

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147484.9 - phase: 0
(125 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
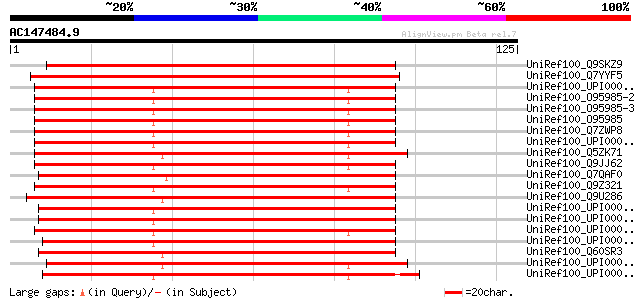
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SKZ9 Putative DNA topoisomerase III beta [Arabidopsi... 136 9e-32
UniRef100_Q7YYF5 DNA topoisomerase III beta-1, probable [Cryptos... 115 2e-25
UniRef100_UPI000036C837 UPI000036C837 UniRef100 entry 96 1e-19
UniRef100_O95985-2 Splice isoform 2 of O95985 [Homo sapiens] 96 1e-19
UniRef100_O95985-3 Splice isoform 3 of O95985 [Homo sapiens] 96 1e-19
UniRef100_O95985 DNA topoisomerase III beta-1 [Homo sapiens] 96 1e-19
UniRef100_Q7ZWP8 MGC53016 protein [Xenopus laevis] 94 5e-19
UniRef100_UPI00001D07C4 UPI00001D07C4 UniRef100 entry 94 7e-19
UniRef100_Q5ZK71 Hypothetical protein [Gallus gallus] 94 7e-19
UniRef100_Q9JJ62 Mus musculus brain cDNA, clone MNCb-2041, simil... 94 7e-19
UniRef100_Q7QAF0 ENSANGP00000018921 [Anopheles gambiae str. PEST] 94 7e-19
UniRef100_Q9Z321 DNA topoisomerase III beta-1 [Mus musculus] 94 7e-19
UniRef100_Q9U286 Hypothetical protein Y48C3A.14 [Caenorhabditis ... 93 2e-18
UniRef100_UPI000043447E UPI000043447E UniRef100 entry 92 3e-18
UniRef100_UPI000043447D UPI000043447D UniRef100 entry 92 3e-18
UniRef100_UPI000035EF98 UPI000035EF98 UniRef100 entry 91 5e-18
UniRef100_UPI000043447F UPI000043447F UniRef100 entry 91 8e-18
UniRef100_Q60SR3 Hypothetical protein CBG20786 [Caenorhabditis b... 91 8e-18
UniRef100_UPI00003AAF77 UPI00003AAF77 UniRef100 entry 89 2e-17
UniRef100_UPI000035EF99 UPI000035EF99 UniRef100 entry 89 3e-17
>UniRef100_Q9SKZ9 Putative DNA topoisomerase III beta [Arabidopsis thaliana]
Length = 836
Score = 136 bits (343), Expect = 9e-32
Identities = 66/86 (76%), Positives = 74/86 (85%)
Query: 10 VAEKPSIALSIATVLSHGKMNTRRGSTEVHEFDGKFRKAPARFKVTSVIGHVFSVDFPGK 69
VAEKPSIALSIA+VLSHG+M+TRRGSTEVHEFDG FR A ++VTSVIGHVFSVDFP K
Sbjct: 40 VAEKPSIALSIASVLSHGQMSTRRGSTEVHEFDGMFRGFKAHYRVTSVIGHVFSVDFPEK 99
Query: 70 YQDWAATDPLDLFQAQVIKNESNPKL 95
YQ+WA DP DLF A +IK ESNPK+
Sbjct: 100 YQNWATIDPQDLFDAPIIKKESNPKV 125
>UniRef100_Q7YYF5 DNA topoisomerase III beta-1, probable [Cryptosporidium parvum]
Length = 835
Score = 115 bits (289), Expect = 2e-25
Identities = 54/91 (59%), Positives = 70/91 (76%)
Query: 6 KVLMVAEKPSIALSIATVLSHGKMNTRRGSTEVHEFDGKFRKAPARFKVTSVIGHVFSVD 65
KVLMVAEKPSI+ +I+ +LS+GK++TRRG T VHEF+G F +F+VTSV GHVF +D
Sbjct: 5 KVLMVAEKPSISDTISRILSNGKLDTRRGKTPVHEFNGTFMGMNVQFRVTSVAGHVFEID 64
Query: 66 FPGKYQDWAATDPLDLFQAQVIKNESNPKLV 96
FP Y +W TDP+ LF A +IKNES+ K+V
Sbjct: 65 FPQSYSNWEKTDPVSLFDAPIIKNESSSKMV 95
>UniRef100_UPI000036C837 UPI000036C837 UniRef100 entry
Length = 862
Score = 96.3 bits (238), Expect = 1e-19
Identities = 50/93 (53%), Positives = 63/93 (66%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G +++ +G + VHE+ G F P RFK+TSV GHV +
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGSLSSHKGLNGACSVHEYTGTFAGQPVRFKMTSVCGHVMT 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY W DP +LF QA K E+NPKL
Sbjct: 64 LDFLGKYNKWDKVDPAELFSQAPTEKKEANPKL 96
>UniRef100_O95985-2 Splice isoform 2 of O95985 [Homo sapiens]
Length = 730
Score = 96.3 bits (238), Expect = 1e-19
Identities = 50/93 (53%), Positives = 63/93 (66%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G +++ +G + VHE+ G F P RFK+TSV GHV +
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGSLSSHKGLNGACSVHEYTGTFAGQPVRFKMTSVCGHVMT 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY W DP +LF QA K E+NPKL
Sbjct: 64 LDFLGKYNKWDKVDPAELFSQAPTEKKEANPKL 96
>UniRef100_O95985-3 Splice isoform 3 of O95985 [Homo sapiens]
Length = 707
Score = 96.3 bits (238), Expect = 1e-19
Identities = 50/93 (53%), Positives = 63/93 (66%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G +++ +G + VHE+ G F P RFK+TSV GHV +
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGSLSSHKGLNGACSVHEYTGTFAGQPVRFKMTSVCGHVMT 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY W DP +LF QA K E+NPKL
Sbjct: 64 LDFLGKYNKWDKVDPAELFSQAPTEKKEANPKL 96
>UniRef100_O95985 DNA topoisomerase III beta-1 [Homo sapiens]
Length = 862
Score = 96.3 bits (238), Expect = 1e-19
Identities = 50/93 (53%), Positives = 63/93 (66%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G +++ +G + VHE+ G F P RFK+TSV GHV +
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGSLSSHKGLNGACSVHEYTGTFAGQPVRFKMTSVCGHVMT 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY W DP +LF QA K E+NPKL
Sbjct: 64 LDFLGKYNKWDKVDPAELFSQAPTEKKEANPKL 96
>UniRef100_Q7ZWP8 MGC53016 protein [Xenopus laevis]
Length = 858
Score = 94.4 bits (233), Expect = 5e-19
Identities = 50/93 (53%), Positives = 63/93 (66%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G ++R+G + VHE+ G F RFK+TSV GHV S
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGNASSRKGLNGACSVHEYTGSFMGQNVRFKMTSVCGHVMS 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY +W DP +LF +A K E+NPKL
Sbjct: 64 LDFIGKYNNWDKVDPAELFSKAPTEKKEANPKL 96
>UniRef100_UPI00001D07C4 UPI00001D07C4 UniRef100 entry
Length = 947
Score = 94.0 bits (232), Expect = 7e-19
Identities = 49/93 (52%), Positives = 62/93 (65%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G M++ +G + VH++ G F P FK+TSV GHV +
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGNMSSHKGLNGACSVHKYTGTFAGQPVHFKMTSVCGHVMT 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY W DP +LF QA K E+NPKL
Sbjct: 64 LDFLGKYNKWDKVDPAELFSQAPTEKKEANPKL 96
>UniRef100_Q5ZK71 Hypothetical protein [Gallus gallus]
Length = 862
Score = 94.0 bits (232), Expect = 7e-19
Identities = 51/96 (53%), Positives = 63/96 (65%), Gaps = 4/96 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRGST---EVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G M++R+G VHE+ G F A FK+TSV GHV +
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGNMSSRKGLNGVCSVHEYTGSFIGQSAHFKMTSVCGHVMT 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKLVDV 98
+DF GKY W DP +LF +A K E+NPKL V
Sbjct: 64 LDFIGKYNSWDKVDPAELFSKAPTEKKEANPKLTMV 99
>UniRef100_Q9JJ62 Mus musculus brain cDNA, clone MNCb-2041, similar to Mus musculus
topoisomerase (DNA) III beta-1 mRNA [Mus musculus]
Length = 464
Score = 94.0 bits (232), Expect = 7e-19
Identities = 49/93 (52%), Positives = 62/93 (65%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G M++ +G + VH++ G F P FK+TSV GHV +
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGNMSSHKGLNGACSVHKYTGTFAGQPVHFKMTSVCGHVMT 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY W DP +LF QA K E+NPKL
Sbjct: 64 LDFLGKYNKWDKVDPAELFSQAPTEKKEANPKL 96
>UniRef100_Q7QAF0 ENSANGP00000018921 [Anopheles gambiae str. PEST]
Length = 839
Score = 94.0 bits (232), Expect = 7e-19
Identities = 46/91 (50%), Positives = 60/91 (65%), Gaps = 3/91 (3%)
Query: 8 LMVAEKPSIALSIATVLSHGKMNTRRGSTE---VHEFDGKFRKAPARFKVTSVIGHVFSV 64
LMVAEKPS+A S+A++LS+GK + R+GS VHE+ G+FR RFK+TSV GH+ +
Sbjct: 5 LMVAEKPSLAASLASILSNGKCSVRKGSNSACSVHEWVGQFRGETTRFKMTSVCGHIMGL 64
Query: 65 DFPGKYQDWAATDPLDLFQAQVIKNESNPKL 95
+F GKY W DP +LF K ES P L
Sbjct: 65 EFVGKYNSWDRVDPAELFACPTEKKESTPNL 95
>UniRef100_Q9Z321 DNA topoisomerase III beta-1 [Mus musculus]
Length = 862
Score = 94.0 bits (232), Expect = 7e-19
Identities = 49/93 (52%), Positives = 62/93 (65%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G M++ +G + VH++ G F P FK+TSV GHV +
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGNMSSHKGLNGACSVHKYTGTFAGQPVHFKMTSVCGHVMT 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY W DP +LF QA K E+NPKL
Sbjct: 64 LDFLGKYNKWDKVDPAELFSQAPTEKKEANPKL 96
>UniRef100_Q9U286 Hypothetical protein Y48C3A.14 [Caenorhabditis elegans]
Length = 855
Score = 92.8 bits (229), Expect = 2e-18
Identities = 47/94 (50%), Positives = 62/94 (65%), Gaps = 3/94 (3%)
Query: 5 PKVLMVAEKPSIALSIATVLSHGKMNTRRGST---EVHEFDGKFRKAPARFKVTSVIGHV 61
P VLMVAEKP +A SIA +LS+G+ R+G V E+DG+F ARFKVTS GHV
Sbjct: 3 PTVLMVAEKPLLADSIANLLSNGQAKKRKGWNGVCSVSEYDGQFNGRAARFKVTSTCGHV 62
Query: 62 FSVDFPGKYQDWAATDPLDLFQAQVIKNESNPKL 95
S+DFP K+ +W DP +L+ A + E+NPK+
Sbjct: 63 MSLDFPPKFNNWERVDPAELYSAPTQRIEANPKM 96
>UniRef100_UPI000043447E UPI000043447E UniRef100 entry
Length = 664
Score = 92.0 bits (227), Expect = 3e-18
Identities = 45/91 (49%), Positives = 61/91 (66%), Gaps = 3/91 (3%)
Query: 8 LMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFSV 64
LMVAEKPS+A S+A +LS+G+ +R+G S VHE+ G+FR FK+TSV GHV ++
Sbjct: 3 LMVAEKPSLAASLANILSNGRCTSRKGLNGSCSVHEWIGQFRSETVNFKMTSVFGHVMTL 62
Query: 65 DFPGKYQDWAATDPLDLFQAQVIKNESNPKL 95
DF GKY +W DP+ LF K E+ P+L
Sbjct: 63 DFIGKYNNWDKVDPVKLFSCPTEKKEAVPRL 93
>UniRef100_UPI000043447D UPI000043447D UniRef100 entry
Length = 307
Score = 92.0 bits (227), Expect = 3e-18
Identities = 45/91 (49%), Positives = 61/91 (66%), Gaps = 3/91 (3%)
Query: 8 LMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFSV 64
LMVAEKPS+A S+A +LS+G+ +R+G S VHE+ G+FR FK+TSV GHV ++
Sbjct: 5 LMVAEKPSLAASLANILSNGRCTSRKGLNGSCSVHEWIGQFRSETVNFKMTSVFGHVMTL 64
Query: 65 DFPGKYQDWAATDPLDLFQAQVIKNESNPKL 95
DF GKY +W DP+ LF K E+ P+L
Sbjct: 65 DFIGKYNNWDKVDPVKLFSCPTEKKEAVPRL 95
>UniRef100_UPI000035EF98 UPI000035EF98 UniRef100 entry
Length = 703
Score = 91.3 bits (225), Expect = 5e-18
Identities = 49/93 (52%), Positives = 63/93 (67%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G ++R+G + V+E+ G F A FK+TSV GHV S
Sbjct: 4 VLMVAEKPSLAQSIAKILSKGNCSSRKGLNGACSVYEYTGSFLGQAAHFKMTSVCGHVMS 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY +W DP +LF +A K E+NPKL
Sbjct: 64 LDFIGKYNNWDKVDPAELFSKAPTEKKEANPKL 96
>UniRef100_UPI000043447F UPI000043447F UniRef100 entry
Length = 680
Score = 90.5 bits (223), Expect = 8e-18
Identities = 44/90 (48%), Positives = 60/90 (65%), Gaps = 3/90 (3%)
Query: 9 MVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFSVD 65
MVAEKPS+A S+A +LS+G+ +R+G S VHE+ G+FR FK+TSV GHV ++D
Sbjct: 1 MVAEKPSLAASLANILSNGRCTSRKGLNGSCSVHEWIGQFRSETVNFKMTSVFGHVMTLD 60
Query: 66 FPGKYQDWAATDPLDLFQAQVIKNESNPKL 95
F GKY +W DP+ LF K E+ P+L
Sbjct: 61 FIGKYNNWDKVDPVKLFSCPTEKKEAVPRL 90
>UniRef100_Q60SR3 Hypothetical protein CBG20786 [Caenorhabditis briggsae]
Length = 591
Score = 90.5 bits (223), Expect = 8e-18
Identities = 45/91 (49%), Positives = 60/91 (65%), Gaps = 3/91 (3%)
Query: 8 LMVAEKPSIALSIATVLSHGKMNTRRGST---EVHEFDGKFRKAPARFKVTSVIGHVFSV 64
LMVAEKP +A SIA +LS+G+ R+G V E+DG+F A+FKVTS GHV S+
Sbjct: 1 LMVAEKPMLADSIANLLSNGQARKRKGWNGVCSVSEYDGQFNGRNAKFKVTSTCGHVMSL 60
Query: 65 DFPGKYQDWAATDPLDLFQAQVIKNESNPKL 95
DFP KY +W DP +L+ A +K E+N K+
Sbjct: 61 DFPSKYNNWERVDPAELYSAPAVKIEANAKM 91
>UniRef100_UPI00003AAF77 UPI00003AAF77 UniRef100 entry
Length = 792
Score = 89.0 bits (219), Expect = 2e-17
Identities = 48/93 (51%), Positives = 60/93 (63%), Gaps = 4/93 (4%)
Query: 10 VAEKPSIALSIATVLSHGKMNTRRGST---EVHEFDGKFRKAPARFKVTSVIGHVFSVDF 66
VAEKPS+A SIA +LS G M++R+G VHE+ G F A FK+TSV GHV ++DF
Sbjct: 1 VAEKPSLAQSIAKILSRGNMSSRKGLNGVCSVHEYTGSFIGQSAHFKMTSVCGHVMTLDF 60
Query: 67 PGKYQDWAATDPLDLF-QAQVIKNESNPKLVDV 98
GKY W DP +LF +A K E+NPKL V
Sbjct: 61 IGKYNSWDKVDPAELFSKAPTEKKEANPKLTMV 93
>UniRef100_UPI000035EF99 UPI000035EF99 UniRef100 entry
Length = 662
Score = 88.6 bits (218), Expect = 3e-17
Identities = 48/97 (49%), Positives = 65/97 (66%), Gaps = 5/97 (5%)
Query: 9 MVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFSVD 65
MVAEKPS+A SIA +LS G ++R+G + V+E+ G F A FK+TSV GHV S+D
Sbjct: 1 MVAEKPSLAQSIAKILSKGNCSSRKGLNGACSVYEYTGSFLGQAAHFKMTSVCGHVMSLD 60
Query: 66 FPGKYQDWAATDPLDLF-QAQVIKNESNPKLVDVVIW 101
F GKY +W DP +LF +A K E+NPKL +++ W
Sbjct: 61 FIGKYNNWDKVDPAELFSKAPTEKKEANPKL-NMIKW 96
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.323 0.137 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 194,783,020
Number of Sequences: 2790947
Number of extensions: 7076359
Number of successful extensions: 13250
Number of sequences better than 10.0: 91
Number of HSP's better than 10.0 without gapping: 71
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 13114
Number of HSP's gapped (non-prelim): 93
length of query: 125
length of database: 848,049,833
effective HSP length: 101
effective length of query: 24
effective length of database: 566,164,186
effective search space: 13587940464
effective search space used: 13587940464
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 67 (30.4 bits)
Medicago: description of AC147484.9


