

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147538.3 - phase: 0
(123 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
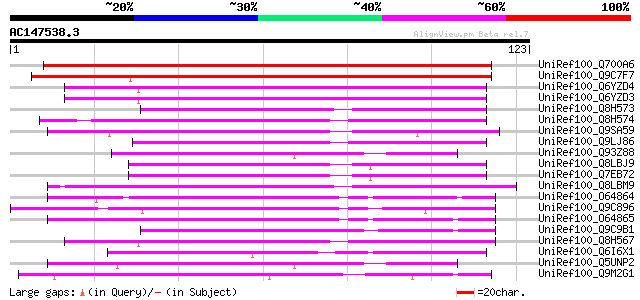
Sequences producing significant alignments: (bits) Value
UniRef100_Q700A6 Putative lipid transfer protein GPI-anchored [C... 188 2e-47
UniRef100_Q9C7F7 Uncharacterized GPI-anchored protein At1g27950 ... 112 3e-24
UniRef100_Q6YZD4 Lipid transfer protein-like [Oryza sativa] 87 9e-17
UniRef100_Q6YZD3 Lipid transfer protein-like [Oryza sativa] 87 9e-17
UniRef100_Q8H573 Protease inhibitor-like protein [Oryza sativa] 68 4e-11
UniRef100_Q8H574 Protease inhibitor-like protein [Oryza sativa] 58 6e-08
UniRef100_Q9SA59 F10O3.7 [Arabidopsis thaliana] 58 6e-08
UniRef100_Q9LJ86 Gb|AAD48513.1 [Arabidopsis thaliana] 56 2e-07
UniRef100_Q93Z88 Nonspecific lipid transfer protein [Bromus iner... 56 2e-07
UniRef100_Q8LBJ9 Hypothetical protein [Arabidopsis thaliana] 56 2e-07
UniRef100_Q7EB72 Hypothetical protein At2g48130 [Arabidopsis tha... 56 2e-07
UniRef100_Q8LBM9 Hypothetical protein [Arabidopsis thaliana] 55 3e-07
UniRef100_O64864 Hypothetical protein At2g44290 [Arabidopsis tha... 55 3e-07
UniRef100_Q9C896 Hypothetical protein F7A10.16 [Arabidopsis thal... 55 3e-07
UniRef100_O64865 Hypothetical protein At2g44300 [Arabidopsis tha... 55 4e-07
UniRef100_Q9C9B1 Hypothetical protein F2P9.24 [Arabidopsis thali... 55 5e-07
UniRef100_Q8H567 Protease inhibitor-like protein [Oryza sativa] 54 6e-07
UniRef100_Q6I6X1 Xylogen protein 1 [Zinnia elegans] 54 1e-06
UniRef100_Q5UNP2 Non-specific lipid transfer protein 6 [Hordeum ... 54 1e-06
UniRef100_Q9M2G1 Hypothetical protein F14P22.140 [Arabidopsis th... 54 1e-06
>UniRef100_Q700A6 Putative lipid transfer protein GPI-anchored [Cicer arietinum]
Length = 185
Score = 188 bits (478), Expect = 2e-47
Identities = 88/106 (83%), Positives = 94/106 (88%)
Query: 9 FMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATD 68
F+C+ L +IIGG GAEDLA KCG VVQKVIPCLDFATGKA TPKKECCDAANSIK TD
Sbjct: 5 FVCVLGLIMIIGGSEGAEDLAQKCGQVVQKVIPCLDFATGKALTPKKECCDAANSIKETD 64
Query: 69 PECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCP 114
PECLCYIIQQTHKGSPESKS+GIQEDKLLQLPTVC V AN++DCP
Sbjct: 65 PECLCYIIQQTHKGSPESKSLGIQEDKLLQLPTVCKVKNANLTDCP 110
>UniRef100_Q9C7F7 Uncharacterized GPI-anchored protein At1g27950 precursor
[Arabidopsis thaliana]
Length = 193
Score = 112 bits (279), Expect = 3e-24
Identities = 52/111 (46%), Positives = 70/111 (62%), Gaps = 2/111 (1%)
Query: 6 HQMFMCLCVLALIIGGCNGAED--LASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANS 63
H + + + ++A I A LA +C QKV CLDFATGKA P K+CCDA
Sbjct: 7 HLVLVTMTIVASIAAAAPAAPGGALADECNQDFQKVTLCLDFATGKATIPSKKCCDAVED 66
Query: 64 IKATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCP 114
IK DP+CLC++IQQ G K +G+QEDKL+QLPT C ++ A+I++CP
Sbjct: 67 IKERDPKCLCFVIQQAKTGGQALKDLGVQEDKLIQLPTSCQLHNASITNCP 117
>UniRef100_Q6YZD4 Lipid transfer protein-like [Oryza sativa]
Length = 179
Score = 87.0 bits (214), Expect = 9e-17
Identities = 42/102 (41%), Positives = 58/102 (56%), Gaps = 2/102 (1%)
Query: 14 VLALIIGGCNGAEDLA--SKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPEC 71
V+A + A D A SKC K+ C+D+ATG P CC ++ + PEC
Sbjct: 12 VVAWCVAAAAAAPDAALQSKCQQDFTKLTDCMDYATGHEEAPSSTCCGDMSATQQARPEC 71
Query: 72 LCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDC 113
LCYIIQQ H G E +S+G++ D+LL +PT C + AN+S C
Sbjct: 72 LCYIIQQVHGGRNEVQSLGLRFDRLLAMPTACKLPNANVSLC 113
>UniRef100_Q6YZD3 Lipid transfer protein-like [Oryza sativa]
Length = 178
Score = 87.0 bits (214), Expect = 9e-17
Identities = 42/102 (41%), Positives = 58/102 (56%), Gaps = 2/102 (1%)
Query: 14 VLALIIGGCNGAEDLA--SKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPEC 71
V+A + A D A SKC K+ C+D+ATG P CC ++ + PEC
Sbjct: 12 VVAWCVAAAAAAPDAALQSKCQQDFTKLTDCMDYATGHEEAPSSTCCGDMSATQQARPEC 71
Query: 72 LCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDC 113
LCYIIQQ H G E +S+G++ D+LL +PT C + AN+S C
Sbjct: 72 LCYIIQQVHGGRNEVQSLGLRFDRLLAMPTACKLPNANVSLC 113
>UniRef100_Q8H573 Protease inhibitor-like protein [Oryza sativa]
Length = 181
Score = 68.2 bits (165), Expect = 4e-11
Identities = 31/82 (37%), Positives = 44/82 (52%), Gaps = 4/82 (4%)
Query: 32 CGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSMGI 91
C SV+ + PCLD+ +GK+P P+ CC + +DP CLC ++ GS S + I
Sbjct: 37 CSSVMMTLSPCLDYISGKSPIPEFTCCTTLAGVVQSDPRCLCMVLD----GSAASFGISI 92
Query: 92 QEDKLLQLPTVCHVNGANISDC 113
+ L+LP VC V IS C
Sbjct: 93 NHTRALELPGVCKVQAPPISQC 114
>UniRef100_Q8H574 Protease inhibitor-like protein [Oryza sativa]
Length = 171
Score = 57.8 bits (138), Expect = 6e-08
Identities = 33/106 (31%), Positives = 48/106 (45%), Gaps = 7/106 (6%)
Query: 8 MFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKAT 67
MF + V+A G S C S + + PCLD+ G A P CC A +S+ +
Sbjct: 10 MFAAVVVVA---GALVAGAAAQSGCTSEMVSLAPCLDYMQGNASRPTASCCAALSSVVKS 66
Query: 68 DPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDC 113
PECLC ++ G S + + + L+LP C V S+C
Sbjct: 67 RPECLCAVL----GGGASSLGVTVNTTRALELPAACGVKTPPPSEC 108
>UniRef100_Q9SA59 F10O3.7 [Arabidopsis thaliana]
Length = 129
Score = 57.8 bits (138), Expect = 6e-08
Identities = 35/111 (31%), Positives = 55/111 (49%), Gaps = 9/111 (8%)
Query: 10 MCLCVLALIIGGC--NGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKAT 67
M L +LAL+I GA + + C + + PCL + G + +P CC +++ +
Sbjct: 1 MILAILALVIATFLYGGATTVQAGCRDTLTSLSPCLYYLNGGSSSPSWSCCRQFSTVVQS 60
Query: 68 DPECLCYIIQQTHKGSPESKSMGIQEDK--LLQLPTVCHVNGANISDCPSK 116
PECLC ++ S ES G + ++ L LPT C+V + S C SK
Sbjct: 61 SPECLCSVV-----NSNESSFYGFKFNRTLALNLPTACNVQTPSPSLCNSK 106
>UniRef100_Q9LJ86 Gb|AAD48513.1 [Arabidopsis thaliana]
Length = 170
Score = 56.2 bits (134), Expect = 2e-07
Identities = 22/84 (26%), Positives = 44/84 (52%), Gaps = 4/84 (4%)
Query: 30 SKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
S C + + + PCL++ TG + +P ++CC+ + + + P+CLC ++ G +
Sbjct: 26 SSCTNALISMSPCLNYITGNSTSPNQQCCNQLSRVVQSSPDCLCQVL----NGGGSQLGI 81
Query: 90 GIQEDKLLQLPTVCHVNGANISDC 113
+ + + L LP C+V +S C
Sbjct: 82 NVNQTQALGLPRACNVQTPPVSRC 105
>UniRef100_Q93Z88 Nonspecific lipid transfer protein [Bromus inermis]
Length = 124
Score = 55.8 bits (133), Expect = 2e-07
Identities = 30/87 (34%), Positives = 42/87 (47%), Gaps = 10/87 (11%)
Query: 25 AEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKA-----TDPECLCYIIQQT 79
AE S CG VV + PC+ +ATG+AP P CC S+ A D + C ++Q
Sbjct: 28 AEGAVSNCGQVVSYLAPCITYATGRAPGPNGGCCSGVRSLNAAASTTADRQATCSCLKQQ 87
Query: 80 HKGSPESKSMGIQEDKLLQLPTVCHVN 106
G GI+ D + +P+ C VN
Sbjct: 88 TSGMG-----GIKPDLVAGIPSKCGVN 109
>UniRef100_Q8LBJ9 Hypothetical protein [Arabidopsis thaliana]
Length = 183
Score = 55.8 bits (133), Expect = 2e-07
Identities = 25/86 (29%), Positives = 44/86 (51%), Gaps = 6/86 (6%)
Query: 29 ASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSP-ESK 87
+S C S + + PCL + TG + TP + CC +S+ + P+C+C + SP +
Sbjct: 27 SSSCVSTLTTLSPCLSYITGNSTTPSQPCCSRLDSVIKSSPQCICSAV-----NSPIPNI 81
Query: 88 SMGIQEDKLLQLPTVCHVNGANISDC 113
+ I + LQLP C++ ++ C
Sbjct: 82 GLNINRTQALQLPNACNIQTPPLTQC 107
>UniRef100_Q7EB72 Hypothetical protein At2g48130 [Arabidopsis thaliana]
Length = 183
Score = 55.8 bits (133), Expect = 2e-07
Identities = 25/86 (29%), Positives = 44/86 (51%), Gaps = 6/86 (6%)
Query: 29 ASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSP-ESK 87
+S C S + + PCL + TG + TP + CC +S+ + P+C+C + SP +
Sbjct: 27 SSSCVSTLTTLSPCLSYITGNSTTPSQPCCSRLDSVIKSSPQCICSAV-----NSPIPNI 81
Query: 88 SMGIQEDKLLQLPTVCHVNGANISDC 113
+ I + LQLP C++ ++ C
Sbjct: 82 GLNINRTQALQLPNACNIQTPPLTQC 107
>UniRef100_Q8LBM9 Hypothetical protein [Arabidopsis thaliana]
Length = 156
Score = 55.5 bits (132), Expect = 3e-07
Identities = 31/111 (27%), Positives = 47/111 (41%), Gaps = 5/111 (4%)
Query: 10 MCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDP 69
MCL +L + + S C +V+ + PCL F T P ++CC+ +
Sbjct: 5 MCL-ILFIALMRVMSIVSAQSSCTNVLISMAPCLSFITQNTSLPSQQCCNQLAHVVRYSS 63
Query: 70 ECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCPSKSKTD 120
ECLC ++ G + + E + L LP CHV S C S S +
Sbjct: 64 ECLCQVLD----GGGSQLGINVNETQALALPKACHVQTPPASRCHSGSSVN 110
>UniRef100_O64864 Hypothetical protein At2g44290 [Arabidopsis thaliana]
Length = 205
Score = 55.5 bits (132), Expect = 3e-07
Identities = 31/108 (28%), Positives = 58/108 (53%), Gaps = 7/108 (6%)
Query: 10 MCLCVLALII--GGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKAT 67
+ L ++A+++ G + +D +C + + + CL + GKA +P +CC + +
Sbjct: 13 IALIMVAMVVDAAGADKGKD-KEECTAQLVGMATCLPYVQGKAKSPTPDCCSGLKQVINS 71
Query: 68 DPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCPS 115
D +CLC IIQ+ + P+ + + L LP+VCH A+I+ CP+
Sbjct: 72 DMKCLCMIIQE--RNDPD-LGLQVNVSLALALPSVCHAT-ADITKCPA 115
>UniRef100_Q9C896 Hypothetical protein F7A10.16 [Arabidopsis thaliana]
Length = 184
Score = 55.5 bits (132), Expect = 3e-07
Identities = 31/120 (25%), Positives = 61/120 (50%), Gaps = 12/120 (10%)
Query: 1 MKNQQHQMFMCLCVLALIIGGCNGAEDLAS---KCGSVVQKVIPCLDFATGKAPTPKKEC 57
M+ +F+ + + ++++G G DLA +C + + ++ C+ + G A P K+C
Sbjct: 1 MEKSTRTLFITIVITSMLLGF--GNSDLAQDREECTNQLIELSTCIPYVGGDAKAPTKDC 58
Query: 58 CDAANSIKATDPECLCYIIQQTHKGSPESKSMGIQEDKLL--QLPTVCHVNGANISDCPS 115
C + +C+C +++ K P+ +GI+ + L LP+ CH+ NI+DC S
Sbjct: 59 CAGFGQVIRKSEKCVCILVRD--KDDPQ---LGIKINATLAAHLPSACHITAPNITDCIS 113
>UniRef100_O64865 Hypothetical protein At2g44300 [Arabidopsis thaliana]
Length = 204
Score = 55.1 bits (131), Expect = 4e-07
Identities = 28/106 (26%), Positives = 53/106 (49%), Gaps = 4/106 (3%)
Query: 10 MCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDP 69
+ L V+A+++ + +C + + CL + G+A +P +CC + ++
Sbjct: 13 IALIVVAMVVAAADDKTKDKEECTEQLVGMATCLPYVQGQAKSPTPDCCSGLKQVLNSNK 72
Query: 70 ECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCPS 115
+CLC IIQ + P+ + I L LP+VCH A+++ CP+
Sbjct: 73 KCLCVIIQD--RNDPD-LGLQINVSLALALPSVCHA-AADVTKCPA 114
>UniRef100_Q9C9B1 Hypothetical protein F2P9.24 [Arabidopsis thaliana]
Length = 193
Score = 54.7 bits (130), Expect = 5e-07
Identities = 30/84 (35%), Positives = 42/84 (49%), Gaps = 4/84 (4%)
Query: 32 CGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSMGI 91
C S + + PC F G A P + CCD+ N I + + CLC + T SP + I
Sbjct: 30 CASRLLSLAPCGPFVQGFAQLPAQPCCDSLNQIYSQEATCLCLFLNNTSTLSP---AFPI 86
Query: 92 QEDKLLQLPTVCHVNGANISDCPS 115
+ LQLP +C++ AN S C S
Sbjct: 87 NQTLALQLPPLCNI-PANSSTCSS 109
>UniRef100_Q8H567 Protease inhibitor-like protein [Oryza sativa]
Length = 170
Score = 54.3 bits (129), Expect = 6e-07
Identities = 29/103 (28%), Positives = 44/103 (42%), Gaps = 5/103 (4%)
Query: 14 VLALIIGGCNGAEDLA-SKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECL 72
V+AL+ GG + S C + + PCL++ TG P CC + + PECL
Sbjct: 16 VVALMAGGAAAQPPSSTSGCTQTLLSMSPCLNYLTGNETAPSASCCGKLGEVVKSQPECL 75
Query: 73 CYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCPS 115
C + + + I + L LP C V +S+C S
Sbjct: 76 CVAL----NADTAALGLSINRTRALGLPDACKVQTPPVSNCKS 114
>UniRef100_Q6I6X1 Xylogen protein 1 [Zinnia elegans]
Length = 183
Score = 53.5 bits (127), Expect = 1e-06
Identities = 28/92 (30%), Positives = 45/92 (48%), Gaps = 7/92 (7%)
Query: 24 GAEDLASKCGSVVQKVIPCLDFATGKAPTPKKE--CCDAANSIKATDPECLCYIIQQTHK 81
GA A+ C +V+ + CL + T + K E CC ++ TD ECLC + K
Sbjct: 29 GAPAPAADCSTVILNMADCLSYVTAGSTVKKPEGTCCSGLKTVLKTDAECLC----EAFK 84
Query: 82 GSPESKSMGIQEDKLLQLPTVCHVNGANISDC 113
S + + + K L LP+ CH+N + ++C
Sbjct: 85 NSAQ-LGVSLNITKALALPSACHINAPSATNC 115
>UniRef100_Q5UNP2 Non-specific lipid transfer protein 6 [Hordeum vulgare var.
distichum]
Length = 124
Score = 53.5 bits (127), Expect = 1e-06
Identities = 32/103 (31%), Positives = 48/103 (46%), Gaps = 11/103 (10%)
Query: 10 MCLCVLALIIGGCNG-AEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKA-- 66
M L V+AL++ G AE + CG VV + PC+ +A G+ P CC S+ A
Sbjct: 12 MALLVVALVLSGAPAPAEGAVASCGQVVSYLAPCIGYAMGRERAPGGGCCTGVRSLNAAA 71
Query: 67 ---TDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
D + C ++Q G GI+ D + +P+ C VN
Sbjct: 72 ATPADRQATCTCLKQQTSGMG-----GIKPDLVAGIPSKCGVN 109
>UniRef100_Q9M2G1 Hypothetical protein F14P22.140 [Arabidopsis thaliana]
Length = 177
Score = 53.5 bits (127), Expect = 1e-06
Identities = 34/117 (29%), Positives = 51/117 (43%), Gaps = 11/117 (9%)
Query: 3 NQQHQMF-MCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDA- 60
NQ QM +C+ V + +G + C + + CL F T KA P CC
Sbjct: 8 NQNPQMLALCITVAVMFLGVRSELSQDIKGCQDAMSDLYSCLPFVTNKAKAPDSTCCSTL 67
Query: 61 -ANSIKATDPECLCYIIQQTHKGSPESKSMGIQED--KLLQLPTVCHVNGANISDCP 114
K +CLC +++ + +G + D + + LP+ CHV ANIS CP
Sbjct: 68 KVKIDKGQTRKCLCTLVKDR-----DDPGLGFKVDANRAMSLPSACHV-PANISQCP 118
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.134 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 202,289,421
Number of Sequences: 2790947
Number of extensions: 7392036
Number of successful extensions: 14183
Number of sequences better than 10.0: 231
Number of HSP's better than 10.0 without gapping: 87
Number of HSP's successfully gapped in prelim test: 144
Number of HSP's that attempted gapping in prelim test: 14048
Number of HSP's gapped (non-prelim): 241
length of query: 123
length of database: 848,049,833
effective HSP length: 99
effective length of query: 24
effective length of database: 571,746,080
effective search space: 13721905920
effective search space used: 13721905920
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC147538.3


