

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
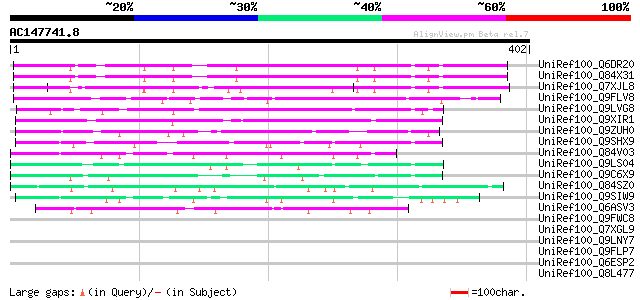
Query= AC147741.8 - phase: 0
(402 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6DR20 Hypothetical protein [Arabidopsis thaliana] 180 6e-44
UniRef100_Q84X31 Hypothetical protein At2g17036/F6P23.4 [Arabido... 178 2e-43
UniRef100_Q7XJL8 At2g17030 protein [Arabidopsis thaliana] 159 1e-37
UniRef100_Q9FLV8 Gb|AAD22308.1 [Arabidopsis thaliana] 98 4e-19
UniRef100_Q9LVG8 Gb|AAF18598.1 [Arabidopsis thaliana] 97 8e-19
UniRef100_Q9XIR1 F13O11.14 protein [Arabidopsis thaliana] 87 1e-15
UniRef100_Q9ZUH0 Expressed protein [Arabidopsis thaliana] 77 7e-13
UniRef100_Q9SHX9 F1E22.13 [Arabidopsis thaliana] 68 4e-10
UniRef100_Q84V03 Hypothetical protein At2g16365/F16F14.28 [Arabi... 65 3e-09
UniRef100_Q9LS04 Gb|AAC14516.2 [Arabidopsis thaliana] 65 4e-09
UniRef100_Q9C6X9 Hypothetical protein T7O23.22 [Arabidopsis thal... 52 2e-05
UniRef100_Q84SZ0 Hypothetical protein OSJNBa0087C10.17 [Oryza sa... 51 7e-05
UniRef100_Q9SIW9 Hypothetical protein At2g16300 [Arabidopsis tha... 49 3e-04
UniRef100_Q6ASV3 Expressed protein [Oryza sativa] 49 3e-04
UniRef100_Q9FWC8 Hypothetical protein OSJNBb0018B10.12 [Oryza sa... 47 0.001
UniRef100_Q7XGL9 Hypothetical protein [Oryza sativa] 46 0.002
UniRef100_Q9LNY7 F9C16.31 [Arabidopsis thaliana] 45 0.004
UniRef100_Q9FLP7 Dbj|BAA95762.1 [Arabidopsis thaliana] 45 0.004
UniRef100_Q6ESP2 Hypothetical protein P0684A08.40 [Oryza sativa] 45 0.004
UniRef100_Q8L477 Hypothetical protein B1147B04.22 [Oryza sativa] 45 0.005
>UniRef100_Q6DR20 Hypothetical protein [Arabidopsis thaliana]
Length = 404
Score = 180 bits (457), Expect = 6e-44
Identities = 135/410 (32%), Positives = 203/410 (48%), Gaps = 50/410 (12%)
Query: 4 VDWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSVPKH---HHKFRFQFPLLKFPF 60
+DW+ LP +LL+LIS+ + DLI FRSVCS+WRS++ PK H F FP PF
Sbjct: 2 MDWATLPKDLLDLISKCLESSFDLIQFRSVCSSWRSAAGPKRLLWAHNLPF-FPSDDKPF 60
Query: 61 NADFINKNRSFCYITKQNIFLIKPPQQEQTLTLRPLLIRICQN--ARGKSKLYHPLLQNG 118
++ I + + Q+I LIKP + + L ++++ N K L PL +
Sbjct: 61 LSNVILR------VAHQSILLIKPNEPQCEADLFGWIVKVWDNIYVSRKMTLLKPLSSSR 114
Query: 119 YKFPHTF--MLDFNKLSILHLGSNFFMIDFDFTFNDKLYNPDDYMYPKKVVVVTCHGK-- 174
FP + D +K ++ L KLY+PD Y P + GK
Sbjct: 115 NYFPQHLPRIFDMSKFTVRELCREV-----------KLYHPDYYCVPGHTALELELGKTV 163
Query: 175 ------KPLIVGTLTSPPQPLLMKCGDEKWNVIPD-MSTKFGDICLFKGQSYAVDKIGKT 227
++ T+ + + + D +W VI D + ++ D+ +F G+ +A+D G+T
Sbjct: 164 VKYLNDDKFVLLTILEYGKLAVFRSWDREWTVINDYIPSRCQDLIMFDGRFFAIDYNGRT 223
Query: 228 VVVGPDS-SVQLVAEPLIGGGDMKFLVESEGDLLLADVYNCL-----FTDLCNPDHNDCV 281
VVV S + L A PLIGGGD KFL+ES G++ L D+ CL FT N+
Sbjct: 224 VVVDYSSFKLTLAANPLIGGGDKKFLIESCGEMFLVDIEFCLNEKPEFTGGFYSYFNETT 283
Query: 282 ---RIDLFKLNEKEKKWVKLTSLGDRVLFLGLGLVCSFSACASDL---CVAKGNSVIFNN 335
+ FKL E+EK+WV++ LGD++ FLG +FSA +D+ CV G+ V F
Sbjct: 284 VSYKFKFFKLVEREKRWVEVEDLGDKMFFLGDD--STFSASTADIIPRCVGTGSFVFF-- 339
Query: 336 YIFESHRPLESECKEYVLDLDLGRHSLLSDYPEYSNLFWPPPEWIRPRYD 385
Y E + + V D G+ L++ PEY+ LFWPPP WI ++
Sbjct: 340 YTHEESLVVMDDRNLGVFDFRSGKTELVNKLPEYAKLFWPPPPWITTSHE 389
>UniRef100_Q84X31 Hypothetical protein At2g17036/F6P23.4 [Arabidopsis thaliana]
Length = 404
Score = 178 bits (452), Expect = 2e-43
Identities = 134/410 (32%), Positives = 202/410 (48%), Gaps = 50/410 (12%)
Query: 4 VDWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSVPKH---HHKFRFQFPLLKFPF 60
+DW+ LP +LL+ IS+ + DLI FRSVCS+WRS++ PK H F FP PF
Sbjct: 2 MDWATLPKDLLDXISKCLESSFDLIQFRSVCSSWRSAAGPKRLLWAHNLPF-FPSDDKPF 60
Query: 61 NADFINKNRSFCYITKQNIFLIKPPQQEQTLTLRPLLIRICQN--ARGKSKLYHPLLQNG 118
++ I + + Q+I LIKP + + L ++++ N K L PL +
Sbjct: 61 LSNVILR------VAHQSILLIKPNEPQCEADLFGWIVKVWDNIYVSRKMTLLKPLSSSR 114
Query: 119 YKFPHTF--MLDFNKLSILHLGSNFFMIDFDFTFNDKLYNPDDYMYPKKVVVVTCHGK-- 174
FP + D +K ++ L KLY+PD Y P + GK
Sbjct: 115 NYFPQHLPRIFDMSKFTVRELCREV-----------KLYHPDYYCVPGHTALELELGKTV 163
Query: 175 ------KPLIVGTLTSPPQPLLMKCGDEKWNVIPD-MSTKFGDICLFKGQSYAVDKIGKT 227
++ T+ + + + D +W VI D + ++ D+ +F G+ +A+D G+T
Sbjct: 164 VKYLNDDKFVLLTILEYGKLAVFRSWDREWTVINDYIPSRCQDLIMFDGRFFAIDYNGRT 223
Query: 228 VVVGPDS-SVQLVAEPLIGGGDMKFLVESEGDLLLADVYNCL-----FTDLCNPDHNDCV 281
VVV S + L A PLIGGGD KFL+ES G++ L D+ CL FT N+
Sbjct: 224 VVVDYSSFKLTLAANPLIGGGDKKFLIESCGEMFLVDIEFCLNEKPEFTGGFYSYFNETT 283
Query: 282 ---RIDLFKLNEKEKKWVKLTSLGDRVLFLGLGLVCSFSACASDL---CVAKGNSVIFNN 335
+ FKL E+EK+WV++ LGD++ FLG +FSA +D+ CV G+ V F
Sbjct: 284 VSYKFKFFKLVEREKRWVEVEDLGDKMFFLGDD--STFSASTADIIPRCVGTGSFVFF-- 339
Query: 336 YIFESHRPLESECKEYVLDLDLGRHSLLSDYPEYSNLFWPPPEWIRPRYD 385
Y E + + V D G+ L++ PEY+ LFWPPP WI ++
Sbjct: 340 YTHEESLVVMDDRNLGVFDFRSGKTELVNKLPEYAKLFWPPPPWITTSHE 389
>UniRef100_Q7XJL8 At2g17030 protein [Arabidopsis thaliana]
Length = 1169
Score = 159 bits (403), Expect = 1e-37
Identities = 125/386 (32%), Positives = 188/386 (48%), Gaps = 50/386 (12%)
Query: 30 FRSVCSTWRSSSVPKH---HHKFRFQFPLLKFPFNADFINKNRSFCYITKQNIFLIKPPQ 86
FRSVCS+WRS++ PK H F FP PF ++ I + + Q+I LIKP +
Sbjct: 768 FRSVCSSWRSAAGPKRLLWAHNLPF-FPSDDKPFLSNVILR------VAHQSILLIKPNE 820
Query: 87 QEQTLTLRPLLIRICQN--ARGKSKLYHPLLQNGYKFPHTF--MLDFNKLSILHLGSNFF 142
+ L ++++ N K L PL + FP + D +K ++ L
Sbjct: 821 PQCEADLFGWIVKVWDNIYVSRKMTLLKPLSSSRNYFPQHLPRIFDMSKFTVRELCREV- 879
Query: 143 MIDFDFTFNDKLYNPDDYMYPKKVVVVTCHGKKPL--------IVGTLTSPPQPLLMKCG 194
KLY+PD Y P + GK + ++ T+ + + +
Sbjct: 880 ----------KLYHPDYYCVPGHTALELELGKTVVKYLNDDKFVLLTILEYGKLAVFRSW 929
Query: 195 DEKWNVIPD-MSTKFGDICLFKGQSYAVDKIGKTVVVGPDS-SVQLVAEPLIGGGDMKFL 252
D +W VI D + ++ D+ +F G+ +A+D G+TVVV S + L A PLIGGGD KFL
Sbjct: 930 DREWTVINDYIPSRCQDLIMFDGRFFAIDYNGRTVVVDYSSFKLTLAANPLIGGGDKKFL 989
Query: 253 VESEGDLLLADVYNCL-----FTDLCNPDHNDCV---RIDLFKLNEKEKKWVKLTSLGDR 304
+ES G++ L D+ CL FT N+ + FKL E+EK+WV++ LGD+
Sbjct: 990 IESCGEMFLVDIEFCLNEKPEFTGGFYSYFNETTVSYKFKFFKLVEREKRWVEVEDLGDK 1049
Query: 305 VLFLGLGLVCSFSACASDL---CVAKGNSVIFNNYIFESHRPLESECKEYVLDLDLGRHS 361
+ FLG +FSA +D+ CV G+ V F Y E + + V D G+
Sbjct: 1050 MFFLGDD--STFSASTADIIPRCVGTGSFVFF--YTHEESLVVMDDRNLGVFDFRSGKTE 1105
Query: 362 LLSDYPEYSNLFWPPPEWIRPRYDVS 387
L++ PEY+ LFWPPP WI ++VS
Sbjct: 1106 LVNKLPEYAKLFWPPPPWITTSHEVS 1131
Score = 114 bits (284), Expect = 6e-24
Identities = 92/277 (33%), Positives = 140/277 (50%), Gaps = 26/277 (9%)
Query: 4 VDWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSVPKH----HHKFRFQFPLLKFP 59
VDWS LP +LL+LIS+ + DLI FRSVCS+WRS++ PK HH P+L P
Sbjct: 2 VDWSTLPKDLLDLISKSLESSFDLIQFRSVCSSWRSAAEPKSPLPTHH-----LPIL--P 54
Query: 60 FNADFINKNRSFCY-ITKQNIFLIKPPQQEQTLTLRPLLIRICQ--NARGKSKLYHPLLQ 116
N + + + + +++++I LIKP + LI++ + N K L PL
Sbjct: 55 DNGGSLFPDSAVGFRLSQRSILLIKPHEPCIESDSFGWLIKVEEDLNVPRKVTLLDPLCD 114
Query: 117 NGYKFPHTF--MLDFNKLSILHLGSNFFMIDFDFTFNDKLYNPDDYMYPKKVVV--VTCH 172
P F +LD +K + LG F + F+ T D + + +Y +K VV + C
Sbjct: 115 TRNSIPENFPRVLDMSKFKVRELGREFKLHYFN-TVGDIV----ESLYLEKAVVKYLDCD 169
Query: 173 GKKPLIVGTLTSPPQPLLMKCGDEKWNVIPDMSTKFGDICLFKGQSYAVDKIGKTVVVGP 232
G ++ T+ + + + D W VI DM + + D+ LF G+ +AVD G+TVVV
Sbjct: 170 GDYKFVLLTIHVSGKLAVFRSWDRAWTVINDMPSPYDDVMLFDGRFFAVDNNGRTVVVDY 229
Query: 233 DS-SVQLVAEPLIGGGD--MKFLVESEGDLLLADVYN 266
S + LVA P+ GG +K ++E L VYN
Sbjct: 230 SSLKLTLVASPVFGGDKNVLKIILEKHHPNLEKHVYN 266
>UniRef100_Q9FLV8 Gb|AAD22308.1 [Arabidopsis thaliana]
Length = 373
Score = 98.2 bits (243), Expect = 4e-19
Identities = 113/401 (28%), Positives = 172/401 (42%), Gaps = 57/401 (14%)
Query: 4 VDWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSVPKHHHKFRFQFPLLKFPFNAD 63
V WS+LP E+L+LIS +I DLIHFRSVCS WRSSS+ K H + PL P
Sbjct: 2 VKWSELPPEILHLISLKIDNPFDLIHFRSVCSFWRSSSLLKFRHMTSLRCPLPLDPGGCG 61
Query: 64 FINKNRSFCYITKQNIFLIKPPQQEQTLTLRPLLIRICQNARGKSKLYHPLLQN-----G 118
C+I ++L+K P +++ + + ++ G+ L+ L+ G
Sbjct: 62 ------DDCHILSSRVYLLKCPNRDRP---QYWMFKLQAKENGEVVLHSLFLRRNSSAYG 112
Query: 119 YKFPHTFMLDFNKLSILHLG-------SNFFMIDFDFTFNDKLYNPDDYMYPKKVVVVTC 171
+P + +D I L S +F + F K +M +
Sbjct: 113 CLYP-SLSIDLLNCQIFELAQEHVACYSEWFEL---FECISKCEERIGFMGLNR------ 162
Query: 172 HGKKPLIVGTLTSPPQPLLMKCGDEKW---NVIPDMSTKFGDICLFKGQSYAVDKIGKTV 228
+ +I+G L S + + D +W + PD + F I FKG+ YA+D+ GKT
Sbjct: 163 EKNEYMILGKL-SFNGLAMYRSVDNRWTELEITPD--SFFEGIVPFKGKFYAIDRTGKTT 219
Query: 229 VVGPDSSVQLVAEPLIGGGDMKFLVESEGD-LLLADVYNCLFTDLCNPD-HNDCVRIDLF 286
VV P V K + GD L+L ++ D P+ + ++
Sbjct: 220 VVDPTLEVNTFQRSRPCDKTRKRWLLMTGDKLILVEMCTKSRYDFHIPNIREKKIWFEIS 279
Query: 287 KLNEKEKKWVKLTSLGDRVLFLGLGLVCSFSACASDLCVAKGNSVIF-------NNYIFE 339
+LNE+ W ++ + RVLF L C+FS A+++ + NS+IF N+Y E
Sbjct: 280 ELNEERNDWDQVEDMDGRVLF--LEHYCTFSCLATEIPGFRANSIIFMDLWGGSNSYELE 337
Query: 340 SHRPLESECKEYVLDLDLGRHSLLSDYPEYSNLFWPPPEWI 380
S E K G S L D EY LF PP W+
Sbjct: 338 SILVYEFNEK--------GVRS-LRDKLEYIELFPFPPGWV 369
>UniRef100_Q9LVG8 Gb|AAF18598.1 [Arabidopsis thaliana]
Length = 374
Score = 97.1 bits (240), Expect = 8e-19
Identities = 101/348 (29%), Positives = 160/348 (45%), Gaps = 37/348 (10%)
Query: 6 WSKLPTELLNLISQRI-YEEVDLIHF---RSVCSTWRSSSVPKHHHKFRFQFPLLKFPFN 61
WS LP ++L LIS R+ ++ D IH RSVC+TWR S +P + R L KFP
Sbjct: 11 WSDLPLDILELISDRLDHDSSDTIHLLCLRSVCATWRLS-LPLSNKNNR----LSKFPKY 65
Query: 62 ADFINKNRS---FCYITKQNIFLIKPPQQEQTLTLRPLLIRICQNARGKSKLYHPLLQNG 118
F + + S F + + N++ ++ P L R L+++ + G ++ +
Sbjct: 66 LPFWSSSSSSSGFFTLKQSNVYKLEAP-----LNPRTCLVKLQETTPGIMRVLDLFSNDR 120
Query: 119 YKF-PHTF--MLDFNKLSILHLGSNFFMIDFDFTFNDKLYNPDDYMYPKKVVVVTCHGKK 175
F P F +D + + + + M ++ N P + VV+ G+
Sbjct: 121 ICFLPENFPSKIDLQEFHVRLVRRTYRM---EYANNGGGAVPCFWSLHSDKVVILSSGED 177
Query: 176 PLIVGTLTSPPQPLLMKCGDEKWNVIPDM-STKFGDICLFK-GQSYAVDKIGKTVVVGPD 233
I+ + L DEKW ++ + + + DI L+K + VD GKTV+ D
Sbjct: 178 SAIIAIHSGGKLGFLKSGNDEKWKILDNSWNVIYEDIMLYKDNRCIVVDDKGKTVIYDVD 237
Query: 234 SSVQLVAEPLIGGGD-MKFLVE-SEGDLLLADVYNCLFTDLCNPD-HNDCVRIDLFKLNE 290
V +AE L+GGG K LVE S G++ L D Y + T C D V ++ L
Sbjct: 238 FKVSDLAEGLVGGGGHKKHLVECSGGEVFLVDKY--VKTVWCKSDISKSVVEFRVYNLKR 295
Query: 291 KEKKWVKLTSLGDRVLFLGLGLVCSFSA--CASDLCVAKGNSVIFNNY 336
+EK+W ++ LGD LF+G CSFS A DL G + +++Y
Sbjct: 296 EEKRWEEVRDLGDVALFIGDD--CSFSVQNPAGDLA---GGFIFYSDY 338
>UniRef100_Q9XIR1 F13O11.14 protein [Arabidopsis thaliana]
Length = 383
Score = 86.7 bits (213), Expect = 1e-15
Identities = 91/340 (26%), Positives = 147/340 (42%), Gaps = 25/340 (7%)
Query: 5 DWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSS-SVPKHHHKFRFQFP-LLKFPFNA 62
+WS+LP ELLNLIS+ + D++H RS+C +WRS+ P + P KFP
Sbjct: 3 NWSQLPEELLNLISKNLDNCFDVVHARSICRSWRSAFPFPSSLSTLSYSLPTFAKFPL-- 60
Query: 63 DFINKNRSFCYITKQNIFLIKPPQQEQTLTLRPLLIRICQNARGKSKLYHPL-LQNGYKF 121
++ C + K IFL + + L + +L PL KF
Sbjct: 61 ----VSKDLCTLKKIQIFLFRARNPAADIPEYFLGGIDQDQSNDHMELPSPLQCSVKVKF 116
Query: 122 PHT--FMLDFNKLSILHLGSNFFMIDFDFTFNDKLYNPDDYMYPKKVVVVTCHGKKPLIV 179
P + F+++ I+ LG + MI +D Y ++ KK +G +V
Sbjct: 117 PQSDPFLVNMLDYQIIPLGFQYIMIGWDPESLANGYVGVAFLPVKK------NGGDEFVV 170
Query: 180 GTLTSPPQPLLMKCGDEKWNVIPDMS-TKFGDICLFKGQSYAVDKIGKTVVVGPDSSVQL 238
L L+++ + +W + S + F+G+ Y G V P S Q
Sbjct: 171 -LLRYRNHLLVLRSSEMRWMKVKKTSIASCKGLVSFRGRFYVTFLNGDIYVFDPYSLEQT 229
Query: 239 VAEPLIGGGDMKFLVESEGD-LLLADVYNCL-FTDLCNPDHNDCVRIDLFKLNEKEKKWV 296
+ P K+L+ + D L L + +N D+ + C + +L+E+ +WV
Sbjct: 230 LLMPSEPLRSSKYLIPNGSDELFLVEKFNPFPEADVLDLSRFAC---RVSRLDEEAGQWV 286
Query: 297 KLTSLGDRVLFLG-LGLVCSFSACASDLCVAKGNSVIFNN 335
++ LGDRVLF+G G VC + D C GNS++F N
Sbjct: 287 EVIDLGDRVLFIGHFGNVCCSAKELPDGCGVSGNSILFTN 326
>UniRef100_Q9ZUH0 Expressed protein [Arabidopsis thaliana]
Length = 374
Score = 77.4 bits (189), Expect = 7e-13
Identities = 95/341 (27%), Positives = 146/341 (41%), Gaps = 46/341 (13%)
Query: 5 DWSKLPTELLNLISQRIYEEV-DLIHFRSVCSTWRSSSVPKHHHKFRFQFPLLKFPFNAD 63
DWS+LP ELL++IS + + D +H RSVC +WRS+ P R + L FP +
Sbjct: 15 DWSQLPEELLHIISTHLEDHYFDAVHARSVCRSWRST-FPFPSSLLRQSYSLPAFPLESK 73
Query: 64 FINKNRSFCYITKQNIFLIK--PPQQEQTLTLRPLLIRICQNARGKSKLYHPLLQNGYKF 121
+ C + K +FL + P + L + Q+ LQ K
Sbjct: 74 DL-----CCTLEKVPLFLFRVLTPPDAADASSEYFLGGLGQDKSNDHVELPSPLQCSVKV 128
Query: 122 --PHTFMLDFNKLS--ILHLGSNF-FMIDFDFTFNDKLYNPDDYMYPKKVVVVTCHGKKP 176
P T + N L I+ LG + MI NP++Y + + G
Sbjct: 129 NVPGTEPILMNMLDCQIIPLGHKYRLMIGC---------NPEEYS--AAFLPLNEQGGGG 177
Query: 177 LIVGTLTSPPQPLLMKCGDEKWNVIPDMST-KFGDICLFKGQSYAVDKIGKTVVVGPDSS 235
V L L+++ + +W + ST ++ F+G+ YA G T V+ P S
Sbjct: 178 EFVALLDCTDLFLVLRSTEMRWIRLEKPSTASCKELFTFRGRFYATFFNGDTFVIDPSS- 236
Query: 236 VQLVAEPLIGGGDMKFLVESEGDLLLADVYNCLFTDLCNPDHNDCVRIDLFKLNEKEKKW 295
L A PL D FLV S + L D +R + +L+E+ +W
Sbjct: 237 --LEATPLTPHIDSNFLVPSGNEELFLV-------------KTDFLRCRVSRLDEEAAEW 281
Query: 296 VKLTSLGDRVLFLGLGLVCSFSACASDL---CVAKGNSVIF 333
V+++ LGDRVLFLG G + +F A +L C G+S++F
Sbjct: 282 VEVSDLGDRVLFLG-GHLGNFYCSAKELPHGCGLTGDSILF 321
>UniRef100_Q9SHX9 F1E22.13 [Arabidopsis thaliana]
Length = 360
Score = 68.2 bits (165), Expect = 4e-10
Identities = 87/351 (24%), Positives = 152/351 (42%), Gaps = 50/351 (14%)
Query: 5 DWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSVPKHHHK--FRFQFPLLKFPFNA 62
DWS LP +LLN+I+ R++ ++L FRS+C +WR SSVP K FR + +L P
Sbjct: 3 DWSTLPVDLLNMIAGRLFSNIELKRFRSICRSWR-SSVPGAGKKNPFRTRPLILLNP--- 58
Query: 63 DFINKNRSFCYITKQNIFLIKPPQQEQTLTLRP----LLIRICQNARGKSKLYHPLLQNG 118
N N+ ++ FL + TL+ P L+ + GK L PL
Sbjct: 59 ---NPNKPLTDHRRRGEFLSRSAFFRVTLSSSPSQGWLIKSDVDVSSGKLHLLDPL---- 111
Query: 119 YKFPHTFMLDFNKLSILHLGSNFFMIDFDFTFNDKLYNPDDYMYPKKVVVVTCHGKKPLI 178
++L + H + +F T + Y D+ K+ + K+ +
Sbjct: 112 -----------SRLPMEHSRKRVDLSEFTITEIREAYQVHDWRTRKETRPIF---KRVAL 157
Query: 179 VGTLTSPPQPLLMKCGDEK--WNV----IPDMSTKFGDICLFKGQSYAVDKIGKTVVVGP 232
V Q L ++ + W++ + +F DI + KGQ+YA+D IG +V
Sbjct: 158 VKDKEGDNQVLGIRSTGKMMYWDIKTWKAKEEGYEFSDIIVHKGQTYALDSIG--IVYWI 215
Query: 233 DSSVQLVA-EPLIGG--GDMKFLVESEGDLLLADVYNCLFT-----DLCNPDHNDCVRID 284
S ++ + PL+G GD + LVE G+ + + T D ++ V
Sbjct: 216 RSDLKFIRFGPLVGDWTGDRR-LVECCGEFYIVERLVGESTWKRKADDTGYEYAKTVGFK 274
Query: 285 LFKLNEKEKKWVKLTSLGDRVLFLGLGLVCSFSACASDLCVAKGNSVIFNN 335
++K ++++ K +++ SLGD+ + FS A + N++ F +
Sbjct: 275 VYKFDDEQGKMMEVKSLGDKAFVIATD--TCFSVLAHEFYGCLENAIYFTD 323
>UniRef100_Q84V03 Hypothetical protein At2g16365/F16F14.28 [Arabidopsis thaliana]
Length = 332
Score = 65.1 bits (157), Expect = 3e-09
Identities = 79/326 (24%), Positives = 134/326 (40%), Gaps = 39/326 (11%)
Query: 1 MATVDWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSVPKHHHKFRFQFPLLKFPF 60
+ DW LP +LL LI + ++ FRSVCS+WR S VP H F
Sbjct: 19 LTMADWPLLPNDLLELIMGHLETSFEIFLFRSVCSSWR-SVVPPLDHSRCLGIKTHDISF 77
Query: 61 NADFINKNR---SFCYITKQNIFLIK--PPQQEQTLTLRPLLIRICQNARGKSK-LYHPL 114
N F R +C + K I+L+K P + LL + + G+ K L PL
Sbjct: 78 NVGFSFNGRPTNEYCTLKKIPIYLVKFWTPFGDDY-----LLAEMRERNDGEPKLLLSPL 132
Query: 115 LQNGYKFPHTFMLDFNKLSILHLGSNF--FMIDFDFTFNDKLYNPDDYMYPKKV------ 166
NG K+ + NK+ L S F ++ T+ +K + + +P K+
Sbjct: 133 SSNGIKYG----MGINKVLFNSLTSPIIPFGQYYEITYIEKRPSEYRFGFPYKLEWVEIT 188
Query: 167 -----VVVTCHGKKPLIVGTLTSPPQPLLMKCGDEKWNVIPDMSTKFG--DICLFKGQSY 219
+ + + V ++ + + W + + K+ D+ FKG+ Y
Sbjct: 189 ERVEFLKLDSEDSRDFAVLFAGRMCNLVMYRSRNMSWTQVVEHPEKYAYQDLVAFKGKFY 248
Query: 220 AVDKIGKTVVVGPDSSVQLVAEPLIGGGDM---KFLVESEGDLLLADVYNCL---FTDLC 273
AVD G+ V + S ++ P +GG + LV+S +LLL + + + +
Sbjct: 249 AVDSSGRGRVFVVELSFEVTEIPSVGGSQQSSKESLVQSGEELLLVQRFTPVGRRYDEYI 308
Query: 274 NPDHNDCVRIDLFKLNEKEKKWVKLT 299
++C +DL K E+E + +T
Sbjct: 309 YIHGSEC--LDLMKKEERESGFKLMT 332
>UniRef100_Q9LS04 Gb|AAC14516.2 [Arabidopsis thaliana]
Length = 377
Score = 64.7 bits (156), Expect = 4e-09
Identities = 84/366 (22%), Positives = 142/366 (37%), Gaps = 59/366 (16%)
Query: 1 MATVDWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSS-SVPKHHHKFRFQFPLLKFP 59
M +WS LP ELL+LI+ R +D++ RS C +WRS+ ++ K +FRF+ L
Sbjct: 1 MEKTEWSDLPEELLDLIANRYSSNIDVLRIRSTCKSWRSAVAMSKERLQFRFERYLP--- 57
Query: 60 FNADFINKNRSFCYITKQNIFLIKPPQQEQTLTLRPLLIRICQNARGKSKLYHPLLQNGY 119
+ + +++ F I P + + L+R Q ++ K+ +G
Sbjct: 58 -----TSNKKIKAHLSPTTFFRITLP---SSCPNKGWLVRTRQASKMYRKITLLCPLSGE 109
Query: 120 KFPHTFM-LDFNKLSILHLGSNFFMIDFDFTFNDKL---------YNPDDYMYPKKV--- 166
+ + LD K+ + + ++ + FD ++K+ Y + P V
Sbjct: 110 RITRSHQTLDLLKVGVSEIRQSYEIQIFDGLKDEKIPLDSEIFSNYIKNSDKIPSGVFFR 169
Query: 167 --------VVVTCHGKKPLIVGTLTSPPQPLLMKCGDEKWNVIPDMSTKFGDICLFKGQS 218
V H K T + G W I + F DI L G+
Sbjct: 170 VAFLDNIFFAVNYHDNKIWCCKT----------REGSRSWTKIKNQVEDFSDIILHMGRI 219
Query: 219 YAVD--------KIGKTVVVGPDSSVQLVAEPLIGGGDMKFLVESEGDLLLADVYNCLFT 270
YAVD + + +V SS L D + LVE GDL + +
Sbjct: 220 YAVDLKGAIWWISLSQLTIVQQTSSTPLDYYKYDSCQDTR-LVEYCGDLCIVHELSITRN 278
Query: 271 DLCNPDHNDCVRIDLFKLNEKEKKWVKLTSLGDRVLFLGLGLVCSFSACASDLCVAKGNS 330
+ V ++K++E KWV+++ LGD L + F+ AS+ NS
Sbjct: 279 HI-----QRTVGFKVYKMDEDLAKWVEVSCLGDNTLIVACN--SCFTVVASEYHGCLKNS 331
Query: 331 VIFNNY 336
+ F+ Y
Sbjct: 332 IYFSYY 337
>UniRef100_Q9C6X9 Hypothetical protein T7O23.22 [Arabidopsis thaliana]
Length = 347
Score = 52.4 bits (124), Expect = 2e-05
Identities = 79/356 (22%), Positives = 132/356 (36%), Gaps = 69/356 (19%)
Query: 1 MATVDWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSVPKH--HHKFRFQFPLLKF 58
M V WS L +L++L++ + ++L+ FRS+C WRS+ K H+ F P K
Sbjct: 1 MGKVGWSDLHEDLIDLLANNLSSNINLLRFRSICKPWRSTVATKKRLHNHFERNLPTFK- 59
Query: 59 PFNADFINKNRSFCYIT-----KQNIFLIKPPQQEQTLTLRPLLIRICQNARGKSKLYHP 113
+ +F +T + +LIK Q ++ L + K
Sbjct: 60 --KKKTVVSPSTFFRVTLPSPCRNKGWLIKNRQVSESSKNNLLSPLSGKTITPSDKTLDL 117
Query: 114 LLQNGYKFPHTFMLDFNKLSILHLGSNFFMIDFDFTFNDKLYNPDDYMYPKKVVVVTCHG 173
L ++ L + ++ L + FF++DF K + C
Sbjct: 118 LKVECFRDSSILQLFADSDRVVFLDNVFFVVDF------------------KNEIWCC-- 157
Query: 174 KKPLIVGTLTSPPQPLLMKCGDE--KWNVIPDMSTK-FGDICLFKGQSYAVDKIG----- 225
K G+E W I + K F DI L KG+ YA+D G
Sbjct: 158 ------------------KSGEETRHWTRINNEEAKGFLDIILHKGKIYALDLTGAIWWI 199
Query: 226 -----KTVVVGPDSSVQLVAEPLIGGGDMKFLVESEGDLLLAD-VYNCLFTDLCNPDHND 279
GP + V I K LVE G+L + Y +
Sbjct: 200 SLSELSIYQYGPSTPVDFYE---IDNCKEKRLVEYCGELCVVHRFYKKFCVKRVLTERTV 256
Query: 280 CVRIDLFKLNEKEKKWVKLTSLGDRVLFLGLGLVCSFSACASDLCVAKGNSVIFNN 335
C ++ +K+++ +WV+++SLGD+ L + F AS+ N++ FN+
Sbjct: 257 CFKV--YKMDKNLVEWVEVSSLGDKALIVATD--NCFLVLASEYYGCLENAIYFND 308
>UniRef100_Q84SZ0 Hypothetical protein OSJNBa0087C10.17 [Oryza sativa]
Length = 531
Score = 50.8 bits (120), Expect = 7e-05
Identities = 94/416 (22%), Positives = 162/416 (38%), Gaps = 44/416 (10%)
Query: 1 MATVDWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSVPKHH--HKFRFQFPLLKF 58
M + LP +LL I R+ E DL+ SVCS+W S+ H R Q P L +
Sbjct: 115 MVEAELPHLPEDLLVQILSRL-EIPDLLRASSVCSSWHSAYTTLHSLGQYKRHQTPCLFY 173
Query: 59 PFNADFINKNRSFCYITKQN----IFLIKPPQQEQTLTLRPLLIRICQNARGKSKLYHPL 114
++ KN Y + I L PP +++ L + + + + L +P+
Sbjct: 174 --TSESAGKNVGCIYSLAEQRTYKITLPDPPIRDRYLIGSSDGWLVTIDDKCEMHLLNPV 231
Query: 115 LQNGYKFPHTFML-------DFNKLSILHLGSNFFMID---FDFTFNDKLYNP--DDYMY 162
+ P + D + + + + F D F + +P
Sbjct: 232 TREQMALPPVITMEQVNPTYDESGAIVKYENRSQFWHDGVMFSSRSMGSIISPRWQQLFL 291
Query: 163 PKKVVVVTCHGKKPLIVGTLTSP-PQPLLMKCGDEKWNVIPDMSTKFGDICLFKGQSYAV 221
+ V + L+V + +P Q + GD++W+ +P+ ++ D G YAV
Sbjct: 292 TGRAFVFSETSTGKLLVVLIRNPFGQLSFARVGDDEWDYLPEYG-RYEDCTYKDGLLYAV 350
Query: 222 DKIGKTVVV---GPDSSVQLVAEPL--IGGGDMK--FLVESEGDLLLADVYNCLFTDLCN 274
+G+ + GP + V++V + IG GD L GD+L ++ D +
Sbjct: 351 TTLGEIHAIDLSGPIAMVKVVMGKVMDIGDGDRNTYILHAPWGDVL--QIWKTEEDDYIH 408
Query: 275 PDHND-------CVRIDLFKLNEKEKKWVKLTSLGDRVLFLGLGLVCSFSACASDLCVAK 327
P +D I+++K + E+K VK+ L D VLF+G A + K
Sbjct: 409 PSEDDYDAILKNTASIEVYKSDLVEEKLVKINRLQDHVLFVGHNQTLCLR--AEEFPSLK 466
Query: 328 GNSVIFNNYIFESHRPLESECKEY-VLDLDLGRHSLLSDYPEYSNLFWPPPEWIRP 382
N F + ++ ++ V +L+ L +SN WP P WI P
Sbjct: 467 ANHAYFTDDSQNWITEFKNNRRDIGVFNLEDNSRDELGSPQLWSN--WPSPVWITP 520
>UniRef100_Q9SIW9 Hypothetical protein At2g16300 [Arabidopsis thaliana]
Length = 467
Score = 48.9 bits (115), Expect = 3e-04
Identities = 96/410 (23%), Positives = 157/410 (37%), Gaps = 82/410 (20%)
Query: 5 DWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSVPKHHHKFRFQFPLLKFPFNADF 64
DWS LP +LL LI + +++ FRSVCS+WR S VP FNA F
Sbjct: 3 DWSLLPNDLLELIVGHLETSFEIVLFRSVCSSWR-SVVPPQDQSRCLSIKTHDISFNAGF 61
Query: 65 INKNRSFCY------ITKQNIFLIK--PPQQEQTLTLRPLLIRICQNARGKSK-LYHPLL 115
+ + +TK I+L+K P + LL + + G+ L PL
Sbjct: 62 SFSGQPTDHPLFGSTLTKIPIYLVKFWTPFGDDY-----LLAEMREREGGEPMFLLSPLS 116
Query: 116 QNGYKFPHTFMLDFNKLSILHLGSNF--FMIDFDFTFNDKLYNPDDYMYPKKVVVVTCHG 173
N + + NK+ L S F ++ T+++K P Y Y + CH
Sbjct: 117 SNRI----IYGMGINKVLFNSLTSPIIPFGQYYEITYSEK--QPIRYRYG-----LPCHL 165
Query: 174 KKPLIVGTLTSPPQPLLMKCGDEK----------------------WNVIPDMSTKFG-- 209
L +T + L + D + W + + K+
Sbjct: 166 WDKLEWVEITERVEFLKLDSEDSRDFAVLFAGRMCNLVMYRSRNMSWTQVVEHPEKYAYQ 225
Query: 210 DICLFKGQSYAVDKIGKTVVVGPDSSVQLVAEPLIGGGDM---KFLVESEGDLLLADVYN 266
D+ FKG+ YA+D G+ V + S++++ P + G + LV S +LLL +
Sbjct: 226 DLVAFKGKFYALDSSGRGRVFVVELSLEVMEIPSVRGSQQSSKENLVLSGEELLLVQRFT 285
Query: 267 CLFTDLCNPDHNDCVRIDLFKLNEKEKKWVKLTSLGDRVLFLGLGLVCSFS--ACASDLC 324
++ + D +F+L+E+E + + L CS CA + C
Sbjct: 286 ---PEVRHYDEYLYTWFRVFRLDEEEGRETQ------------TNLCCSVHKLPCAKENC 330
Query: 325 VA---KGNSVIFNN-----YIFESHRPLE--SECKEYVLDLDLGRHSLLS 364
+ N I +N + ++ R +ECK Y+ SLLS
Sbjct: 331 IVFIESNNCFIIDNDSVLLFDLKTRRTSTAFNECKGYMGVFGANLESLLS 380
Score = 36.6 bits (83), Expect = 1.3
Identities = 19/48 (39%), Positives = 27/48 (55%), Gaps = 1/48 (2%)
Query: 6 WSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSVPKHHHK-FRFQ 52
WS L +LL LI ++++ RSVCS+WRS P + FR+Q
Sbjct: 402 WSLLLNDLLKLIVGHFETSFEIVNLRSVCSSWRSVVPPLDNFPLFRYQ 449
>UniRef100_Q6ASV3 Expressed protein [Oryza sativa]
Length = 468
Score = 48.9 bits (115), Expect = 3e-04
Identities = 75/324 (23%), Positives = 135/324 (41%), Gaps = 44/324 (13%)
Query: 21 IYEEVDLIHFRSVCSTWRSSSVPKHH---HKFRFQFPLLKFPFNA--DFINKNRSFCYIT 75
+ E DL+ SVC++WRS+ +K Q P L + + D + S
Sbjct: 76 LLEIPDLVRAGSVCNSWRSAYNGMRSLGIYKLS-QTPCLLYTSESAGDSVVSLYSLVEKR 134
Query: 76 KQNIFLIKPPQQEQTLTLRPLLIRICQNARGKSKLYHPLLQNGYKFPHTFMLD-----FN 130
+ I L +PP + + L L + + + L +P+ P ++ FN
Sbjct: 135 EYKITLPEPPVRSRFLIGSSLGCLVTVDDVSEMHLVNPITGEQIALPSVITIEHVNPIFN 194
Query: 131 KLSILHLGSNFFMIDFDFTFNDKLYNPD----------DYMYPKKVVVVTCHGKKPLIVG 180
+ +H M ++ + ++Y+ + +Y+ K V + L+V
Sbjct: 195 ESGAIH------MYEYSWYSASRVYHSEPSIFSLDELREYLLDKAFVFSDTSTENYLVVL 248
Query: 181 TLTSPPQPLLMKCGDEKWNVIPDMSTKFGDICLFK-GQSYAVDKIGKTVVV---GPDSSV 236
Q + GD+KW +P T + D C++K G YAV+K+G+ GP ++
Sbjct: 249 IHNPHSQLSFARVGDDKWTWLPP-HTHYAD-CIYKDGILYAVNKVGEIHAFDLSGPVVTM 306
Query: 237 QLVAEPLIGGG-DMKFLVESE-GDLLLA-----DVYNCLFTDLCNPDH----NDCVRIDL 285
+ + E + G D ++V++ GDLL + DL + D + I +
Sbjct: 307 KTIIEMVPGYACDKMYIVQAPWGDLLQVWRSYEYIEGDYEADLHDADPAISVENTAEIKI 366
Query: 286 FKLNEKEKKWVKLTSLGDRVLFLG 309
F ++ EKK V++ +L VLFLG
Sbjct: 367 FVVDTVEKKRVEIENLDGHVLFLG 390
>UniRef100_Q9FWC8 Hypothetical protein OSJNBb0018B10.12 [Oryza sativa]
Length = 579
Score = 46.6 bits (109), Expect = 0.001
Identities = 18/43 (41%), Positives = 28/43 (64%)
Query: 1 MATVDWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSVP 43
MA WS LP +LL +S R+Y + D +H VC+ WR++++P
Sbjct: 1 MAGGGWSSLPADLLREVSGRLYSDADHLHIHQVCTHWRAATLP 43
>UniRef100_Q7XGL9 Hypothetical protein [Oryza sativa]
Length = 450
Score = 45.8 bits (107), Expect = 0.002
Identities = 17/38 (44%), Positives = 28/38 (72%)
Query: 4 VDWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSS 41
+DWS LP +LL L S R+++ +D + FR+VC WR+++
Sbjct: 1 MDWSGLPGDLLRLFSGRLHDPLDFLRFRAVCRAWRAAT 38
Score = 38.9 bits (89), Expect = 0.26
Identities = 21/57 (36%), Positives = 35/57 (60%), Gaps = 4/57 (7%)
Query: 279 DCVRIDLFKLN--EKEKKWVKLTSLGDRVLFLGLGLVCSFSACASDLCVAKGNSVIF 333
DC +++ +L ++ +WV++ S+GDR+LF +GL FS A+D +GN V F
Sbjct: 255 DCRALEVHRLEIAGEKSRWVQMRSIGDRMLF--VGLYQGFSLRAADFAGLEGNCVYF 309
Score = 36.6 bits (83), Expect = 1.3
Identities = 20/57 (35%), Positives = 34/57 (59%), Gaps = 4/57 (7%)
Query: 279 DCVRIDLFKLN--EKEKKWVKLTSLGDRVLFLGLGLVCSFSACASDLCVAKGNSVIF 333
DC +++ +L ++ +WV++ S+ DR+LF +GL FS A+D +GN V F
Sbjct: 356 DCCALEVHRLEIAGEKSRWVQMRSIRDRMLF--VGLYQGFSLRAADFAGLEGNCVYF 410
>UniRef100_Q9LNY7 F9C16.31 [Arabidopsis thaliana]
Length = 311
Score = 45.1 bits (105), Expect = 0.004
Identities = 20/59 (33%), Positives = 32/59 (53%), Gaps = 2/59 (3%)
Query: 1 MATVDWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSVPKH--HHKFRFQFPLLK 57
M V WS L +L++L++ + ++L+ FRS+C WRS+ K H+ F P K
Sbjct: 1 MGKVGWSDLHEDLIDLLANNLSSNINLLRFRSICKPWRSTVATKKRLHNHFERNLPTFK 59
Score = 35.8 bits (81), Expect = 2.2
Identities = 42/158 (26%), Positives = 67/158 (41%), Gaps = 21/158 (13%)
Query: 192 KCGDEK--WNVIPDMSTK-FGDICLFKGQSYAVDKIGKTVVV----------GPDSSVQL 238
K G+E W I + K F DI L KG+ YA+D G + GP + V
Sbjct: 122 KSGEETRHWTRINNEEAKGFLDIILHKGKIYALDLTGAIWWISLSELSIYQYGPSTPVDF 181
Query: 239 VAEPLIGGGDMKFLVESEGDLLLAD-VYNCLFTDLCNPDHNDCVRIDLFKLNEKEKKWVK 297
I K LVE G+L + Y + C ++ +K+++ +WV+
Sbjct: 182 YE---IDNCKEKRLVEYCGELCVVHRFYKKFCVKRVLTERTVCFKV--YKMDKNLVEWVE 236
Query: 298 LTSLGDRVLFLGLGLVCSFSACASDLCVAKGNSVIFNN 335
++SLGD+ L + F AS+ N++ FN+
Sbjct: 237 VSSLGDKALIVATD--NCFLVLASEYYGCLENAIYFND 272
>UniRef100_Q9FLP7 Dbj|BAA95762.1 [Arabidopsis thaliana]
Length = 340
Score = 45.1 bits (105), Expect = 0.004
Identities = 34/145 (23%), Positives = 62/145 (42%), Gaps = 10/145 (6%)
Query: 194 GDEKWNVIPDMSTKFGDICLFKGQSYAVDKIGKTV-----VVGPDSSVQLVAEPLIGGGD 248
GD++W + +++ DI G +A+D++G+ P ++ P
Sbjct: 152 GDKQWTDLESVASSVDDIVFCNGVFFAIDRLGEIYHCELSANNPKATPLCSTSPFRYDSC 211
Query: 249 MKFLVESEGDLLLADVYNCLFTDLCNPDHNDCVRIDLFKLNEKEKKWVKLTSLGDRVLFL 308
K+L ES+ D L + D C+ + ++++ N + +W K+ SL + LFL
Sbjct: 212 KKYLAESDYDELWVVLKKLELNDDCDFE----TSFEIYEFNRETNEWTKVMSLRGKALFL 267
Query: 309 GLGLVCSFSACASDLCVAKGNSVIF 333
C + A + K NSV F
Sbjct: 268 SPQGRC-IAVLAGERGFFKDNSVYF 291
Score = 34.3 bits (77), Expect = 6.4
Identities = 13/36 (36%), Positives = 23/36 (63%)
Query: 6 WSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSS 41
WS+ ELLN + + + D+++ +VCS+W+ SS
Sbjct: 8 WSEFLPELLNTVFHNLNDARDILNCATVCSSWKDSS 43
>UniRef100_Q6ESP2 Hypothetical protein P0684A08.40 [Oryza sativa]
Length = 435
Score = 45.1 bits (105), Expect = 0.004
Identities = 20/41 (48%), Positives = 29/41 (69%)
Query: 1 MATVDWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSS 41
MA WS LP+ELL+ I+ + E D+ FRSVCS+WR+++
Sbjct: 78 MAATGWSDLPSELLSEIAGGLVELGDIARFRSVCSSWRAAA 118
>UniRef100_Q8L477 Hypothetical protein B1147B04.22 [Oryza sativa]
Length = 340
Score = 44.7 bits (104), Expect = 0.005
Identities = 19/47 (40%), Positives = 28/47 (59%)
Query: 4 VDWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSVPKHHHKFR 50
+DW+ LP +LL IS + E D + FR+VC WR++ K H F+
Sbjct: 1 MDWASLPADLLFSISSHLREPEDFVRFRAVCPQWRAAVSHKEHAFFQ 47
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.142 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 757,279,323
Number of Sequences: 2790947
Number of extensions: 34017673
Number of successful extensions: 63947
Number of sequences better than 10.0: 86
Number of HSP's better than 10.0 without gapping: 45
Number of HSP's successfully gapped in prelim test: 41
Number of HSP's that attempted gapping in prelim test: 63819
Number of HSP's gapped (non-prelim): 133
length of query: 402
length of database: 848,049,833
effective HSP length: 130
effective length of query: 272
effective length of database: 485,226,723
effective search space: 131981668656
effective search space used: 131981668656
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 76 (33.9 bits)
Medicago: description of AC147741.8


