

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
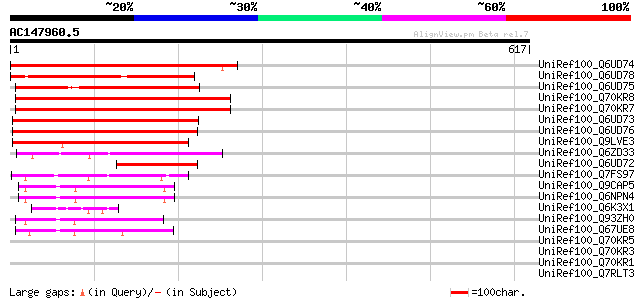
Query= AC147960.5 - phase: 0
(617 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6UD74 LysM domain-containing receptor-like kinase 7 [... 457 e-127
UniRef100_Q6UD78 LysM domain-containing receptor-like kinase 1 [... 285 2e-75
UniRef100_Q6UD75 LysM domain-containing receptor-like kinase 6 [... 251 6e-65
UniRef100_Q70KR8 Nod-facor receptor 1a [Lotus japonicus] 236 1e-60
UniRef100_Q70KR7 Nod-facor receptor 1b [Lotus japonicus] 236 1e-60
UniRef100_Q6UD73 LysM domain-containing receptor-like kinase 3 [... 221 6e-56
UniRef100_Q6UD76 LysM domain-containing receptor-like kinase 4 [... 217 7e-55
UniRef100_Q9LVE3 Similarity to receptor protein kinase [Arabidop... 214 8e-54
UniRef100_Q6ZD33 Receptor protein kinase PERK1-like protein [Ory... 172 3e-41
UniRef100_Q6UD72 LysM domain-containing receptor-like kinase 4 [... 119 2e-25
UniRef100_Q7FS97 Hypothetical protein [Sorghum bicolor] 108 6e-22
UniRef100_Q9CAP5 Hypothetical protein T5M16.22 [Arabidopsis thal... 58 7e-07
UniRef100_Q6NPN4 At1g77630 [Arabidopsis thaliana] 58 7e-07
UniRef100_Q6K3X1 Receptor protein kinase PERK1-like [Oryza sativa] 56 4e-06
UniRef100_Q93ZH0 LysM-domain GPI-anchored protein 1 precursor [A... 54 1e-05
UniRef100_Q67UE8 Putative Erwinia induced protein 1 [Oryza sativa] 49 3e-04
UniRef100_Q70KR5 SYM10 protein [Pisum sativum] 47 0.001
UniRef100_Q70KR3 SYM10 protein [Pisum sativum] 47 0.001
UniRef100_Q70KR1 Nod-factor receptor 5 [Lotus japonicus] 40 0.16
UniRef100_Q7RLT3 Unnamed protein product [Plasmodium yoelii yoelii] 39 0.59
>UniRef100_Q6UD74 LysM domain-containing receptor-like kinase 7 [Medicago truncatula]
Length = 620
Score = 457 bits (1176), Expect = e-127
Identities = 234/280 (83%), Positives = 244/280 (86%), Gaps = 10/280 (3%)
Query: 1 MKPIKFRLSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKP 60
MKPIKFRLSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKP
Sbjct: 1 MKPIKFRLSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKP 60
Query: 61 EDIVSYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYS 120
EDIVSYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYS
Sbjct: 61 EDIVSYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYS 120
Query: 121 NLTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELIS 180
NLTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELIS
Sbjct: 121 NLTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELIS 180
Query: 181 NKTEIDAELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHTRSIRWSYNWNICWSSSRT 240
NKTEIDAELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFH + S S
Sbjct: 181 NKTEIDAELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHISTGGLSGGVITGISVGAV 240
Query: 241 NSIIILHLC-YV-------LPKEE--DKEARISFGRIQRD 270
+I+L C YV + K+E +E+ FG+++ D
Sbjct: 241 AGLILLSFCIYVTYYRKKKIRKQEFLSEESSAIFGQVKND 280
>UniRef100_Q6UD78 LysM domain-containing receptor-like kinase 1 [Medicago truncatula]
Length = 590
Score = 285 bits (730), Expect = 2e-75
Identities = 140/219 (63%), Positives = 171/219 (77%), Gaps = 8/219 (3%)
Query: 1 MKPIKFRLSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKP 60
MKPIKF LS L MLLAS S AESKCSKTC++ALASYY+ + TNLTY+SNIMQSN+V+KP
Sbjct: 1 MKPIKFILSLLLMLLASSS--AESKCSKTCDLALASYYIWEGTNLTYISNIMQSNVVSKP 58
Query: 61 EDIVSYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYS 120
DI SYNTDT+ N D ++ +R+NVPFPCDCI+DEFLGH F Y+ ++TY S+A +S
Sbjct: 59 LDIFSYNTDTLPNLDMLRFSSRLNVPFPCDCINDEFLGHTFLYEFHPRETYASIAELTFS 118
Query: 121 NLTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELIS 180
NLT EW++ N +PD+ +NVTVNCSCG+ VSKDYGLFITYPL ED+LE I+
Sbjct: 119 NLTNKEWMEKVN------VPDSVKVNVTVNCSCGDKMVSKDYGLFITYPLSSEDTLESIA 172
Query: 181 NKTEIDAELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVP 219
T++ ELLQKY PGVNFS+GSGLV+IPGKD+N YVP
Sbjct: 173 KHTKVKPELLQKYTPGVNFSKGSGLVFIPGKDKNGVYVP 211
>UniRef100_Q6UD75 LysM domain-containing receptor-like kinase 6 [Medicago truncatula]
Length = 574
Score = 251 bits (640), Expect = 6e-65
Identities = 122/219 (55%), Positives = 160/219 (72%), Gaps = 10/219 (4%)
Query: 7 RLSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKPEDIVSY 66
++SF F++L ESKC++ C++ALASY L +NLTY+SNIM+SN+++KP+DI+
Sbjct: 2 KISFSFIVLILLIASTESKCNEGCSLALASYTLNHVSNLTYISNIMKSNVLSKPQDIIIN 61
Query: 67 NTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTTSE 126
N NK R NVPFPC+CI+ EFL + F Y++ +TY SVA ++SNLTT
Sbjct: 62 NDK---NK-------RANVPFPCNCINGEFLAYTFLYELQPGETYTSVAEESFSNLTTDV 111
Query: 127 WLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELISNKTEID 186
W+QNFN Y +IPD + VTVNCSCGN +VS DYGLFITYPLR ED+LE I+ EI+
Sbjct: 112 WMQNFNVYRPTNIPDFAMIKVTVNCSCGNKEVSMDYGLFITYPLRSEDTLESIAKGAEIE 171
Query: 187 AELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHTRSI 225
AELLQ+YNPGVNFS+GSGLV+IPGKDQN +Y+P H ++
Sbjct: 172 AELLQRYNPGVNFSKGSGLVFIPGKDQNGSYLPLHPSTV 210
>UniRef100_Q70KR8 Nod-facor receptor 1a [Lotus japonicus]
Length = 621
Score = 236 bits (602), Expect = 1e-60
Identities = 121/258 (46%), Positives = 172/258 (65%), Gaps = 3/258 (1%)
Query: 8 LSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTN-LTYVSNIMQSNLVTKPEDIVSY 66
L F +LL F ES C K C++ALASYY+ L ++ MQS +V+ + I SY
Sbjct: 8 LLFFILLLGHVCFHVESNCLKGCDLALASYYILPGVFILQNITTFMQSEIVSSNDAITSY 67
Query: 67 NTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTTSE 126
N D I N +QSF R+N+PFPCDCI EFLGH+F+Y + DTY ++A+ Y+NLTT +
Sbjct: 68 NKDKILNDINIQSFQRLNIPFPCDCIGGEFLGHVFEYSASKGDTYETIANLYYANLTTVD 127
Query: 127 WLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELISNKTEID 186
L+ FNSY +IP +NVTVNCSCGNS VSKDYGLFITYP+RP D+L+ I+N++ +D
Sbjct: 128 LLKRFNSYDPKNIPVNAKVNVTVNCSCGNSQVSKDYGLFITYPIRPGDTLQDIANQSSLD 187
Query: 187 AELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHTRSIRWSYNWNICWSSSRTNSIIIL 246
A L+Q +NP VNFS+ SG+ +IPG+ +N YVP + R+ + + S + T +++L
Sbjct: 188 AGLIQSFNPSVNFSKDSGIAFIPGRYKNGVYVPLYHRTAGLASGAAVGISIAGTFVLLLL 247
Query: 247 HLC-YV-LPKEEDKEARI 262
C YV K+E+++A++
Sbjct: 248 AFCMYVRYQKKEEEKAKL 265
>UniRef100_Q70KR7 Nod-facor receptor 1b [Lotus japonicus]
Length = 623
Score = 236 bits (602), Expect = 1e-60
Identities = 121/258 (46%), Positives = 172/258 (65%), Gaps = 3/258 (1%)
Query: 8 LSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTN-LTYVSNIMQSNLVTKPEDIVSY 66
L F +LL F ES C K C++ALASYY+ L ++ MQS +V+ + I SY
Sbjct: 8 LLFFILLLGHVCFHVESNCLKGCDLALASYYILPGVFILQNITTFMQSEIVSSNDAITSY 67
Query: 67 NTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTTSE 126
N D I N +QSF R+N+PFPCDCI EFLGH+F+Y + DTY ++A+ Y+NLTT +
Sbjct: 68 NKDKILNDINIQSFQRLNIPFPCDCIGGEFLGHVFEYSASKGDTYETIANLYYANLTTVD 127
Query: 127 WLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELISNKTEID 186
L+ FNSY +IP +NVTVNCSCGNS VSKDYGLFITYP+RP D+L+ I+N++ +D
Sbjct: 128 LLKRFNSYDPKNIPVNAKVNVTVNCSCGNSQVSKDYGLFITYPIRPGDTLQDIANQSSLD 187
Query: 187 AELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHTRSIRWSYNWNICWSSSRTNSIIIL 246
A L+Q +NP VNFS+ SG+ +IPG+ +N YVP + R+ + + S + T +++L
Sbjct: 188 AGLIQSFNPSVNFSKDSGIAFIPGRYKNGVYVPLYHRTAGLASGAAVGISIAGTFVLLLL 247
Query: 247 HLC-YV-LPKEEDKEARI 262
C YV K+E+++A++
Sbjct: 248 AFCMYVRYQKKEEEKAKL 265
>UniRef100_Q6UD73 LysM domain-containing receptor-like kinase 3 [Medicago truncatula]
Length = 620
Score = 221 bits (562), Expect = 6e-56
Identities = 109/223 (48%), Positives = 148/223 (65%), Gaps = 2/223 (0%)
Query: 4 IKFRLSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLV-TKPED 62
+K L + L F ESKC K C++ALASYY+ L +SN MQS +V T D
Sbjct: 3 LKNGLLLFILFLDCVFFKVESKCVKGCDVALASYYIIPSIQLRNISNFMQSKIVLTNSFD 62
Query: 63 IV-SYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSN 121
++ SYN D + +K + S+TR+NVPFPC+CI EFLGH+F+Y D Y +A+ Y++
Sbjct: 63 VIMSYNRDVVFDKSGLISYTRINVPFPCECIGGEFLGHVFEYTTKEGDDYDLIANTYYAS 122
Query: 122 LTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELISN 181
LTT E L+ FNSY N IP +NVTV CSCGNS +SKDYGLF+TYPLR +D+L I+
Sbjct: 123 LTTVELLKKFNSYDPNHIPVKAKINVTVICSCGNSQISKDYGLFVTYPLRSDDTLAKIAT 182
Query: 182 KTEIDAELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHTRS 224
K +D L+Q +N NFS GSG+V+IPG+DQN ++ P ++R+
Sbjct: 183 KAGLDEGLIQNFNQDANFSIGSGIVFIPGRDQNGHFFPLYSRT 225
>UniRef100_Q6UD76 LysM domain-containing receptor-like kinase 4 [Medicago truncatula]
Length = 624
Score = 217 bits (553), Expect = 7e-55
Identities = 108/222 (48%), Positives = 147/222 (65%), Gaps = 2/222 (0%)
Query: 4 IKFRLSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLV-TKPED 62
+K L + L F E+KC K C++ALASYY+ L VSN +QS +V T D
Sbjct: 3 LKNGLLLFILFLDCVFFKVETKCVKGCDVALASYYIMPSIQLINVSNFIQSKIVLTNSFD 62
Query: 63 IV-SYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSN 121
++ SYN + +K + S+TR+NVPFPC+CI EFLGH+F+Y D Y +A+ Y++
Sbjct: 63 VIMSYNRVVVFDKSGLISYTRINVPFPCECIGGEFLGHVFEYTTKEGDDYDLIANTYYAS 122
Query: 122 LTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELISN 181
LTT E L+ FNSY N IP +NVTV CSCGNS +SKD+GLF+TYPLR +D+L I+
Sbjct: 123 LTTVELLKKFNSYDPNHIPVKAKINVTVICSCGNSQISKDFGLFVTYPLRSDDTLAKIAT 182
Query: 182 KTEIDAELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHTR 223
K ++D LLQ +N NFS+GSG+V+IPG+D+N YVP +R
Sbjct: 183 KADLDEGLLQNFNQDANFSKGSGIVFIPGRDENGVYVPLPSR 224
>UniRef100_Q9LVE3 Similarity to receptor protein kinase [Arabidopsis thaliana]
Length = 603
Score = 214 bits (544), Expect = 8e-54
Identities = 106/213 (49%), Positives = 148/213 (68%), Gaps = 5/213 (2%)
Query: 4 IKFRLSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKPE-- 61
+K L +LL S F ESKC +C +ALASYYL++ T L+ ++ + S++ +
Sbjct: 3 LKISLIAPILLLFSFFFAVESKCRTSCPLALASYYLENGTTLSVINQNLNSSIAPYDQIN 62
Query: 62 --DIVSYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNY 119
I+ YN++ I +KD +Q +RV VPFPC+C +FLGH F Y V +DTY VA +NY
Sbjct: 63 FDPILRYNSN-IKDKDRIQMGSRVLVPFPCECQPGDFLGHNFSYSVRQEDTYERVAISNY 121
Query: 120 SNLTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELI 179
+NLTT E LQ N +P+ +IP + TLNV VNCSCG+ VSKD+GLF+TYPLRPEDSL I
Sbjct: 122 ANLTTMESLQARNPFPATNIPLSATLNVLVNCSCGDESVSKDFGLFVTYPLRPEDSLSSI 181
Query: 180 SNKTEIDAELLQKYNPGVNFSQGSGLVYIPGKD 212
+ + + A++LQ+YNPGVNF+ G+G+VY+PG+D
Sbjct: 182 ARSSGVSADILQRYNPGVNFNSGNGIVYVPGRD 214
>UniRef100_Q6ZD33 Receptor protein kinase PERK1-like protein [Oryza sativa]
Length = 594
Score = 172 bits (436), Expect = 3e-41
Identities = 99/260 (38%), Positives = 152/260 (58%), Gaps = 18/260 (6%)
Query: 9 SFLFMLLASKSFIAESK-------CSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKPE 61
S L ++LA+ +F A + CS C++ALAS+Y+ + N+T ++++
Sbjct: 6 SLLVLVLAAAAFAAGTVTEAAGDGCSAGCDLALASFYVTPNQNVTNMADLFGIGAANY-R 64
Query: 62 DIVSYNTDTITNKDFVQSFTRVNVPFPCDCIH------DEFLGHIFQYQVATKDTYLSVA 115
+ YN + I N DF+ RVNV F C C +L F +Q++ Y SVA
Sbjct: 65 SLAPYNPN-IPNLDFINVGGRVNVYFTCGCRSLPGSPGATYLAGAFPFQMSRGQIYTSVA 123
Query: 116 SNNYSNLTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDS 175
+N Y+NLTT+EWLQ NSYP+N+IPDT +N TVNCSCG++ +S DYGLF+TYPLR ED+
Sbjct: 124 AN-YNNLTTAEWLQATNSYPANNIPDTAVINATVNCSCGDASISPDYGLFLTYPLRAEDT 182
Query: 176 LELISNKTEIDAEL--LQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHTRSIRWSYNWNI 233
L ++ + ++L +++YNPG+ + GSG+VYIP KD N +Y+P + R + +
Sbjct: 183 LASVAATYGLSSQLDVVRRYNPGMESATGSGIVYIPVKDPNGSYLPLKSPGRRKAKQATL 242
Query: 234 CWSSSRTNSIIILHLCYVLP 253
SS + + + + V P
Sbjct: 243 LQSSEDSTQLGTISMDKVTP 262
>UniRef100_Q6UD72 LysM domain-containing receptor-like kinase 4 [Medicago truncatula]
Length = 496
Score = 119 bits (299), Expect = 2e-25
Identities = 54/96 (56%), Positives = 71/96 (73%)
Query: 128 LQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELISNKTEIDA 187
L+ FNSY N IP +NVTV CSCGNS +SKD+GLF+TYPLR +D+L I+ K ++D
Sbjct: 1 LKKFNSYDPNHIPVKAKINVTVICSCGNSQISKDFGLFVTYPLRSDDTLAKIATKADLDE 60
Query: 188 ELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHTR 223
LLQ +N NFS+GSG+V+IPG+D+N YVP +R
Sbjct: 61 GLLQNFNQDANFSKGSGIVFIPGRDENGVYVPLPSR 96
>UniRef100_Q7FS97 Hypothetical protein [Sorghum bicolor]
Length = 302
Score = 108 bits (269), Expect = 6e-22
Identities = 82/231 (35%), Positives = 123/231 (52%), Gaps = 28/231 (12%)
Query: 3 PIKFRLSFLFMLLAS---KSFIAESKCS--KTCNIALASYYLQDDTNLTYVSNIMQSNLV 57
P + L +LLA+ ++ A + CS C++AL SYY+ + NLTY+S++ +
Sbjct: 2 PQSLIICLLLLLLAAAAPQTAAAGADCSVGMGCDLALGSYYISRNQNLTYISSLFG---I 58
Query: 58 TKPEDIVSYNTDTITNKDFVQSFTRVNVPFPCDCI------HDEFLGHIFQYQVATKDTY 111
+ YNT T+ DF+Q +R+NV F C C+ +L F Y V+ +TY
Sbjct: 59 DDYHTLAPYNTG-YTSLDFIQVGSRINVYFRCGCLTLPFAPFSTYLAGSFPYVVSQGETY 117
Query: 112 LSVASNNYSNLTTSEWLQNFNSYPS--NDIPDTGT-LNVTVNCSCGNSDVS-KDYGLFIT 167
SVA+ + NLTT+ LQ S + N + D GT +NVTV CSC N+DVS DY +T
Sbjct: 118 ASVAA-EFHNLTTATLLQPAPSSDNDFNGVLDAGTVVNVTVTCSCANADVSAPDYRFLLT 176
Query: 168 YPLRPEDSLELI------SNKTEIDAELLQKYNPGVNFSQGSGLVYIPGKD 212
YPL ++ + S+ E+D L ++YNP + + +VYIP KD
Sbjct: 177 YPLGDGETPNSVAASHGLSSPAELD--LFRRYNPRADSVKEGEVVYIPLKD 225
>UniRef100_Q9CAP5 Hypothetical protein T5M16.22 [Arabidopsis thaliana]
Length = 409
Score = 58.2 bits (139), Expect = 7e-07
Identities = 54/207 (26%), Positives = 90/207 (43%), Gaps = 26/207 (12%)
Query: 11 LFMLLAS--------KSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKPED 62
LF++LAS KS I TCN +L Y L D +T V+++ Q + P
Sbjct: 10 LFLILASSLASMATAKSTIEPCSSKDTCN-SLLGYTLYTDLKVTEVASLFQVD----PVS 64
Query: 63 IVSYNTDTITNKDF----VQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNN 118
++ N+ I+ D + + + +P C C+ Y+ T DT S+A +
Sbjct: 65 MLLSNSIDISYPDVENHVLPAKLFLKIPITCSCVDGIRKSLSTHYKTRTSDTLGSIADSV 124
Query: 119 YSNLTTSEWLQNFNSYPSNDIPDTGT-LNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLE 177
Y L + E +Q NS + D GT L + + C+C N L+++Y +R D++
Sbjct: 125 YGGLVSPEQIQVANSETDLSVLDVGTKLVIPLPCACFNGTDESLPALYLSYVVRGIDTMA 184
Query: 178 LISNK--------TEIDAELLQKYNPG 196
I+ + T ++A NPG
Sbjct: 185 GIAKRFSTSVTDLTNVNAMGAPDINPG 211
>UniRef100_Q6NPN4 At1g77630 [Arabidopsis thaliana]
Length = 423
Score = 58.2 bits (139), Expect = 7e-07
Identities = 54/207 (26%), Positives = 90/207 (43%), Gaps = 26/207 (12%)
Query: 11 LFMLLAS--------KSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKPED 62
LF++LAS KS I TCN +L Y L D +T V+++ Q + P
Sbjct: 10 LFLILASSLASMATAKSTIEPCSSKDTCN-SLLGYTLYTDLKVTEVASLFQVD----PVS 64
Query: 63 IVSYNTDTITNKDF----VQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNN 118
++ N+ I+ D + + + +P C C+ Y+ T DT S+A +
Sbjct: 65 MLLSNSIDISYPDVENHVLPAKLFLKIPITCSCVDGIRKSLSTHYKTRTSDTLGSIADSV 124
Query: 119 YSNLTTSEWLQNFNSYPSNDIPDTGT-LNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLE 177
Y L + E +Q NS + D GT L + + C+C N L+++Y +R D++
Sbjct: 125 YGGLVSPEQIQVANSETDLSVLDVGTKLVIPLPCACFNGTDESLPALYLSYVVRGIDTMA 184
Query: 178 LISNK--------TEIDAELLQKYNPG 196
I+ + T ++A NPG
Sbjct: 185 GIAKRFSTSVTDLTNVNAMGAPDINPG 211
>UniRef100_Q6K3X1 Receptor protein kinase PERK1-like [Oryza sativa]
Length = 377
Score = 55.8 bits (133), Expect = 4e-06
Identities = 39/115 (33%), Positives = 61/115 (52%), Gaps = 15/115 (13%)
Query: 26 CSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKPEDIVSYNTDTITNKDFVQSFTRVNV 85
C C++A+A+YY + +NLT+++ I + ++ YN ITN D+V + RV V
Sbjct: 75 CRAGCSLAIAAYYFSEGSNLTFIATIFAIG-GGGYQALLPYN-PAITNPDYVVTGDRVFV 132
Query: 86 PFPCDCI------HDEFLGHIFQYQVATK-----DTYLSVASNNYSNLTTSEWLQ 129
FPC C+ FL Y + DTY +VA+ N++NLTT+ WL+
Sbjct: 133 -FPCSCLGLPAAPASTFLAGAIPYPLPLPRGGGGDTYDAVAA-NFANLTTAAWLE 185
>UniRef100_Q93ZH0 LysM-domain GPI-anchored protein 1 precursor [Arabidopsis thaliana]
Length = 416
Score = 54.3 bits (129), Expect = 1e-05
Identities = 48/188 (25%), Positives = 84/188 (44%), Gaps = 18/188 (9%)
Query: 8 LSFLFMLLAS--------KSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTK 59
L F+ ++LAS KS I + TCN AL Y L D ++ V+++ Q +
Sbjct: 10 LIFVSLILASSLTFTATAKSTIEPCSSNDTCN-ALLGYTLYTDLKVSEVASLFQVD---- 64
Query: 60 PEDIVSYNTDTITNKD----FVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVA 115
P I+ N I+ D + S + +P C C+ Y+ D S+A
Sbjct: 65 PISILLANAIDISYPDVENHILPSKLFLKIPITCSCVDGIRKSVSTHYKTRPSDNLGSIA 124
Query: 116 SNNYSNLTTSEWLQNFNSYPSNDIPDTGT-LNVTVNCSCGNSDVSKDYGLFITYPLRPED 174
+ Y L ++E +Q NS + D GT L + + C+C N + ++++Y ++ D
Sbjct: 125 DSVYGGLVSAEQIQEANSVNDPSLLDVGTSLVIPLPCACFNGTDNSLPAVYLSYVVKEID 184
Query: 175 SLELISNK 182
+L I+ +
Sbjct: 185 TLVGIARR 192
>UniRef100_Q67UE8 Putative Erwinia induced protein 1 [Oryza sativa]
Length = 401
Score = 49.3 bits (116), Expect = 3e-04
Identities = 50/203 (24%), Positives = 84/203 (40%), Gaps = 20/203 (9%)
Query: 8 LSFLFMLLASKSFIA--------ESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTK 59
L L +L A+ + +A ES S T AL SY L D L ++ + ++
Sbjct: 5 LLLLLLLAAAAAAVAPARSKSTLESCSSSTACPALLSYTLYADLKLAELAALFSAD---- 60
Query: 60 PEDIVSYNTDTITNKD----FVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVA 115
P I++ N+ D + + + VP PC C +Y DT SVA
Sbjct: 61 PLAILAANSIDFAVPDPADRILPAGLPLRVPVPCACSDGIRRVTTVRYVARPGDTLASVA 120
Query: 116 SNNYSNLTTSEWLQNFN----SYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLR 171
S+ Y LTT +W+ + N + P + TL V ++C+C + +++TY
Sbjct: 121 SSVYGGLTTPDWISDSNGILGAKPDAAVDAGTTLFVPLHCACFGGVDNGLPAVYLTYVAG 180
Query: 172 PEDSLELISNKTEIDAELLQKYN 194
D++ ++ + A L N
Sbjct: 181 KGDTVAAVAQRYRTTATDLMSVN 203
>UniRef100_Q70KR5 SYM10 protein [Pisum sativum]
Length = 594
Score = 47.4 bits (111), Expect = 0.001
Identities = 28/100 (28%), Positives = 49/100 (49%), Gaps = 3/100 (3%)
Query: 85 VPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTTSEWLQNFN-SYPSNDIPDTG 143
+P C C + + + F Y + D Y V++ +Y NLT ++NFN + N +P
Sbjct: 97 IPVTCGCTRNRYFAN-FTYTIKLGDNYFIVSTTSYQNLTNYVEMENFNPNLSPNLLPPEI 155
Query: 144 TLNVTVNCSC-GNSDVSKDYGLFITYPLRPEDSLELISNK 182
+ V + C C + +SK ITY + D++ +S+K
Sbjct: 156 KVVVPLFCKCPSKNQLSKGIKHLITYVWQANDNVTRVSSK 195
>UniRef100_Q70KR3 SYM10 protein [Pisum sativum]
Length = 594
Score = 47.4 bits (111), Expect = 0.001
Identities = 28/100 (28%), Positives = 49/100 (49%), Gaps = 3/100 (3%)
Query: 85 VPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTTSEWLQNFN-SYPSNDIPDTG 143
+P C C + + + F Y + D Y V++ +Y NLT ++NFN + N +P
Sbjct: 97 IPVTCGCTRNRYFAN-FTYTIKLGDNYFIVSTTSYQNLTNYVEMENFNPNLSPNLLPPEI 155
Query: 144 TLNVTVNCSC-GNSDVSKDYGLFITYPLRPEDSLELISNK 182
+ V + C C + +SK ITY + D++ +S+K
Sbjct: 156 KVVVPLFCKCPSKNQLSKGIKHLITYVWQANDNVTRVSSK 195
>UniRef100_Q70KR1 Nod-factor receptor 5 [Lotus japonicus]
Length = 595
Score = 40.4 bits (93), Expect = 0.16
Identities = 35/129 (27%), Positives = 59/129 (45%), Gaps = 5/129 (3%)
Query: 85 VPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTTSEWLQNFN-SYPSNDIPDTG 143
VP C C + + YQ+ D+Y VA+ Y NLT +Q N +P+
Sbjct: 97 VPVTCGCAGNHSSANT-SYQIQLGDSYDFVATTLYENLTNWNIVQASNPGVNPYLLPERV 155
Query: 144 TLNVTVNCSC-GNSDVSKDYGLFITYPLRPEDSLELISNKTEID-AELLQKYNPGVNFSQ 201
+ + C C + ++K ITY +P D++ L+S K A++L + G +F+
Sbjct: 156 KVVFPLFCRCPSKNQLNKGIQYLITYVWKPNDNVSLVSAKFGASPADILTENRYGQDFTA 215
Query: 202 GSGL-VYIP 209
+ L + IP
Sbjct: 216 ATNLPILIP 224
>UniRef100_Q7RLT3 Unnamed protein product [Plasmodium yoelii yoelii]
Length = 893
Score = 38.5 bits (88), Expect = 0.59
Identities = 40/159 (25%), Positives = 67/159 (41%), Gaps = 12/159 (7%)
Query: 44 NLTYVSNIMQSNLVTKPEDIVSYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQY 103
N + +N++Q KP +I SY D SF + N+ F ++ ++G+ F Y
Sbjct: 329 NSFFNNNVIQEYKCNKPLNIESYADGNTRINDHRSSFNKNNINFVNKLSYNNYIGNNFLY 388
Query: 104 QVAT------KDTYLSVASNNYSNLTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGN-S 156
Q + D Y + NY +L + + + +NS ND T +N N S N
Sbjct: 389 QDPSFLLPDKIDIYKNYYDKNYKDL--NMFKEYYNSTYINDF-YTNLINGNYNLSLRNPC 445
Query: 157 DVSKDYGLFITY--PLRPEDSLELISNKTEIDAELLQKY 193
+ D G TY P ++ +T++ A +L Y
Sbjct: 446 ALYIDVGFSHTYILPYIEYKVIDYAILRTKVSASILNTY 484
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.351 0.153 0.577
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 933,749,152
Number of Sequences: 2790947
Number of extensions: 35795161
Number of successful extensions: 137135
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 26
Number of HSP's that attempted gapping in prelim test: 137086
Number of HSP's gapped (non-prelim): 51
length of query: 617
length of database: 848,049,833
effective HSP length: 133
effective length of query: 484
effective length of database: 476,853,882
effective search space: 230797278888
effective search space used: 230797278888
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 78 (34.7 bits)
Medicago: description of AC147960.5


