

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147959.10 + phase: 1 /pseudo
(378 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
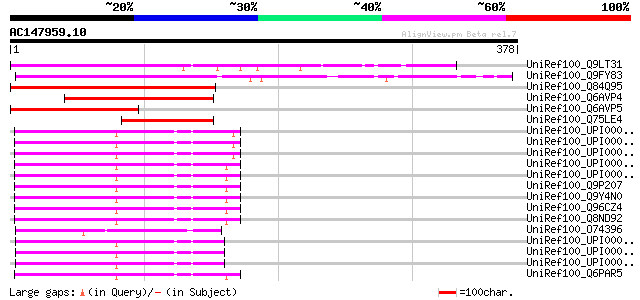
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LT31 Similarity to vacuolar protein sorting-associat... 197 4e-49
UniRef100_Q9FY83 Hypothetical protein T5E8_120 [Arabidopsis thal... 191 4e-47
UniRef100_Q84Q95 Hypothetical protein OJ1041F02.10 [Oryza sativa] 189 8e-47
UniRef100_Q6AVP4 Putative Vacuolar sorting protein [Oryza sativa] 160 5e-38
UniRef100_Q6AVP5 Putative Vacuolar sorting protein [Oryza sativa] 137 5e-31
UniRef100_Q75LE4 Putative vacuolar sorting protein , 5'-partial ... 105 3e-21
UniRef100_UPI00003AB258 UPI00003AB258 UniRef100 entry 95 3e-18
UniRef100_UPI00003AB257 UPI00003AB257 UniRef100 entry 95 3e-18
UniRef100_UPI00003AB256 UPI00003AB256 UniRef100 entry 95 3e-18
UniRef100_UPI00001CF0D0 UPI00001CF0D0 UniRef100 entry 95 4e-18
UniRef100_UPI0000368FB4 UPI0000368FB4 UniRef100 entry 95 4e-18
UniRef100_Q9P207 Hypothetical protein KIAA1521 [Homo sapiens] 95 4e-18
UniRef100_Q9Y4N0 Hypothetical protein DKFZp434C212 [Homo sapiens] 95 4e-18
UniRef100_Q96CZ4 Hypothetical protein [Homo sapiens] 95 4e-18
UniRef100_Q8ND92 Hypothetical protein DKFZp434J231 [Homo sapiens] 95 4e-18
UniRef100_O74396 Vps901 protein [Schizosaccharomyces pombe] 94 8e-18
UniRef100_UPI00004390C0 UPI00004390C0 UniRef100 entry 93 1e-17
UniRef100_UPI00004390BF UPI00004390BF UniRef100 entry 93 1e-17
UniRef100_UPI00004390BE UPI00004390BE UniRef100 entry 93 1e-17
UniRef100_Q6PAR5 RIKEN cDNA 4432404J10 [Mus musculus] 93 1e-17
>UniRef100_Q9LT31 Similarity to vacuolar protein sorting-associated protein VPS9
[Arabidopsis thaliana]
Length = 520
Score = 197 bits (501), Expect = 4e-49
Identities = 141/353 (39%), Positives = 192/353 (53%), Gaps = 28/353 (7%)
Query: 1 DGRRVQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPEDAKID 60
D VQ+FF ME A R HPLW+ +EE++D A +GLEKY+MTKLF+R FA++ E+ D
Sbjct: 46 DCAMVQEFFSKMEAAFRAHPLWSGCSEEELDSAGDGLEKYVMTKLFTRVFASNTEEVIAD 105
Query: 61 HEISEKISLLQTFLKPEHLDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRV 120
++ +K+SL+Q F+ PE+LDI P NE+SWLLA+KELQKIN +KAP++KL I+NCC+V
Sbjct: 106 EKLFQKMSLVQQFISPENLDIQPTFQNESSWLLAQKELQKINMYKAPRDKLVCILNCCKV 165
Query: 121 INNLLLNA--AMSEYVPAGADDFIPVLIYVTIKAR------LALLVQTSNSLSCIEGK-- 170
INNLLLNA A +E P GAD+F+PVLIYVTIKA L +Q S + G+
Sbjct: 166 INNLLLNASIASNENAP-GADEFLPVLIYVTIKANPPQLHSNLLYIQRYRRESKLVGEAA 224
Query: 171 ---QNSSQKLSIISQI---LCQLRHL*LN*ILNPSQLMKSNLRSACKQPNW----PRKSQ 220
N S IS I L + ++ S L S Q P + +
Sbjct: 225 YFFTNILSAESFISNIDAKSISLDEAEFEKNMESARARISGLDSQTYQTGHGSAPPPRDE 284
Query: 221 VNYTLHVKLNKK*RTRAMFQTKCITNLILETK*TILGSLTKLSLKK*KIFEL*FQPLSQS 280
LN K R +FQ+K +L + + S T + K I +L + +
Sbjct: 285 STLQKTQSLNPK-RENTLFQSKSSDSLSGTNELLNINSETPMK-KAESISDL--ENKGAT 340
Query: 281 HLNYHAEFHVLQHGTNYPYMEAESKDLAMEDVDILLNHYKDLVAKYTIICKAI 333
L V Q YPY+ A + DL + DV+ LLN YK LV KY + K +
Sbjct: 341 LLKDTEPSKVFQ---EYPYIFASAGDLRIGDVEGLLNSYKQLVFKYVCLTKGL 390
>UniRef100_Q9FY83 Hypothetical protein T5E8_120 [Arabidopsis thaliana]
Length = 712
Score = 191 bits (484), Expect = 4e-47
Identities = 140/386 (36%), Positives = 206/386 (53%), Gaps = 37/386 (9%)
Query: 5 VQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPEDAKIDHEIS 64
VQDFF ME A R HPLW+ +++++D A +GLEKY+MTKLF R FA++ ED D ++
Sbjct: 47 VQDFFYKMESAFRAHPLWSGCSDDELDNAGDGLEKYVMTKLFPRVFASNTEDVISDEKLF 106
Query: 65 EKISLLQTFLKPEHLDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRVINNL 124
+KISL+Q F+ PE+LDI P N+ SWLLA+KELQKIN + AP++KL I+ CC+VINNL
Sbjct: 107 QKISLVQQFISPENLDIQPTFQNQTSWLLAQKELQKINMYNAPRDKLMCILRCCKVINNL 166
Query: 125 LLNAAM-SEYVPAGADDFIPVLIYVTIKARLALLVQTSNSLSCIEGKQNSSQKLS----I 179
LLNA++ S GAD F+PVLIYVTIKA Q ++L I+ + S+ + +
Sbjct: 167 LLNASIASNQNEPGADQFLPVLIYVTIKANPP---QFHSNLLYIQRYRRQSKLVGEAGYL 223
Query: 180 ISQILCQ---LRHL*LN*ILNPSQLMKSNLRSACKQPNWPRKSQVNYTLHVKLNKK*RTR 236
+ IL + ++ + ++ ++SA + + P L T+
Sbjct: 224 FTNILSAESFISNIDAKSLSMDEADFETKMKSAHARLSGPGSQSYQTDHGAALPTAHNTK 283
Query: 237 AMFQTKCITNLILETK*TILGSLTKLSLKK*KIFEL*FQPLSQ-------SHLNYHAEFH 289
N++L TK T S T +L + I + P++ + LN +E
Sbjct: 284 R-------ENMLLHTKSTDSFSGTNETLSETPIKKA--DPITDLENKGAATLLNDRSE-- 332
Query: 290 VLQHGTNYPYMEAESKDLAMEDVDILLNHYKDLVAKYTIICKAINYLSMSEKEPLLHQLE 349
+ YPYM A DL + DV+ LLN+YK LV KY + K + + P + L
Sbjct: 333 ATKIFQEYPYMFASVGDLKIGDVEDLLNNYKQLVFKYVCLSKGLG--DATSLTPCISPL- 389
Query: 350 MQGEGSMLSECHGINTNTNDRTTSHE 375
+ S +SE H T ++D T E
Sbjct: 390 ---QASKVSENH--TTLSSDFQTKSE 410
>UniRef100_Q84Q95 Hypothetical protein OJ1041F02.10 [Oryza sativa]
Length = 559
Score = 189 bits (481), Expect = 8e-47
Identities = 93/154 (60%), Positives = 121/154 (78%), Gaps = 1/154 (0%)
Query: 1 DGRRVQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPEDAKID 60
D VQ+F +ME A R H WA ++EE+++ A EGLEKY+MTKLF+R FA+ PED K D
Sbjct: 138 DSAAVQEFLENMEGAFRAHTPWAGSSEEELESAGEGLEKYVMTKLFNRVFASVPEDVKSD 197
Query: 61 HEISEKISLLQTFLKPEHLDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRV 120
E+ EK+SLLQ F++PE+LDI P +E SWLLA+KELQKIN +KAP++KL+ I+NCC+V
Sbjct: 198 EELFEKMSLLQQFIRPENLDIKPEYQSETSWLLAQKELQKINMYKAPRDKLACILNCCKV 257
Query: 121 INNLLLNAA-MSEYVPAGADDFIPVLIYVTIKAR 153
INNLLLNA+ +S P GAD+F+PVLIYVTIK +
Sbjct: 258 INNLLLNASIVSNENPPGADEFLPVLIYVTIKKK 291
Score = 38.1 bits (87), Expect = 0.41
Identities = 17/37 (45%), Positives = 24/37 (63%)
Query: 297 YPYMEAESKDLAMEDVDILLNHYKDLVAKYTIICKAI 333
YP++ A S DL + DV+ LLN YK LV KY + + +
Sbjct: 441 YPFLFARSGDLTVADVENLLNSYKQLVLKYVALSQGM 477
>UniRef100_Q6AVP4 Putative Vacuolar sorting protein [Oryza sativa]
Length = 218
Score = 160 bits (405), Expect = 5e-38
Identities = 79/112 (70%), Positives = 95/112 (84%), Gaps = 1/112 (0%)
Query: 42 MTKLFSRTFAASPEDAKIDHEISEKISLLQTFLKPEHLDIPPVLHNEASWLLAEKELQKI 101
MTKLF R FA+S ED K D EISEKI LLQ F++P HLDIP +LHNEA+WLLA KELQKI
Sbjct: 1 MTKLFDRAFASSAEDVKSDMEISEKIGLLQHFVRPHHLDIPKLLHNEAAWLLAVKELQKI 60
Query: 102 NAFKAPQEKLSTIMNCCRVINNLLLNAAMS-EYVPAGADDFIPVLIYVTIKA 152
N+FK+P+EKLS IM+CC+VINNLLLN +MS + +GADDF+P+LIY+TIKA
Sbjct: 61 NSFKSPREKLSCIMSCCQVINNLLLNVSMSNDRTLSGADDFLPILIYITIKA 112
>UniRef100_Q6AVP5 Putative Vacuolar sorting protein [Oryza sativa]
Length = 145
Score = 137 bits (345), Expect = 5e-31
Identities = 65/96 (67%), Positives = 77/96 (79%)
Query: 1 DGRRVQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPEDAKID 60
DG RVQ FF ME AIR+HPLWA AT ++ID A+EGLEKYIMTKLF R FA+S ED K D
Sbjct: 50 DGGRVQTFFAEMETAIRDHPLWANATNQEIDNALEGLEKYIMTKLFDRAFASSAEDVKSD 109
Query: 61 HEISEKISLLQTFLKPEHLDIPPVLHNEASWLLAEK 96
EISEKI LLQ F++P HLDIP +LHNEA+WL+ ++
Sbjct: 110 MEISEKIGLLQHFVRPHHLDIPKLLHNEAAWLVRQQ 145
>UniRef100_Q75LE4 Putative vacuolar sorting protein , 5'-partial [Oryza sativa]
Length = 177
Score = 105 bits (261), Expect = 3e-21
Identities = 50/70 (71%), Positives = 63/70 (89%), Gaps = 1/70 (1%)
Query: 84 VLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRVINNLLLNAAMS-EYVPAGADDFI 142
+LHNEA+WLLA KELQKIN+FK+P+EKLS IM+CC+VINNLLLN +MS + +GADDF+
Sbjct: 2 LLHNEAAWLLAVKELQKINSFKSPREKLSCIMSCCQVINNLLLNVSMSNDRTLSGADDFL 61
Query: 143 PVLIYVTIKA 152
P+LIY+TIKA
Sbjct: 62 PILIYITIKA 71
>UniRef100_UPI00003AB258 UPI00003AB258 UniRef100 entry
Length = 804
Score = 95.1 bits (235), Expect = 3e-18
Identities = 59/177 (33%), Positives = 95/177 (53%), Gaps = 10/177 (5%)
Query: 4 RVQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPE-DAKIDHE 62
+V+DF + A+ +W A+EE + A +E+ +M ++F F + + D D
Sbjct: 610 QVEDFLQFLYGAMAQDAIWQNASEEQLQDAQLAIERSVMNRIFKLAFYPNQDGDILRDQV 669
Query: 63 ISEKISLLQTFLKPEH--LDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRV 120
+ E I L + H L IP V EA W A+ E++ I+A+K P++K+ I+ C
Sbjct: 670 LHEHIQRLSKVVTANHKALQIPEVYLREAPWPSAQSEIRTISAYKTPRDKVQCILRMCST 729
Query: 121 INNLLLNAAMSEYVPAGADDFIPVLIYVTIKARLALLVQTSNSLS-----CIEGKQN 172
I N LL+ A + VP GADDF+PVL++V IKA L+ T +S C+ G+++
Sbjct: 730 IMN-LLSLANEDSVP-GADDFVPVLVFVLIKANPPCLLSTVQYISSFYANCLSGEES 784
>UniRef100_UPI00003AB257 UPI00003AB257 UniRef100 entry
Length = 1281
Score = 95.1 bits (235), Expect = 3e-18
Identities = 59/177 (33%), Positives = 95/177 (53%), Gaps = 10/177 (5%)
Query: 4 RVQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPE-DAKIDHE 62
+V+DF + A+ +W A+EE + A +E+ +M ++F F + + D D
Sbjct: 1087 QVEDFLQFLYGAMAQDAIWQNASEEQLQDAQLAIERSVMNRIFKLAFYPNQDGDILRDQV 1146
Query: 63 ISEKISLLQTFLKPEH--LDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRV 120
+ E I L + H L IP V EA W A+ E++ I+A+K P++K+ I+ C
Sbjct: 1147 LHEHIQRLSKVVTANHKALQIPEVYLREAPWPSAQSEIRTISAYKTPRDKVQCILRMCST 1206
Query: 121 INNLLLNAAMSEYVPAGADDFIPVLIYVTIKARLALLVQTSNSLS-----CIEGKQN 172
I N LL+ A + VP GADDF+PVL++V IKA L+ T +S C+ G+++
Sbjct: 1207 IMN-LLSLANEDSVP-GADDFVPVLVFVLIKANPPCLLSTVQYISSFYANCLSGEES 1261
>UniRef100_UPI00003AB256 UPI00003AB256 UniRef100 entry
Length = 1448
Score = 95.1 bits (235), Expect = 3e-18
Identities = 59/177 (33%), Positives = 95/177 (53%), Gaps = 10/177 (5%)
Query: 4 RVQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPE-DAKIDHE 62
+V+DF + A+ +W A+EE + A +E+ +M ++F F + + D D
Sbjct: 1254 QVEDFLQFLYGAMAQDAIWQNASEEQLQDAQLAIERSVMNRIFKLAFYPNQDGDILRDQV 1313
Query: 63 ISEKISLLQTFLKPEH--LDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRV 120
+ E I L + H L IP V EA W A+ E++ I+A+K P++K+ I+ C
Sbjct: 1314 LHEHIQRLSKVVTANHKALQIPEVYLREAPWPSAQSEIRTISAYKTPRDKVQCILRMCST 1373
Query: 121 INNLLLNAAMSEYVPAGADDFIPVLIYVTIKARLALLVQTSNSLS-----CIEGKQN 172
I N LL+ A + VP GADDF+PVL++V IKA L+ T +S C+ G+++
Sbjct: 1374 IMN-LLSLANEDSVP-GADDFVPVLVFVLIKANPPCLLSTVQYISSFYANCLSGEES 1428
>UniRef100_UPI00001CF0D0 UPI00001CF0D0 UniRef100 entry
Length = 1472
Score = 94.7 bits (234), Expect = 4e-18
Identities = 60/177 (33%), Positives = 95/177 (52%), Gaps = 10/177 (5%)
Query: 4 RVQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPE-DAKIDHE 62
+V+DF + A+ +W A+EE + A +E+ +M ++F F + + D D
Sbjct: 1278 QVEDFLQFLYGAMAQDVIWQNASEEQLQDAQLAIERSVMNRIFKLAFYPNQDGDILRDQV 1337
Query: 63 ISEKISLLQTFLKPEH--LDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRV 120
+ E I L + H L IP V EA W A+ E++ I+A+K P++K+ I+ C
Sbjct: 1338 LHEHIQRLSKVVTANHRALQIPEVYLREAPWPSAQSEIRTISAYKTPRDKVQCILRMCST 1397
Query: 121 INNLLLNAAMSEYVPAGADDFIPVLIYVTIKARLALLVQT-----SNSLSCIEGKQN 172
I N LL+ A + VP GADDF+PVL++V IKA L+ T S SC+ G+++
Sbjct: 1398 IMN-LLSLANEDSVP-GADDFVPVLVFVLIKANPPCLLSTVQYISSFYASCLSGEES 1452
>UniRef100_UPI0000368FB4 UPI0000368FB4 UniRef100 entry
Length = 814
Score = 94.7 bits (234), Expect = 4e-18
Identities = 60/177 (33%), Positives = 95/177 (52%), Gaps = 10/177 (5%)
Query: 4 RVQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPE-DAKIDHE 62
+V+DF + A+ +W A+EE + A +E+ +M ++F F + + D D
Sbjct: 620 QVEDFLQFLYGAMAQDVIWQNASEEQLQDAQLAIERSVMNRIFKLAFYPNQDGDILRDQV 679
Query: 63 ISEKISLLQTFLKPEH--LDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRV 120
+ E I L + H L IP V EA W A+ E++ I+A+K P++K+ I+ C
Sbjct: 680 LHEHIQRLSKVVTANHRALQIPEVYLREAPWPSAQSEIRTISAYKTPRDKVQCILRMCST 739
Query: 121 INNLLLNAAMSEYVPAGADDFIPVLIYVTIKARLALLVQT-----SNSLSCIEGKQN 172
I N LL+ A + VP GADDF+PVL++V IKA L+ T S SC+ G+++
Sbjct: 740 IMN-LLSLANEDSVP-GADDFVPVLVFVLIKANPPCLLSTVQYISSFYASCLSGEES 794
>UniRef100_Q9P207 Hypothetical protein KIAA1521 [Homo sapiens]
Length = 1272
Score = 94.7 bits (234), Expect = 4e-18
Identities = 60/177 (33%), Positives = 95/177 (52%), Gaps = 10/177 (5%)
Query: 4 RVQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPE-DAKIDHE 62
+V+DF + A+ +W A+EE + A +E+ +M ++F F + + D D
Sbjct: 1078 QVEDFLQFLYGAMAQDVIWQNASEEQLQDAQLAIERSVMNRIFKLAFYPNQDGDILRDQV 1137
Query: 63 ISEKISLLQTFLKPEH--LDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRV 120
+ E I L + H L IP V EA W A+ E++ I+A+K P++K+ I+ C
Sbjct: 1138 LHEHIQRLSKVVTANHRALQIPEVYLREAPWPSAQSEIRTISAYKTPRDKVQCILRMCST 1197
Query: 121 INNLLLNAAMSEYVPAGADDFIPVLIYVTIKARLALLVQT-----SNSLSCIEGKQN 172
I N LL+ A + VP GADDF+PVL++V IKA L+ T S SC+ G+++
Sbjct: 1198 IMN-LLSLANEDSVP-GADDFVPVLVFVLIKANPPCLLSTVQYISSFYASCLSGEES 1252
>UniRef100_Q9Y4N0 Hypothetical protein DKFZp434C212 [Homo sapiens]
Length = 813
Score = 94.7 bits (234), Expect = 4e-18
Identities = 60/177 (33%), Positives = 95/177 (52%), Gaps = 10/177 (5%)
Query: 4 RVQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPE-DAKIDHE 62
+V+DF + A+ +W A+EE + A +E+ +M ++F F + + D D
Sbjct: 619 QVEDFLQFLYGAMAQDVIWQNASEEQLQDAQLAIERSVMNRIFKLAFYPNQDGDILRDQV 678
Query: 63 ISEKISLLQTFLKPEH--LDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRV 120
+ E I L + H L IP V EA W A+ E++ I+A+K P++K+ I+ C
Sbjct: 679 LHEHIQRLSKVVTANHRALQIPEVYLREAPWPSAQSEIRTISAYKTPRDKVQCILRMCST 738
Query: 121 INNLLLNAAMSEYVPAGADDFIPVLIYVTIKARLALLVQT-----SNSLSCIEGKQN 172
I N LL+ A + VP GADDF+PVL++V IKA L+ T S SC+ G+++
Sbjct: 739 IMN-LLSLANEDSVP-GADDFVPVLVFVLIKANPPCLLSTVQYISSFYASCLSGEES 793
>UniRef100_Q96CZ4 Hypothetical protein [Homo sapiens]
Length = 549
Score = 94.7 bits (234), Expect = 4e-18
Identities = 60/177 (33%), Positives = 95/177 (52%), Gaps = 10/177 (5%)
Query: 4 RVQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPE-DAKIDHE 62
+V+DF + A+ +W A+EE + A +E+ +M ++F F + + D D
Sbjct: 355 QVEDFLQFLYGAMAQDVIWQNASEEQLQDAQLAIERSVMNRIFKLAFYPNQDGDILRDQV 414
Query: 63 ISEKISLLQTFLKPEH--LDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRV 120
+ E I L + H L IP V EA W A+ E++ I+A+K P++K+ I+ C
Sbjct: 415 LHEHIQRLSKVVTANHRALQIPEVYLREAPWPSAQSEIRTISAYKTPRDKVQCILRMCST 474
Query: 121 INNLLLNAAMSEYVPAGADDFIPVLIYVTIKARLALLVQT-----SNSLSCIEGKQN 172
I N LL+ A + VP GADDF+PVL++V IKA L+ T S SC+ G+++
Sbjct: 475 IMN-LLSLANEDSVP-GADDFVPVLVFVLIKANPPCLLSTVQYISSFYASCLSGEES 529
>UniRef100_Q8ND92 Hypothetical protein DKFZp434J231 [Homo sapiens]
Length = 329
Score = 94.7 bits (234), Expect = 4e-18
Identities = 60/177 (33%), Positives = 95/177 (52%), Gaps = 10/177 (5%)
Query: 4 RVQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPE-DAKIDHE 62
+V+DF + A+ +W A+EE + A +E+ +M ++F F + + D D
Sbjct: 135 QVEDFLQFLYGAMAQDVIWQNASEEQLQDAQLAIERSVMNRIFKLAFYPNQDGDILRDQV 194
Query: 63 ISEKISLLQTFLKPEH--LDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRV 120
+ E I L + H L IP V EA W A+ E++ I+A+K P++K+ I+ C
Sbjct: 195 LHEHIQRLSKVVTANHRALQIPEVYLREAPWPSAQSEIRTISAYKTPRDKVQCILRMCST 254
Query: 121 INNLLLNAAMSEYVPAGADDFIPVLIYVTIKARLALLVQT-----SNSLSCIEGKQN 172
I N LL+ A + VP GADDF+PVL++V IKA L+ T S SC+ G+++
Sbjct: 255 IMN-LLSLANEDSVP-GADDFVPVLVFVLIKANPPCLLSTVQYISSFYASCLSGEES 309
>UniRef100_O74396 Vps901 protein [Schizosaccharomyces pombe]
Length = 537
Score = 93.6 bits (231), Expect = 8e-18
Identities = 54/166 (32%), Positives = 93/166 (55%), Gaps = 17/166 (10%)
Query: 5 VQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAAS----------- 53
++DF + I + WA+ ++ +ID A EG+EK ++ +L++ F+
Sbjct: 155 IRDFLKFINEKIEQYEPWASGSQAEIDNAKEGMEKLVLNRLYTSLFSPEIAKSGIPLSSE 214
Query: 54 -PEDAKIDHEISEKISLLQTFLKPEHLDIPPVLHNEASWLLAEKELQKINAFKAPQEKLS 112
+D + D +SEK+ L Q ++ E+LDI + + LA EL++IN + AP++K+
Sbjct: 215 HSDDVEEDRVLSEKMELFQ-WITEENLDIKKQKSSSKFFKLAADELRRINDYHAPRDKII 273
Query: 113 TIMNCCRVINNLLLNAAMSEYVPAGADDFIPVLIYVTIKARLALLV 158
++NCC+VI + L N E AD F+P+LI+V ++AR A LV
Sbjct: 274 CLLNCCKVIFSYLRNVVKEE----SADMFVPILIFVVLQARPAHLV 315
>UniRef100_UPI00004390C0 UPI00004390C0 UniRef100 entry
Length = 1043
Score = 93.2 bits (230), Expect = 1e-17
Identities = 57/159 (35%), Positives = 86/159 (53%), Gaps = 5/159 (3%)
Query: 5 VQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPE-DAKIDHEI 63
V+DF + A+ + +W A+EE + A +E+ +M ++F F + + D D
Sbjct: 850 VEDFLRYLYGAMAHDAIWQYASEEQLQDAQMAIERSVMNRIFKLAFYPNQDGDIHRDELF 909
Query: 64 SEKISLLQTFLKPEH--LDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRVI 121
E I L + H L IP V EA W A+ E++ INA+K P++K+ I+ C I
Sbjct: 910 HEHIQRLSRVVSANHKALQIPEVYLKEAPWPSAQAEIKTINAYKTPRDKVQCILRMCSTI 969
Query: 122 NNLLLNAAMSEYVPAGADDFIPVLIYVTIKARLALLVQT 160
N LL+ A + VP GADDF+PVL++V IKA L+ T
Sbjct: 970 MN-LLSLANEDAVP-GADDFVPVLVFVLIKANPPCLLST 1006
>UniRef100_UPI00004390BF UPI00004390BF UniRef100 entry
Length = 1033
Score = 93.2 bits (230), Expect = 1e-17
Identities = 57/159 (35%), Positives = 86/159 (53%), Gaps = 5/159 (3%)
Query: 5 VQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPE-DAKIDHEI 63
V+DF + A+ + +W A+EE + A +E+ +M ++F F + + D D
Sbjct: 840 VEDFLRYLYGAMAHDAIWQYASEEQLQDAQMAIERSVMNRIFKLAFYPNQDGDIHRDELF 899
Query: 64 SEKISLLQTFLKPEH--LDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRVI 121
E I L + H L IP V EA W A+ E++ INA+K P++K+ I+ C I
Sbjct: 900 HEHIQRLSRVVSANHKALQIPEVYLKEAPWPSAQAEIKTINAYKTPRDKVQCILRMCSTI 959
Query: 122 NNLLLNAAMSEYVPAGADDFIPVLIYVTIKARLALLVQT 160
N LL+ A + VP GADDF+PVL++V IKA L+ T
Sbjct: 960 MN-LLSLANEDAVP-GADDFVPVLVFVLIKANPPCLLST 996
>UniRef100_UPI00004390BE UPI00004390BE UniRef100 entry
Length = 784
Score = 93.2 bits (230), Expect = 1e-17
Identities = 57/159 (35%), Positives = 86/159 (53%), Gaps = 5/159 (3%)
Query: 5 VQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPE-DAKIDHEI 63
V+DF + A+ + +W A+EE + A +E+ +M ++F F + + D D
Sbjct: 591 VEDFLRYLYGAMAHDAIWQYASEEQLQDAQMAIERSVMNRIFKLAFYPNQDGDIHRDELF 650
Query: 64 SEKISLLQTFLKPEH--LDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRVI 121
E I L + H L IP V EA W A+ E++ INA+K P++K+ I+ C I
Sbjct: 651 HEHIQRLSRVVSANHKALQIPEVYLKEAPWPSAQAEIKTINAYKTPRDKVQCILRMCSTI 710
Query: 122 NNLLLNAAMSEYVPAGADDFIPVLIYVTIKARLALLVQT 160
N LL+ A + VP GADDF+PVL++V IKA L+ T
Sbjct: 711 MN-LLSLANEDAVP-GADDFVPVLVFVLIKANPPCLLST 747
>UniRef100_Q6PAR5 RIKEN cDNA 4432404J10 [Mus musculus]
Length = 1437
Score = 93.2 bits (230), Expect = 1e-17
Identities = 59/177 (33%), Positives = 94/177 (52%), Gaps = 10/177 (5%)
Query: 4 RVQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPE-DAKIDHE 62
+V+DF + + +W A+EE + A +E+ +M ++F F + + D D
Sbjct: 1243 QVEDFLQFLYGVMAQDVIWQNASEEQLQDAQLAIERSVMNRIFKLAFYPNQDGDILRDQV 1302
Query: 63 ISEKISLLQTFLKPEH--LDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRV 120
+ E I L + H L IP V EA W A+ E++ I+A+K P++K+ I+ C
Sbjct: 1303 LHEHIQRLSKVVTANHRALQIPEVYLREAPWPSAQSEIRTISAYKTPRDKVQCILRMCST 1362
Query: 121 INNLLLNAAMSEYVPAGADDFIPVLIYVTIKARLALLVQT-----SNSLSCIEGKQN 172
I N LL+ A + VP GADDF+PVL++V IKA L+ T S SC+ G+++
Sbjct: 1363 IMN-LLSLANEDSVP-GADDFVPVLVFVLIKANPPCLLSTVQYISSFYASCLSGEES 1417
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.328 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 560,629,262
Number of Sequences: 2790947
Number of extensions: 20859304
Number of successful extensions: 60772
Number of sequences better than 10.0: 133
Number of HSP's better than 10.0 without gapping: 54
Number of HSP's successfully gapped in prelim test: 79
Number of HSP's that attempted gapping in prelim test: 60566
Number of HSP's gapped (non-prelim): 150
length of query: 378
length of database: 848,049,833
effective HSP length: 129
effective length of query: 249
effective length of database: 488,017,670
effective search space: 121516399830
effective search space used: 121516399830
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 76 (33.9 bits)
Medicago: description of AC147959.10


