

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148241.10 - phase: 1 /pseudo
(611 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
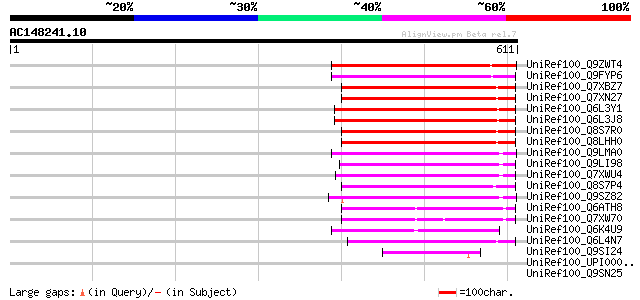
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9ZWT4 Transposase [Ipomoea purpurea] 230 1e-58
UniRef100_Q9FYP6 Putative transposable element Tip100 protein [O... 169 2e-40
UniRef100_Q7XBZ7 Putative transposase [Oryza sativa] 156 1e-36
UniRef100_Q7XN27 OSJNBb0016D16.6 protein [Oryza sativa] 156 1e-36
UniRef100_Q6L3Y1 Putative hAT family dimerisation domain contain... 156 1e-36
UniRef100_Q6L3J8 Putative transposase [Solanum demissum] 156 1e-36
UniRef100_Q8S7R0 Putative transposase protein, 5'-partial [Oryza... 156 1e-36
UniRef100_Q8LHH0 Putative transposase [Oryza sativa] 156 1e-36
UniRef100_Q9LMA0 T29M8.13 protein [Arabidopsis thaliana] 151 5e-35
UniRef100_Q9LI98 Similarity to transposase [Arabidopsis thaliana] 150 8e-35
UniRef100_Q7XWU4 OSJNBa0065B15.7 protein [Oryza sativa] 150 8e-35
UniRef100_Q8S7P4 Putative transposase [Oryza sativa] 150 1e-34
UniRef100_Q9SZ82 Hypothetical protein F17A8.10 [Arabidopsis thal... 146 2e-33
UniRef100_Q6ATH8 Hypothetical protein OSJNBa0093E24.3 [Oryza sat... 135 3e-30
UniRef100_Q7XW70 OSJNBa0019J05.24 protein [Oryza sativa] 132 4e-29
UniRef100_Q6K4U9 Putative hAT dimerisation domain-containing pro... 100 2e-19
UniRef100_Q6L4N7 Hypothetical protein P0473H02.5 [Oryza sativa] 88 8e-16
UniRef100_Q9SI24 Hypothetical protein At2g05020 [Arabidopsis tha... 72 6e-11
UniRef100_UPI0000248927 UPI0000248927 UniRef100 entry 45 0.006
UniRef100_Q9SN25 Hypothetical protein F28M11.120 [Arabidopsis th... 42 0.053
>UniRef100_Q9ZWT4 Transposase [Ipomoea purpurea]
Length = 808
Score = 230 bits (586), Expect = 1e-58
Identities = 111/222 (50%), Positives = 163/222 (73%), Gaps = 1/222 (0%)
Query: 389 MTASREVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQV 448
+ +S+EV +H+FF KL ++NVVG+S KR+D+L+AA + I +LL E+ +G+G NQ+
Sbjct: 410 VASSKEVIPVHQFFTKLNSIINVVGASCKRNDQLKAAHASNISHLLSIDELESGRGLNQI 469
Query: 449 GTVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDF 508
G+++R GDTRW SH SI SL+ M+ ATC VL I +D A R DAD++Y L SF+F
Sbjct: 470 GSLQRPGDTRWSSHLKSISSLMRMFSATCEVLLNIIEDGTTHAHRGDADAAYEVLTSFEF 529
Query: 509 IFILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSF 568
+FI+HLMK+++ +++LCQALQ QSQD++NA+ LV STK LIQ LR++GWD+L A+V SF
Sbjct: 530 VFIMHLMKKVLEISNMLCQALQLQSQDILNAMHLVSSTKLLIQTLRDSGWDELVASVKSF 589
Query: 569 CEKHDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFFY 610
CE +I VPD D+ R GR+ +++++TI H+++VD FY
Sbjct: 590 CETVNITVPDF-DAQYIARRGRARHQQDELTIGHHYKVDIFY 630
>UniRef100_Q9FYP6 Putative transposable element Tip100 protein [Oryza sativa]
Length = 802
Score = 169 bits (428), Expect = 2e-40
Identities = 86/221 (38%), Positives = 131/221 (58%), Gaps = 2/221 (0%)
Query: 389 MTASREVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQV 448
+ S +I FF + +VN VG+S R D L A + +E GEI TG+G NQ
Sbjct: 390 VAVSTSTPAIADFFNYVPLIVNTVGASCMRKDALLAKHHDVLLEKVENGEITTGRGLNQE 449
Query: 449 GTVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDF 508
++ R GDTRWGSH ++ ++ M+ A VL+I+ KD+ K A ++SFDF
Sbjct: 450 SSLARPGDTRWGSHLKTLLRILVMWEAIIDVLEIVKKDSTKPTFNGGAFGLMGKMQSFDF 509
Query: 509 IFILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSF 568
+FI+HLM +++ TD L +ALQ++ QD+V A+ L+ K L+QD+RENGW+ L V+SF
Sbjct: 510 VFIMHLMIDMLSITDDLSRALQRKDQDIVEAMSLLIDVKELLQDMRENGWEPLLNRVISF 569
Query: 569 CEKHDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
C KH+I+VP ++ G S ++VT +HY+ V+ +
Sbjct: 570 CNKHEIKVPKMD--KEVNERGTSTHRRHKVTNKHYYHVEIY 608
>UniRef100_Q7XBZ7 Putative transposase [Oryza sativa]
Length = 1283
Score = 156 bits (395), Expect = 1e-36
Identities = 85/210 (40%), Positives = 130/210 (61%), Gaps = 3/210 (1%)
Query: 401 FFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDTRWG 460
FFE+L F++N++G+S K+ + L+ AQ++ I L+ G+I TGKG NQ + R DTRWG
Sbjct: 639 FFEQLRFLLNLLGNSCKKTEMLRVAQAQRIVEELDLGDIETGKGLNQEMGLGRPADTRWG 698
Query: 461 SHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKEIMG 520
SH+ ++ ++S+Y + VL ++ KD A+ A+A + +SF+F+F+ HLM+ ++G
Sbjct: 699 SHYKTVMHVLSLYPSIQKVLIMVGKDRSFGAECANAQTVLTIFQSFEFVFMAHLMQTVLG 758
Query: 521 TTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLREN-GWDKLFANVVSFCEKHDIEVPDL 579
T L ALQK+ QD+VNAV L+ TK +Q LRE+ GWD V SFC KH I++PD+
Sbjct: 759 FTSDLNHALQKRDQDIVNAVGLILLTKFQLQQLREDPGWDDFLQEVQSFCVKHKIKIPDM 818
Query: 580 NDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
+ + GR ++ H F VD F
Sbjct: 819 DSFYRPV--GRDRRFFIKIKNMHRFHVDMF 846
>UniRef100_Q7XN27 OSJNBb0016D16.6 protein [Oryza sativa]
Length = 897
Score = 156 bits (395), Expect = 1e-36
Identities = 85/210 (40%), Positives = 130/210 (61%), Gaps = 3/210 (1%)
Query: 401 FFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDTRWG 460
FFE+L F++N++G+S K+ + L+ AQ++ I L+ G+I TGKG NQ + R DTRWG
Sbjct: 499 FFEQLRFLLNLLGNSCKKTEMLRVAQAQRIVEELDLGDIETGKGLNQEMGLGRPADTRWG 558
Query: 461 SHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKEIMG 520
SH+ ++ ++S+Y + VL ++ KD A+ A+A + +SF+F+F+ HLM+ ++G
Sbjct: 559 SHYKTVMHVLSLYPSIQKVLIMVGKDRSFGAECANAQTVLTIFQSFEFVFMAHLMQTVLG 618
Query: 521 TTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLREN-GWDKLFANVVSFCEKHDIEVPDL 579
T L ALQK+ QD+VNAV L+ TK +Q LRE+ GWD V SFC KH I++PD+
Sbjct: 619 FTSDLNHALQKRDQDIVNAVGLILLTKFQLQQLREDPGWDDFLQEVQSFCVKHKIKIPDM 678
Query: 580 NDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
+ + GR ++ H F VD F
Sbjct: 679 DSFYRPV--GRDRRFFIKIKNMHRFHVDMF 706
>UniRef100_Q6L3Y1 Putative hAT family dimerisation domain containing protein [Solanum
demissum]
Length = 805
Score = 156 bits (395), Expect = 1e-36
Identities = 83/218 (38%), Positives = 132/218 (60%), Gaps = 2/218 (0%)
Query: 392 SREVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTV 451
S++ + K + + NV+GSS KR D L+ +Q I+ L+ GE+ TG G NQ +
Sbjct: 410 SKKCVEVGKLVVLISNIFNVLGSSFKRMDNLRDSQKSTIQVALDMGELTTGSGLNQQLGL 469
Query: 452 KRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFI 511
RA DTRWGSH+ S + I M+G+ VL+ +A DA+ + +RA A ++F+ F+
Sbjct: 470 SRACDTRWGSHYKSFNNFIIMFGSILEVLESLALDARSMDERAKAMGHLEACQTFEIAFM 529
Query: 512 LHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEK 571
LHLM++++ T+ L + LQK+ QD+ NA+LLV K +Q LR++ WD L A V +FC K
Sbjct: 530 LHLMRDVLAITNELNKCLQKKEQDIANAMLLVEVAKRRLQVLRDDEWDSLIAKVSTFCIK 589
Query: 572 HDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
H+I +P+ + + ++ RS + TI H++RV+ F
Sbjct: 590 HNILIPNFEEPYVSSL--RSRRKLANYTILHHYRVEVF 625
>UniRef100_Q6L3J8 Putative transposase [Solanum demissum]
Length = 705
Score = 156 bits (395), Expect = 1e-36
Identities = 83/218 (38%), Positives = 132/218 (60%), Gaps = 2/218 (0%)
Query: 392 SREVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTV 451
S++ + K + + NV+GSS KR D L+ +Q I+ L+ GE+ TG G NQ +
Sbjct: 310 SKKCVEVGKLVVLISNIFNVLGSSFKRMDNLRDSQKSTIQVALDMGELTTGSGLNQQLGL 369
Query: 452 KRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFI 511
RA DTRWGSH+ S + I M+G+ VL+ +A DA+ + +RA A ++F+ F+
Sbjct: 370 SRACDTRWGSHYKSFNNFIIMFGSILEVLESLALDARSMDERAKAMGHLEACQTFEIAFM 429
Query: 512 LHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEK 571
LHLM++++ T+ L + LQK+ QD+ NA+LLV K +Q LR++ WD L A V +FC K
Sbjct: 430 LHLMRDVLAITNELNKCLQKKEQDIANAMLLVEVAKRRLQVLRDDEWDSLIAKVSTFCIK 489
Query: 572 HDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
H+I +P+ + + ++ RS + TI H++RV+ F
Sbjct: 490 HNILIPNFEEPYVSSL--RSRRKLANYTILHHYRVEVF 525
>UniRef100_Q8S7R0 Putative transposase protein, 5'-partial [Oryza sativa]
Length = 881
Score = 156 bits (395), Expect = 1e-36
Identities = 85/210 (40%), Positives = 130/210 (61%), Gaps = 3/210 (1%)
Query: 401 FFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDTRWG 460
FFE+L F++N++G+S K+ + L+ AQ++ I L+ G+I TGKG NQ + R DTRWG
Sbjct: 133 FFEQLRFLLNLLGNSCKKTEMLRVAQAQRIVEELDLGDIETGKGLNQEMGLGRPADTRWG 192
Query: 461 SHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKEIMG 520
SH+ ++ ++S+Y + VL ++ KD A+ A+A + +SF+F+F+ HLM+ ++G
Sbjct: 193 SHYKTVMHVLSLYPSIQKVLIMVGKDRSFGAECANAQTVLTIFQSFEFVFMAHLMQTVLG 252
Query: 521 TTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLREN-GWDKLFANVVSFCEKHDIEVPDL 579
T L ALQK+ QD+VNAV L+ TK +Q LRE+ GWD V SFC KH I++PD+
Sbjct: 253 FTSDLNHALQKRDQDIVNAVGLILLTKFQLQQLREDPGWDDFLQEVQSFCVKHKIKIPDM 312
Query: 580 NDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
+ + GR ++ H F VD F
Sbjct: 313 DSFYRPV--GRDRRFFIKIKNMHRFHVDMF 340
>UniRef100_Q8LHH0 Putative transposase [Oryza sativa]
Length = 897
Score = 156 bits (395), Expect = 1e-36
Identities = 85/210 (40%), Positives = 130/210 (61%), Gaps = 3/210 (1%)
Query: 401 FFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDTRWG 460
FFE+L F++N++G+S K+ + L+ AQ++ I L+ G+I TGKG NQ + R DTRWG
Sbjct: 499 FFEQLRFLLNLLGNSCKKTEMLRVAQAQRIVEELDLGDIETGKGLNQEMGLGRPADTRWG 558
Query: 461 SHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKEIMG 520
SH+ ++ ++S+Y + VL ++ KD A+ A+A + +SF+F+F+ HLM+ ++G
Sbjct: 559 SHYKTVMHVLSLYPSIQKVLIMVGKDRSFGAECANAQTVLTIFQSFEFVFMAHLMQTVLG 618
Query: 521 TTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLREN-GWDKLFANVVSFCEKHDIEVPDL 579
T L ALQK+ QD+VNAV L+ TK +Q LRE+ GWD V SFC KH I++PD+
Sbjct: 619 FTSDLNHALQKRDQDIVNAVGLILLTKFQLQQLREDPGWDDFLQEVQSFCVKHKIKIPDM 678
Query: 580 NDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
+ + GR ++ H F VD F
Sbjct: 679 DSFYRPV--GRDRRFFIKIKNMHRFHVDMF 706
>UniRef100_Q9LMA0 T29M8.13 protein [Arabidopsis thaliana]
Length = 811
Score = 151 bits (382), Expect = 5e-35
Identities = 76/222 (34%), Positives = 133/222 (59%), Gaps = 3/222 (1%)
Query: 389 MTASREVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQV 448
M +++ + +FF + ++NVVG+S KR D+++ +++E + GEI TG G NQ
Sbjct: 415 MAVAKKHVEVGEFFYMISVLLNVVGASCKRKDKIREIHRQKVEEKISNGEIKTGTGLNQE 474
Query: 449 GTVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDF 508
+++R G+TRWGSH+ ++ L ++ + VL+ I + +R A + +FDF
Sbjct: 475 LSLQRPGNTRWGSHYKTLLRLEELFSSIVIVLEYIQDEGTDTTKRQQAYGILKYFHTFDF 534
Query: 509 IFILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSF 568
+F L LM +MG TD L +ALQ++ QD++NA+ LV++TK +Q +R++GWD A V SF
Sbjct: 535 VFYLELMLLVMGLTDSLSKALQRKDQDILNAISLVKTTKCQLQKVRDDGWDAFMAKVSSF 594
Query: 569 CEKHDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFFY 610
EK++ + + + +R R ++ +T H+++VD FY
Sbjct: 595 SEKNNTGMLKMEEEFVDSRRPR---KKTGITNLHHYKVDCFY 633
>UniRef100_Q9LI98 Similarity to transposase [Arabidopsis thaliana]
Length = 571
Score = 150 bits (380), Expect = 8e-35
Identities = 78/212 (36%), Positives = 127/212 (59%), Gaps = 3/212 (1%)
Query: 398 IHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDT 457
I FF+ + ++NVVG+S KR D ++ K++E + GEI TGKG NQ +++R G+T
Sbjct: 315 IGDFFDMISVLINVVGASCKRKDRVRDEFRKKLEERINQGEIKTGKGLNQKLSLQRPGNT 374
Query: 458 RWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKE 517
RWG+H+ ++ L+ ++ VL+ I D +R A+ + +FDF+F L LM
Sbjct: 375 RWGTHYTTLLRLVDLFSVIIKVLEWIEDDGTDSTKRRQANGLLKYFNTFDFVFYLQLMLL 434
Query: 518 IMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEKHDIEVP 577
I+G T+ L ALQ++ QD++NA+ LV+STK + LR++GWD V SFC+ HDIE
Sbjct: 435 ILGLTNSLSVALQRKDQDILNAMSLVKSTKQQLFKLRDDGWDSFLNEVFSFCKDHDIEFV 494
Query: 578 DLNDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
++ R R +++ +T H+++V+ F
Sbjct: 495 IMDGEFVDPRKPR---KKSNMTNLHHYQVECF 523
>UniRef100_Q7XWU4 OSJNBa0065B15.7 protein [Oryza sativa]
Length = 639
Score = 150 bits (380), Expect = 8e-35
Identities = 77/217 (35%), Positives = 127/217 (58%), Gaps = 3/217 (1%)
Query: 393 REVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVK 452
R+ + FF K+ ++NVVG S+KR D ++ KE+ L +G++ TG NQ ++
Sbjct: 254 RKHKDVSDFFTKISILLNVVGGSSKRRDLIRDINVKEMSKALGSGQLQTGTRLNQEQCLQ 313
Query: 453 RAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFIL 512
R GDTRW SH+ ++ SL+ M+ VL+I+ KD R + + +SFDF+F L
Sbjct: 314 RPGDTRWSSHYKTLKSLVGMFATIVKVLEIVEKDKNDWKIRDQGSNLLEYFQSFDFVFYL 373
Query: 513 HLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEKH 572
HLM I+ T+ L ALQ++ QD+VNA+ V+ST+ + +LR W+K+ V FC+ +
Sbjct: 374 HLMLTILTITNSLSLALQRKDQDIVNAMKCVKSTRLNLDELRREKWEKVLDEVSDFCDNY 433
Query: 573 DIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
DI ++ D++ + R ++ +T +HY++VD F
Sbjct: 434 DIVKLEIEDTYIDPKKRR---HKSGITNKHYYQVDCF 467
>UniRef100_Q8S7P4 Putative transposase [Oryza sativa]
Length = 811
Score = 150 bits (378), Expect = 1e-34
Identities = 85/210 (40%), Positives = 121/210 (57%), Gaps = 3/210 (1%)
Query: 401 FFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDTRWG 460
FF ++ ++N+VG+S KRHD L+ ++++I LE GEI +G G NQ + R GDTRWG
Sbjct: 424 FFTQVSHLLNIVGTSCKRHDMLRDVRAQKIMEALELGEIESGVGLNQEMGLARPGDTRWG 483
Query: 461 SHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKEIMG 520
SH+ +I +I MY VL + KD + + +SFDF+F HLM I+G
Sbjct: 484 SHYKTILHIIGMYPTIHEVLITLGKDPTQRDDWPRIHAVVGAFESFDFVFSAHLMLVILG 543
Query: 521 TTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLF-ANVVSFCEKHDIEVPDL 579
T+ LC LQK+ QD+VNA+ LV K +Q LR GW++ F VVSFC KH I+VP L
Sbjct: 544 YTNELCLCLQKRDQDIVNAMSLVTLAKERMQKLRSEGWEEFFQGTVVSFCNKHSIQVPTL 603
Query: 580 NDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
+ + GRS T + +FR + +
Sbjct: 604 DGKY--VPHGRSPRFYPDQTNDDHFRREVY 631
>UniRef100_Q9SZ82 Hypothetical protein F17A8.10 [Arabidopsis thaliana]
Length = 664
Score = 146 bits (368), Expect = 2e-33
Identities = 78/232 (33%), Positives = 137/232 (58%), Gaps = 9/232 (3%)
Query: 385 EENGMTAS--REVASIH----KFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGE 438
E NG+ + +E+A H +FF+ + ++NVVG+S R D+++ +++E + GE
Sbjct: 307 EFNGLRSLILKEIAKKHVEVGEFFDMISVLLNVVGASCTRKDKIREIHRQKVEEKISNGE 366
Query: 439 IVTGKGKNQVGTVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADS 498
I TG NQ +++R G+TRWGSH+ ++ L ++ + VL+ I + +R A
Sbjct: 367 IKTGTRLNQELSLQRPGNTRWGSHYKTLLRLEELFSSIVIVLEYIQDEGTDTTKRQQAYG 426
Query: 499 SYNHLKSFDFIFILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGW 558
+ +FDF+F L LM +MG TD L +ALQ++ QD++N + LV++TK +Q +R++GW
Sbjct: 427 ILKYFHTFDFVFYLELMLLVMGLTDSLSKALQRKDQDILNVISLVKTTKCQLQKVRDDGW 486
Query: 559 DKLFANVVSFCEKHDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFFY 610
D A V SF EK++ + + + +R R +++ +T H+++VD FY
Sbjct: 487 DAFMAEVSSFSEKNNTAMLKMEEEFVDSRRPR---KKSGITNLHHYKVDCFY 535
>UniRef100_Q6ATH8 Hypothetical protein OSJNBa0093E24.3 [Oryza sativa]
Length = 774
Score = 135 bits (341), Expect = 3e-30
Identities = 74/210 (35%), Positives = 121/210 (57%), Gaps = 4/210 (1%)
Query: 401 FFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDTRWG 460
FF +L +++NV+G S K+ L+ AQ++ + L+ GEI +G+G NQ + R GDTRWG
Sbjct: 433 FFGQLAYLLNVLGMSCKKIRMLRIAQAEYMIEALKLGEIESGQGLNQEMGLARPGDTRWG 492
Query: 461 SHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKEIMG 520
SH+ ++ ++ +Y + +L + K+ + A+ A + +SF+ +F+LH+M EI G
Sbjct: 493 SHYKTVMHVMLLYPSIKKLLFKVGKECNE-AEAIGAQTMLQVFQSFELVFLLHMMNEIFG 551
Query: 521 TTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLREN-GWDKLFANVVSFCEKHDIEVPDL 579
T C ALQ++ QD+VNA+ L+ TKA + LRE+ GW + V SFC KH ++V D+
Sbjct: 552 YTSDFCNALQRREQDIVNAMDLLEFTKAELDVLREDCGWKEFLGKVTSFCVKHKVKVVDM 611
Query: 580 NDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
+ + + R + H F D F
Sbjct: 612 DGKYKPIQRSRKFFRD--AINYHRFHADMF 639
>UniRef100_Q7XW70 OSJNBa0019J05.24 protein [Oryza sativa]
Length = 629
Score = 132 bits (331), Expect = 4e-29
Identities = 75/210 (35%), Positives = 120/210 (56%), Gaps = 5/210 (2%)
Query: 401 FFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDTRWG 460
FF +L +++NV+G S K+ L AQ++ + L+ GEI +G+G NQ + R GDTRWG
Sbjct: 339 FFGQLAYLLNVLGMSCKKFHMLCIAQAEYMIEALKLGEIQSGQGLNQEMGLARPGDTRWG 398
Query: 461 SHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKEIMG 520
SH+ ++ ++ +Y + L + K+ A+ A + +SF+F+F H+M EI
Sbjct: 399 SHYKTVMHVMFLYPSIKKFLFKVGKECNG-AEAIGAQTMLQVFQSFEFVFSWHMMNEIFE 457
Query: 521 TTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLREN-GWDKLFANVVSFCEKHDIEVPDL 579
T+D C ALQ++ QD+VNA+ L+ TKA + LRE+ GW + V SFC KH ++V D+
Sbjct: 458 TSD-FCNALQRREQDIVNAMDLLEFTKAELDVLREDCGWKEFLGKVTSFCVKHKVKVVDM 516
Query: 580 NDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
+ + + R ++ T H F D F
Sbjct: 517 DGKYKPIQRSRKFFKD--ATNYHRFHADMF 544
>UniRef100_Q6K4U9 Putative hAT dimerisation domain-containing protein [Oryza sativa]
Length = 505
Score = 99.8 bits (247), Expect = 2e-19
Identities = 59/203 (29%), Positives = 106/203 (52%), Gaps = 4/203 (1%)
Query: 389 MTASREVAS-IHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQ 447
+T SR +S IH FF + ++ S + D+ + N LE+GEI++ ++
Sbjct: 299 ITVSRCSSSLIHDFFVSISLIITTTIESCQMMDKSTEKHHQTTLNKLESGEILSEGSNHK 358
Query: 448 VGTVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFD 507
+ G TRWGS++ ++ + M+ + VL I+ + + Q + A ++SF+
Sbjct: 359 EKNLASLGGTRWGSYYTTLGRINMMWDSVLDVLMIVHQHGR---QPSRAGGLIQTMESFE 415
Query: 508 FIFILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVS 567
F+FIL +M ++ T+ L LQ++ + ++ AV + K +Q LR NGW+ LF +V S
Sbjct: 416 FVFILKMMLKLFTITNELSLVLQRKDEGIIQAVGQLTDVKECLQTLRNNGWESLFEDVKS 475
Query: 568 FCEKHDIEVPDLNDSHSTTRFGR 590
FC + I VP++++ T R
Sbjct: 476 FCATNWIPVPNMDEHIRATGHSR 498
>UniRef100_Q6L4N7 Hypothetical protein P0473H02.5 [Oryza sativa]
Length = 717
Score = 87.8 bits (216), Expect = 8e-16
Identities = 55/202 (27%), Positives = 94/202 (46%), Gaps = 2/202 (0%)
Query: 408 VVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDTRWGSHFNSIC 467
+V VG S + D + +I + G+ + G + + TRWGS+ ++
Sbjct: 341 IVRAVGDSCRTKDAMLQELYGKIREKIVRGDALPKIGMHPENDLSGPAHTRWGSYSTTLL 400
Query: 468 SLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKEIMGTTDLLCQ 527
+L++M+ L + + ++SF+F+F+LHLM ++ TD L
Sbjct: 401 NLLTMWDVVLDTLVTMCDKGIYPEPGSIPSDMIEQMESFEFVFVLHLMIRVLIWTDDLSC 460
Query: 528 ALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEKHDIEVPDLNDSHSTTR 587
L+++ + +VN + L+ S K ++QDL+EN W L V FC I VP ++DS
Sbjct: 461 LLERKGKYIVNPLELITSVKNILQDLKENQWVDLLEEVKRFCILKSIPVPSMDDSIPVR- 519
Query: 588 FGRSHLEENQVTIEHYFRVDFF 609
GRS VT + V+ F
Sbjct: 520 -GRSRHRGLVVTCHQQYYVETF 540
>UniRef100_Q9SI24 Hypothetical protein At2g05020 [Arabidopsis thaliana]
Length = 370
Score = 71.6 bits (174), Expect = 6e-11
Identities = 37/124 (29%), Positives = 67/124 (53%), Gaps = 6/124 (4%)
Query: 450 TVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFI 509
++++ +TRWG+H+ ++ +I + ++ I + +R A+ + +FDF
Sbjct: 127 SLQKPANTRWGTHYKTLLRIIEFFSCIIEYIEYIQDEDVDNIKRRQANGLLKYFHTFDFA 186
Query: 510 FILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQ------DLRENGWDKLFA 563
F L LM I+G D+L QALQ++ QD++N + LV STK + D++ ++ LF
Sbjct: 187 FYLQLMLHILGFADILSQALQRRDQDILNVMSLVASTKREVDYYYNVLDMQLQAFNDLFD 246
Query: 564 NVVS 567
V S
Sbjct: 247 EVNS 250
>UniRef100_UPI0000248927 UPI0000248927 UniRef100 entry
Length = 717
Score = 45.1 bits (105), Expect = 0.006
Identities = 48/227 (21%), Positives = 90/227 (39%), Gaps = 28/227 (12%)
Query: 396 ASIH--KFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKR 453
+SIH FF + + STKRH Q L G+ +V +++
Sbjct: 334 SSIHFVTFFSLVEKLYVFCTGSTKRHTAFLKCQQS-----LYPGQ--------RVVELQK 380
Query: 454 AGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFIL- 512
DT+W S+ +L + A +L + + DA +L++ DF F+L
Sbjct: 381 LSDTQWACRERSLKALNKVLKALIKLLTYLCESDPPDTAAGDAKM---YLRAIDFEFLLC 437
Query: 513 -HLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLR-ENGWDKLFANVVSFCE 570
+ + T + ALQK+ D+ A + +++LR E + +F V E
Sbjct: 438 SEITTTVFQITGVTSDALQKKDLDLSTAYTITDGVLDTVKNLRSEEEFKTIFQKAVEKAE 497
Query: 571 KHDIEVPDLNDSHSTTR-------FGRSHLEENQVTIEHYFRVDFFY 610
I++P + H R + +++ T+E ++R ++
Sbjct: 498 DVGIDIPTVPPGHGRKRKVPTRYLHSATAAQDSHTTVEEFYRAKVYF 544
>UniRef100_Q9SN25 Hypothetical protein F28M11.120 [Arabidopsis thaliana]
Length = 733
Score = 42.0 bits (97), Expect = 0.053
Identities = 25/100 (25%), Positives = 52/100 (52%), Gaps = 4/100 (4%)
Query: 480 LKIIAKDAKKIAQRADAD----SSYNHLKSFDFIFILHLMKEIMGTTDLLCQALQKQSQD 535
L +A++ R+DAD S + + F+F+F + + ++ T + + +ALQ ++ D
Sbjct: 426 LDYLAENCDDPKARSDADCLATSETHRIGGFEFLFGMVIWYNLLFTMNTVSKALQSENID 485
Query: 536 VVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEKHDIE 575
+ A++ ++ + +Q+ RE G+ + A E DIE
Sbjct: 486 IELALVQLKGLVSYLQNYRETGFQEAKAEATLIAESMDIE 525
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.359 0.157 0.535
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 784,476,124
Number of Sequences: 2790947
Number of extensions: 26771359
Number of successful extensions: 143761
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 143714
Number of HSP's gapped (non-prelim): 32
length of query: 611
length of database: 848,049,833
effective HSP length: 133
effective length of query: 478
effective length of database: 476,853,882
effective search space: 227936155596
effective search space used: 227936155596
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 78 (34.7 bits)
Medicago: description of AC148241.10


