

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148098.5 + phase: 0
(189 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
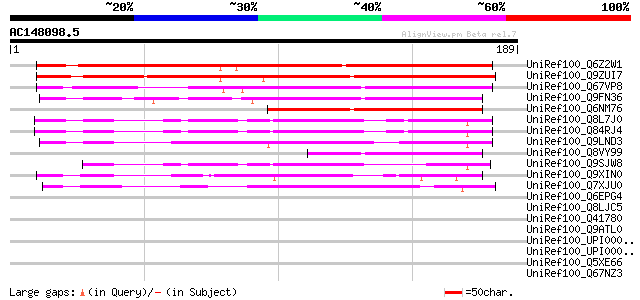
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6Z2W1 Hypothetical protein OJ1734_E02.2 [Oryza sativa] 198 5e-50
UniRef100_Q9ZUI7 T2K10.11 protein [Arabidopsis thaliana] 192 4e-48
UniRef100_Q67VP8 Hypothetical protein OSJNBb0061B07.12 [Oryza sa... 170 2e-41
UniRef100_Q9FN36 Gb|AAF34833.1 [Arabidopsis thaliana] 109 4e-23
UniRef100_Q6NM76 At3g15240 [Arabidopsis thaliana] 83 3e-15
UniRef100_Q8L7J0 Hypothetical protein At2g31280/F16D14.12 [Arabi... 72 1e-11
UniRef100_Q84RJ4 Hypothetical protein At2g31280/F16D14.12 [Arabi... 71 1e-11
UniRef100_Q9LND3 T21E18.20 protein [Arabidopsis thaliana] 69 5e-11
UniRef100_Q8VY99 Hypothetical protein At5g53900 [Arabidopsis tha... 63 4e-09
UniRef100_Q9SJW8 Hypothetical protein At2g31280 [Arabidopsis tha... 56 5e-07
UniRef100_Q9XIN0 Expressed protein [Arabidopsis thaliana] 51 2e-05
UniRef100_Q7XJU0 BHLH transcription factor [Arabidopsis thaliana] 47 3e-04
UniRef100_Q6EPG4 Transcription factor-like [Oryza sativa] 41 0.014
UniRef100_Q8LJC5 P0505D12.9 protein [Oryza sativa] 39 0.069
UniRef100_Q41780 Regulatory protein [Zea mays] 39 0.091
UniRef100_Q9ATL0 Anthocyanin regulatory B protein [Zea mays] 39 0.091
UniRef100_UPI000034B9A7 UPI000034B9A7 UniRef100 entry 34 2.2
UniRef100_UPI000049A559 UPI000049A559 UniRef100 entry 32 6.5
UniRef100_Q5XE66 Hypothetical protein [Streptococcus pyogenes] 32 6.5
UniRef100_Q67NZ3 Branched-chain amino acid ABC transporter perme... 32 6.5
>UniRef100_Q6Z2W1 Hypothetical protein OJ1734_E02.2 [Oryza sativa]
Length = 336
Score = 198 bits (504), Expect = 5e-50
Identities = 99/184 (53%), Positives = 124/184 (66%), Gaps = 19/184 (10%)
Query: 11 LLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVW 70
LLQHTLR LC+ S+WVYAVFWR++PR +PPP+W+ G DRS+GN+RNWI+ W
Sbjct: 13 LLQHTLRGLCT----QGDSQWVYAVFWRILPRNYPPPKWDLQGGVYDRSRGNRRNWILAW 68
Query: 71 EDGFCDF--NECEQR------------KSGCLNERFGADVFFKMSHEVYSYGEGLVGKVA 116
EDGFC+F + C+Q +G + ++FFKMSH++Y+YGEGLVGKVA
Sbjct: 69 EDGFCNFAASACDQEDTPAAAGYTDYAAAGHEVKGLQPELFFKMSHDIYNYGEGLVGKVA 128
Query: 117 ADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKEGLVQLGS 176
AD+GHKWV S N E N V WN D PR WE Q SGI+TIA+IAV+EG+VQLGS
Sbjct: 129 ADHGHKWV-SQEANEHEINLVTSWNNPADSHPRTWEAQFQSGIKTIALIAVREGVVQLGS 187
Query: 177 FDKV 180
KV
Sbjct: 188 MKKV 191
>UniRef100_Q9ZUI7 T2K10.11 protein [Arabidopsis thaliana]
Length = 386
Score = 192 bits (488), Expect = 4e-48
Identities = 95/189 (50%), Positives = 129/189 (67%), Gaps = 24/189 (12%)
Query: 11 LLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVW 70
LLQHTLR+LC +S+WVYAVFWR++PR +PPP+W+ G + DRS+GN+RNWI+VW
Sbjct: 13 LLQHTLRSLCIHE----NSQWVYAVFWRILPRNYPPPKWD-GQGAYDRSRGNRRNWILVW 67
Query: 71 EDGFCDF-------NECEQRKSGCLNERFG-----------ADVFFKMSHEVYSYGEGLV 112
EDGFC+F + E G + +G ++FFKMSHE+Y+YGEGL+
Sbjct: 68 EDGFCNFAASAAEMSSGEGSGGGGGSAAYGNSDFQQYQGLQPELFFKMSHEIYNYGEGLI 127
Query: 113 GKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKEGLV 172
GKVAAD+ HKW+Y + N E N++ W+ S D PR WE Q SGI+TIA+I+V+EG+V
Sbjct: 128 GKVAADHSHKWIYKE-PNDQEINFLSAWHNSADSYPRTWEAQFQSGIKTIALISVREGVV 186
Query: 173 QLGSFDKVL 181
QLG+ KV+
Sbjct: 187 QLGAVHKVI 195
>UniRef100_Q67VP8 Hypothetical protein OSJNBb0061B07.12 [Oryza sativa]
Length = 353
Score = 170 bits (430), Expect = 2e-41
Identities = 90/188 (47%), Positives = 112/188 (58%), Gaps = 40/188 (21%)
Query: 11 LLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVW 70
LLQHTLR+LC TS S+WVYAVFWR++PR +PPP+ I+ W
Sbjct: 13 LLQHTLRSLC---TSGDDSQWVYAVFWRILPRNYPPPK------------------ILAW 51
Query: 71 EDGFCDFN-----------------ECEQRKS-GCLNERFGADVFFKMSHEVYSYGEGLV 112
EDGFC+F ECE+ K G ++FFKMSH++Y+YGEGL+
Sbjct: 52 EDGFCNFAATSAACGDGAAAAYAAAECEETKQVGVAGGGLQPELFFKMSHDIYNYGEGLI 111
Query: 113 GKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKEGLV 172
GKVAAD+ HKWV+ + Q E N + WN D PR WE Q SGIQTIA+IAV+EG+V
Sbjct: 112 GKVAADHSHKWVFKEPQEQ-EINLISSWNNPADSHPRTWEAQFQSGIQTIALIAVREGVV 170
Query: 173 QLGSFDKV 180
QLGS KV
Sbjct: 171 QLGSMKKV 178
>UniRef100_Q9FN36 Gb|AAF34833.1 [Arabidopsis thaliana]
Length = 377
Score = 109 bits (272), Expect = 4e-23
Identities = 69/173 (39%), Positives = 89/173 (50%), Gaps = 25/173 (14%)
Query: 12 LQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFG-GTSLDRSKGNKRNWIIVW 70
L LR++C +S W+Y+VFW + PR PR G G + G+ +++W
Sbjct: 21 LHEALRSVCF------NSDWIYSVFWTIRPR----PRVRGGNGCKIGDESGSL---MLMW 67
Query: 71 EDGFCDFNECEQRKSGCLN-------ERFGADVFFKMSHEVYSYGEGLVGKVAADNGHKW 123
EDGFC E CL E F KMS ++Y+YGEGL+GKVA+D HKW
Sbjct: 68 EDGFCGGGRSEDL---CLETDIEGHEEDLVRKAFSKMSIQLYNYGEGLMGKVASDKCHKW 124
Query: 124 VYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKEGLVQLGS 176
V+ + E N W +S D P W Q SGIQTIAVI GL+QLGS
Sbjct: 125 VFKEPSES-EPNLANYWQSSFDALPPEWTDQFESGIQTIAVIQAGHGLLQLGS 176
>UniRef100_Q6NM76 At3g15240 [Arabidopsis thaliana]
Length = 264
Score = 83.2 bits (204), Expect = 3e-15
Identities = 42/80 (52%), Positives = 53/80 (65%), Gaps = 1/80 (1%)
Query: 97 FFKMSHEVYSYGEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLN 156
F KMS ++Y+YGEGL+GKVA+D HKWV+ + Q E+N W +S D P W Q
Sbjct: 11 FSKMSIQLYNYGEGLMGKVASDKCHKWVFKE-QTESESNASSYWQSSFDAIPSEWNDQFE 69
Query: 157 SGIQTIAVIAVKEGLVQLGS 176
SGI+TIAVI GL+QLGS
Sbjct: 70 SGIRTIAVIQAGHGLLQLGS 89
>UniRef100_Q8L7J0 Hypothetical protein At2g31280/F16D14.12 [Arabidopsis thaliana]
Length = 720
Score = 71.6 bits (174), Expect = 1e-11
Identities = 53/172 (30%), Positives = 84/172 (48%), Gaps = 40/172 (23%)
Query: 10 SLLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIV 69
S Q L++ C ++ W YAVFW++ +G++ ++
Sbjct: 3 STSQEILKSFCF------NTDWDYAVFWQL------------------NHRGSRM--VLT 36
Query: 70 WEDGFCDFNECEQRKSGCLNERFGADVFFKMSHEVYSYGEGLVGKVAADNGHKWVYSDTQ 129
ED + D + + ++ G V KMS+ VYS GEG+VG+VA H+WV+ +
Sbjct: 37 LEDAYYDHHGTNMHGA---HDPLGXAVA-KMSYHVYSLGEGIVGQVAVSGEHQWVFPENY 92
Query: 130 NGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKE-GLVQLGSFDKV 180
N C N++ + WE Q+++GI+TI V+AV G+VQLGS KV
Sbjct: 93 NNC--------NSAFEFH-NVWESQISAGIKTILVVAVGPCGVVQLGSLCKV 135
>UniRef100_Q84RJ4 Hypothetical protein At2g31280/F16D14.12 [Arabidopsis thaliana]
Length = 720
Score = 71.2 bits (173), Expect = 1e-11
Identities = 53/172 (30%), Positives = 84/172 (48%), Gaps = 40/172 (23%)
Query: 10 SLLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIV 69
S Q L++ C ++ W YAVFW++ +G++ ++
Sbjct: 3 STSQEILKSFCF------NTDWDYAVFWQL------------------NHRGSRM--VLT 36
Query: 70 WEDGFCDFNECEQRKSGCLNERFGADVFFKMSHEVYSYGEGLVGKVAADNGHKWVYSDTQ 129
ED + D + + ++ G V KMS+ VYS GEG+VG+VA H+WV+ +
Sbjct: 37 LEDAYYDHHGTNMHGA---HDPLGLAVA-KMSYHVYSLGEGIVGQVAVSGEHQWVFPENY 92
Query: 130 NGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKE-GLVQLGSFDKV 180
N C N++ + WE Q+++GI+TI V+AV G+VQLGS KV
Sbjct: 93 NNC--------NSAFEFH-NVWESQISAGIKTILVVAVGPCGVVQLGSLCKV 135
>UniRef100_Q9LND3 T21E18.20 protein [Arabidopsis thaliana]
Length = 1322
Score = 69.3 bits (168), Expect = 5e-11
Identities = 51/174 (29%), Positives = 78/174 (44%), Gaps = 41/174 (23%)
Query: 12 LQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVWE 71
LQ LR++CS ++ W YAVFW++ + ++ E
Sbjct: 5 LQQILRSICS------NTDWNYAVFWKL---------------------NHHSPMVLTLE 37
Query: 72 DGFCDFNECEQRKSGCLNERFGAD----VFFKMSHEVYSYGEGLVGKVAADNGHKWVYSD 127
D +C +E R D KMS+ V+S GEG+VG+VA H+W++S+
Sbjct: 38 DVYCVNHERGLMPESLHGGRHAHDPLGLAVAKMSYHVHSLGEGIVGQVAISGQHQWIFSE 97
Query: 128 TQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKE-GLVQLGSFDKV 180
N + WE Q+++GI+TI ++AV G+VQLGS KV
Sbjct: 98 YLNDSHSTL---------QVHNGWESQISAGIKTILIVAVGSCGVVQLGSLCKV 142
>UniRef100_Q8VY99 Hypothetical protein At5g53900 [Arabidopsis thaliana]
Length = 265
Score = 62.8 bits (151), Expect = 4e-09
Identities = 32/65 (49%), Positives = 38/65 (58%), Gaps = 1/65 (1%)
Query: 112 VGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKEGL 171
+GKVA+D HKWV+ + E N W +S D P W Q SGIQTIAVI GL
Sbjct: 1 MGKVASDKCHKWVFKEPSES-EPNLANYWQSSFDALPPEWTDQFESGIQTIAVIQAGHGL 59
Query: 172 VQLGS 176
+QLGS
Sbjct: 60 LQLGS 64
>UniRef100_Q9SJW8 Hypothetical protein At2g31280 [Arabidopsis thaliana]
Length = 723
Score = 55.8 bits (133), Expect = 5e-07
Identities = 44/153 (28%), Positives = 67/153 (43%), Gaps = 48/153 (31%)
Query: 28 SSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVWEDGFCDFNECEQRKSGC 87
++ W YAVFW++ +G++ ++ ED + D + +
Sbjct: 41 NTDWDYAVFWQL------------------NHRGSRM--VLTLEDAYYDHHGTNMHGA-- 78
Query: 88 LNERFGADVFFKMSHEVYSYGEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQ 147
++ G V KMS+ VYS GEG+VG+VA H+WV+ + N C
Sbjct: 79 -HDPLGLAVA-KMSYHVYSLGEGIVGQVAVSGEHQWVFPENYNNC--------------- 121
Query: 148 PRAWEFQLNSGIQTIAVIAVKE-GLVQLGSFDK 179
NS +TI V+AV G+VQLGS K
Sbjct: 122 --------NSAFETILVVAVGPCGVVQLGSLCK 146
>UniRef100_Q9XIN0 Expressed protein [Arabidopsis thaliana]
Length = 650
Score = 50.8 bits (120), Expect = 2e-05
Identities = 46/181 (25%), Positives = 78/181 (42%), Gaps = 57/181 (31%)
Query: 11 LLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVW 70
LL+ LR++C +++W YAVFW++ G + + +++W
Sbjct: 4 LLREALRSMC------VNNQWSYAVFWKI---------------------GCQNSSLLIW 36
Query: 71 EDGFCDFNECEQRKSGCLNERFGADVF-----------FKMSHEVYSYGEGLVGKVAADN 119
E+ C +NE E + G D +++ + GEGLVG+ A
Sbjct: 37 EE--C-YNETESSSNPRRLCGLGVDTQGNEKVQLLTNRMMLNNRIILVGEGLVGRAAFTG 93
Query: 120 GHKWVYSDTQNGCEANYVGPWNASIDHQPRAWE---FQLNSGIQTIAVI-AVKEGLVQLG 175
H+W+ +++ +N + H P Q ++GIQT+AV V G+VQLG
Sbjct: 94 HHQWILANS-----------FNRDV-HPPEVINEMLLQFSAGIQTVAVFPVVPHGVVQLG 141
Query: 176 S 176
S
Sbjct: 142 S 142
>UniRef100_Q7XJU0 BHLH transcription factor [Arabidopsis thaliana]
Length = 527
Score = 47.0 bits (110), Expect = 3e-04
Identities = 43/170 (25%), Positives = 69/170 (40%), Gaps = 47/170 (27%)
Query: 13 QHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVWED 72
+H L++LC S W YAVFWR P + I+ +E+
Sbjct: 6 KHILKSLC------LSHGWSYAVFWRYDPI---------------------NSMILRFEE 38
Query: 73 GFCDFNECEQRKSGCLNERFGADVFFKMSHEVYSYGEGLVGKVAADNGHKWVYSDTQNGC 132
+ N+ + M + G+G+VG+VA+ H+W++SDT
Sbjct: 39 AY--------------NDEQSVALVDDMVLQAPILGQGIVGEVASSGNHQWLFSDTLFQW 84
Query: 133 EANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAV-KEGLVQLGSFDKVL 181
E + + R + + QTIA+I + G+VQLGS K+L
Sbjct: 85 EHEFQNQFLCGFKILIRQFTY-----TQTIAIIPLGSSGVVQLGSTQKIL 129
>UniRef100_Q6EPG4 Transcription factor-like [Oryza sativa]
Length = 792
Score = 41.2 bits (95), Expect = 0.014
Identities = 28/87 (32%), Positives = 44/87 (50%), Gaps = 3/87 (3%)
Query: 95 DVFFKMSHEVYSYGEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQ 154
+V KM ++V+ GEG++G+ +W+ SDT A N + W+ Q
Sbjct: 45 NVVEKMLNQVHVVGEGIIGRALVSGECQWI-SDTSFSF-AQTSDADNQDLFQGYTWWQHQ 102
Query: 155 LNSGIQTIAVIAVKE-GLVQLGSFDKV 180
GI+TIAVI + + G+ Q GS K+
Sbjct: 103 FLCGIKTIAVIPIADLGVAQFGSMQKI 129
>UniRef100_Q8LJC5 P0505D12.9 protein [Oryza sativa]
Length = 979
Score = 38.9 bits (89), Expect = 0.069
Identities = 49/196 (25%), Positives = 73/196 (37%), Gaps = 62/196 (31%)
Query: 12 LQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWE--FGGTSLDRSKGNKRNWIIV 69
L+ +LR LC T W YAVFWR R R + +G +R+ G
Sbjct: 184 LRDSLRRLC------TDVGWSYAVFWRAT-RAADSQRLKLVWGDGHYERAAGAPSI---- 232
Query: 70 WEDGFCDFNECEQRKSGCL-----------------------NERFGADVFFKMSHEVYS 106
GF + + K+ L ++R A V M+ +V+
Sbjct: 233 --SGFEAMDLLLKEKAAALRSGTGRGGGGGEGHAADGAAGHSHDRVDALVHKAMAQQVHV 290
Query: 107 YGEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIA 166
GEG++G+ A H+W+ D + CE +TIAVI
Sbjct: 291 VGEGVIGQAALTGLHRWIVHDIVDECEEE-----------------------DETIAVIP 327
Query: 167 V-KEGLVQLGSFDKVL 181
V G++QLGS V+
Sbjct: 328 VLPRGVIQLGSTKMVM 343
>UniRef100_Q41780 Regulatory protein [Zea mays]
Length = 562
Score = 38.5 bits (88), Expect = 0.091
Identities = 52/202 (25%), Positives = 78/202 (37%), Gaps = 39/202 (19%)
Query: 3 SGLPLLHSLLQHTLRNLCS-FPTSSTSSKWVYAVFWRVV----PRIFPPPRWEFGGTSLD 57
S P LLQ R L ++ S W YA+FW + PR+ + G
Sbjct: 4 SASPAQEELLQPAGRPLRKQLAAAARSINWSYALFWSISSTQRPRVLTWTDGFYNGEVKT 63
Query: 58 RSKGNK----------------RNWIIVWEDGFCDFNECEQRKSGCLN-ERFGADVFFKM 100
R + R G CD R G L+ E G ++ +
Sbjct: 64 RKISHSVELTADQLLMQRSEQLRELYEALRSGECDRRGA--RPVGSLSPEDLGDTEWYYV 121
Query: 101 SHEVYSY--GEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSG 158
Y++ G+GL G+ +A N H W+ C A+ G + PRA ++
Sbjct: 122 ICMTYAFLPGQGLPGRSSASNEHVWL-------CNAHLAGSKDF-----PRAL-LAKSAS 168
Query: 159 IQTIAVIAVKEGLVQLGSFDKV 180
IQTI I + G+++LG+ DKV
Sbjct: 169 IQTIVCIPLMGGVLELGTTDKV 190
>UniRef100_Q9ATL0 Anthocyanin regulatory B protein [Zea mays]
Length = 191
Score = 38.5 bits (88), Expect = 0.091
Identities = 52/202 (25%), Positives = 79/202 (38%), Gaps = 39/202 (19%)
Query: 3 SGLPLLHSLLQHTLRNLCS-FPTSSTSSKWVYAVFWRVV----PRIFPPPRWEFGGTSLD 57
S P LLQ R L ++ S W YA+FW + PR+ + G
Sbjct: 4 SACPAQEELLQPAGRPLRKQLAAAARSINWSYALFWSISSTQRPRVLTWTDRFYNGEVKT 63
Query: 58 RSKGNK----------------RNWIIVWEDGFCDFNECEQRKSGCLN-ERFGADVFFKM 100
R + R + G CD R G L+ E G ++ +
Sbjct: 64 RKISHSVELTADQLLMQRSEQLRELYEALQSGECDRRAA--RPVGSLSPEDLGDTEWYYV 121
Query: 101 SHEVYSY--GEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSG 158
Y++ G+GL G+ +A N H W+ C A+ G + PRA ++
Sbjct: 122 ICMTYAFLPGQGLPGRSSASNEHVWL-------CNAHLAGSKDF-----PRAL-LAKSAC 168
Query: 159 IQTIAVIAVKEGLVQLGSFDKV 180
IQTI I + G+++LG+ DKV
Sbjct: 169 IQTIVCIPLMGGVLELGTTDKV 190
>UniRef100_UPI000034B9A7 UPI000034B9A7 UniRef100 entry
Length = 310
Score = 33.9 bits (76), Expect = 2.2
Identities = 24/84 (28%), Positives = 35/84 (41%), Gaps = 2/84 (2%)
Query: 107 YGEGLVGKVAADNGHKWVYSDTQNGCEANYVG--PWNASIDHQPRAWEFQLNSGIQTIAV 164
+G+G+ GK D V NG EAN V ++ + + A E + SGI +
Sbjct: 29 FGDGIFGKALEDGNIVEVSYILSNGSEANGVSNVSFSGKLTYNRNAVENTITSGISLLTA 88
Query: 165 IAVKEGLVQLGSFDKVLLFFPSKY 188
G ++ S D V F P Y
Sbjct: 89 TNPSSGGDEIESVDSVKKFAPQIY 112
>UniRef100_UPI000049A559 UPI000049A559 UniRef100 entry
Length = 219
Score = 32.3 bits (72), Expect = 6.5
Identities = 33/138 (23%), Positives = 53/138 (37%), Gaps = 23/138 (16%)
Query: 14 HTLRNLCSFPTSSTSS----------KWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKG-- 61
+ + N + PT TSS ++ + W ++ R+ W +G T KG
Sbjct: 43 NAINNWGNSPTKETSSPENVEMPPEEQYKMPLSWIILKRVVQGRDWHYGRTDKKEEKGVV 102
Query: 62 ---NKRNWIIV-WEDGFCDFNECEQRKSGCLNERFGADVF----FKMSHEVYSYGEGLVG 113
NK I V W+DG D C K G + ++ + S +V+ L+G
Sbjct: 103 FSVNKDGQITVRWDDG--DKTHCNWGKDGNFDVTVISNALDMTPVEPSEQVHKPISWLIG 160
Query: 114 KVAADNGHKWVYSDTQNG 131
+ G W +SD G
Sbjct: 161 R-KVKRGTDWKWSDQDGG 177
>UniRef100_Q5XE66 Hypothetical protein [Streptococcus pyogenes]
Length = 221
Score = 32.3 bits (72), Expect = 6.5
Identities = 28/126 (22%), Positives = 49/126 (38%), Gaps = 11/126 (8%)
Query: 25 SSTSSKWVYAVFWRVVPRIFPPPR-----WEFGGTSLDRSKGNKRNWIIVWEDGFCDFNE 79
+ST +WVY F +P+ P+ +G ++ + W WE+ +FN+
Sbjct: 71 NSTKEQWVYPDFSVFLPKGVKAPKEITYEHHYGNGTVHSETRSNTQWHYNWENQQTNFNQ 130
Query: 80 CEQRKSGCLNERFGADVFFKMSHEVYSYGEGLVGKVAADN----GHKWVYSDTQNGCEAN 135
+ G D F+K+ ++ + LV + HK V+ + E N
Sbjct: 131 EFDKFPGYTGWSTSLDKFYKLKND-GKFSHVLVDTYGRQSHTYFSHKMVWK-FKTELEDN 188
Query: 136 YVGPWN 141
Y WN
Sbjct: 189 YKDKWN 194
>UniRef100_Q67NZ3 Branched-chain amino acid ABC transporter permease protein
[Symbiobacterium thermophilum]
Length = 296
Score = 32.3 bits (72), Expect = 6.5
Identities = 18/53 (33%), Positives = 28/53 (51%), Gaps = 2/53 (3%)
Query: 4 GLPLLHSLLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSL 56
G P + +L+ T + F TS +V+ +R VPR+ P RWE GG ++
Sbjct: 91 GAPRIAALI--TAIGVSLFIEYFTSLNFVFGSDYRNVPRVLPDRRWELGGITV 141
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.138 0.457
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 364,394,886
Number of Sequences: 2790947
Number of extensions: 15448463
Number of successful extensions: 32133
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 32087
Number of HSP's gapped (non-prelim): 24
length of query: 189
length of database: 848,049,833
effective HSP length: 120
effective length of query: 69
effective length of database: 513,136,193
effective search space: 35406397317
effective search space used: 35406397317
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 71 (32.0 bits)
Medicago: description of AC148098.5


