

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147484.3 - phase: 0 /pseudo
(526 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
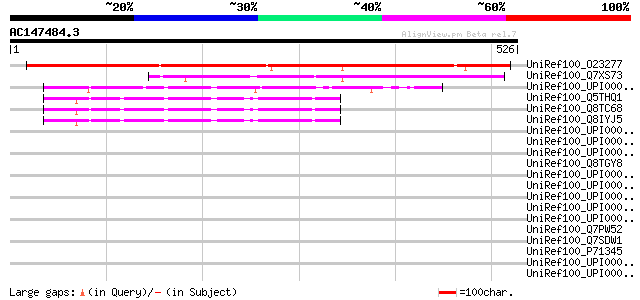
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O23277 Hypothetical protein dl3130w [Arabidopsis thali... 489 e-137
UniRef100_Q7XS73 OSJNBa0020I02.9 protein [Oryza sativa] 227 7e-58
UniRef100_UPI000021D525 UPI000021D525 UniRef100 entry 49 3e-04
UniRef100_Q5THQ1 OTTHUMP00000028520 [Homo sapiens] 48 6e-04
UniRef100_Q8TC68 MGC40042 protein [Homo sapiens] 48 6e-04
UniRef100_Q8IYJ5 MGC40042 protein [Homo sapiens] 48 6e-04
UniRef100_UPI000036C9AE UPI000036C9AE UniRef100 entry 47 0.001
UniRef100_UPI000042CC32 UPI000042CC32 UniRef100 entry 41 0.098
UniRef100_UPI00002A3B67 UPI00002A3B67 UniRef100 entry 41 0.098
UniRef100_Q8TGY8 Lhr-like Superfamily II helicase [Methanopyrus ... 38 0.83
UniRef100_UPI00001CFDC7 UPI00001CFDC7 UniRef100 entry 37 1.1
UniRef100_UPI00004532F5 UPI00004532F5 UniRef100 entry 36 2.4
UniRef100_UPI0000281841 UPI0000281841 UniRef100 entry 36 2.4
UniRef100_UPI00003A97DE UPI00003A97DE UniRef100 entry 35 4.1
UniRef100_UPI00002C5FCC UPI00002C5FCC UniRef100 entry 35 4.1
UniRef100_Q7PW52 ENSANGP00000016531 [Anopheles gambiae str. PEST] 35 5.4
UniRef100_Q7SDW1 Hypothetical protein [Neurospora crassa] 35 5.4
UniRef100_P71345 Branched-chain amino acid transport system carr... 35 7.0
UniRef100_UPI0000499F7B UPI0000499F7B UniRef100 entry 34 9.1
UniRef100_UPI0000498657 UPI0000498657 UniRef100 entry 34 9.1
>UniRef100_O23277 Hypothetical protein dl3130w [Arabidopsis thaliana]
Length = 1268
Score = 489 bits (1259), Expect = e-137
Identities = 268/533 (50%), Positives = 356/533 (66%), Gaps = 35/533 (6%)
Query: 18 CSQGHRSSLNLQTNQGGTICLVCFSNLISNPSSPTIHVSYALSQLSRSLSIPQFLHSIVT 77
C+ GHRS+++L+ +QGGT CL+CFSNL+S+P PT+HVSYAL QLS ++S P FL ++++
Sbjct: 31 CANGHRSTISLRDDQGGTFCLICFSNLVSDPRIPTVHVSYALHQLSIAISEPIFLRTLLS 90
Query: 78 FHPHFLVSPLVAALSSFDDEPIAEQLVHLILNLSASSDPSVCREFVARVSDRVSSGALGW 137
H HFLVSPLV ALSS DD PIA Q++ +I L + + S+ +FV R+SD++SSGALGW
Sbjct: 91 SHIHFLVSPLVHALSSIDDAPIAIQIMDMISLLCSVEESSIGEDFVERISDQLSSGALGW 150
Query: 138 SSRQLHSLHCLGVLLNCEKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLYKLF 197
S RQLH LHC GVL++CE +++ HI+D +L+ LV GLQLPSEEIRGE+LF LYK
Sbjct: 151 SRRQLHMLHCFGVLMSCE-NININSHIRDKEALVCQLVEGLQLPSEEIRGEILFALYKFS 209
Query: 198 ALHSTSDEGDGSDMLIPFCPKLLYLLGDVLLKTQNDDVRLNCIALLTMLAQRQLLREEPA 257
AL T DG ++L CPKLL L + L KTQ DDVRLNC+ALLT+LAQ+ LL +
Sbjct: 210 ALQFTEQNVDGIEVLSLLCPKLLCLSLEALAKTQRDDVRLNCVALLTILAQQGLLANSHS 269
Query: 258 YDTGNMSLSGEVN---FKETEDVPEGASLVNLFAEAIKGPLLSSVSQVQIGALDLLFHYL 314
+MSL EV+ + E+V L LFAEAIKGPLLS+ S+VQI LDL+FHY+
Sbjct: 270 NSASSMSLD-EVDDDPMQTAENVAARPCLNVLFAEAIKGPLLSTDSEVQIKTLDLIFHYI 328
Query: 315 SSVGTSGNQIQVLVEENIADYLFEILRLS-------------------------VPFHPV 349
S T QIQV+VEEN+ADY+FEILRLS VP HP
Sbjct: 329 SQESTPSKQIQVMVEENVADYIFEILRLSAEHSFRKRLVIGFPSVIRVLHYVGEVPCHPF 388
Query: 350 QYETLKLIYECISECPGAVSTSQMEELVLVLIKMLTKHSDGEMGMIPETFIMACSVFVAL 409
Q +TLKLI CIS+ PG S+SQ++E+ LVL KML ++ EMG+ P+ F + CSVFV+L
Sbjct: 389 QIQTLKLISSCISDFPGIASSSQVQEIALVLKKMLERYYSQEMGLFPDAFAIICSVFVSL 448
Query: 410 MRSPSCNGALDLSKSIEEAVKQAILACLYVSERNINQILQCFYLLKEAYAYSHDENSTHI 469
M++PS D+ S++E+++ +ILA L + E++ QIL YLL E Y Y ST I
Sbjct: 449 MKTPSFGETADVLTSLQESLRHSILASLSLPEKDSTQILHAVYLLNEVYVYC--TASTSI 506
Query: 470 SK---LELRSGILDICRTHLLPWLATGIMNGEEELFLGLLEIFHSILLLHCSI 519
+K +ELR ++D+C +HLLPW + + EE LG++E FHSILL + I
Sbjct: 507 NKTICIELRHCVIDVCTSHLLPWFLSDVNEVNEEATLGIMETFHSILLQNSDI 559
>UniRef100_Q7XS73 OSJNBa0020I02.9 protein [Oryza sativa]
Length = 1176
Score = 227 bits (578), Expect = 7e-58
Identities = 167/446 (37%), Positives = 226/446 (50%), Gaps = 86/446 (19%)
Query: 145 LHCLGVLLNCEKEDDLHVHIKDVYSLISVLVTGLQLP----------------------- 181
LHCLG+LLN K D +I D SL LV L+LP
Sbjct: 5 LHCLGILLNSTK--DAATYIGDKQSLYLNLVNNLRLPRLIPLHIDTFLALRITLSDSILN 62
Query: 182 ----SEEIRGEVLFVLYKLFALHST--SDEGDGSDM-LIPFCPKLLYLLGDVLLKTQNDD 234
S+EIRGE+LFVLYKL L++T D D ++ L LL +VLLKTQNDD
Sbjct: 63 LFWYSDEIRGEILFVLYKLSLLNATPWDDICDNDNVDLSAIGRSLLQFSLEVLLKTQNDD 122
Query: 235 VRLNCIALLTMLAQRQLLREEPAYDTGNMSLSGEVNFKETED-VPEGASLVNLFAEAIKG 293
VRLNCIALL LA++ A+D +S +N E ED VP SLV LFAEA+KG
Sbjct: 123 VRLNCIALLLTLAKKG------AFDILLLSDPSLINSAEAEDNVPLNDSLVILFAEAVKG 176
Query: 294 PLLSSVSQVQIGALDLLFHYLSSVGTSGNQIQVLVEENIADYLFEILRLS---------- 343
LLS+ +VQ G L+L+FH+LSS + ++ L+++N+ADY+FE+LRLS
Sbjct: 177 SLLSTNIEVQTGTLELIFHFLSS-DANIFVLKTLIDQNVADYVFEVLRLSGNNDPLVISS 235
Query: 344 ----------------------------------VPFHPVQYETLKLIYECISECPGAVS 369
+PFHPVQ + L+L+ I C G +S
Sbjct: 236 IKVLSILANSEERFKEKLAIAVSTLLPVLHYVSEIPFHPVQSQVLRLVCISIINCSGILS 295
Query: 370 TSQMEELVLVLIKMLTKHSDGEMGMIPETFIMACSVFVALMRSPSCNGALDLSKSIEEAV 429
SQ E++ L +L +H +GE+GM ETF + CS+ V +++ PS + L I EA
Sbjct: 296 LSQEEQIACTLSAILRRHGNGELGMSSETFALVCSMLVEILKLPSADDIQKLPSFIVEAS 355
Query: 430 KQAILACLYVSERNINQILQCFYLLKEAYAYSHDENSTHI-SKLELRSGILDICRTHLLP 488
K AI + I LLKEA + + N I K L I++ C T+LLP
Sbjct: 356 KHAISLTFSHEYDCLFLIPHSLLLLKEALIFCLEGNKDQILRKKSLEDSIIETCETYLLP 415
Query: 489 WLATGIMNG-EEELFLGLLEIFHSIL 513
WL + I++G +EE G+L+IF IL
Sbjct: 416 WLESAIVDGNDEETLSGILQIFQIIL 441
>UniRef100_UPI000021D525 UPI000021D525 UniRef100 entry
Length = 566
Score = 49.3 bits (116), Expect = 3e-04
Identities = 94/442 (21%), Positives = 180/442 (40%), Gaps = 68/442 (15%)
Query: 36 ICLVCFSNLISNPSSPTIHVSYALSQLSRSL-SIPQFLHSIVTFHP----HFLVSPLVAA 90
+CL C L+ P + + L +L + +V P HF V L
Sbjct: 46 VCLACALELLPEPGVSLVRKKHVLFCFQDALVRHTSLVTQLVAQDPRVCIHF-VRVLFGL 104
Query: 91 LSSFDDEPIAEQLVHLILNLSASSDPSVCREFVARVSDR--VSSGALGWSSRQLHSLHCL 148
LSS +D +A+ + +++ L+ + + + R + ++G S L + L
Sbjct: 105 LSSVEDGSVADLCIEVLIQLTTQPNMEQTIQCLLNECHRELCNLPSMGGS---LATTTLL 161
Query: 149 GVLLNC--EKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLYKLFALHSTSDEG 206
G L+N + +DL + + +L+ L+ GL P+E ++ + ++ KL++ + ++
Sbjct: 162 GKLVNAIPDLAEDL---VMEYGNLMEHLLRGLVYPNEGVQASICYLYGKLYSSPTAAEML 218
Query: 207 DGSDMLIPFCPKLLYLLGDVLLKTQNDDVRLNCIALLTMLAQRQLLR------------- 253
G F KL L L Q D+++NC+ LL L + L
Sbjct: 219 SGH-----FREKLCALFLSTLDNAQTRDLQINCLGLLRQLLKYDLFVSVIMSKSVPVEGA 273
Query: 254 ---EEPAYDTGNMSLSGEVNFKETEDVPEGASLVNLFAEAIKGPLLSSVSQVQIGALDLL 310
E+P+ +T ++ L + ++V + AS + A + P +V+ + + L
Sbjct: 274 ESVEQPSRET-SLPLVLKKFLLSRDEVLQVASSHCITAVLVHSPAKHAVAFIYADIPEFL 332
Query: 311 FHYLSSVGTSGNQIQVLVEENIADYLFEILRLSVPFHPVQYETLKLIYECISECPGAVST 370
F +LSS ++LV + Y IL P + T+ I + G++
Sbjct: 333 FEHLSSSS------EILVWSS---YNCLILLAEEPLFFSKCHTVYGIEAVVRSLQGSLQM 383
Query: 371 SQME---ELVLVLIKMLTKHSDGEMGMIPETFIMACSVFVALMRSPSCNGALDLSKSIEE 427
+ E + +L+ ++LT+ PE + S S C D S++++E
Sbjct: 384 TNTELHKQGLLLFAEILTRQ--------PEEIRLFTS-------SAMCR---DASRALQE 425
Query: 428 AVKQAILACLYVSERNINQILQ 449
AV +LA + R I+ L+
Sbjct: 426 AVSSPVLAVAAEALRAISAFLR 447
>UniRef100_Q5THQ1 OTTHUMP00000028520 [Homo sapiens]
Length = 1280
Score = 48.1 bits (113), Expect = 6e-04
Identities = 70/317 (22%), Positives = 129/317 (40%), Gaps = 32/317 (10%)
Query: 36 ICLVCFSNLISNPSSPTIHVSYALSQLSRSLS-----IPQFLHSIVTFHPHFLVSPLVAA 90
+CL C L+ +P + + LS +L + Q + HF +S L
Sbjct: 43 LCLACALELLPDPGVSLVRKKHMLSCFQDALVRHTSLVTQLVSQDQRVCIHF-ISVLFGL 101
Query: 91 LSSFDDEPIAEQLVHLILNLSASSDPSVCREFVARVSDRVSSGALGWSSRQ--LHSLHCL 148
L S +D + + + +++ ++ + + + D S + L +L L
Sbjct: 102 LCSMEDGSVTDLCIEVLIQITTQLK---LEQTIRCLLDECHKELCNMPSMRGSLATLTLL 158
Query: 149 GVLLNC--EKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLYKLFALHSTSDEG 206
G L++ D+L + + +L+ L+ GL PSE I+ V ++ KL++ ++
Sbjct: 159 GKLVDAIPALADEL---VMEHGNLMEHLLRGLVYPSEGIQASVCYLYGKLYSSPVAAEML 215
Query: 207 DGSDMLIPFCPKLLYLLGDVLLKTQNDDVRLNCIALLTMLAQRQLLREEPAYDTGNMSLS 266
G F KL L +L Q ++++NC+ LL RQLL+ YD +
Sbjct: 216 SGH-----FREKLFPLFLSILDGAQTKELQINCLGLL-----RQLLK----YDLFVSMIM 261
Query: 267 GEVNFKETEDVPEGASLVNLFAEAIKGPLLSSVSQVQIGALDLLFHYLSSVGTSGNQIQV 326
+ E+ EG+S +K LLS +Q+ + + L V +
Sbjct: 262 NQDGLGESAKNIEGSSGNTSLPLVLKKLLLSRDETLQVASAHCITAVL--VHSPAKHASA 319
Query: 327 LVEENIADYLFEILRLS 343
+ +I ++LFE L S
Sbjct: 320 FIHADIPEFLFEHLSSS 336
>UniRef100_Q8TC68 MGC40042 protein [Homo sapiens]
Length = 582
Score = 48.1 bits (113), Expect = 6e-04
Identities = 70/317 (22%), Positives = 129/317 (40%), Gaps = 32/317 (10%)
Query: 36 ICLVCFSNLISNPSSPTIHVSYALSQLSRSLS-----IPQFLHSIVTFHPHFLVSPLVAA 90
+CL C L+ +P + + LS +L + Q + HF +S L
Sbjct: 56 LCLACALELLPDPGVSLVRKKHMLSCFQDALVRHTSLVTQLVSQDQRVCIHF-ISVLFGL 114
Query: 91 LSSFDDEPIAEQLVHLILNLSASSDPSVCREFVARVSDRVSSGALGWSSRQ--LHSLHCL 148
L S +D + + + +++ ++ + + + D S + L +L L
Sbjct: 115 LCSMEDGSVTDLCIEVLIQITTQLK---LEQTIRCLLDECHKELCNMPSMRGSLATLTLL 171
Query: 149 GVLLNC--EKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLYKLFALHSTSDEG 206
G L++ D+L + + +L+ L+ GL PSE I+ V ++ KL++ ++
Sbjct: 172 GKLVDAIPALADEL---VMEHGNLMEHLLRGLVYPSEGIQASVCYLYGKLYSSPVAAEML 228
Query: 207 DGSDMLIPFCPKLLYLLGDVLLKTQNDDVRLNCIALLTMLAQRQLLREEPAYDTGNMSLS 266
G F KL L +L Q ++++NC+ LL RQLL+ YD +
Sbjct: 229 SGH-----FREKLFPLFLSILDGAQTKELQINCLGLL-----RQLLK----YDLFVSMIM 274
Query: 267 GEVNFKETEDVPEGASLVNLFAEAIKGPLLSSVSQVQIGALDLLFHYLSSVGTSGNQIQV 326
+ E+ EG+S +K LLS +Q+ + + L V +
Sbjct: 275 NQDGLGESAKNIEGSSGNTSLPLVLKKLLLSRDETLQVASAHCITAVL--VHSPAKHASA 332
Query: 327 LVEENIADYLFEILRLS 343
+ +I ++LFE L S
Sbjct: 333 FIHADIPEFLFEHLSSS 349
>UniRef100_Q8IYJ5 MGC40042 protein [Homo sapiens]
Length = 452
Score = 48.1 bits (113), Expect = 6e-04
Identities = 70/317 (22%), Positives = 129/317 (40%), Gaps = 32/317 (10%)
Query: 36 ICLVCFSNLISNPSSPTIHVSYALSQLSRSLS-----IPQFLHSIVTFHPHFLVSPLVAA 90
+CL C L+ +P + + LS +L + Q + HF +S L
Sbjct: 51 LCLACALELLPDPGVSLVRKKHMLSCFQDALVRHTSLVTQLVSQDQRVCIHF-ISVLFGL 109
Query: 91 LSSFDDEPIAEQLVHLILNLSASSDPSVCREFVARVSDRVSSGALGWSSRQ--LHSLHCL 148
L S +D + + + +++ ++ + + + D S + L +L L
Sbjct: 110 LCSMEDGSVTDLCIEVLIQITTQLK---LEQTIRCLLDECHKELCNMPSMRGSLATLTLL 166
Query: 149 GVLLNC--EKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLYKLFALHSTSDEG 206
G L++ D+L + + +L+ L+ GL PSE I+ V ++ KL++ ++
Sbjct: 167 GKLVDAIPALADEL---VMEHGNLMEHLLRGLVYPSEGIQASVCYLYGKLYSSPVAAEML 223
Query: 207 DGSDMLIPFCPKLLYLLGDVLLKTQNDDVRLNCIALLTMLAQRQLLREEPAYDTGNMSLS 266
G F KL L +L Q ++++NC+ LL RQLL+ YD +
Sbjct: 224 SGH-----FREKLFPLFLSILDGAQTKELQINCLGLL-----RQLLK----YDLFVSMIM 269
Query: 267 GEVNFKETEDVPEGASLVNLFAEAIKGPLLSSVSQVQIGALDLLFHYLSSVGTSGNQIQV 326
+ E+ EG+S +K LLS +Q+ + + L V +
Sbjct: 270 NQDGLGESAKNIEGSSGNTSLPLVLKKLLLSRDETLQVASAHCITAVL--VHSPAKHASA 327
Query: 327 LVEENIADYLFEILRLS 343
+ +I ++LFE L S
Sbjct: 328 FIHADIPEFLFEHLSSS 344
>UniRef100_UPI000036C9AE UPI000036C9AE UniRef100 entry
Length = 582
Score = 47.4 bits (111), Expect = 0.001
Identities = 70/317 (22%), Positives = 129/317 (40%), Gaps = 32/317 (10%)
Query: 36 ICLVCFSNLISNPSSPTIHVSYALSQLSRSLS-----IPQFLHSIVTFHPHFLVSPLVAA 90
+CL C L+ +P + + LS +L + Q + HF +S L
Sbjct: 56 LCLACALELLPDPGVSLVRKKHMLSCFQDALVRHTSLVTQLVSQDQRVCIHF-ISVLFGL 114
Query: 91 LSSFDDEPIAEQLVHLILNLSASSDPSVCREFVARVSDRVSSGALGWSSRQ--LHSLHCL 148
L S +D + + +++ ++ + + + D S + L +L L
Sbjct: 115 LCSMEDGSVTNLCIEVLIQITTHLK---LEQTIHCLLDECHKELCNMPSMRGSLATLTLL 171
Query: 149 GVLLNC--EKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLYKLFALHSTSDEG 206
G L++ D+L + + +L+ L+ GL PSE I+ V ++ KL++ ++
Sbjct: 172 GKLVDAIPALADEL---VMEHGNLMEHLLRGLVYPSEGIQASVCYLYGKLYSSPVAAEML 228
Query: 207 DGSDMLIPFCPKLLYLLGDVLLKTQNDDVRLNCIALLTMLAQRQLLREEPAYDTGNMSLS 266
G F KL L +L Q ++++NC+ LL RQLL+ YD +
Sbjct: 229 SGH-----FREKLFPLFLSILDGAQTKELQINCLGLL-----RQLLK----YDLFVSMIM 274
Query: 267 GEVNFKETEDVPEGASLVNLFAEAIKGPLLSSVSQVQIGALDLLFHYLSSVGTSGNQIQV 326
+ +E+ EG+S +K LLS +Q+ + + L V +
Sbjct: 275 NQDGMEESAKNIEGSSGNTSLPLVLKKLLLSRDETLQVASAHCITAVL--VHSPAKHAPA 332
Query: 327 LVEENIADYLFEILRLS 343
+ +I ++LFE L S
Sbjct: 333 FIHADIPEFLFEHLSSS 349
>UniRef100_UPI000042CC32 UPI000042CC32 UniRef100 entry
Length = 864
Score = 40.8 bits (94), Expect = 0.098
Identities = 37/125 (29%), Positives = 57/125 (45%), Gaps = 15/125 (12%)
Query: 102 QLVHLILNLSASSDPS--VCREFVARVSDRVSSGALGWSSRQLHSLHCLGVLLNCEKEDD 159
QLVH L+ ++D + +C F V V+ L +L +L LG + E +
Sbjct: 349 QLVHATLDRETTADDTDLLCEVFKG-VDSMVTQEILAAGMERLATL--LGHAIAAESQQQ 405
Query: 160 LHVHIKDVYSLISVLVTGLQLPSEEI-RGEVLFVLYKLFAL---------HSTSDEGDGS 209
L HIK + L+ +L + LPS+ + VL +KL L + S DGS
Sbjct: 406 LETHIKSIDQLLRILDSMNNLPSQSVTEPSVLNARHKLLDLLALALQTVERNLSTTRDGS 465
Query: 210 DMLIP 214
D+L+P
Sbjct: 466 DLLLP 470
>UniRef100_UPI00002A3B67 UPI00002A3B67 UniRef100 entry
Length = 204
Score = 40.8 bits (94), Expect = 0.098
Identities = 18/44 (40%), Positives = 28/44 (62%)
Query: 439 VSERNINQILQCFYLLKEAYAYSHDENSTHISKLELRSGILDIC 482
+SE I + ++CF LLKEA +HDE + H+ +L I+D+C
Sbjct: 84 LSESVIKRTIKCFDLLKEAEESAHDEKNPHLHELGTSDTIIDVC 127
>UniRef100_Q8TGY8 Lhr-like Superfamily II helicase [Methanopyrus kandleri]
Length = 910
Score = 37.7 bits (86), Expect = 0.83
Identities = 27/97 (27%), Positives = 46/97 (46%), Gaps = 12/97 (12%)
Query: 267 GEVNFKETEDVPEGASLVNLFAEAIKGPLLSSVSQVQIGALDLLFHYLSSVGTSGNQIQV 326
G V +D+ E A + A+KG L +++ G LD+L H L + G+Q+ V
Sbjct: 345 GLVITSNPDDLAEAAVICR---RALKGSL--EPTEIPEGCLDVLAHQLVGLCLDGHQVTV 399
Query: 327 LVEENIADYLFEILRLSVPFHPVQYETLKLIYECISE 363
DY E+ R + P+ + ETL+ + E + +
Sbjct: 400 -------DYALEVFRRAYPYRHLDAETLREVAEFLDD 429
>UniRef100_UPI00001CFDC7 UPI00001CFDC7 UniRef100 entry
Length = 1468
Score = 37.4 bits (85), Expect = 1.1
Identities = 61/276 (22%), Positives = 112/276 (40%), Gaps = 39/276 (14%)
Query: 36 ICLVCFSNLISNPSSPTIHVSYALSQLSRSL-SIPQFLHSIVTFHPHFLVS--PLVAALS 92
+CL C L+ P + + L +L + +V P + ++ L
Sbjct: 38 VCLACALELLPEPGVSLVRKKHVLFCFQDALVRHTSLVTQLVAQDPRVCIHFVRVLFVLI 97
Query: 93 SFDDEPIAEQLVHLILNLSASSDPSVCREFVARVSDRVSSGALGWSSRQLHSLHCLGVLL 152
+P EQ + +LN E + + S G L + LG L+
Sbjct: 98 QLTTQPNMEQTIQCLLN-----------ECHRELCNLPSMGG------SLATTTLLGKLV 140
Query: 153 NC--EKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLYKLFALHSTSDEGDGSD 210
N + +DL + + +L+ L+ GL P+E ++ + ++ KL++ + ++ G
Sbjct: 141 NAIPDLAEDL---VMEYGNLMEHLLRGLVYPNEGVQASICYLYGKLYSSPTAAEMLSGH- 196
Query: 211 MLIPFCPKLLYLLGDVLLKTQNDDVRLNCIALLTMLAQRQLLREEPAYDTGNMSLSGEVN 270
F KL L L Q D+++NC+ LL RQLL+ + + + MS S V
Sbjct: 197 ----FREKLCALFLSTLDNAQTRDLQINCLGLL-----RQLLKYD-LFVSVIMSKSVPVE 246
Query: 271 FKETEDVPEGASLVNLFAEAIKGPLLSSVSQVQIGA 306
E+ + P + + L +K LLS +Q+ +
Sbjct: 247 GAESVEQPSRETSLPL---VLKKFLLSRDEVLQVAS 279
>UniRef100_UPI00004532F5 UPI00004532F5 UniRef100 entry
Length = 681
Score = 36.2 bits (82), Expect = 2.4
Identities = 58/245 (23%), Positives = 103/245 (41%), Gaps = 41/245 (16%)
Query: 248 QRQLLREEPAYDTGNMSLSGEVNFKETEDVPEGASLVN----LFAEAIKGPLLSSVSQV- 302
QR+++ + P+ D +S+S V + V EG +N + + AI LL+S+SQ+
Sbjct: 302 QREVVIQHPSID--QVSISIGVKLEICPLVDEGEIFMNAVDTIISNAIHTKLLNSLSQLA 359
Query: 303 ---QIGALDLLFHYLSSVGTSGNQIQVLVEENIADYLFEILRLSVPFHPVQYETLKLIYE 359
++ ++ F+ S N ++ +E +F + I +
Sbjct: 360 CDEELTSISWDFYNSSRENCGWNTFSIVSDEKNWRAVFRLG----------------IQQ 403
Query: 360 CISECPGAVSTSQMEELVLVLIKMLTKHSDGEMGMIPETFIMACSVFVALMRSPSCNGAL 419
+S C +S + EE+VL+ I K ++ E+G P+ V L+ C G++
Sbjct: 404 IVSICNTKMSMEEFEEIVLITISDYKKSAEEELGDDPKL------VLDGLIDDWLC-GSI 456
Query: 420 DLSKSIEEAVKQAILACLYVSERNINQILQC-FYLLKEAYAYSHDENSTHISKLELRSGI 478
LSK KQ V +R +I+QC L Y DEN H +K +G
Sbjct: 457 PLSK------KQEYQLFCKVVDRINPEIIQCRCKALFSHIIYYFDEND-HYNKESSFTGC 509
Query: 479 LDICR 483
+ + +
Sbjct: 510 IFVSK 514
>UniRef100_UPI0000281841 UPI0000281841 UniRef100 entry
Length = 307
Score = 36.2 bits (82), Expect = 2.4
Identities = 33/110 (30%), Positives = 50/110 (45%), Gaps = 9/110 (8%)
Query: 371 SQMEELVLVLIKMLTKHSDGEMGMIPETFIMACSVFVALMRSPSCNGALDLSKSIEEAVK 430
SQ E + I ++T + E ++ +TF S V + R+ S N +++SK E+ K
Sbjct: 33 SQFYENKTIPILIITNYEVSERSLLEKTFSNKESKRVVIKRAKSIN-EINISKLAEKNAK 91
Query: 431 QAILACLYVSERNINQI--------LQCFYLLKEAYAYSHDENSTHISKL 472
QA+ L S+ N N I L + L E Y SH + S I L
Sbjct: 92 QALTQKLIQSDTNNNLIENLAKKFKLNNYIDLIEVYDNSHIQGSDSIGAL 141
>UniRef100_UPI00003A97DE UPI00003A97DE UniRef100 entry
Length = 438
Score = 35.4 bits (80), Expect = 4.1
Identities = 38/175 (21%), Positives = 80/175 (45%), Gaps = 16/175 (9%)
Query: 169 SLISVLVTGLQLPSEEIRGEVLFVLYKLFALHSTSDEGDGSDMLIPFCPKLLYLLGDVLL 228
+L+ L+ GL P+E ++ V ++ KL++ +++ + + F +L L +
Sbjct: 62 NLVDHLLVGLMYPNEGVKAAVCYLYGKLYSSPVGTEK-----LSVHFEERLCGLFLNTFG 116
Query: 229 KTQNDDVRLNCIALLTMLAQRQLLREEPAYDTGNMSLSGEVNFKETEDVPEGASLVNLFA 288
+ Q ++++NC+ LL ++LL+ + + + M S E D+ EG + + L
Sbjct: 117 QAQTKELQVNCLGLL-----KELLKSD-RFVSILMDNSKPEEESENSDLLEGENPLPLI- 169
Query: 289 EAIKGPLLSSVSQVQIGALDLLFHYLSSVGTSGNQIQVLVEENIADYLFEILRLS 343
+K LLS +Q + + L V + + ++ ++LFE L S
Sbjct: 170 --LKKLLLSRDEMLQAASSHCMAAVL--VHSPSRYAPAFIHADVPEFLFECLMCS 220
>UniRef100_UPI00002C5FCC UPI00002C5FCC UniRef100 entry
Length = 303
Score = 35.4 bits (80), Expect = 4.1
Identities = 31/110 (28%), Positives = 50/110 (45%), Gaps = 9/110 (8%)
Query: 371 SQMEELVLVLIKMLTKHSDGEMGMIPETFIMACSVFVALMRSPSCNGALDLSKSIEEAVK 430
SQ E + I +LT + E ++ +TF + + + + ++ S N + +SK E+ K
Sbjct: 34 SQFYENKTIPILILTNYEVSEKTLLEKTFSIKENKQIVIKQAKSKN-EISISKLAEKNAK 92
Query: 431 QAILACLYVSERNINQILQCFYLLK--------EAYAYSHDENSTHISKL 472
QA+ L S+ NIN I + K E Y SH + S I L
Sbjct: 93 QALTQKLIQSDTNINLIENLAKIFKLNNNIDLIEVYDNSHIQGSDSIGAL 142
>UniRef100_Q7PW52 ENSANGP00000016531 [Anopheles gambiae str. PEST]
Length = 544
Score = 35.0 bits (79), Expect = 5.4
Identities = 41/176 (23%), Positives = 77/176 (43%), Gaps = 30/176 (17%)
Query: 346 FHPVQYETLKLIYECISECPGAVSTSQMEELVLVLIKMLTKHSDGEMGMIPETFIMACSV 405
F+P Q ++ E ++ G V S + V ++++ + G++ +P++F+
Sbjct: 79 FYPSQTSYSSIVGETLAAGLGVVGFSWICSPVCTELEVIMMNWLGQLLNLPKSFL----- 133
Query: 406 FVALMRSPSCNGALDLSKSIEEAVKQAILACLYVSERNINQILQCFYLLKEA------YA 459
NG + S E++ A+LA E+ + ++ L EA A
Sbjct: 134 -----NCDEGNGGGIIQGSASESILVAVLAA---REQAVRRLRTQHPELTEADIRGRLVA 185
Query: 460 YSHDENSTHISKLELRSGILDICRTHLLPWLATGIMNG-------EEELFLGLLEI 508
Y+ D++++ + K SGIL + LLP TGI+ G EE++ GL +
Sbjct: 186 YTSDQSNSAVEK----SGILGAIKMRLLPADETGILRGSTFIQAVEEDVAKGLFPV 237
>UniRef100_Q7SDW1 Hypothetical protein [Neurospora crassa]
Length = 560
Score = 35.0 bits (79), Expect = 5.4
Identities = 20/53 (37%), Positives = 30/53 (55%), Gaps = 1/53 (1%)
Query: 469 ISKLELRSGILDICRTHLLPWLATGIMNGEEELFLGLLEIFHSILLLHCSIIL 521
++ L + S IL + W ATG +NGE + FLG+L FH I L C++ +
Sbjct: 209 LTHLNVVSNILQMADVDGRYWSATGGLNGEGDKFLGVLPFFH-IYGLTCALFM 260
>UniRef100_P71345 Branched-chain amino acid transport system carrier protein
[Haemophilus influenzae]
Length = 436
Score = 34.7 bits (78), Expect = 7.0
Identities = 17/72 (23%), Positives = 32/72 (43%)
Query: 336 LFEILRLSVPFHPVQYETLKLIYECISECPGAVSTSQMEELVLVLIKMLTKHSDGEMGMI 395
L + LR +P Y + L+ C S C + + E + L+K S+G ++
Sbjct: 356 LLQFLRKKLPSIKFTYNSTLLVTVCFSLCDSLNNVKMLPESINSLLKHFPLSSEGMAWLV 415
Query: 396 PETFIMACSVFV 407
P ++ S+F+
Sbjct: 416 PTLVMLVASIFI 427
>UniRef100_UPI0000499F7B UPI0000499F7B UniRef100 entry
Length = 447
Score = 34.3 bits (77), Expect = 9.1
Identities = 18/68 (26%), Positives = 38/68 (55%)
Query: 151 LLNCEKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLYKLFALHSTSDEGDGSD 210
+L E E+++ H + ++ LV ++ + E++ VLY +FALHS + + +G++
Sbjct: 196 ILKIENENNIVNHRNVIKRILYCLVAIDKIKYTQGMNELVSVLYYVFALHSNNQDFEGAE 255
Query: 211 MLIPFCPK 218
+ +C K
Sbjct: 256 VSSYYCMK 263
>UniRef100_UPI0000498657 UPI0000498657 UniRef100 entry
Length = 729
Score = 34.3 bits (77), Expect = 9.1
Identities = 35/170 (20%), Positives = 68/170 (39%), Gaps = 40/170 (23%)
Query: 42 SNLISNPSSPTIHVSYALSQLSR---------------------------SLSIPQFLHS 74
+ LIS+PSS +S+ L ++S+ +L + F+
Sbjct: 26 NTLISDPSSRENGLSFILKRISQPMFREFLLRVHPQLYTHLLKSSNDDFGTLQVSLFIVH 85
Query: 75 IVTFHPHFL----------VSPLVAALSSFDDEPIAEQLVHLILNLSASSDPSVCREFVA 124
++ +PHF+ L + L S DD I + ++LN S S + + +
Sbjct: 86 ALSIYPHFINLLVKYDITVFEDLKSLLQSRDDPSITAHTLEIVLNASTSIED---KTLLQ 142
Query: 125 RVSDRVSSGALGWSSRQLHSLHCLGVLLNCEKEDDLHVHIKDVYSLISVL 174
+ D + S G + + CL +LL C V + +Y+++++L
Sbjct: 143 SLIDPLLSFFRGNVQHRNLAGKCLCILLTCSANQISFVKQEGIYAILNIL 192
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.137 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 844,599,964
Number of Sequences: 2790947
Number of extensions: 34417610
Number of successful extensions: 72256
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 16
Number of HSP's that attempted gapping in prelim test: 72234
Number of HSP's gapped (non-prelim): 26
length of query: 526
length of database: 848,049,833
effective HSP length: 132
effective length of query: 394
effective length of database: 479,644,829
effective search space: 188980062626
effective search space used: 188980062626
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 77 (34.3 bits)
Medicago: description of AC147484.3


