--NLPNLLSESRTMALGEGDRVFVVRAARKLLQVTSSNVAEHSGGESSRIGMFLG 167
Query: 292 WIMAAIYTCSRIPQIWLNIKRGSVEGLNPF 321
W MAAIY R+PQI LN++RG VE L F
Sbjct: 168 WAMAAIYMGGRLPQICLNMRRGHVEILLQF 197
>UniRef100_Q6FL26 Candida glabrata strain CBS138 chromosome L complete sequence
[Candida glabrata]
Length = 329
Score = 66.2 bits (160), Expect = 1e-09
Identities = 32/93 (34%), Positives = 53/93 (56%)
Query: 288 QYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESI 347
Q G++ A +Y SRIPQI LN KR S EG++ F+FA + N +++ S+L + + +
Sbjct: 237 QIFGYLSAVLYLGSRIPQILLNFKRKSCEGVSLLFFLFACLGNINFILSVLAVSVSKKYL 296
Query: 348 KANLPWLLDATVCVALDFFIISQYIYYRYFRSS 380
N WL+ + + LD I +Q+ Y + + S
Sbjct: 297 LVNASWLIGSAGTLLLDMTIFAQFFIYTHSKDS 329
Score = 55.8 bits (133), Expect = 2e-06
Identities = 31/78 (39%), Positives = 44/78 (55%), Gaps = 6/78 (7%)
Query: 29 RDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLEPA 88
R +S G +S+ W V PQI FR KSS G+SL F++ W+ GDI N+VG +++
Sbjct: 10 RRTVSEIAGSISIACWVVVFAPQIYENFRRKSSDGLSLMFIILWLIGDIFNIVGAIMQ-N 68
Query: 89 TLPTQ-----FYTALTVI 101
LPT +YT +I
Sbjct: 69 LLPTMIILAAYYTVADII 86
>UniRef100_UPI000021A472 UPI000021A472 UniRef100 entry
Length = 325
Score = 65.1 bits (157), Expect = 3e-09
Identities = 36/105 (34%), Positives = 58/105 (54%), Gaps = 1/105 (0%)
Query: 284 STYGQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTE 343
+ +G +G++ AA+Y +R+PQI+ N K S EGL F+ +L N +Y S++ + E
Sbjct: 218 TAFGLTMGYVSAALYLLARLPQIYKNYKEKSCEGLALLFFMLSLTGNLTYGVSLVAYSQE 277
Query: 344 FESIKANLPWLLDATVCVALDFFIISQY-IYYRYFRSSESSDDGE 387
I +PWLL + + D I Q+ +Y SS+SSD+ E
Sbjct: 278 KSYIIKTIPWLLGSLGTIVEDIVIFFQFRLYSTPKESSKSSDENE 322
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.137 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 661,650,650
Number of Sequences: 2790947
Number of extensions: 26837990
Number of successful extensions: 58710
Number of sequences better than 10.0: 119
Number of HSP's better than 10.0 without gapping: 98
Number of HSP's successfully gapped in prelim test: 21
Number of HSP's that attempted gapping in prelim test: 58362
Number of HSP's gapped (non-prelim): 322
length of query: 390
length of database: 848,049,833
effective HSP length: 129
effective length of query: 261
effective length of database: 488,017,670
effective search space: 127372611870
effective search space used: 127372611870
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC148241.11


