

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147712.6 + phase: 0
(80 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
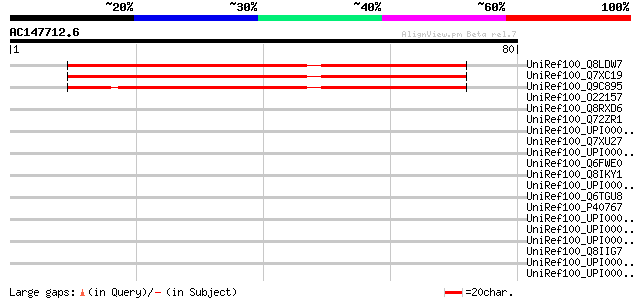
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8LDW7 Hypothetical protein [Arabidopsis thaliana] 67 1e-10
UniRef100_Q7XC19 Hypothetical protein [Oryza sativa] 63 2e-09
UniRef100_Q9C895 Hypothetical protein F7A10.17 [Arabidopsis thal... 62 3e-09
UniRef100_O22157 Hypothetical protein At2g44950 [Arabidopsis tha... 41 0.007
UniRef100_Q8RXD6 Hypothetical protein At2g44950:At2g44960 [Arabi... 41 0.007
UniRef100_Q72ZR1 DNA recombinase, putative [Bacillus cereus] 40 0.019
UniRef100_UPI000032CA6C UPI000032CA6C UniRef100 entry 38 0.056
UniRef100_Q7XU27 OSJNBb0034G17.7 protein [Oryza sativa] 37 0.12
UniRef100_UPI000049883E UPI000049883E UniRef100 entry 37 0.16
UniRef100_Q6FWE0 Candida glabrata strain CBS138 chromosome D com... 36 0.21
UniRef100_Q8IKY1 Hypothetical protein [Plasmodium falciparum] 36 0.28
UniRef100_UPI000049975E UPI000049975E UniRef100 entry 35 0.36
UniRef100_Q6TGU8 Ribosome binding protein 1 homolog 180kDa [Brac... 35 0.36
UniRef100_P40767 Hypothetical protein yvcE [Bacillus subtilis] 35 0.47
UniRef100_UPI00002BC7C3 UPI00002BC7C3 UniRef100 entry 35 0.61
UniRef100_UPI000049A011 UPI000049A011 UniRef100 entry 34 0.80
UniRef100_UPI000028F98B UPI000028F98B UniRef100 entry 34 0.80
UniRef100_Q8IIG7 Hypothetical protein [Plasmodium falciparum] 34 0.80
UniRef100_UPI0000499FE0 UPI0000499FE0 UniRef100 entry 34 1.0
UniRef100_UPI000049875E UPI000049875E UniRef100 entry 34 1.0
>UniRef100_Q8LDW7 Hypothetical protein [Arabidopsis thaliana]
Length = 365
Score = 67.0 bits (162), Expect = 1e-10
Identities = 32/63 (50%), Positives = 47/63 (73%), Gaps = 2/63 (3%)
Query: 10 DLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVFSDRP 69
+LD +R +KKLEEELM +N ++ EL+SE+ E + +L +EE++ CKN++KC V DRP
Sbjct: 263 ELDDERREKKKLEEELMELNKELEELDSESVEAAIVRL--QEEVKNCKNILKCGVCFDRP 320
Query: 70 KEV 72
KEV
Sbjct: 321 KEV 323
>UniRef100_Q7XC19 Hypothetical protein [Oryza sativa]
Length = 789
Score = 63.2 bits (152), Expect = 2e-09
Identities = 31/63 (49%), Positives = 46/63 (72%), Gaps = 2/63 (3%)
Query: 10 DLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVFSDRP 69
+L+ +R+ + KLEEE V N+++EL SET ET +Q+L ++EI+ CK ++KC V DRP
Sbjct: 687 ELERERNERIKLEEEYEEVKNEVSELTSETEETTIQKL--QDEIKECKAILKCGVCFDRP 744
Query: 70 KEV 72
KEV
Sbjct: 745 KEV 747
>UniRef100_Q9C895 Hypothetical protein F7A10.17 [Arabidopsis thaliana]
Length = 899
Score = 62.4 bits (150), Expect = 3e-09
Identities = 32/63 (50%), Positives = 46/63 (72%), Gaps = 3/63 (4%)
Query: 10 DLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVFSDRP 69
+LD +R +KKLEEELM +N ++ EL SE+ E + +L +EE++ CKN++KC V DRP
Sbjct: 798 ELDDERE-KKKLEEELMELNKELEELGSESVEAAIVRL--QEEVKNCKNILKCGVCFDRP 854
Query: 70 KEV 72
KEV
Sbjct: 855 KEV 857
>UniRef100_O22157 Hypothetical protein At2g44950 [Arabidopsis thaliana]
Length = 518
Score = 41.2 bits (95), Expect = 0.007
Identities = 22/68 (32%), Positives = 42/68 (61%), Gaps = 3/68 (4%)
Query: 6 ASEKDLDSKRSSQKKLEEELMNVNNQIAELNSET-GETVVQQLEEEEEIRVCKNMIKCTV 64
A E +L+ +R +++++EEE+ +++ L S G + +Q+L +E + K ++KC
Sbjct: 411 ALELELEIERFNRRRIEEEMEIAKKKVSRLRSLIEGSSAIQKLRQE--LSEFKEILKCKA 468
Query: 65 FSDRPKEV 72
+DRPKEV
Sbjct: 469 CNDRPKEV 476
>UniRef100_Q8RXD6 Hypothetical protein At2g44950:At2g44960 [Arabidopsis thaliana]
Length = 878
Score = 41.2 bits (95), Expect = 0.007
Identities = 22/68 (32%), Positives = 42/68 (61%), Gaps = 3/68 (4%)
Query: 6 ASEKDLDSKRSSQKKLEEELMNVNNQIAELNSET-GETVVQQLEEEEEIRVCKNMIKCTV 64
A E +L+ +R +++++EEE+ +++ L S G + +Q+L +E + K ++KC
Sbjct: 771 ALELELEIERFNRRRIEEEMEIAKKKVSRLRSLIEGSSAIQKLRQE--LSEFKEILKCKA 828
Query: 65 FSDRPKEV 72
+DRPKEV
Sbjct: 829 CNDRPKEV 836
>UniRef100_Q72ZR1 DNA recombinase, putative [Bacillus cereus]
Length = 526
Score = 39.7 bits (91), Expect = 0.019
Identities = 25/81 (30%), Positives = 39/81 (47%), Gaps = 8/81 (9%)
Query: 1 MSVASASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCK--- 57
+S+ +K+L RS+ +KLE E + +N QI ELN E Q++ +EE + K
Sbjct: 381 LSIEKRLKKELSDLRSANRKLETEAIELNQQIEELNREIKLHYEQEIIQEEVLEFMKILM 440
Query: 58 -----NMIKCTVFSDRPKEVK 73
M + V D E+K
Sbjct: 441 SHRSNKMHELNVIKDNQDELK 461
>UniRef100_UPI000032CA6C UPI000032CA6C UniRef100 entry
Length = 183
Score = 38.1 bits (87), Expect = 0.056
Identities = 21/58 (36%), Positives = 36/58 (61%), Gaps = 1/58 (1%)
Query: 8 EKDLDSKRSSQKKLEEELMNVNNQIAELNSETGET-VVQQLEEEEEIRVCKNMIKCTV 64
+K+L +KK++ ++ +NQ+ +L E E +VQ ++EEEEIR+ N IK T+
Sbjct: 3 KKELTDFTFYKKKVDIQIKKSSNQLLKLAKENAEGYIVQNIKEEEEIRLLLNQIKQTL 60
>UniRef100_Q7XU27 OSJNBb0034G17.7 protein [Oryza sativa]
Length = 883
Score = 37.0 bits (84), Expect = 0.12
Identities = 17/63 (26%), Positives = 40/63 (62%), Gaps = 1/63 (1%)
Query: 10 DLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVFSDRP 69
+L+ +R S+K++E++L ++ + + L ++ E+ V + + E++ + ++KC + DR
Sbjct: 780 ELEKERFSKKRIEDDLEVMSRKASSLRAKARESAVLE-KLRHEVKEYRGILKCGICHDRQ 838
Query: 70 KEV 72
KEV
Sbjct: 839 KEV 841
>UniRef100_UPI000049883E UPI000049883E UniRef100 entry
Length = 146
Score = 36.6 bits (83), Expect = 0.16
Identities = 22/74 (29%), Positives = 44/74 (58%), Gaps = 8/74 (10%)
Query: 8 EKDLDSKRSS--QKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEI-----RVCKNMI 60
+K++++K ++K +EE++ + NQI EL + E ++++EEE+ I R K ++
Sbjct: 21 QKEIENKEEKIEEEKEKEEILALQNQITELKEKLKELSMKRMEEEQAISEEMMRKAKEIV 80
Query: 61 KCTVFSDRPKEVKT 74
+ F + KE+KT
Sbjct: 81 R-KEFEEEIKEMKT 93
>UniRef100_Q6FWE0 Candida glabrata strain CBS138 chromosome D complete sequence
[Candida glabrata]
Length = 1980
Score = 36.2 bits (82), Expect = 0.21
Identities = 17/60 (28%), Positives = 34/60 (56%)
Query: 2 SVASASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIK 61
++ S +K+++ ++ LE EL + +I + N + E + E+E+EI+V KN +K
Sbjct: 1475 TIKSEYDKEINILEDKKEVLESELSDKKQEIIDYNQKIKEQETKATEKEKEIQVAKNALK 1534
>UniRef100_Q8IKY1 Hypothetical protein [Plasmodium falciparum]
Length = 1288
Score = 35.8 bits (81), Expect = 0.28
Identities = 19/59 (32%), Positives = 34/59 (57%), Gaps = 3/59 (5%)
Query: 8 EKDLDSKRSSQKKLEEELMNVNNQ---IAELNSETGETVVQQLEEEEEIRVCKNMIKCT 63
EK+ + +++ +KK +E+ N N+ I+ N +T + V+ Q EE+EE C + K T
Sbjct: 452 EKNDEEEKNDEKKKNDEMNNKQNRQENISINNIQTSDAVLVQKEEKEEFSECNQLFKNT 510
>UniRef100_UPI000049975E UPI000049975E UniRef100 entry
Length = 143
Score = 35.4 bits (80), Expect = 0.36
Identities = 14/45 (31%), Positives = 31/45 (68%)
Query: 9 KDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEI 53
K++++ + ++K +EE++ + NQI EL + E ++++EEE+ I
Sbjct: 48 KEIENDKEEEQKDKEEILALKNQITELKEKVKEFSMKRMEEEQAI 92
>UniRef100_Q6TGU8 Ribosome binding protein 1 homolog 180kDa [Brachydanio rerio]
Length = 978
Score = 35.4 bits (80), Expect = 0.36
Identities = 20/73 (27%), Positives = 42/73 (57%), Gaps = 1/73 (1%)
Query: 2 SVASASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIK 61
+V + ++++ RSS K+ EE+L ++ ++ +L E ETV + EE + RV + +
Sbjct: 539 TVQKINSEEVEQLRSSLKEREEQLTSMEAELTQLREEL-ETVKRAQAEETQNRVNEADTR 597
Query: 62 CTVFSDRPKEVKT 74
C ++ +++KT
Sbjct: 598 CREYTTEIQQLKT 610
>UniRef100_P40767 Hypothetical protein yvcE [Bacillus subtilis]
Length = 473
Score = 35.0 bits (79), Expect = 0.47
Identities = 18/62 (29%), Positives = 36/62 (58%)
Query: 2 SVASASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIK 61
S A EK+L + +Q K+E+EL ++N++ + +++ + + + +EEI+ K IK
Sbjct: 49 SSIEAKEKELTELQENQSKIEKELKDINDKALDTSNKIEDKKEENDKTKEEIKKLKKEIK 108
Query: 62 CT 63
T
Sbjct: 109 ET 110
>UniRef100_UPI00002BC7C3 UPI00002BC7C3 UniRef100 entry
Length = 372
Score = 34.7 bits (78), Expect = 0.61
Identities = 18/55 (32%), Positives = 31/55 (55%), Gaps = 2/55 (3%)
Query: 8 EKDLDSKRSSQKKLEEELMNVNNQIAE--LNSETGETVVQQLEEEEEIRVCKNMI 60
E ++ S+ +Q ++EE+ VNNQ E L ++ E +V+ +EEEI + I
Sbjct: 253 ENEITSETINQSGVQEEINQVNNQYHEELLKNQQAENIVEDTSKEEEISFAQEAI 307
>UniRef100_UPI000049A011 UPI000049A011 UniRef100 entry
Length = 157
Score = 34.3 bits (77), Expect = 0.80
Identities = 21/74 (28%), Positives = 44/74 (59%), Gaps = 8/74 (10%)
Query: 8 EKDLDSKRSS--QKKLEEELMNVNNQIAELNSETGETVVQQLEE-----EEEIRVCKNMI 60
++++++K ++K +EE++ + NQI EL + E ++++EE EE +R K ++
Sbjct: 21 QEEIENKEEKIEEEKDKEEIIALKNQITELKEKVKEFSMEKMEEDQAISEEMMRKAKEIV 80
Query: 61 KCTVFSDRPKEVKT 74
+ F + KE+KT
Sbjct: 81 R-KEFEEEIKEMKT 93
>UniRef100_UPI000028F98B UPI000028F98B UniRef100 entry
Length = 305
Score = 34.3 bits (77), Expect = 0.80
Identities = 16/54 (29%), Positives = 33/54 (60%), Gaps = 2/54 (3%)
Query: 6 ASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNM 59
A++ ++D+K++S LE++L NN+I LN + ++ EE+ ++V K +
Sbjct: 168 ANQNNIDNKKTSLNSLEQQLAKSNNEIQALNDNKKDLSIK--IEEQALKVAKQL 219
>UniRef100_Q8IIG7 Hypothetical protein [Plasmodium falciparum]
Length = 964
Score = 34.3 bits (77), Expect = 0.80
Identities = 20/73 (27%), Positives = 38/73 (51%), Gaps = 2/73 (2%)
Query: 3 VASASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEE--EEIRVCKNMI 60
+ E+ + + K+++EE+ V +I E E E + ++++EE EEI+ K +
Sbjct: 610 IKEVKEEIKEEVKEEIKEVKEEIKEVKEEIKEEIKEVKEEIKEEVKEEIKEEIKEIKEEL 669
Query: 61 KCTVFSDRPKEVK 73
K + S+ KE K
Sbjct: 670 KNDISSETTKEEK 682
>UniRef100_UPI0000499FE0 UPI0000499FE0 UniRef100 entry
Length = 322
Score = 33.9 bits (76), Expect = 1.0
Identities = 20/66 (30%), Positives = 38/66 (57%), Gaps = 6/66 (9%)
Query: 14 KRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEI-----RVCKNMIKCTVFSDR 68
++ ++K +EE++ + NQI EL + E ++++EEE+ I R K ++K F +
Sbjct: 29 EKIEEEKEKEEILALKNQITELKEKVKEFSMEKMEEEQAISEEMMRKAKEIVK-KEFEEE 87
Query: 69 PKEVKT 74
E+KT
Sbjct: 88 ITEMKT 93
>UniRef100_UPI000049875E UPI000049875E UniRef100 entry
Length = 515
Score = 33.9 bits (76), Expect = 1.0
Identities = 20/66 (30%), Positives = 38/66 (57%), Gaps = 6/66 (9%)
Query: 14 KRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEI-----RVCKNMIKCTVFSDR 68
++ ++K +EE++ + NQI EL + E ++++EEE+ I R K ++K F +
Sbjct: 222 EKIEEEKDKEEILALKNQITELQEKVKELSMEKMEEEQAISEEMMRKAKEIVK-KEFEEE 280
Query: 69 PKEVKT 74
E+KT
Sbjct: 281 ITEMKT 286
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.304 0.122 0.303
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 114,051,666
Number of Sequences: 2790947
Number of extensions: 3838911
Number of successful extensions: 26760
Number of sequences better than 10.0: 230
Number of HSP's better than 10.0 without gapping: 152
Number of HSP's successfully gapped in prelim test: 80
Number of HSP's that attempted gapping in prelim test: 26414
Number of HSP's gapped (non-prelim): 463
length of query: 80
length of database: 848,049,833
effective HSP length: 56
effective length of query: 24
effective length of database: 691,756,801
effective search space: 16602163224
effective search space used: 16602163224
T: 11
A: 40
X1: 16 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.9 bits)
S2: 68 (30.8 bits)
Medicago: description of AC147712.6


