

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147482.6 + phase: 2 /pseudo
(675 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
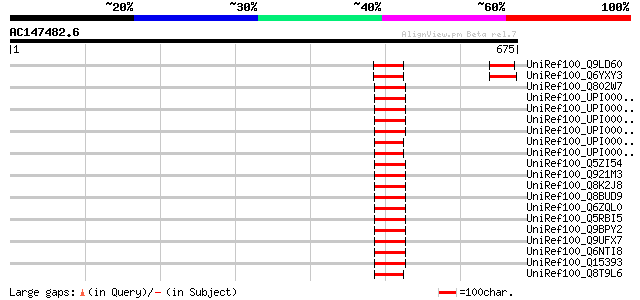
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LD60 Spliceosomal-like protein [Arabidopsis thaliana] 87 2e-15
UniRef100_Q6YXY3 Putative splicing factor 3b, subunit 3, 130kDa ... 80 2e-13
UniRef100_Q802W7 Zgc:55440 [Brachydanio rerio] 59 6e-07
UniRef100_UPI00001D1488 UPI00001D1488 UniRef100 entry 58 1e-06
UniRef100_UPI00003AA5C2 UPI00003AA5C2 UniRef100 entry 58 1e-06
UniRef100_UPI00003AA5C1 UPI00003AA5C1 UniRef100 entry 58 1e-06
UniRef100_UPI00003AA5C0 UPI00003AA5C0 UniRef100 entry 58 1e-06
UniRef100_UPI0000348B4A UPI0000348B4A UniRef100 entry 58 1e-06
UniRef100_UPI000027C433 UPI000027C433 UniRef100 entry 58 1e-06
UniRef100_Q5ZI54 Hypothetical protein [Gallus gallus] 58 1e-06
UniRef100_Q921M3 Splicing factor 3b, subunit 3, 130kDa (Mus musc... 58 1e-06
UniRef100_Q8K2J8 Sf3b3 protein [Mus musculus] 58 1e-06
UniRef100_Q8BUD9 Mus musculus 10 days lactation, adult female ma... 58 1e-06
UniRef100_Q6ZQL0 MKIAA0017 protein [Mus musculus] 58 1e-06
UniRef100_Q5RBI5 Hypothetical protein DKFZp469G1915 [Pongo pygma... 58 1e-06
UniRef100_Q9BPY2 SF3B3 protein [Homo sapiens] 58 1e-06
UniRef100_Q9UFX7 Hypothetical protein DKFZp434P041 [Homo sapiens] 58 1e-06
UniRef100_Q6NTI8 Splicing factor 3b, subunit 3, 130kDa [Homo sap... 58 1e-06
UniRef100_Q15393 Splicing factor 3B subunit 3 [Homo sapiens] 58 1e-06
UniRef100_Q8T9L6 GM01240p [Drosophila melanogaster] 54 1e-05
>UniRef100_Q9LD60 Spliceosomal-like protein [Arabidopsis thaliana]
Length = 1214
Score = 86.7 bits (213), Expect = 2e-15
Identities = 37/40 (92%), Positives = 39/40 (97%)
Query: 485 CKYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG 524
CKYR DENQLYIFADDCVPRWLTAS+H+DFDTMAGADKFG
Sbjct: 1010 CKYRRDENQLYIFADDCVPRWLTASHHVDFDTMAGADKFG 1049
Score = 63.5 bits (153), Expect = 2e-08
Identities = 30/33 (90%), Positives = 31/33 (93%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEE 672
LCEQFPTLPMDLQRKIADELDRT EILKKLE+
Sbjct: 1176 LCEQFPTLPMDLQRKIADELDRTPAEILKKLED 1208
>UniRef100_Q6YXY3 Putative splicing factor 3b, subunit 3, 130kDa [Oryza sativa]
Length = 1234
Score = 80.5 bits (197), Expect = 2e-13
Identities = 36/40 (90%), Positives = 37/40 (92%)
Query: 485 CKYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG 524
CKYR DENQLYIFADD VPRWLTA+ HIDFDTMAGADKFG
Sbjct: 1030 CKYRRDENQLYIFADDSVPRWLTAANHIDFDTMAGADKFG 1069
Score = 61.6 bits (148), Expect = 7e-08
Identities = 28/35 (80%), Positives = 32/35 (91%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQFP+LP D+QRKIADELDRT GEILKKLE+ +
Sbjct: 1196 LCEQFPSLPADMQRKIADELDRTPGEILKKLEDIR 1230
>UniRef100_Q802W7 Zgc:55440 [Brachydanio rerio]
Length = 1217
Score = 58.5 bits (140), Expect = 6e-07
Identities = 27/42 (64%), Positives = 33/42 (78%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+YR +ENQL IFADD PRW+T + +D+DTMA ADKFG IC
Sbjct: 1013 RYRRNENQLIIFADDTYPRWITTACLLDYDTMASADKFGNIC 1054
Score = 37.7 bits (86), Expect = 1.1
Identities = 16/35 (45%), Positives = 25/35 (70%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ Q+ +++ELDRT E+ KKLE+ +
Sbjct: 1178 LCEQFNSMDPHKQKSVSEELDRTPPEVSKKLEDIR 1212
>UniRef100_UPI00001D1488 UPI00001D1488 UniRef100 entry
Length = 1003
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 799 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 840
Score = 38.5 bits (88), Expect = 0.66
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 964 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 998
>UniRef100_UPI00003AA5C2 UPI00003AA5C2 UniRef100 entry
Length = 1189
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 990 RYKRNENQLIIFADDTYPRWVTTATLLDYDTVAGADKFGNIC 1031
Score = 36.6 bits (83), Expect = 2.5
Identities = 16/31 (51%), Positives = 23/31 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKL 670
LCEQF ++ + Q+ +A+ELDRT E+ KKL
Sbjct: 1159 LCEQFNSMEPNKQKNVAEELDRTPPEVSKKL 1189
>UniRef100_UPI00003AA5C1 UPI00003AA5C1 UniRef100 entry
Length = 1209
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 1008 RYKRNENQLIIFADDTYPRWVTTATLLDYDTVAGADKFGNIC 1049
Score = 39.7 bits (91), Expect = 0.29
Identities = 17/35 (48%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +A+ELDRT E+ KKLE+ +
Sbjct: 1173 LCEQFNSMEPNKQKNVAEELDRTPPEVSKKLEDIR 1207
>UniRef100_UPI00003AA5C0 UPI00003AA5C0 UniRef100 entry
Length = 1124
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 920 RYKRNENQLIIFADDTYPRWVTTATLLDYDTVAGADKFGNIC 961
Score = 39.7 bits (91), Expect = 0.29
Identities = 17/35 (48%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +A+ELDRT E+ KKLE+ +
Sbjct: 1085 LCEQFNSMEPNKQKNVAEELDRTPPEVSKKLEDIR 1119
>UniRef100_UPI0000348B4A UPI0000348B4A UniRef100 entry
Length = 312
Score = 57.8 bits (138), Expect = 1e-06
Identities = 24/39 (61%), Positives = 32/39 (81%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG 524
KY+ DE +YIFADD PR++TA+ +D+DT+AGADKFG
Sbjct: 264 KYKSDEGSIYIFADDTKPRYITATLPLDYDTLAGADKFG 302
>UniRef100_UPI000027C433 UPI000027C433 UniRef100 entry
Length = 1171
Score = 57.8 bits (138), Expect = 1e-06
Identities = 25/39 (64%), Positives = 31/39 (79%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG 524
+YR +ENQL IFADD PRW+T + +D+DTMA ADKFG
Sbjct: 1070 RYRRNENQLIIFADDTYPRWVTTACLLDYDTMASADKFG 1108
>UniRef100_Q5ZI54 Hypothetical protein [Gallus gallus]
Length = 503
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 299 RYKRNENQLIIFADDTYPRWVTTATLLDYDTVAGADKFGNIC 340
Score = 39.7 bits (91), Expect = 0.29
Identities = 17/35 (48%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +A+ELDRT E+ KKLE+ +
Sbjct: 464 LCEQFNSMEPNKQKNVAEELDRTPPEVSKKLEDIR 498
>UniRef100_Q921M3 Splicing factor 3b, subunit 3, 130kDa (Mus musculus 2 days neonate
thymus thymic cells cDNA, RIKEN full-length enriched
library, clone:E430008P19 product:SPLICING FACTOR 3B
SUBUNIT 3 (SPLICEOSOME ASSOCIATED PROTEIN 130) (SAP 130)
(SF3B130) (PRE-MRNA SPLICING FACTOR SF3B 130 kDa SUBUNIT)
homolog) [Mus musculus]
Length = 1217
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 1013 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 1054
Score = 38.5 bits (88), Expect = 0.66
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 1178 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 1212
>UniRef100_Q8K2J8 Sf3b3 protein [Mus musculus]
Length = 494
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 290 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 331
Score = 38.5 bits (88), Expect = 0.66
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 455 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 489
>UniRef100_Q8BUD9 Mus musculus 10 days lactation, adult female mammary gland cDNA,
RIKEN full-length enriched library, clone:D730019G02
product:SPLICING FACTOR 3B SUBUNIT 3 (SPLICEOSOME
ASSOCIATED PROTEIN 130) (SAP 130) (SF3B130) (PRE-MRNA
SPLICING FACTOR SF3B 130 kDa SUBUNIT) homolog [Mus
musculus]
Length = 463
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 259 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 300
Score = 38.5 bits (88), Expect = 0.66
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 424 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 458
>UniRef100_Q6ZQL0 MKIAA0017 protein [Mus musculus]
Length = 1122
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 918 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 959
Score = 38.5 bits (88), Expect = 0.66
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 1083 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 1117
>UniRef100_Q5RBI5 Hypothetical protein DKFZp469G1915 [Pongo pygmaeus]
Length = 1217
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 1013 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 1054
Score = 38.9 bits (89), Expect = 0.50
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 1178 LCEQFNSMEPNKQKNVSEELDRTPPEVTKKLEDIR 1212
>UniRef100_Q9BPY2 SF3B3 protein [Homo sapiens]
Length = 399
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 195 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 236
Score = 38.5 bits (88), Expect = 0.66
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 360 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 394
>UniRef100_Q9UFX7 Hypothetical protein DKFZp434P041 [Homo sapiens]
Length = 215
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 11 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 52
Score = 38.5 bits (88), Expect = 0.66
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 176 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 210
>UniRef100_Q6NTI8 Splicing factor 3b, subunit 3, 130kDa [Homo sapiens]
Length = 1217
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 1013 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 1054
Score = 38.5 bits (88), Expect = 0.66
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 1178 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 1212
>UniRef100_Q15393 Splicing factor 3B subunit 3 [Homo sapiens]
Length = 1217
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 1013 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 1054
Score = 38.5 bits (88), Expect = 0.66
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 1178 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 1212
>UniRef100_Q8T9L6 GM01240p [Drosophila melanogaster]
Length = 688
Score = 54.3 bits (129), Expect = 1e-05
Identities = 25/39 (64%), Positives = 31/39 (79%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG 524
+YR ENQL IFADD PRW+TA+ +D+DT+A ADKFG
Sbjct: 484 RYRRAENQLIIFADDTHPRWVTATTLLDYDTIAIADKFG 522
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.360 0.160 0.619
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 979,716,752
Number of Sequences: 2790947
Number of extensions: 35308785
Number of successful extensions: 145089
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 47
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 144998
Number of HSP's gapped (non-prelim): 91
length of query: 675
length of database: 848,049,833
effective HSP length: 134
effective length of query: 541
effective length of database: 474,062,935
effective search space: 256468047835
effective search space used: 256468047835
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 78 (34.7 bits)
Medicago: description of AC147482.6


