

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147496.2 - phase: 0 /pseudo
(550 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
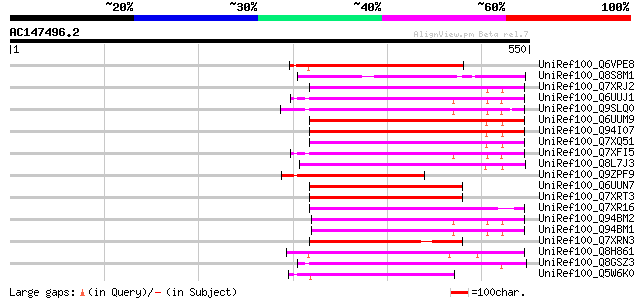
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6VPE8 Putative gag-pol polyprotein [Petunia hybrida] 178 3e-43
UniRef100_Q8S8M1 Putative retroelement integrase [Arabidopsis th... 178 4e-43
UniRef100_Q7XRJ2 OSJNBa0042D13.18 protein [Oryza sativa] 177 5e-43
UniRef100_Q6UUJ1 Putative gag-pol polyprotein [Oryza sativa] 177 7e-43
UniRef100_Q9SLQ0 Gag-pol polyprotein [Oryza sativa] 175 3e-42
UniRef100_Q6UUM9 Putative gag-pol polyprotein [Oryza sativa] 175 3e-42
UniRef100_Q94I07 Putative retroelement [Oryza sativa] 175 3e-42
UniRef100_Q7XQ51 OSJNBb0046P18.9 protein [Oryza sativa] 174 5e-42
UniRef100_Q7XFI5 Putative retroelement [Oryza sativa] 172 2e-41
UniRef100_Q8L7J3 Gag-pol polyprotein [Zea mays] 171 4e-41
UniRef100_Q9ZPF9 F5K24.1 protein [Arabidopsis thaliana] 171 7e-41
UniRef100_Q6UUN7 Putative gag-pol polyprotein [Oryza sativa] 168 3e-40
UniRef100_Q7XRT3 OSJNBa0042F21.10 protein [Oryza sativa] 167 7e-40
UniRef100_Q7XR16 OSJNBb0022F23.14 protein [Oryza sativa] 167 1e-39
UniRef100_Q94BM2 Gag-pol polyprotein [Hordeum vulgare] 155 3e-36
UniRef100_Q94BM1 Gag-pol polyprotein [Hordeum vulgare] 155 3e-36
UniRef100_Q7XRN3 OSJNBa0024J22.19 protein [Oryza sativa] 146 1e-33
UniRef100_Q8H861 Putative polyprotein [Oryza sativa] 139 2e-31
UniRef100_Q8GSZ3 P0466H10.33 protein [Oryza sativa] 138 4e-31
UniRef100_Q5W6K0 Retrotransposon protein, putative, Ty3-gypsy su... 137 8e-31
>UniRef100_Q6VPE8 Putative gag-pol polyprotein [Petunia hybrida]
Length = 803
Score = 178 bits (452), Expect = 3e-43
Identities = 96/191 (50%), Positives = 123/191 (64%), Gaps = 7/191 (3%)
Query: 297 VENHEGYVEEEAEEIPSGD-----LFMIRRFLGNQAKEE-ESNQRETLFHTRCLVQGKVC 350
VE EG VEE+ E+ D ++RR LG E+ E+ QRE LFH RC V+G VC
Sbjct: 421 VEGDEG-VEEDGEDDTVEDEEPHTFLVVRRSLGAMIAEDGETLQRENLFHARCRVKGVVC 479
Query: 351 FLIIDGGSRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKYED 410
+IID GS TNV S L+++++L T HPKPY+LQWL+E E+ V KQ + FK+G Y D
Sbjct: 480 SMIIDSGSCTNVVSQSLITQLKLPTMKHPKPYRLQWLSECGELKVVKQAMIKFKMGSYVD 539
Query: 411 DVLCDVVPMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQKK 470
+ L DVVPM+A HLL GRP QFDR V+H+GR N Y+ H G+K L PL+P +V DD +
Sbjct: 540 EHLFDVVPMQACHLLFGRPWQFDRQVVHNGRVNNYTLQHEGRKFVLKPLTPKQVSDDYNR 599
Query: 471 RKEKYEKEKRK 481
KE E K K
Sbjct: 600 MKELREAAKAK 610
>UniRef100_Q8S8M1 Putative retroelement integrase [Arabidopsis thaliana]
Length = 1215
Score = 178 bits (451), Expect = 4e-43
Identities = 109/243 (44%), Positives = 145/243 (58%), Gaps = 19/243 (7%)
Query: 306 EEAEEIPS-GDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVAS 364
+E EE+P+ G+L + RR L Q K +E QR+ LFHTRC V GKVC LIIDGGS TNVAS
Sbjct: 200 KENEELPAQGELLVARRTLSVQTKTDEQEQRKNLFHTRCHVHGKVCSLIIDGGSCTNVAS 259
Query: 365 TRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHL 424
+V K+ L+ WLN++ ++ V QV V IGKYED++LCDV+PMEA H+
Sbjct: 260 ETMVKKLGLK-----------WLNDSGKMRVKNQVVVPIVIGKYEDEILCDVLPMEAGHI 308
Query: 425 LLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQKKRKEKYEKEKRKIKR 484
LLGRP Q DR V+HDG TN++SF G K L ++P EV DQ K+K K ++ +
Sbjct: 309 LLGRPWQSDRKVMHDGFTNRHSFEFKGGKTILVSMTPHEVYQDQIHLKQK----KEQVVK 364
Query: 485 KEREFA*KKKKRRVSRNSHEQTAPIFTTLQKVALLTN-TQNFPSCTKFLLQECEDVFPKK 543
+ FA K S S +Q +F + + LTN PS LLQ+ +DVFP+
Sbjct: 365 QPNFFA--KSGEVKSAYSSKQPMLLFVFKEALTSLTNFAPVLPSEMTSLLQDYKDVFPED 422
Query: 544 VPQ 546
P+
Sbjct: 423 NPK 425
>UniRef100_Q7XRJ2 OSJNBa0042D13.18 protein [Oryza sativa]
Length = 2241
Score = 177 bits (450), Expect = 5e-43
Identities = 99/239 (41%), Positives = 143/239 (59%), Gaps = 11/239 (4%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+V+ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 553 IVQRVLSAQMEKAEQNQRHTLFQTKCVVKERCCRMIIDGGSCNNLASSEMVEKLALSTKP 612
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP PY +QWLN + + V K V + F IG Y D V CDVVPM+A ++LLGRP QFDR +
Sbjct: 613 HPHPYCIQWLNNSGKAKVTKLVHINFAIGNYHDVVECDVVPMQACNILLGRPWQFDRDSM 672
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEV-RDD-QKKRKEKYEKEKRKIKRKEREFA*KKKK 495
H GR+N+YSF++ +KI L P+SP ++ RDD K K K E +K+ ++ K
Sbjct: 673 HHGRSNQYSFLYHDKKIVLHPMSPEDILRDDVAKAAKSKCESDKKAQSDGKKPETINLKP 732
Query: 496 RRVSRNSHE------QTAPIFTTLQKVALLT---NTQNFPSCTKFLLQECEDVFPKKVP 545
R + + + + + K AL++ + P +LQE DVFPK+VP
Sbjct: 733 RYLLATKSDINELIASPSVAYALVCKDALISLHDMQHSLPPAVANILQEYSDVFPKEVP 791
>UniRef100_Q6UUJ1 Putative gag-pol polyprotein [Oryza sativa]
Length = 1616
Score = 177 bits (449), Expect = 7e-43
Identities = 103/276 (37%), Positives = 150/276 (54%), Gaps = 33/276 (11%)
Query: 298 ENHEGYVEEEAEEIPSGDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGG 357
E H G E AE S +++R L Q + E NQR TLF T+C+++ + C +IIDGG
Sbjct: 455 EEHIG--AEAAEHYES---LVVQRVLSAQMERAEQNQRHTLFQTKCVIKERSCRVIIDGG 509
Query: 358 SRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVV 417
S N+AS +V K+ L T+PHP+PY +QWLN + ++ V + V V F IG Y D + CDVV
Sbjct: 510 SCNNLASAEMVEKLALSTQPHPQPYYIQWLNSSGKVKVTRLVRVHFAIGSYHDSINCDVV 569
Query: 418 PMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQ-----KKRK 472
PM+A +LLGRP QFD+ LH G++N+YSF+H+G+K+ L P+SP + D+ K++
Sbjct: 570 PMQACSMLLGRPWQFDKDSLHFGKSNQYSFVHNGKKLVLHPMSPEVILKDELARASKQKN 629
Query: 473 EKYEKEKRKIKRKEREFA*KKKKRRVSRNSHE--------------------QTAPIFTT 512
+++ + + I E E KK V N +E T +
Sbjct: 630 QEHTRSEHLIAANELEKHKKKPTNSVQNNKNEIKLKGSCFIATKSDLDEVDTDTVVCYAL 689
Query: 513 LQKVALL---TNTQNFPSCTKFLLQECEDVFPKKVP 545
+ K L + P LLQE D+FPK+VP
Sbjct: 690 VCKETLFPIEDTPISLPPPVTNLLQEYADIFPKEVP 725
>UniRef100_Q9SLQ0 Gag-pol polyprotein [Oryza sativa]
Length = 1587
Score = 175 bits (444), Expect = 3e-42
Identities = 104/287 (36%), Positives = 155/287 (53%), Gaps = 33/287 (11%)
Query: 288 DDLGE*NVEVENHEGYVEEEAEEIPSGDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQG 347
++ GE + ++ E E AE S +++R L Q + E NQ TLF T+C+++
Sbjct: 443 NNAGEGDAPHQDEEHIRAEAAEHYES---LVVQRVLSAQMERAEQNQLHTLFQTKCVIKE 499
Query: 348 KVCFLIIDGGSRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGK 407
+ C +IIDGGS N+AS +V K+ L T+PHP+PY +QWLN + ++ V + V V F IG
Sbjct: 500 RSCHVIIDGGSCNNLASAEMVEKLALSTQPHPQPYYIQWLNSSGKVKVTRLVRVHFAIGS 559
Query: 408 YEDDVLCDVVPMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDD 467
Y D + CDVVPM+A +LLGRP QFD+ LH G++N+YSF+H+G+K+ L P+SP + D
Sbjct: 560 YHDSINCDVVPMQACSMLLGRPWQFDKDSLHFGKSNQYSFVHNGKKLVLHPMSPEVILKD 619
Query: 468 Q-----KKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHE------------------ 504
+ K++ +++ + + I E E KK V N +E
Sbjct: 620 ELARASKQKNQEHTRSEHLIAANELEKHKKKPTNSVQNNKNEIKLKGSCFIATKSDLDEV 679
Query: 505 --QTAPIFTTLQKVALL----TNTQNFPSCTKFLLQECEDVFPKKVP 545
T + + K L T P T LLQE D+FPK+VP
Sbjct: 680 DTDTVFCYALVCKETLFPIEDTPISLRPPVTN-LLQEYADIFPKEVP 725
>UniRef100_Q6UUM9 Putative gag-pol polyprotein [Oryza sativa]
Length = 1619
Score = 175 bits (444), Expect = 3e-42
Identities = 101/239 (42%), Positives = 145/239 (60%), Gaps = 11/239 (4%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+V+ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 438 IVQRVLSAQMEKAEQNQRHTLFQTKCVVKERCCRMIIDGGSCNNLASSEMVEKLALSTKP 497
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP Y +QWLN + + V K V + F IG Y D V CDVVPM+A ++LLGRP QFDR +
Sbjct: 498 HPHSYYIQWLNNSGKAKVTKLVHINFAIGNYHDVVECDVVPMQACNILLGRPWQFDRDSM 557
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEV-RDD-QKKRKEKYEKEKR-KIKRKEREFA*KKK 494
H GR+N+YSF++ +KI L P+SP ++ RDD K K K E +K+ + K+ E K
Sbjct: 558 HHGRSNQYSFLYHDKKIVLHPMSPEDILRDDVAKAAKSKCESDKKAQSDGKKPETINLKP 617
Query: 495 KRRVSRNSH-----EQTAPIFTTLQKVALLT---NTQNFPSCTKFLLQECEDVFPKKVP 545
K ++ S + + + K AL++ + P +LQE DVFPK+VP
Sbjct: 618 KCLLATKSDINELIASPSVAYALVCKDALISLHDMQHSLPPAVANILQEYSDVFPKEVP 676
>UniRef100_Q94I07 Putative retroelement [Oryza sativa]
Length = 2447
Score = 175 bits (444), Expect = 3e-42
Identities = 101/239 (42%), Positives = 145/239 (60%), Gaps = 11/239 (4%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+V+ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 459 IVQRVLSAQMEKAEQNQRHTLFQTKCVVKERCCRMIIDGGSCNNLASSEMVEKLALSTKP 518
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP PY +QWLN + + V K V + F IG Y D V CDVVPM+A ++LLGRP QFDR +
Sbjct: 519 HPHPYYIQWLNNSGKAKVTKLVHINFAIGNYHDVVECDVVPMQACNILLGRPWQFDRDSM 578
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEV-RDD-QKKRKEKYEKEKR-KIKRKEREFA*KKK 494
H GR+N+YSF++ +KI L +SP ++ RDD K K K E +K+ + K+ E K
Sbjct: 579 HHGRSNQYSFLYHDKKIVLHSMSPEDILRDDVAKAAKSKCESDKKAQSDGKKPETINLKP 638
Query: 495 KRRVSRNSH-----EQTAPIFTTLQKVALLT---NTQNFPSCTKFLLQECEDVFPKKVP 545
K ++ S + + + K AL++ + P +LQE DVFPK+VP
Sbjct: 639 KCLLATKSDINELIASPSVAYALVCKDALISLHDMQHSLPPAVANILQEYSDVFPKEVP 697
>UniRef100_Q7XQ51 OSJNBb0046P18.9 protein [Oryza sativa]
Length = 1134
Score = 174 bits (442), Expect = 5e-42
Identities = 99/239 (41%), Positives = 143/239 (59%), Gaps = 11/239 (4%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+V+ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 642 IVQRVLSTQMEKAEQNQRHTLFQTKCVVKERCCRMIIDGGSCNNLASSEMVEKLALSTKP 701
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP PY +QWLN + + V K V + F IG Y D V CDVVPM+A ++LLGRP QFD+ L
Sbjct: 702 HPHPYYIQWLNNSGKAKVTKLVHINFAIGNYHDVVECDVVPMQACNILLGRPWQFDKDSL 761
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEVRDDQ--KKRKEKYEKEKR-KIKRKEREFA*KKK 494
H GR+N+YSF++ +KI L P+S ++ D K K K E +K+ + K+ E K
Sbjct: 762 HHGRSNQYSFLYHDKKIVLHPMSSEDILHDDVAKAAKSKCESDKKAQSDGKKPETINLKP 821
Query: 495 KRRVSRNSH-----EQTAPIFTTLQKVALLT---NTQNFPSCTKFLLQECEDVFPKKVP 545
K ++ S + + + K AL++ + P +LQE DVFPK+VP
Sbjct: 822 KCLLATKSDINELIASPSVAYALVCKDALISLHDMQHSLPPAIANILQEYSDVFPKEVP 880
>UniRef100_Q7XFI5 Putative retroelement [Oryza sativa]
Length = 1708
Score = 172 bits (437), Expect = 2e-41
Identities = 101/276 (36%), Positives = 148/276 (53%), Gaps = 33/276 (11%)
Query: 298 ENHEGYVEEEAEEIPSGDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGG 357
E H G E AE S +++R L Q + E NQR TLF T+C+++ + C +IID G
Sbjct: 455 EEHIG--AEAAEHYES---LVVQRVLSAQMERAEQNQRHTLFQTKCVIKERSCRVIIDRG 509
Query: 358 SRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVV 417
S N+AS +V K+ L T+PHP+PY +QWLN + ++ V + V V F IG Y D + CDVV
Sbjct: 510 SCNNLASAEMVEKLALSTQPHPQPYYIQWLNSSGKVKVTRLVRVHFAIGSYHDSINCDVV 569
Query: 418 PMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQ-----KKRK 472
PM+A + LGRP QFD+ LH G++N+YSF+H+G+K+ L P+SP + D+ K++
Sbjct: 570 PMQACSIFLGRPWQFDKDSLHFGKSNQYSFVHNGKKLVLHPMSPEVILKDELARASKQKN 629
Query: 473 EKYEKEKRKIKRKEREFA*KKKKRRVSRNSHE--------------------QTAPIFTT 512
+++ + + I E E KK V N +E T +
Sbjct: 630 QEHTRSEHLIAANELEKHKKKPTNSVQNNKNEIKLKGSCFIATKSDLDEVDTDTVVCYAL 689
Query: 513 LQKVALL---TNTQNFPSCTKFLLQECEDVFPKKVP 545
+ K L + P LLQE D+FPK+VP
Sbjct: 690 VCKETLFPIEDTPISLPPPVTNLLQEYADIFPKEVP 725
>UniRef100_Q8L7J3 Gag-pol polyprotein [Zea mays]
Length = 1618
Score = 171 bits (434), Expect = 4e-41
Identities = 99/250 (39%), Positives = 145/250 (57%), Gaps = 11/250 (4%)
Query: 308 AEEIPSGDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRL 367
A++ + +++R L Q ++ E NQR TLF T+C+++ + C LIIDGGS N+AS+ +
Sbjct: 452 ADDAEHYESLIVQRVLSAQMEKAEQNQRHTLFQTKCVIKERSCRLIIDGGSCNNLASSDM 511
Query: 368 VSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLG 427
V K+ L TKPHP PY +QWLN + ++ V K V + F IG Y D V CDVVPM+A ++LLG
Sbjct: 512 VEKLALTTKPHPHPYHIQWLNNSGKVKVTKLVRINFAIGSYRDVVDCDVVPMDACNILLG 571
Query: 428 RP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSE-VRDD-QKKRKEKYEKEKR-KIKR 484
RP QFD +H GR+N+YS +H +KI L P+SP VRDD K K K E K K+
Sbjct: 572 RPWQFDSDCMHHGRSNQYSLIHHDKKIILLPMSPEAIVRDDVAKATKAKTENNKNIKVVG 631
Query: 485 KEREFA*KKKKRRVSRNS-----HEQTAPIFTTLQKVALLT---NTQNFPSCTKFLLQEC 536
++ K ++ + T + + K AL++ + P +LQE
Sbjct: 632 NNKDGIKLKGHCLLATKTDVNELFASTTVAYALVCKDALISIQDMQHSLPPVITNILQEY 691
Query: 537 EDVFPKKVPQ 546
DVFP ++P+
Sbjct: 692 SDVFPSEIPE 701
>UniRef100_Q9ZPF9 F5K24.1 protein [Arabidopsis thaliana]
Length = 1138
Score = 171 bits (432), Expect = 7e-41
Identities = 88/151 (58%), Positives = 105/151 (69%), Gaps = 3/151 (1%)
Query: 289 DLGE*NVEVENHEGYVEEEAEEIPSGDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGK 348
D GE E E E E + EE P G+L + R L K EE QRE LFHTRCL++GK
Sbjct: 303 DSGEVESEDEKPE---ESDVEEAPKGELLVTMRVLSVLNKAEEQAQRENLFHTRCLIKGK 359
Query: 349 VCFLIIDGGSRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKY 408
VC LIIDGGS TNVAS +V K+ LE PHPKPYKLQWLNE+ E+ V +QV+V IGKY
Sbjct: 360 VCSLIIDGGSCTNVASETMVQKLGLEEFPHPKPYKLQWLNESGEMAVTRQVQVPLAIGKY 419
Query: 409 EDDVLCDVVPMEASHLLLGRP*QFDRSVLHD 439
ED++LCD++P+EASH+LLGRP Q DR D
Sbjct: 420 EDEILCDILPLEASHVLLGRPWQSDRKDYQD 450
>UniRef100_Q6UUN7 Putative gag-pol polyprotein [Oryza sativa]
Length = 1234
Score = 168 bits (426), Expect = 3e-40
Identities = 83/165 (50%), Positives = 115/165 (69%), Gaps = 2/165 (1%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+V+ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 404 IVQRVLSAQMEKAEQNQRHTLFQTKCVVKERCCRMIIDGGSCKNLASSEMVEKLALSTKP 463
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP PY +QWLN + + V K V + F IG Y D V CDVVPM+A ++LLGRP QFDR +
Sbjct: 464 HPHPYYIQWLNNSGKAKVTKLVHINFAIGNYHDVVECDVVPMQACNILLGRPWQFDRDSM 523
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEV-RDD-QKKRKEKYEKEKR 480
H GR+N+YSF++ +KI L P+SP ++ RDD K K K E +K+
Sbjct: 524 HHGRSNQYSFLYHDKKIVLHPMSPEDILRDDVAKAAKSKCESDKK 568
>UniRef100_Q7XRT3 OSJNBa0042F21.10 protein [Oryza sativa]
Length = 2388
Score = 167 bits (423), Expect = 7e-40
Identities = 82/165 (49%), Positives = 114/165 (68%), Gaps = 2/165 (1%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+V+ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 350 IVQRVLSAQMEKAEQNQRHTLFQTKCVVKERCCRMIIDGGSCNNLASSEMVEKLALSTKP 409
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP PY +QWLN + + V K V + F IG Y D V CDVVPM+ ++LLGRP QFDR +
Sbjct: 410 HPHPYYIQWLNNSGKAKVTKLVHINFAIGNYHDVVECDVVPMQVCNILLGRPWQFDRDSM 469
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEV-RDD-QKKRKEKYEKEKR 480
H GR+N+YSF++ +KI L P+SP ++ RDD K K K E +K+
Sbjct: 470 HHGRSNQYSFLYHDKKIVLHPMSPEDILRDDVAKAAKSKCESDKK 514
>UniRef100_Q7XR16 OSJNBb0022F23.14 protein [Oryza sativa]
Length = 1431
Score = 167 bits (422), Expect = 1e-39
Identities = 94/230 (40%), Positives = 133/230 (56%), Gaps = 18/230 (7%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+V+ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 459 IVQRVLSAQMEKAEQNQRHTLFQTKCVVKERCCRMIIDGGSCNNLASSEMVEKLALSTKP 518
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP PY +QWLN + + V V + F IG Y D V CDV+PM+A ++LLGRP QFDR +
Sbjct: 519 HPHPYYIQWLNNSGKAKVTNLVHINFAIGNYHDVVECDVLPMQACNILLGRPWQFDRDSM 578
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEV-RDD-QKKRKEKYEKEKRKIKRKEREFA*KKKK 495
H GR+N+YSF++ +KI L P+SP ++ RDD K K K E +K+ ++ K
Sbjct: 579 HHGRSNQYSFLYHDKKIVLHPMSPEDILRDDVAKAAKSKCESDKKAQSDGKKPETINLKP 638
Query: 496 RRVSRNSHEQTAPIFTTLQKVALLTNTQNFPSCTKFLLQECEDVFPKKVP 545
R + + I + AL E DVFPK+VP
Sbjct: 639 RCLLATKSDINELIASPSVAYAL----------------EYSDVFPKEVP 672
>UniRef100_Q94BM2 Gag-pol polyprotein [Hordeum vulgare]
Length = 1720
Score = 155 bits (392), Expect = 3e-36
Identities = 94/252 (37%), Positives = 132/252 (52%), Gaps = 26/252 (10%)
Query: 320 RRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKPHP 379
+R L Q + E NQR LFHT+ +V+ + +IIDGGS N+AS +V K+ L T+PHP
Sbjct: 505 QRVLSVQVTQVEQNQRHNLFHTKGVVKERSVRVIIDGGSCNNLASMEMVEKLSLTTRPHP 564
Query: 380 KPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVLHD 439
PY +QW N + ++ V + V V F I Y D V CDVVPM+A +LLGRP QFD++ +H
Sbjct: 565 HPYYIQWFNNSGKVKVTRTVRVHFSIATYSDFVDCDVVPMQACSVLLGRPWQFDKNSVHH 624
Query: 440 GRTNKYSFMHSGQKISLAPLSPSEVRDDQ----KKRKEKYEKEKRKIKRKEREFA*KKKK 495
GRTN+Y+ +H Q I+L P+SP + D K K+ K + +I KE E K K
Sbjct: 625 GRTNQYTLVHKDQNITLLPMSPEHIMKDDIARASKAKQDLHKSENQIVAKEFEQHNKPNK 684
Query: 496 RRVSRNSHEQ-------------------TAPIFTTLQKVALLT---NTQNFPSCTKFLL 533
S S + T+ + + K L + + P +L
Sbjct: 685 SSSSVASEIKLKSGCLLATKSDITDLDITTSVCYAFVCKEVLFSFEDMPSSLPPAVINIL 744
Query: 534 QECEDVFPKKVP 545
QE DVFP+ VP
Sbjct: 745 QEFADVFPQDVP 756
>UniRef100_Q94BM1 Gag-pol polyprotein [Hordeum vulgare]
Length = 1717
Score = 155 bits (392), Expect = 3e-36
Identities = 94/252 (37%), Positives = 132/252 (52%), Gaps = 26/252 (10%)
Query: 320 RRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKPHP 379
+R L Q + E NQR LFHT+ +V+ + +IIDGGS N+AS +V K+ L T+PHP
Sbjct: 502 QRVLSVQVTQVEQNQRHNLFHTKGVVKERSVRVIIDGGSCNNLASMEMVEKLSLTTRPHP 561
Query: 380 KPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVLHD 439
PY +QW N + ++ V + V V F I Y D V CDVVPM+A +LLGRP QFD++ +H
Sbjct: 562 HPYYIQWFNNSGKVKVTRTVRVHFSIATYSDFVDCDVVPMQACSVLLGRPWQFDKNSVHH 621
Query: 440 GRTNKYSFMHSGQKISLAPLSPSEVRDDQ----KKRKEKYEKEKRKIKRKEREFA*KKKK 495
GRTN+Y+ +H Q I+L P+SP + D K K+ K + +I KE E K K
Sbjct: 622 GRTNQYTLVHKDQNITLLPMSPEHIMKDDIARASKAKQDLHKSENQIVAKEFEQHNKPNK 681
Query: 496 RRVSRNSHEQ-------------------TAPIFTTLQKVALLT---NTQNFPSCTKFLL 533
S S + T+ + + K L + + P +L
Sbjct: 682 SSSSVASEIKLKSGCLLATKSDITDLDITTSVCYAFVCKEVLFSFEDMPSSLPPAVINIL 741
Query: 534 QECEDVFPKKVP 545
QE DVFP+ VP
Sbjct: 742 QEFADVFPQDVP 753
>UniRef100_Q7XRN3 OSJNBa0024J22.19 protein [Oryza sativa]
Length = 1667
Score = 146 bits (369), Expect = 1e-33
Identities = 78/165 (47%), Positives = 105/165 (63%), Gaps = 13/165 (7%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++ L Q ++ E NQR TLF T+C+V+ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 418 IMQHVLSAQMEKAEQNQRHTLFQTKCVVKERCCRMIIDGGSCNNLASSEMVEKLALSTKP 477
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP PY +QWLN + + V K V + F IG Y D V CDVVPM+A ++LLGRP QFDR
Sbjct: 478 HPHPYYIQWLNNSGKAKVTKLVHINFAIGNYHDVVECDVVPMQACNILLGRPWQFDRDS- 536
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEV-RDD-QKKRKEKYEKEKR 480
MH G+KI L P+SP ++ RDD K K K E +K+
Sbjct: 537 ----------MHHGKKIVLHPMSPEDILRDDVAKAAKSKCESDKK 571
>UniRef100_Q8H861 Putative polyprotein [Oryza sativa]
Length = 928
Score = 139 bits (351), Expect = 2e-31
Identities = 90/270 (33%), Positives = 135/270 (49%), Gaps = 18/270 (6%)
Query: 294 NVEVENHEGYVEEEAEEIPSGD------LFMIRRFLGNQAKEEESNQRETLFHTRCLVQG 347
+VE E E VE+E E+ D +++R L ++ + QR LF ++
Sbjct: 441 DVEEEEREEEVEDEDGEVFGRDDTTDYRTIIVQRVLSTYVQQHDKLQRYNLFQIFFVINN 500
Query: 348 KVCFLIIDGGSRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGK 407
+IIDGGS N+ S+ LV K+ L T+ HP PY +QWLN++ V + V F IG
Sbjct: 501 HRARVIIDGGSCKNLLSSDLVKKLGLTTRTHPHPYHIQWLNDSGRAKVTQVCRVLFSIGS 560
Query: 408 YEDDVLCDVVPMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEV--- 464
Y D V CDVVPM+A LLLG P + D H GR+NKY+F+H+G+K L PL+P+E+
Sbjct: 561 YADSVDCDVVPMQACSLLLGCPWEHDNDATHHGRSNKYTFVHNGKKFILLPLTPAEIVEA 620
Query: 465 -RDDQKKRKEKYEKEKRKIKRKEREFA*KKK-------KRRVSRNSHEQTAPIFTTLQKV 516
++Q+ K + +K K+ + K K K ++ S Q +
Sbjct: 621 ESENQQVAKSVFPPKKDKLSPSSKVEGIKLKDGVMLATKCDIAEISDVDMCYFLICKQAL 680
Query: 517 ALLTN-TQNFPSCTKFLLQECEDVFPKKVP 545
+ + P LLQE E+VF ++P
Sbjct: 681 VSFDDIASSIPPVVTNLLQEYENVFLAEIP 710
>UniRef100_Q8GSZ3 P0466H10.33 protein [Oryza sativa]
Length = 767
Score = 138 bits (348), Expect = 4e-31
Identities = 85/266 (31%), Positives = 133/266 (49%), Gaps = 27/266 (10%)
Query: 306 EEAEEIPSGDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVAST 365
E ++ PS +++ L + K EE NQR LF ++ LV+G +IID GS N+AS
Sbjct: 433 ESLDQYPS---LIVKLALSAKEKCEEQNQRHNLFQSKVLVKGHTVKMIIDSGSCNNLASE 489
Query: 366 RLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLL 425
+V K+ L T PHP PY +QW + ++ V + V++ F IG Y D + CDVV M+A ++
Sbjct: 490 EMVKKLGLTTHPHPHPYHIQWFHTCGKMKVTRLVKIPFSIGSYHDQIECDVVHMQACSVI 549
Query: 426 LGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQKKRKEKYEKEKRKIKRK 485
LGRP Q+DR V+ DGRTN Y FM G+++ L ++P+ D R E +K K + +
Sbjct: 550 LGRPWQYDREVVFDGRTNTYKFMFDGKRVVLHSMTPAAALKDDLARLECEKKAKSENQII 609
Query: 486 EREFA*KKK-KRRVSRNSHEQTAPIFTTLQKVALL-----------------------TN 521
+ A K + ++N + + + + L +
Sbjct: 610 AKHPASTSKIEPSTTKNEIKLKGALLASKYNICALDEPDSCCYALFCKDPILTLATAPST 669
Query: 522 TQNFPSCTKFLLQECEDVFPKKVPQK 547
+ P LLQE EDVF ++P +
Sbjct: 670 LYSLPPAITNLLQEYEDVFHDEIPPR 695
>UniRef100_Q5W6K0 Retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza
sativa]
Length = 651
Score = 137 bits (345), Expect = 8e-31
Identities = 75/187 (40%), Positives = 109/187 (58%), Gaps = 13/187 (6%)
Query: 296 EVENHEGYVEEEAEEIPSGDLF-----------MIRRFLGNQAKEEESNQRETLFHTRCL 344
+VE EG EEE E G++F +++R L Q ++ + Q LF +
Sbjct: 378 DVEEEEG--EEEEVEDEDGEVFGRDDTTDYRTIIVQRVLSAQVQQHDRLQCHNLFEIFFV 435
Query: 345 VQGKVCFLIIDGGSRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFK 404
+ + +IIDGGS N+ S+ LV K+ L T+ HP PY +QWLN++ V + V F
Sbjct: 436 INNRRARVIIDGGSCNNLVSSDLVKKLGLTTRTHPHPYHIQWLNDSGRAKVTQVCRVSFS 495
Query: 405 IGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEV 464
IG Y D V CDVV M+A LLLGR + D H GR+NKY+F+H+G+KI+L PL+P+E+
Sbjct: 496 IGSYADSVDCDVVSMQACSLLLGRLWEHDNDATHHGRSNKYTFVHNGKKITLLPLTPAEI 555
Query: 465 RDDQKKR 471
+ K+R
Sbjct: 556 VEADKER 562
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.358 0.160 0.569
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 764,666,419
Number of Sequences: 2790947
Number of extensions: 29195199
Number of successful extensions: 295017
Number of sequences better than 10.0: 230
Number of HSP's better than 10.0 without gapping: 145
Number of HSP's successfully gapped in prelim test: 95
Number of HSP's that attempted gapping in prelim test: 289550
Number of HSP's gapped (non-prelim): 2400
length of query: 550
length of database: 848,049,833
effective HSP length: 132
effective length of query: 418
effective length of database: 479,644,829
effective search space: 200491538522
effective search space used: 200491538522
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 77 (34.3 bits)
Medicago: description of AC147496.2


