

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
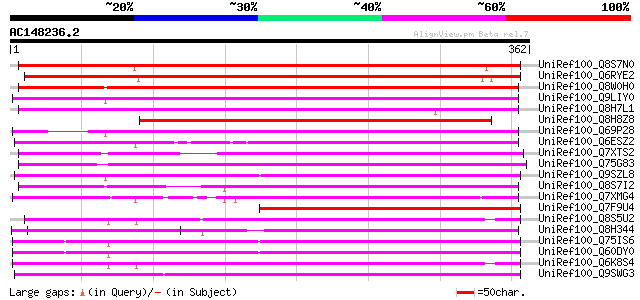
Query= AC148236.2 - phase: 0 /pseudo
(362 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8S7N0 Putative transposase [Oryza sativa] 374 e-102
UniRef100_Q6RYE2 Putative transposase [Triticum monococcum] 363 5e-99
UniRef100_Q8W0H0 P0529E05.9 protein [Oryza sativa] 320 3e-86
UniRef100_Q9LIY0 Similar to Arabidopsis thaliana DNA chromosome ... 291 1e-77
UniRef100_Q8H7L1 Putative transposase [Oryza sativa] 286 6e-76
UniRef100_Q8H8Z8 Putative far-red impaired response protein [Ory... 277 3e-73
UniRef100_Q69P28 Putative far-red impaired response protein [Ory... 266 7e-70
UniRef100_Q6ESZ2 Putative far-red impaired response protein [Ory... 263 7e-69
UniRef100_Q7XTS2 OSJNBa0008M17.13 protein [Oryza sativa] 257 3e-67
UniRef100_Q75G83 Putative transposase [Oryza sativa] 257 4e-67
UniRef100_Q9SZL8 Hypothetical protein F20D10.300 [Arabidopsis th... 244 4e-63
UniRef100_Q8S7I2 Putative transposase [Oryza sativa] 241 3e-62
UniRef100_Q7XMG4 OSJNBa0028I23.18 protein [Oryza sativa] 235 2e-60
UniRef100_Q7F9U4 OSJNBb0006L01.4 protein [Oryza sativa] 224 3e-57
UniRef100_Q8S5U2 Putative far-red impaired response protein [Ory... 218 2e-55
UniRef100_Q8H344 Putative far-red impaired response protein [Ory... 212 1e-53
UniRef100_Q75IS6 Putative Mutator-like transposase [Oryza sativa] 212 1e-53
UniRef100_Q60DY0 Hypothetical protein OSJNBa0090H02.8 [Oryza sat... 212 1e-53
UniRef100_Q6K8S4 Putative far-red impaired response protein [Ory... 210 6e-53
UniRef100_Q9SWG3 Far-red impaired response protein [Arabidopsis ... 207 3e-52
>UniRef100_Q8S7N0 Putative transposase [Oryza sativa]
Length = 486
Score = 374 bits (959), Expect = e-102
Identities = 186/369 (50%), Positives = 240/369 (64%), Gaps = 19/369 (5%)
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTH-RKKDGSVSSCRFVCCKEGMRKPDKR 65
P+V M F S AW FWN+Y G+ GF VRK YT+ RK DG V SCR+VC EG RK DKR
Sbjct: 55 PQVGMEFSSTDEAWMFWNSYGGQKGFEVRKRYTNKRKSDGKVRSCRYVCANEGHRKEDKR 114
Query: 66 DYKTKNPRLETRTNCEARLG--LKNVDGKLMVHDFVEDHNHELLLPETTHMLSSQRKVSE 123
D+ TK P ETRT+C+ R+G L G V D V +HNH L LPET+H++ SQRK+SE
Sbjct: 115 DHLTKCPTAETRTDCQVRMGVVLDLEKGNYKVADLVLEHNHILQLPETSHLMVSQRKISE 174
Query: 124 IHCQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYL 183
+ +IE AD G+ K + +L S +VGG NL + + KNYLR KR+R + G+AG +
Sbjct: 175 LQGFEIETADDAGIGPKAAHELASIQVGGSLNLRYTLREHKNYLRGKRQREMAYGQAGSM 234
Query: 184 LQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHR 243
L YFQ K ENP+F +A Q+D E+QI N+FW DA+ML DY+YFGDVVS D+T+ TN R
Sbjct: 235 LMYFQDKIAENPSFQYALQMDQEEQIANIFWVDAKMLTDYAYFGDVVSFDTTFGTNKESR 294
Query: 244 PLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKA 303
+ GFN R ++FGA L+YDET ES++WLFETFL+AH K P+T++TDQ +M KA
Sbjct: 295 LFGVFVGFNQFRETIVFGAVLMYDETYESFKWLFETFLKAHNGKQPKTIYTDQDFAMGKA 354
Query: 304 LAEVMPEAYHGLCTWHLMQNGIKHLGNL----------------MKGESYFLSDFEKCMY 347
+ EV E +HGLCT+H+MQN KHL + + E L+DF CM+
Sbjct: 355 VKEVFSEVWHGLCTFHIMQNAAKHLAEVDNKEESSTSPEQIVEDNEKEPSILADFSACMF 414
Query: 348 GYEDVEQFE 356
YED E FE
Sbjct: 415 EYEDEETFE 423
>UniRef100_Q6RYE2 Putative transposase [Triticum monococcum]
Length = 751
Score = 363 bits (931), Expect = 5e-99
Identities = 186/397 (46%), Positives = 244/397 (60%), Gaps = 51/397 (12%)
Query: 11 MFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKK-DGSVSSCRFVCCKEGMRKPDKRDYKT 69
M F + AW FW TY G+ GF VRK YT+R+ DG V+SCRFVC EG R DKRD+ T
Sbjct: 1 MEFRNSDEAWAFWLTYSGQKGFEVRKRYTNRRPTDGKVTSCRFVCANEGHRWQDKRDHLT 60
Query: 70 KNPRLETRTNCEARLGLKN--VDGKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIHCQ 127
K PR ETRT C+ + LK G L V + +HNH L LPET H++ SQRK+SE+
Sbjct: 61 KCPRAETRTECQVFMNLKTDRKKGNLKVSELSLEHNHTLHLPETLHLMVSQRKISEVQSF 120
Query: 128 QIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQYF 187
+I+ AD+ G+ K + +L S++VGG NL + D+KNYLR KR+R + GEAG +L+YF
Sbjct: 121 EIDTADAAGIGPKAAQELASRQVGGPLNLSYTLRDRKNYLRTKRQREMAYGEAGSMLKYF 180
Query: 188 QRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPLAI 247
Q K ENP+F +A Q+D E+QI N+FWADA+M++DY++FGDVVS D+T+ TN RP +
Sbjct: 181 QDKVAENPSFQYALQMDCEEQIANIFWADAKMIMDYAHFGDVVSFDTTFGTNKESRPFGV 240
Query: 248 ISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALAEV 307
GFNH R V+FGAAL+YDET +S+EWLF+TFL+AH + P+T +TDQ +M A EV
Sbjct: 241 FVGFNHFRETVVFGAALMYDETFDSFEWLFKTFLKAHNGQHPKTFYTDQDTAMGNAAGEV 300
Query: 308 MPEAYHGLCTWHLMQNGIKHL-----------------GNLMKG---------------- 334
PE +HGLCT+H+MQN IKHL G +G
Sbjct: 301 FPETWHGLCTFHIMQNAIKHLHEEKNEENNEEKTAGKNGEQNEGKNEDKNEGKKQKKHEG 360
Query: 335 ---------------ESYFLSDFEKCMYGYEDVEQFE 356
E LSDF CM+ YED+ +FE
Sbjct: 361 KKKKKNEGKKIEKNEEPSILSDFSACMFEYEDMIEFE 397
>UniRef100_Q8W0H0 P0529E05.9 protein [Oryza sativa]
Length = 773
Score = 320 bits (821), Expect = 3e-86
Identities = 148/350 (42%), Positives = 222/350 (63%), Gaps = 2/350 (0%)
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRKPDKRD 66
P + M F A+EF+N Y G GF +RK + + + + +FVC +EG K + D
Sbjct: 151 PVIGMEFDDEDIAYEFYNRYAGDVGFSIRKFWHDKSSTNVIRTKKFVCSREGFNKRNTSD 210
Query: 67 YKTKNPRLETRTNCEARLGLK-NVDGKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIH 125
+ R +TR C A + +K GK + F HNHEL+ P H+L SQR+++E
Sbjct: 211 -ACQRKRADTRVGCMAEMTIKITPTGKYAIASFSNTHNHELITPSKAHLLRSQRRMTEAQ 269
Query: 126 CQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQ 185
QI++ + GV+ K+ +++S++ GGR +L F R D KNYLR KR +S+ +G+ G +LQ
Sbjct: 270 KAQIDILNDSGVRSKEGHEVMSRQAGGRQSLTFTRKDYKNYLRSKRMKSIQEGDTGAILQ 329
Query: 186 YFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPL 245
Y Q K +ENP+F++A Q+D ++ +TN+FWADAR ++D+ YFGDV+ D+TY TN+ RP
Sbjct: 330 YLQDKQMENPSFFYAIQVDEDEMMTNIFWADARSVLDFDYFGDVICFDTTYRTNNYGRPF 389
Query: 246 AIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALA 305
A+ G NHH+ V+FGAALLYDET ++EWLFETF A K P+T+ TDQ ++ A+
Sbjct: 390 ALFVGVNHHKQTVVFGAALLYDETTSTFEWLFETFKRAMSGKEPRTILTDQCAAIINAIG 449
Query: 306 EVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQF 355
V P + H LC WH+ QN HL ++ +G F +DF KC++ +E+V++F
Sbjct: 450 TVFPNSTHRLCVWHMYQNAAVHLSHIFQGSKTFKNDFGKCVFDFEEVDEF 499
>UniRef100_Q9LIY0 Similar to Arabidopsis thaliana DNA chromosome 4 [Oryza sativa]
Length = 739
Score = 291 bits (746), Expect = 1e-77
Identities = 145/358 (40%), Positives = 210/358 (58%), Gaps = 5/358 (1%)
Query: 3 EDWNPKVDMFFCSLKNAWEFWNTYXGKXGFGVRKD-YTHRKKDGSVSSCRFVCCKEGMRK 61
ED PKV M F S + A++ +N Y G+ GF +R+ Y H + + F C ++G R
Sbjct: 89 EDKVPKVGMKFNSEQEAYDLYNAYAGEKGFSIRRSSYHHVGHTKIIKNMTFCCSRQGTRG 148
Query: 62 PDKR----DYKTKNPRLETRTNCEARLGLKNVDGKLMVHDFVEDHNHELLLPETTHMLSS 117
DKR Y R ETR C+A + + +DG V+ F+ +H+H L H L S
Sbjct: 149 IDKRAEAAGYGDSFSRPETRCKCQACMKISLIDGLYSVYHFMPEHSHNLATKSQAHQLRS 208
Query: 118 QRKVSEIHCQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQ 177
QRK++E +E+A SVG+ K + DL++K+ G NLGF R+D KN L KR Q
Sbjct: 209 QRKINEAQVASVEVAKSVGISTKAAIDLMAKQACGFENLGFTRVDMKNKLYSKRSLQTKQ 268
Query: 178 GEAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYC 237
G+ G +L+Y ++K E+ F+++ Q+D +D ITN+FWAD++M+ DY F DVV D+TY
Sbjct: 269 GDTGGVLEYMEKKASEDVKFFYSIQVDEDDLITNIFWADSKMVSDYEDFCDVVCFDTTYR 328
Query: 238 TNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQA 297
RP ++ G N+H+ +FGA LLYDETA S+ WLF TFL K P+T+FTD+
Sbjct: 329 KLDDGRPFGLLVGVNNHKKTTVFGATLLYDETANSFVWLFNTFLNVMSGKKPKTIFTDED 388
Query: 298 KSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQF 355
+MAKA+ V +H +C WH+ QN KHL +K F +DF+ C+Y E+ E+F
Sbjct: 389 AAMAKAIKIVFKGTHHRICVWHMNQNACKHLAGCVKDYKKFNADFQNCIYDQEEEEEF 446
>UniRef100_Q8H7L1 Putative transposase [Oryza sativa]
Length = 778
Score = 286 bits (732), Expect = 6e-76
Identities = 142/352 (40%), Positives = 206/352 (58%), Gaps = 3/352 (0%)
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRKPDKRD 66
P++ M F A+EF+N Y G GF VRK + + + S FVC +EG RK K
Sbjct: 86 PELSMEFDDEDKAYEFYNRYAGHVGFSVRKSSSDKSAENITRSRTFVCSREGFRKDKKGA 145
Query: 67 YKTKNPRLETRTNCEARLGLK-NVDGKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIH 125
+ K PR ETR C AR+ +K DGK + +FV DHNH+ P T HML SQR ++E+
Sbjct: 146 KEVKRPRPETRIGCPARMSIKITSDGKYRISEFVPDHNHQPAPPSTMHMLRSQRVLTELQ 205
Query: 126 CQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQ 185
+ + ++ + S L K+ + F+ + + L KR++++ G+AG ++
Sbjct: 206 TTEADSSEESATPSRFSSCSLVKQAEVIRHTNFLPAEYRCSLCSKRKKNMQPGDAGVTVK 265
Query: 186 YFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPL 245
Y Q + NP+F++A QLD +D++TN+FWAD++ D+SY+ DVV LD+TY N RPL
Sbjct: 266 YLQSMQLSNPSFFYAVQLDEDDKLTNIFWADSKSRTDFSYYSDVVCLDTTYKINEHSRPL 325
Query: 246 AIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTD--QAKSMAKA 303
+ G NHH+ IFGAALLYDE+ ES++WLF+TF A K P+T+ TD A + A A
Sbjct: 326 TLFLGVNHHKQISIFGAALLYDESEESFKWLFDTFKIAANGKQPKTILTDWSMAATTASA 385
Query: 304 LAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQF 355
+ P H LC W + QN +KHL ++ +G F DF KC+Y Y+D E F
Sbjct: 386 ITAAWPGTVHRLCPWQVYQNSVKHLNHIFQGSKTFAKDFGKCVYDYDDEENF 437
>UniRef100_Q8H8Z8 Putative far-red impaired response protein [Oryza sativa]
Length = 535
Score = 277 bits (709), Expect = 3e-73
Identities = 130/246 (52%), Positives = 172/246 (69%)
Query: 91 GKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIHCQQIELADSVGVQQKKSFDLLSKEV 150
G V D V +HNH L LPET+H++ SQRK+SE+ +IE AD G+ K + L S +V
Sbjct: 10 GNYKVADLVLEHNHILQLPETSHLMVSQRKISELQGFEIETADDAGIGPKAAHQLASIQV 69
Query: 151 GGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQYFQRKTVENPTFYHAYQLDIEDQIT 210
GG NL D KNYLR KR+R +V G+AG +L +FQ K ENP+F +A Q+D E+QI
Sbjct: 70 GGSLNLNCTLRDHKNYLRGKRQREMVYGQAGSMLMHFQDKIAENPSFQYALQMDSEEQIA 129
Query: 211 NVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETA 270
N+FW DA+ML DY+YFGDVVS D+T+ TN RP + GFN R ++FGA LLYDET
Sbjct: 130 NIFWVDAKMLTDYAYFGDVVSFDTTFGTNKESRPFGVFVGFNQFRETMVFGAVLLYDETY 189
Query: 271 ESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGN 330
ES++WLFETFL+AH K P+T++TDQ +M KA+ +V E++HGLCT+H+MQN +KH+
Sbjct: 190 ESFKWLFETFLKAHNGKQPKTIYTDQDSAMGKAIKKVFLESWHGLCTFHIMQNAVKHVAE 249
Query: 331 LMKGES 336
L ES
Sbjct: 250 LEDEES 255
>UniRef100_Q69P28 Putative far-red impaired response protein [Oryza sativa]
Length = 516
Score = 266 bits (680), Expect = 7e-70
Identities = 137/357 (38%), Positives = 198/357 (55%), Gaps = 31/357 (8%)
Query: 3 EDWNPKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRKP 62
ED PK+ M F S + A++F+N Y K+G R
Sbjct: 118 EDLVPKIGMKFSSEQEAYDFYNAYA---------------------------LKKGSRGI 150
Query: 63 DKR----DYKTKNPRLETRTNCEARLGLKNVDGKLMVHDFVEDHNHELLLPETTHMLSSQ 118
DKR Y ETR C+A + + +DG ++ FV +HNH L H L SQ
Sbjct: 151 DKRAEASGYGDSFSWPETRCKCQACMKISIIDGLYSIYHFVPEHNHNLATRSQAHQLRSQ 210
Query: 119 RKVSEIHCQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQG 178
RK++E +E+A SVG+ K + DL++K+ G NLGF R+D KN L KR QG
Sbjct: 211 RKINEAQIASVEVAKSVGISTKAAIDLMAKQACGFENLGFTRVDMKNKLYSKRSLQTKQG 270
Query: 179 EAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCT 238
+ G +L+Y ++K E+ F+++ Q+D + ITN+FWAD++M+ DY FGDVV D+TY
Sbjct: 271 DTGGVLEYMEKKASEDVKFFYSIQVDEDVLITNIFWADSKMVADYEDFGDVVCFDTTYRK 330
Query: 239 NSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAK 298
P ++ G N+H+ +FGAALLYDETA S+ WLF TFL K P+T+ ++
Sbjct: 331 LDDGHPFGLLVGVNNHKKTTVFGAALLYDETANSFVWLFNTFLNVISGKKPKTILPNEDA 390
Query: 299 SMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQF 355
+MAKA+ V+PE +H +C WH+ QN KHL +K F +DF+ C+Y E+ E+F
Sbjct: 391 AMAKAIKIVLPETHHRICVWHMNQNACKHLTGCVKDYKKFNADFQNCIYDQEEEEEF 447
>UniRef100_Q6ESZ2 Putative far-red impaired response protein [Oryza sativa]
Length = 815
Score = 263 bits (671), Expect = 7e-69
Identities = 138/355 (38%), Positives = 215/355 (59%), Gaps = 9/355 (2%)
Query: 4 DWNPKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHR-KKDGSVSSCRFVCCKEGMRKP 62
D+ P + F S A+EF+ Y K GF VR++Y ++ +K G ++S +FVC +EG + P
Sbjct: 47 DYTPWLGQEFASEHEAYEFYRYYAWKLGFSVRREYANKSRKTGEITSRKFVCSREGFKAP 106
Query: 63 DKRDYKTKNPRLETRTNCEARLGL--KNVDGKLMVHDFVEDHNHELLLPETTHMLSSQRK 120
DKR T+ P+ +TRT C A L + KN K V+ F HNH L +P + L Q+K
Sbjct: 107 DKRTNHTRTPQPDTRTGCHANLVIRRKNDTSKYEVYAFEAQHNHPLFIPSCANPL--QKK 164
Query: 121 VSEIHCQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEA 180
+S++ Q + +S V D + + +T + + Q++ L +R+R + GEA
Sbjct: 165 LSDV--QSSDADNSASVTHASEPDSRNSILAEKT-IKSPEISQRS-LHTRRQREIKCGEA 220
Query: 181 GYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNS 240
LL Y Q + +P FYHA QLD ED++TN+FWADA+M++D+ FGDVVS D N
Sbjct: 221 SALLNYLQDQCRADPLFYHAVQLDAEDKVTNIFWADAKMVIDFGQFGDVVSFDIVPRNNM 280
Query: 241 SHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSM 300
+ RP A GFN++ V+ G AL+YD++ ES++WLFETFL A + P+TVF+ Q +
Sbjct: 281 NLRPFASFVGFNNYGETVLLGMALMYDDSLESFQWLFETFLHAMSGQAPRTVFSRQDAIV 340
Query: 301 AKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQF 355
+KA++ VMP+ H +CTW+L Q+ +L +L++G+ F+ +F+ C+ YE+ +F
Sbjct: 341 SKAISLVMPDTCHAICTWNLKQSAKSNLNHLIRGDCGFIKEFKACINDYEEEVEF 395
>UniRef100_Q7XTS2 OSJNBa0008M17.13 protein [Oryza sativa]
Length = 961
Score = 257 bits (657), Expect = 3e-67
Identities = 138/353 (39%), Positives = 194/353 (54%), Gaps = 30/353 (8%)
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRKPDKRD 66
P V M F + + A+E++ +Y G GF VRK + S FVC +EG R ++
Sbjct: 186 PIVGMVFENEEKAYEYYASYAGNVGFSVRKGLWDKTVKNVARSRVFVCSREGFRSKNE-- 243
Query: 67 YKTKNPRLETRTNCEARLGLKNV-DGKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIH 125
K PR ETRT C AR+ +K V +GK V +FVEDHNH+L P ML S+R
Sbjct: 244 --AKRPRPETRTGCPARIAIKLVSNGKYRVAEFVEDHNHQLAAPFDIDMLKSER------ 295
Query: 126 CQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQ 185
+L+K G N I KNY+R K + + G L+
Sbjct: 296 -------------------VLTKVQPGNRNASNIPPGCKNYIRAKSSTHMNSEDLGALMD 336
Query: 186 YFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPL 245
YF+R +NP+FY+A Q+D D+ TNVFWADAR +VDY YF DV+ D+T+ +N +PL
Sbjct: 337 YFRRMKSDNPSFYYAIQVDENDKATNVFWADARSIVDYHYFCDVICFDTTHRSNDCRKPL 396
Query: 246 AIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALA 305
A+ G NHHR +IFG+A LYDET ES++WL ETF A K P+T+ TD++ ++ +AL+
Sbjct: 397 ALFLGMNHHRQTIIFGSAFLYDETVESFKWLLETFKSAMCGKQPKTILTDRSAALKEALS 456
Query: 306 EVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFEDV 358
P H C W + QN K L ++ F DF CM+ ED ++F+++
Sbjct: 457 LTWPGTIHRSCVWQIYQNAFKSLEHVFNTLEEFTHDFHHCMFDIEDDQEFDEI 509
>UniRef100_Q75G83 Putative transposase [Oryza sativa]
Length = 813
Score = 257 bits (656), Expect = 4e-67
Identities = 138/357 (38%), Positives = 209/357 (57%), Gaps = 10/357 (2%)
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRKPDKRD 66
P+V M F S A+E +NTY GK GF +RK R+ +G++ VC K+G ++
Sbjct: 85 PQVGMSFESKDKAYEMYNTYAGKVGFSIRKSNVKRRSNGTIYQKHMVCNKQGQQE----- 139
Query: 67 YKTKNPRLETRTNCEARLGLKNVDGKL-MVHDFVEDHNHELLLPETTHMLSSQRKVSEIH 125
T + TRT C+AR+ ++ MV V +HNH+L+ P +H L SQR+V E
Sbjct: 140 --TSSSLDTTRTCCKARVQFSVCRKEIWMVQKVVLEHNHDLVSPNKSHKLRSQRRVIEAD 197
Query: 126 CQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQ 185
Q I G++ + + + + GG + F ++D N + ++R++ L + LL+
Sbjct: 198 RQLIGQIREAGMKPAQVYGFMKEWYGGADKVPFSKMDCNNEIGRERKKYLESNDTQTLLE 257
Query: 186 YFQRKTVENPTFYHAYQLDIED-QITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRP 244
Y + K +E+PTF++A Q+D ED +I N FWAD + ++DY+ FGD VS D+T+ TN P
Sbjct: 258 YLRNKQLEDPTFFYAIQVDKEDGRIANFFWADGQSIMDYACFGDFVSFDTTFDTNKCEMP 317
Query: 245 LAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKAL 304
A + G NHH+ +IFGAALL+++T ES+ WLFETFL A K P T+FTDQ +MA A+
Sbjct: 318 FAPLLGTNHHKQTIIFGAALLFNQTIESFVWLFETFLTAMSGKHPSTIFTDQDAAMAAAI 377
Query: 305 AEVMPEAYHGLCTWHLMQNGIKHLGNLM-KGESYFLSDFEKCMYGYEDVEQFEDVGH 360
A V H LC WH+ NG K+L +++ K + FL+DF++C+Y F + H
Sbjct: 378 AFVFRNTSHRLCLWHIYLNGGKNLSHVIHKHPNKFLADFKRCVYEDRSEYYFNKMWH 434
>UniRef100_Q9SZL8 Hypothetical protein F20D10.300 [Arabidopsis thaliana]
Length = 788
Score = 244 bits (622), Expect = 4e-63
Identities = 129/358 (36%), Positives = 201/358 (56%), Gaps = 6/358 (1%)
Query: 4 DWNPKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHR-KKDGSVSSCRFVCCKEGMRKP 62
D P + F S + A F+N+Y + GF R + R ++DG++ +FVC KEG R
Sbjct: 70 DLEPYDGLEFESEEAAKAFYNSYARRIGFSTRVSSSRRSRRDGAIIQRQFVCAKEGFRNM 129
Query: 63 DKR---DYKTKNPRLETRTNCEARLGLKNVD-GKLMVHDFVEDHNHELLLPETTHMLSSQ 118
+++ D + K PR TR C+A L +K D GK +V FV+DHNHEL+ P+ H L S
Sbjct: 130 NEKRTKDREIKRPRTITRVGCKASLSVKMQDSGKWLVSGFVKDHNHELVPPDQVHCLRSH 189
Query: 119 RKVSEIHCQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQG 178
R++S I+ + G+ ++ L KE GG + +GF +D +NY+R R++S ++G
Sbjct: 190 RQISGPAKTLIDTLQAAGMGPRRIMSALIKEYGGISKVGFTEVDCRNYMRNNRQKS-IEG 248
Query: 179 EAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCT 238
E LL Y ++ +NP F+++ Q + + NVFWAD + ++D+++FGD V+ D+TY +
Sbjct: 249 EIQLLLDYLRQMNADNPNFFYSVQGSEDQSVGNVFWADPKAIMDFTHFGDTVTFDTTYRS 308
Query: 239 NSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAK 298
N P A +G NHH ++FG A + +ET S+ WLF T+L A P ++ TD
Sbjct: 309 NRYRLPFAPFTGVNHHGQPILFGCAFIINETEASFVWLFNTWLAAMSAHPPVSITTDHDA 368
Query: 299 SMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFE 356
+ A+ V P A H C WH+++ + L ++ F SDF KC+ E VE FE
Sbjct: 369 VIRAAIMHVFPGARHRFCKWHILKKCQEKLSHVFLKHPSFESDFHKCVNLTESVEDFE 426
>UniRef100_Q8S7I2 Putative transposase [Oryza sativa]
Length = 1203
Score = 241 bits (614), Expect = 3e-62
Identities = 133/352 (37%), Positives = 191/352 (53%), Gaps = 28/352 (7%)
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRKPDKRD 66
PKV M F + A++ + TY G GF VRK + + + S +VC KEG R
Sbjct: 414 PKVGMVFLNEDKAYDCYVTYAGTVGFSVRKGWLEKTATNTTKSRAYVCSKEGFRSKSVST 473
Query: 67 YKTKNPRLETRTNCEARLGLK-NVDGKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIH 125
K PR ETRT C A + +K V GK +V ++V DHNH+L P
Sbjct: 474 -DPKKPRPETRTGCLAHMTIKITVSGKYVVTEYVADHNHDLETP---------------- 516
Query: 126 CQQIELADSVGVQQKKSFDLLSK--EVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYL 183
V +Q +S LL+K + + I D KNY+R + + + G+ +
Sbjct: 517 --------LVDIQVLRSHKLLAKLQQPPDPPRVVLIPNDYKNYVRTRHTKDMQLGDTQAI 568
Query: 184 LQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHR 243
+Y QRK E+P+F++A Q+D +DQ+TNVFWAD + ++DY YFGDV+ +D+ Y T+ R
Sbjct: 569 YEYLQRKKGEHPSFFYAIQVDEDDQLTNVFWADVKSILDYHYFGDVLCVDTRYSTSDHSR 628
Query: 244 PLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKA 303
PL + G NHH+ VIFGAAL+YDE+ ES++WLFETF A K P+TV DQ+ ++++A
Sbjct: 629 PLLLFIGVNHHKQPVIFGAALVYDESVESFKWLFETFKSAMSGKQPKTVMIDQSTAISEA 688
Query: 304 LAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQF 355
+A V P HL +N K L + + F DF +YGYE+ F
Sbjct: 689 VASVWPRTTQRFSLIHLYKNATKILRDAFQASETFADDFSMWLYGYEEEGDF 740
>UniRef100_Q7XMG4 OSJNBa0028I23.18 protein [Oryza sativa]
Length = 1002
Score = 235 bits (599), Expect = 2e-60
Identities = 140/363 (38%), Positives = 207/363 (56%), Gaps = 22/363 (6%)
Query: 3 EDWNPKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHR-KKDGSVSSCRFVCCKEGMRK 61
E++ P++ M F S A+EF+ Y K GF VRK+Y ++ KK G ++S +F C +EG R
Sbjct: 201 EEYTPRMGMEFESEHEAYEFYRYYGWKVGFNVRKEYANKSKKTGEITSRKFACSREGYRA 260
Query: 62 PDKRDYKTKNPRLETRTNCEARLGL--KNVDGKLMVHDFVEDHNHELLLPETTHMLSSQR 119
KR P ++RT C A L + K KL V+ F HNH L T + +
Sbjct: 261 NVKRGNHMV-PMPDSRTGCNAHLVIRRKKPGAKLEVYAFQPRHNHPLF---ATSCMPNPL 316
Query: 120 KVSEIHCQQIELADSVGVQQKKSFDLLSK--EVGGRTNL-----GFIRLDQKNYLRKKRE 172
+ + +H L D+V DLL EVGG+ + ++ LR +R+
Sbjct: 317 QPNVVHWTT--LPDAVTPP-----DLLMDGGEVGGQNSTEENSTASAGEGRRQPLRTRRQ 369
Query: 173 RSLVQGEAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSL 232
+ GEAG LL YFQ +++ NP+FYH+ QLD+ED++ NVFWAD RM+ DYS FGDV++
Sbjct: 370 WEMKYGEAGALLNYFQDQSLANPSFYHSVQLDVEDKVANVFWADPRMITDYSQFGDVIAF 429
Query: 233 DSTYCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTV 292
D + S R A G N+ ++FG AL+YDETAES++WLFETFL A + P+T
Sbjct: 430 DVVSRNSISLRHFASFVGCNNFGEPIVFGLALMYDETAESFQWLFETFLHAMSGRAPKTF 489
Query: 293 FTDQAKSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDV 352
F+ Q +AKA++ VMP+ H +C WHL +++ N +K +S F+ +F+ C+ YE+
Sbjct: 490 FSHQDAVVAKAVSIVMPDTTHVICAWHLKHAATRNI-NQLKSDSDFMKEFKACINLYEEE 548
Query: 353 EQF 355
+F
Sbjct: 549 TEF 551
>UniRef100_Q7F9U4 OSJNBb0006L01.4 protein [Oryza sativa]
Length = 513
Score = 224 bits (571), Expect = 3e-57
Identities = 102/182 (56%), Positives = 132/182 (72%)
Query: 175 LVQGEAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDS 234
+ G+AG +L YFQ K ENP+F +A Q+D E+QI N+FWADA+M+ DY +FGDVVS D+
Sbjct: 1 MAYGQAGSMLMYFQDKIAENPSFQYALQMDSEEQIANIFWADAKMISDYVHFGDVVSFDT 60
Query: 235 TYCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFT 294
T+ TN+ RP + GFNH R V+F AAL+YDET +S++WLFETFL+AH K P+T++T
Sbjct: 61 TFGTNNESRPFGVFVGFNHFRETVVFCAALMYDETFDSFKWLFETFLKAHNGKHPKTIYT 120
Query: 295 DQAKSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQ 354
DQ +M KA+ EV P A+HGLCT+H+ QN KHL GES LSD CMY YEDV +
Sbjct: 121 DQDIAMGKAIEEVFPAAWHGLCTFHISQNAAKHLSQGNNGESSILSDLSACMYEYEDVTK 180
Query: 355 FE 356
FE
Sbjct: 181 FE 182
>UniRef100_Q8S5U2 Putative far-red impaired response protein [Oryza sativa]
Length = 611
Score = 218 bits (555), Expect = 2e-55
Identities = 123/352 (34%), Positives = 186/352 (51%), Gaps = 13/352 (3%)
Query: 11 MFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKK-DGSVSSCRFVCCKEGMRKPDKRDY-- 67
M F S + F+N Y + GF VRKDY+ R + G + RF C +EG RK DY
Sbjct: 20 MKFSSEDEGFAFYNQYAKEKGFSVRKDYSRRDRFSGLIFHKRFTCSREGFRKEVYMDYSG 79
Query: 68 KTKNPRLETRTNCEARLGLK--NVDGKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIH 125
+T+ PR T C A +K + G V FV DHNH L + L S R+++
Sbjct: 80 RTREPRALTHCGCNAHFEIKLDEIKGHWYVTRFVADHNHPLCKADEVAFLRSHRRITPAQ 139
Query: 126 -CQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLL 184
+ +EL D +G+ Q + D++ ++ GG GF+ D N+ K +++ + G+A ++
Sbjct: 140 QAKLVELRD-LGLHQHQVMDVIERDHGGFEGAGFVTRDLYNFFVKMKKKRIDGGDADRVI 198
Query: 185 QYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRP 244
+Y Q + ++ FY+ Y+ D + +FWAD + +DY FGDVV DSTY N + P
Sbjct: 199 KYMQARQKDDMDFYYEYETDEAGCLKRLFWADPQSRIDYDAFGDVVVFDSTYRVNKYNLP 258
Query: 245 LAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKAL 304
G NHH VIFG A++ DE +YEW+ + FL QK P++V TD +M +A+
Sbjct: 259 FIPFVGVNHHGSTVIFGCAVVSDERVGTYEWVLKQFLSCMCQKHPKSVITDGDNAMRRAI 318
Query: 305 AEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFE 356
V P + H LCTWH+ QN ++L M LSDF ++ + ++FE
Sbjct: 319 LLVFPNSDHRLCTWHIEQNMARNLSPTM------LSDFRVLVHAPLEEDEFE 364
>UniRef100_Q8H344 Putative far-red impaired response protein [Oryza sativa]
Length = 903
Score = 212 bits (540), Expect = 1e-53
Identities = 114/349 (32%), Positives = 186/349 (52%), Gaps = 17/349 (4%)
Query: 13 FCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRKPDKRDYKTKNP 72
F S A+EF++ Y GK GF VR+ + ++ FVC KEG R+ + + K P
Sbjct: 237 FGSEDEAYEFYSMYAGKIGFNVRRASMTMNAENVITRRMFVCSKEGFREKKRGAKRVKKP 296
Query: 73 RLETRTNCEARLGLK-NVDGKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIHCQQIEL 131
R ETRT C A + ++ +GK V +FV HNH+L + ++++ + ++L
Sbjct: 297 RPETRTGCPACMVIRLTSNGKYHVTEFVTFHNHQLGATVPSDLVATSQSTETGQDDGLDL 356
Query: 132 AD-----SVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQY 186
D ++ Q ++++ + GR+N F +Y G+ G L+Y
Sbjct: 357 VDGSADANIHRQNLIIGNIMATSLEGRSNKRFKCTKVPHY-----------GDVGATLEY 405
Query: 187 FQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPLA 246
Q+ +NP+F+ A + D + +TN W+D++ ++D+ +FGDVV LDSTY RPLA
Sbjct: 406 LQKMQHDNPSFFFAVKSDDDGNLTNFLWSDSKSIMDFVHFGDVVCLDSTYALQGYGRPLA 465
Query: 247 IISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALAE 306
+ +G NHH+ VIF AALLYDE+ E++ WLF+TF A P+T+ TD++ ++++ +A
Sbjct: 466 LFTGVNHHKQTVIFAAALLYDESVEAFRWLFDTFKMAMNGTQPKTLLTDRSDAISEGVAA 525
Query: 307 VMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQF 355
P H C W + QN ++ L G F+KC++ ED +F
Sbjct: 526 SWPATAHRYCVWQIYQNALQQLSQAFHGSKTLDYCFQKCLFDSEDEPEF 574
Score = 83.6 bits (205), Expect = 8e-15
Identities = 43/119 (36%), Positives = 65/119 (54%), Gaps = 1/119 (0%)
Query: 2 NEDWNPKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRK 61
+E PKVDM F K A++F+N Y GF VR+ ++ FVC +EG R+
Sbjct: 97 DEKMLPKVDMLFDGEKEAYDFYNAYAEMVGFFVRRSTLWTTSKNIITRRTFVCSREGFRE 156
Query: 62 PDKRDYKTKNPRLETRTNCEARLGLK-NVDGKLMVHDFVEDHNHELLLPETTHMLSSQR 119
+ + K PR ETR C A + ++ N +GK + +FV +HNH+L T HML +++
Sbjct: 157 KKRGTKEAKCPRPETRIGCPASMTIRLNTNGKYRLTEFVPNHNHQLATASTMHMLKAKK 215
>UniRef100_Q75IS6 Putative Mutator-like transposase [Oryza sativa]
Length = 1510
Score = 212 bits (539), Expect = 1e-53
Identities = 120/363 (33%), Positives = 189/363 (52%), Gaps = 11/363 (3%)
Query: 3 EDWNPKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRKP 62
E P V M F +L + +F+ +Y + GF VR H+K++ + R+ C +EG K
Sbjct: 703 EHLKPMVGMIFDTLTDVEKFYKSYAHEAGFSVRVGQ-HKKQNEEILFKRYYCSREGYIKE 761
Query: 63 DKRDY-------KTKNP-RLETRTNCEARLGLK-NVDGKLMVHDFVEDHNHELLLPETTH 113
+D K K P +ETR CEA + +K D K + + +H+H + P+ H
Sbjct: 762 RVKDVSDESGKKKRKTPYMMETRCGCEAHIVVKLGSDKKYRISSMIGEHSHGFVSPDKRH 821
Query: 114 MLSSQRKVSEIHCQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRER 173
+L S R VSE + + ++F LL GG N+G + +NY R +
Sbjct: 822 LLRSNRTVSERAKSTLFSCHKASIGTSQAFRLLQVSDGGFQNVGCTLRNLQNYYHDLRCK 881
Query: 174 SLVQGEAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLD 233
+ +A + +RK NP F++ + +D + ++ VFWADA +YS FGDV+S+D
Sbjct: 882 -IKDADAQMFVGQLERKKEVNPAFFYEFMVDKQGRLVRVFWADAICRKNYSVFGDVLSVD 940
Query: 234 STYCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVF 293
STY TN + +G NHH +V GA+ L DE ES+ WLF+TFL+A P+ +
Sbjct: 941 STYSTNQYNMKFVPFTGVNHHLQSVFLGASFLADEKIESFVWLFQTFLKATGGVAPRLII 1000
Query: 294 TDQAKSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVE 353
TD+ SM A+A+++P H LC WH+M+ + +G ++ + F KC++G ED +
Sbjct: 1001 TDEDASMKAAIAQILPNTVHRLCMWHIMEKVPEKVGPSIREDGEFWDRLHKCVWGSEDSD 1060
Query: 354 QFE 356
FE
Sbjct: 1061 DFE 1063
>UniRef100_Q60DY0 Hypothetical protein OSJNBa0090H02.8 [Oryza sativa]
Length = 896
Score = 212 bits (539), Expect = 1e-53
Identities = 120/363 (33%), Positives = 189/363 (52%), Gaps = 11/363 (3%)
Query: 3 EDWNPKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRKP 62
E P V M F +L + +F+ +Y + GF VR H+K++ + R+ C +EG K
Sbjct: 94 EHLKPMVGMIFDTLTDVEKFYKSYAHEAGFSVRVGQ-HKKQNEEILFKRYYCSREGYIKE 152
Query: 63 DKRDY-------KTKNP-RLETRTNCEARLGLK-NVDGKLMVHDFVEDHNHELLLPETTH 113
+D K K P +ETR CEA + +K D K + + +H+H + P+ H
Sbjct: 153 RVKDVSDESGKKKRKTPYMMETRCGCEAHIVVKLGSDKKYRISSMIGEHSHGFVSPDKRH 212
Query: 114 MLSSQRKVSEIHCQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRER 173
+L S R VSE + + ++F LL GG N+G + +NY R +
Sbjct: 213 LLRSNRTVSERAKSTLFSCHKASIGTSQAFRLLQVSDGGFQNVGCTLRNLQNYYHDLRCK 272
Query: 174 SLVQGEAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLD 233
+ +A + +RK NP F++ + +D + ++ VFWADA +YS FGDV+S+D
Sbjct: 273 -IKDADAQMFVGQLERKKEVNPAFFYEFMVDKQGRLVRVFWADAICRKNYSVFGDVLSVD 331
Query: 234 STYCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVF 293
STY TN + +G NHH +V GA+ L DE ES+ WLF+TFL+A P+ +
Sbjct: 332 STYSTNQYNMKFVPFTGVNHHLQSVFLGASFLADEKIESFVWLFQTFLKATGGVAPRLII 391
Query: 294 TDQAKSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVE 353
TD+ SM A+A+++P H LC WH+M+ + +G ++ + F KC++G ED +
Sbjct: 392 TDEDASMKAAIAQILPNTVHRLCMWHIMEKVPEKVGPSIREDGEFWDRLHKCVWGSEDSD 451
Query: 354 QFE 356
FE
Sbjct: 452 DFE 454
>UniRef100_Q6K8S4 Putative far-red impaired response protein [Oryza sativa]
Length = 880
Score = 210 bits (534), Expect = 6e-53
Identities = 119/359 (33%), Positives = 186/359 (51%), Gaps = 11/359 (3%)
Query: 3 EDWNPKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKK-DGSVSSCRFVCCKEGMRK 61
E++ M F + K + F+N Y + GF VRK+Y R +V+ ++ C +EG RK
Sbjct: 53 EEYRTMTAMRFKTEKEGFLFYNRYAKEKGFSVRKNYIRRDPITVAVTHRQYQCSREGHRK 112
Query: 62 PDKRDY--KTKNPRLETRTNCEARLGLK--NVDGKLMVHDFVEDHNHELLLPETTHMLSS 117
+ +++ P+ TR C A +K G V +V HNH L + L S
Sbjct: 113 EVYMEAANRSREPKALTRCGCNALFEIKLDKKKGDWFVVKYVAKHNHPLAKSDEVAFLRS 172
Query: 118 QRKVSEIHCQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQ 177
R +S I VG++Q + D++ + GG GF+ D NY + R++ ++
Sbjct: 173 HRTISNAQKANILELKEVGLRQHQVMDVMERHHGGFDATGFVSRDLYNYFTRLRKKHILG 232
Query: 178 GEAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYC 237
G+A +++YFQ + + F+ Y+ D ++ +FW+D + +DY FGDVV DSTY
Sbjct: 233 GDAERVIKYFQWRQKHDMEFFFEYETDEAGRLKRLFWSDPQSRIDYDAFGDVVVFDSTYR 292
Query: 238 TNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQA 297
N + P G NHH +IFG A++ DE +YEW+ + FL+ QK P+ + TD
Sbjct: 293 VNRYNLPFIPFVGVNHHGSTIIFGCAIVADEKVATYEWILKQFLDCMYQKHPRALITDGD 352
Query: 298 KSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFE 356
+M +A+A VMP++ H LCTWH+ QN +HL M LSDF ++ D E+FE
Sbjct: 353 NAMRRAIAAVMPDSEHWLCTWHIEQNMARHLRPDM------LSDFRTLVHAPYDHEEFE 405
>UniRef100_Q9SWG3 Far-red impaired response protein [Arabidopsis thaliana]
Length = 827
Score = 207 bits (528), Expect = 3e-52
Identities = 113/356 (31%), Positives = 193/356 (53%), Gaps = 4/356 (1%)
Query: 4 DWNPKVDMFFCSLKNAWEFWNTYXGKXGFGVR-KDYTHRKKDGSVSSCRFVCCKEGMRKP 62
D P+ + F + + A+ F+ Y GF K+ KK +F C + G+
Sbjct: 48 DLEPRNGIDFDTHEAAYIFYQEYAKSMGFTTSIKNSRRSKKTKDFIDAKFACSRYGVTPE 107
Query: 63 DKRDYKTKNPRLETRTNCEARLGLKN-VDGKLMVHDFVEDHNHELLLPETTHMLSSQRKV 121
+ + +T+C+A + +K DGK ++H+FV+DHNHELL P + QR V
Sbjct: 108 SESSGSSSRRSTVKKTDCKASMHVKRRPDGKWIIHEFVKDHNHELL-PALAYHFRIQRNV 166
Query: 122 SEIHCQQIELADSVGVQQKKSFDLLSKEVGGRTNLG-FIRLDQKNYLRKKRERSLVQGEA 180
I++ +V + KK + +S++ GG N+G ++ D + + K R +L +G++
Sbjct: 167 KLAEKNNIDILHAVSERTKKMYVEMSRQSGGYKNIGSLLQTDVSSQVDKGRYLALEEGDS 226
Query: 181 GYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNS 240
LL+YF+R ENP F++A L+ + ++ N+FWADA+ DY F DVVS D+TY +
Sbjct: 227 QVLLEYFKRIKKENPKFFYAIDLNEDQRLRNLFWADAKSRDDYLSFNDVVSFDTTYVKFN 286
Query: 241 SHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSM 300
PLA+ G NHH ++ G AL+ DE+ E++ WL +T+L A + P+ + TDQ K +
Sbjct: 287 DKLPLALFIGVNHHSQPMLLGCALVADESMETFVWLIKTWLRAMGGRAPKVILTDQDKFL 346
Query: 301 AKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFE 356
A++E++P H WH+++ ++ ++MK FL F KC++ ++F+
Sbjct: 347 MSAVSELLPNTRHCFALWHVLEKIPEYFSHVMKRHENFLLKFNKCIFRSWTDDEFD 402
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 614,069,876
Number of Sequences: 2790947
Number of extensions: 24491489
Number of successful extensions: 52514
Number of sequences better than 10.0: 237
Number of HSP's better than 10.0 without gapping: 172
Number of HSP's successfully gapped in prelim test: 65
Number of HSP's that attempted gapping in prelim test: 52122
Number of HSP's gapped (non-prelim): 286
length of query: 362
length of database: 848,049,833
effective HSP length: 128
effective length of query: 234
effective length of database: 490,808,617
effective search space: 114849216378
effective search space used: 114849216378
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 75 (33.5 bits)
Medicago: description of AC148236.2


